Submitted:
24 March 2026
Posted:
25 March 2026
You are already at the latest version
Abstract
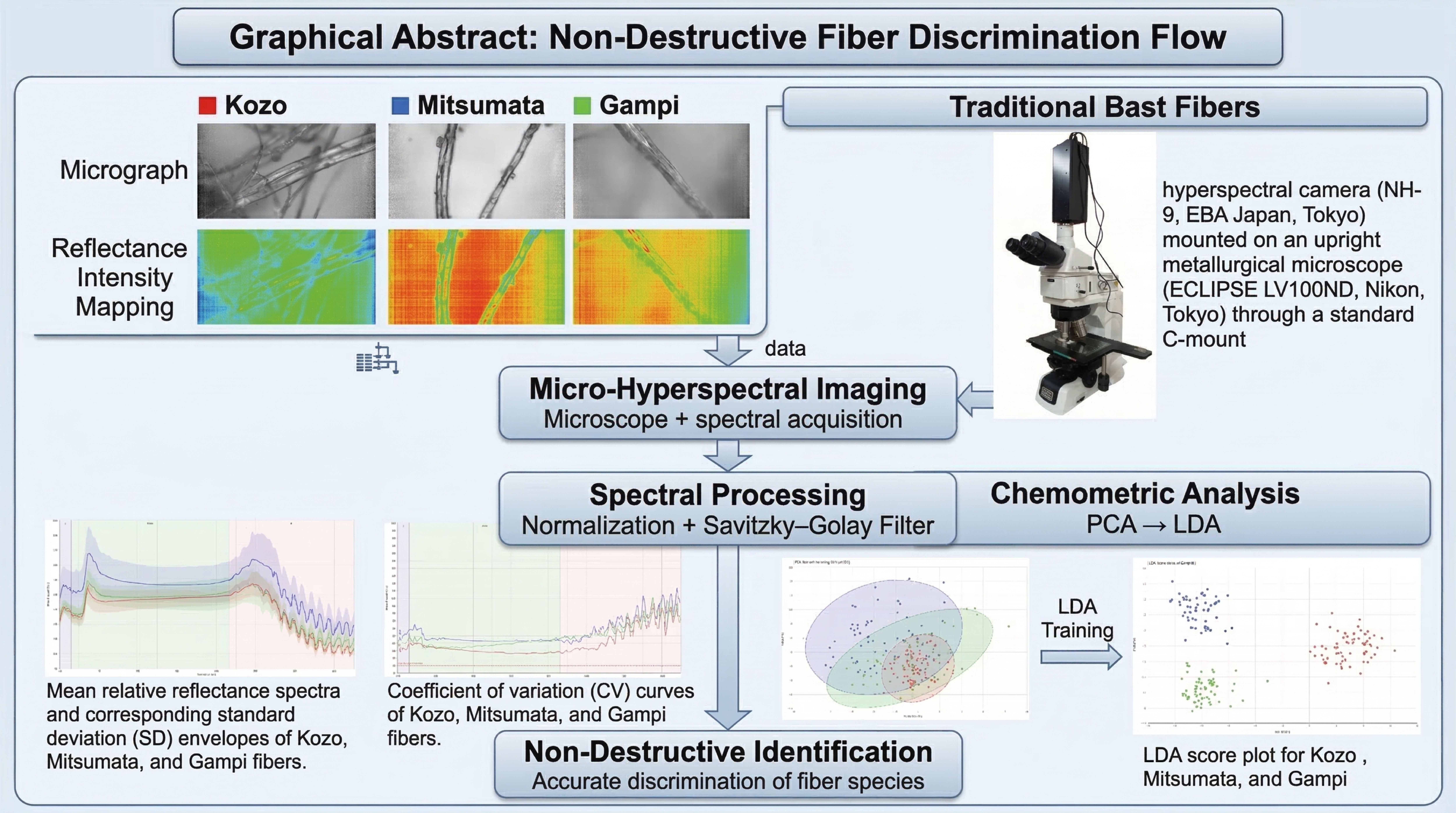
Keywords:
1. Introduction
1.1. The Materiality of Paper Artifacts and the Urgency of Conservation
1.2. Limitations of Traditional Fiber Identification
1.3. Spectroscopic Approaches for Fiber Discrimination
1.4. Hyperspectral Imaging and the Need for Micro-Scale Analysis
1.5. Research Aim
2. Materials and Methods
2.1. Optical and Chemical Basis of Bast Fiber Spectroscopy
2.2. Micro-Hyperspectral Imaging Principles
2.3. Sample Selection and Preparation
2.4. Experimental Setup and Hardware Configuration
2.5. Data Acquisition and Standardization Protocol
2.6. Spectral Pre-Processing and Chemometrics
3. Results
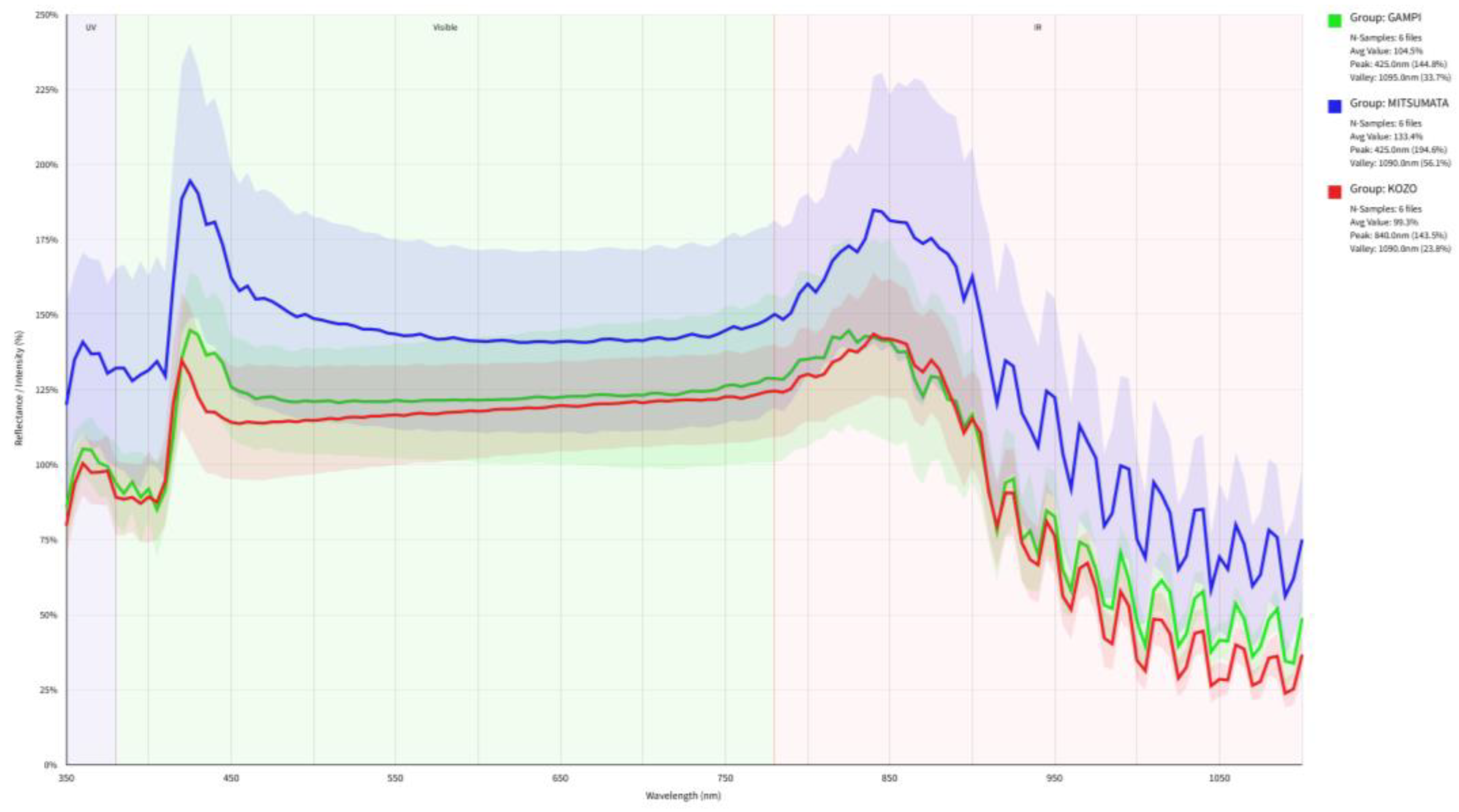
3.1. Raw Spectral Signatures and Baseline Calibration
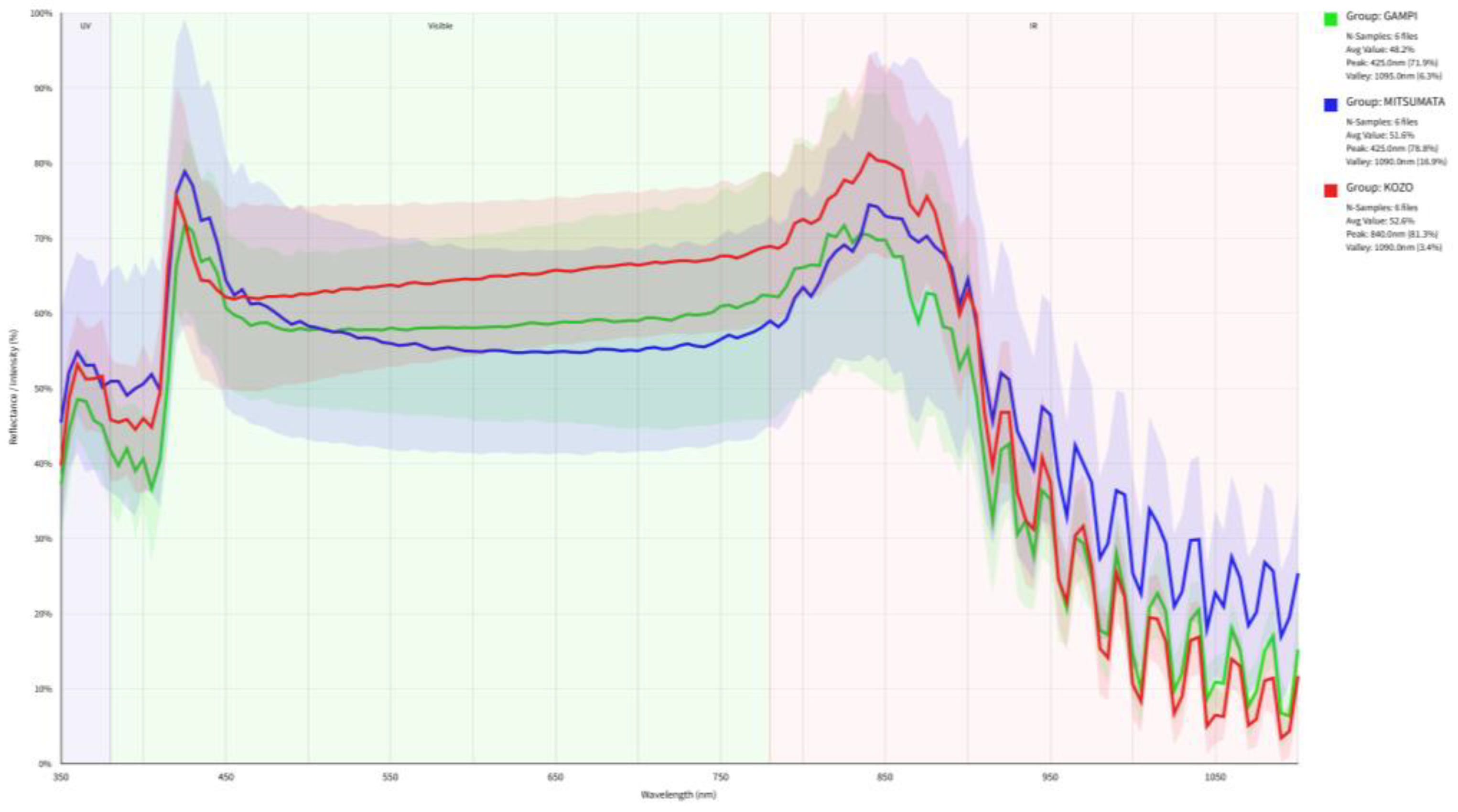
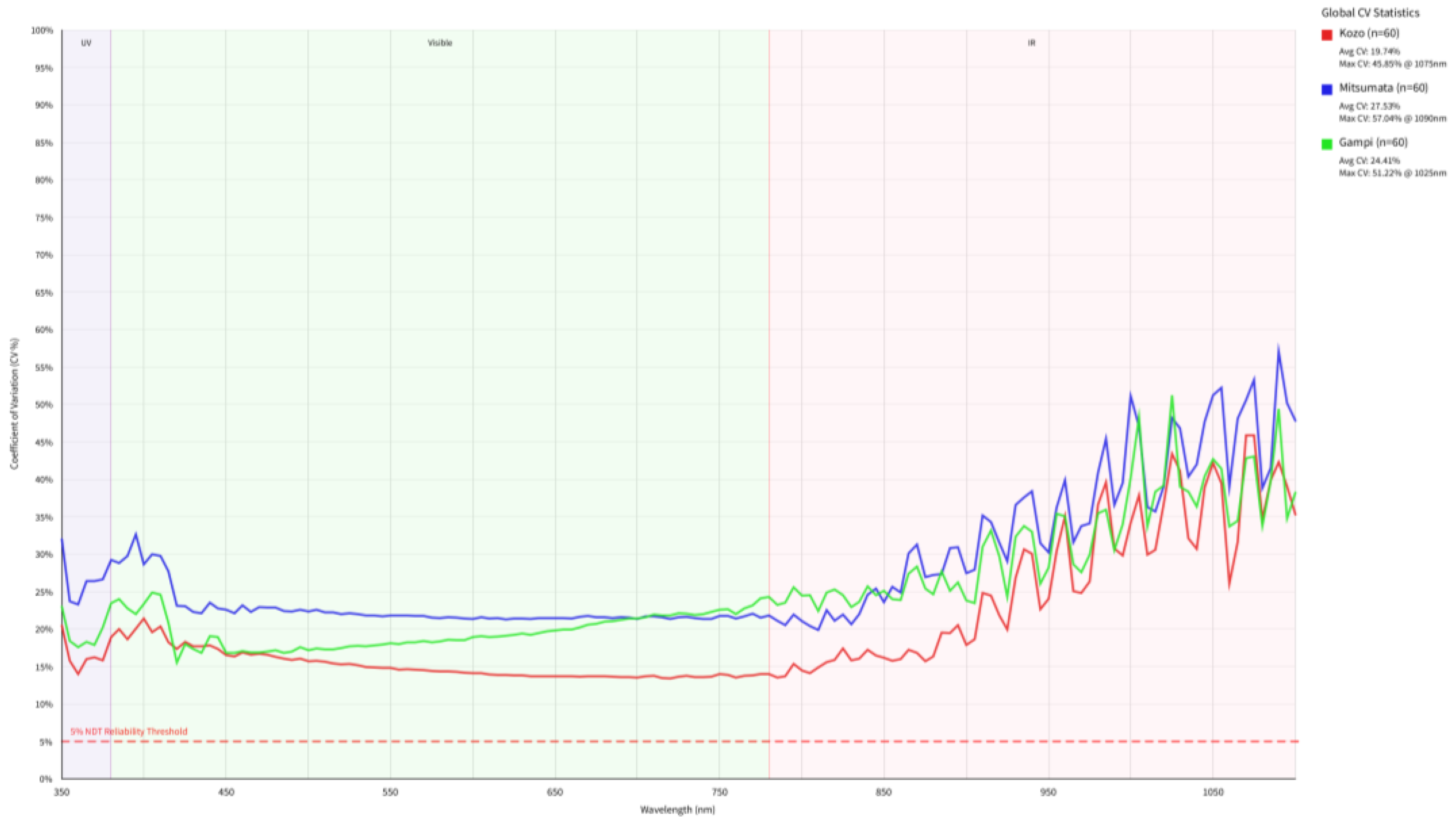
3.2. Physical Scattering Interference and Spectral Normalization
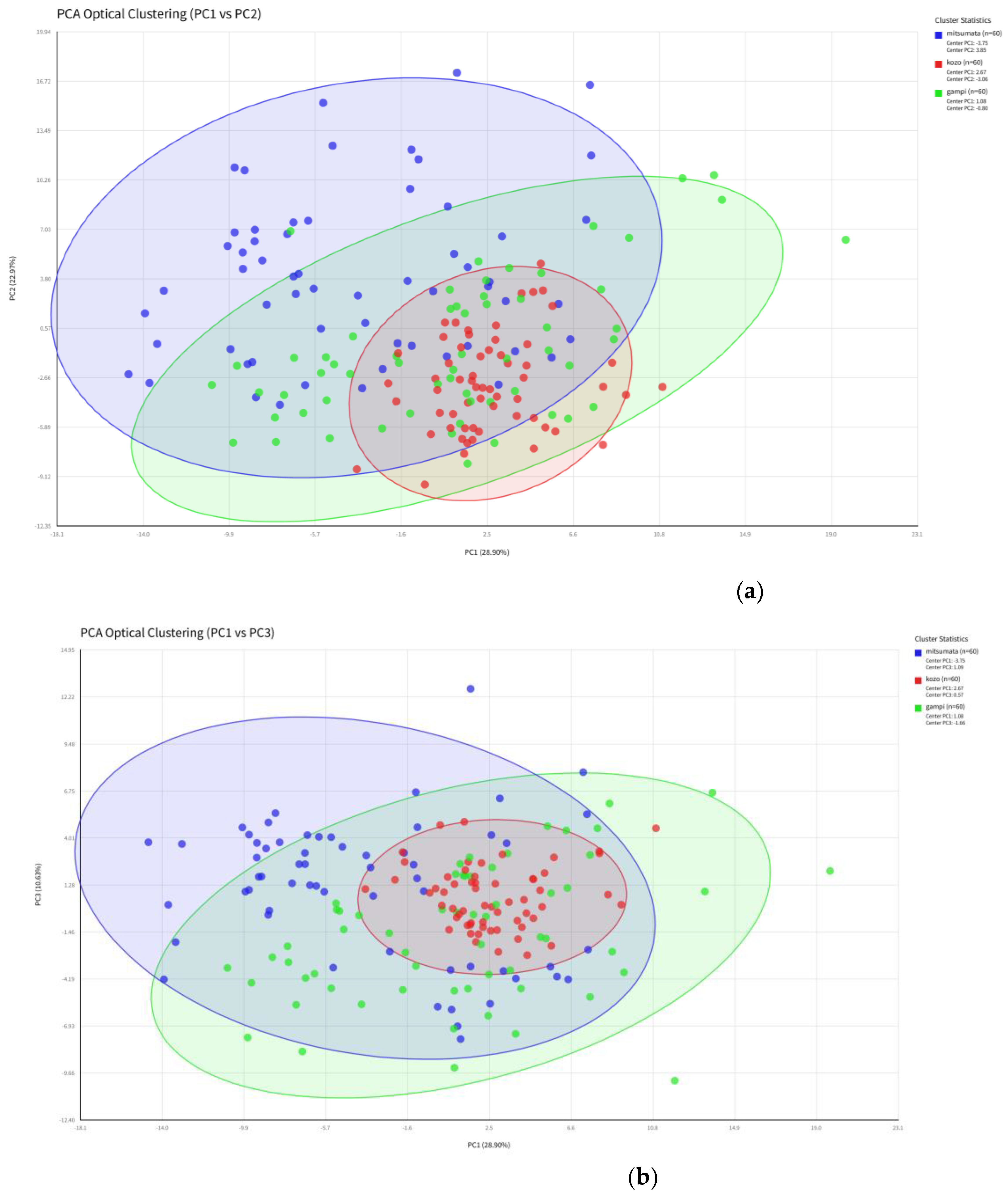
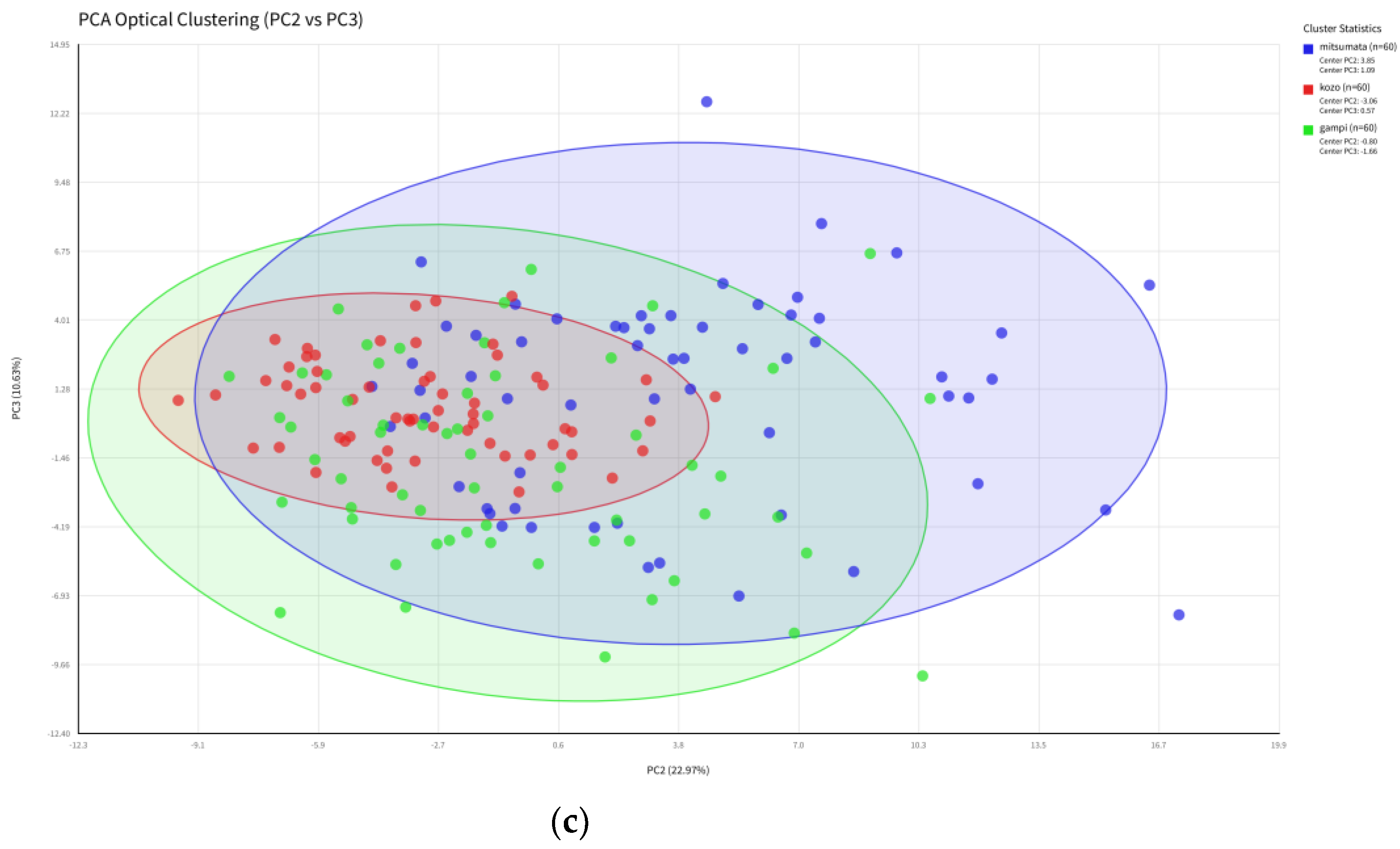
3.3. Unsupervised Dimensionality Reduction and Inter-Class Overlap
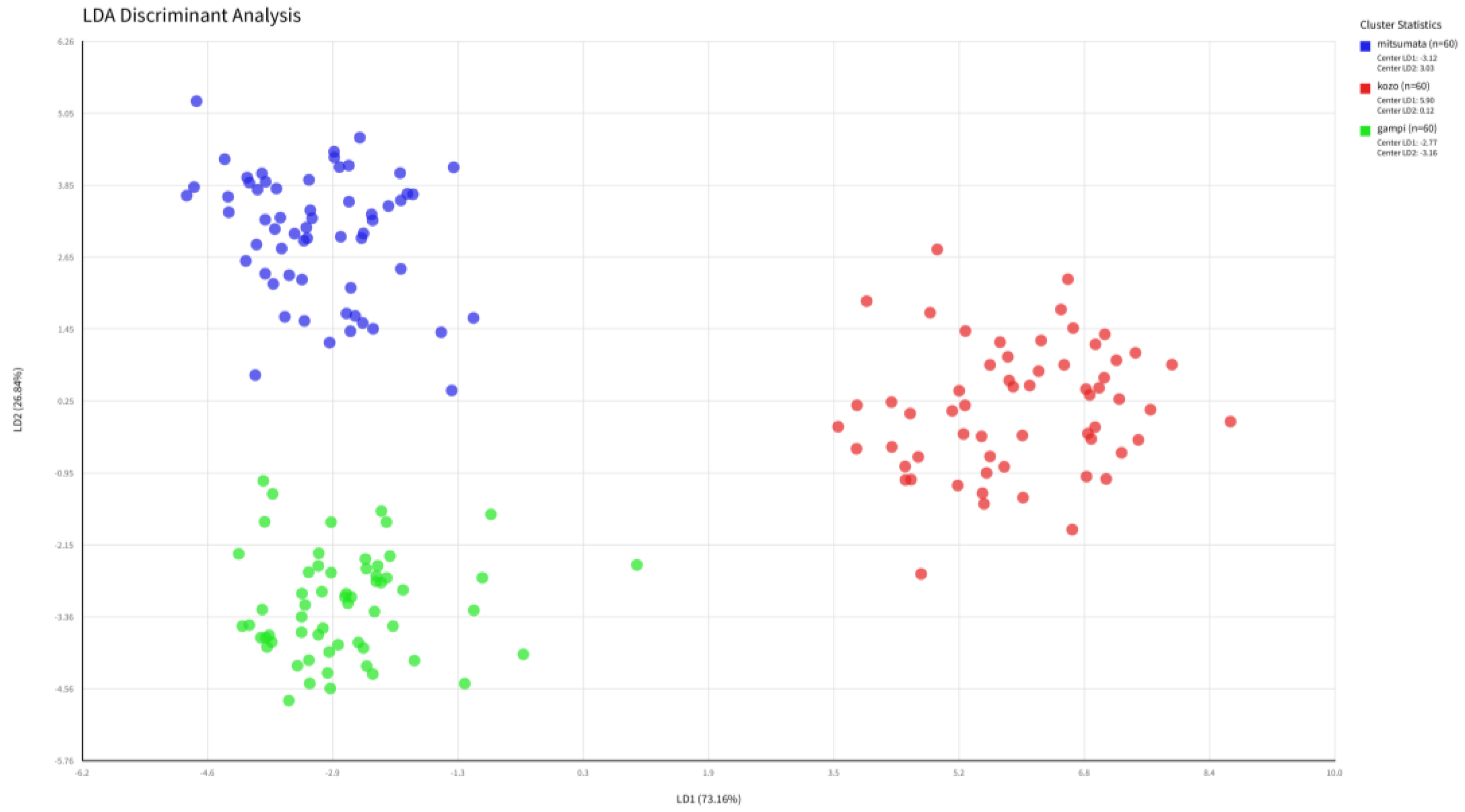
3.4. Supervised Species Discrimination via LDA
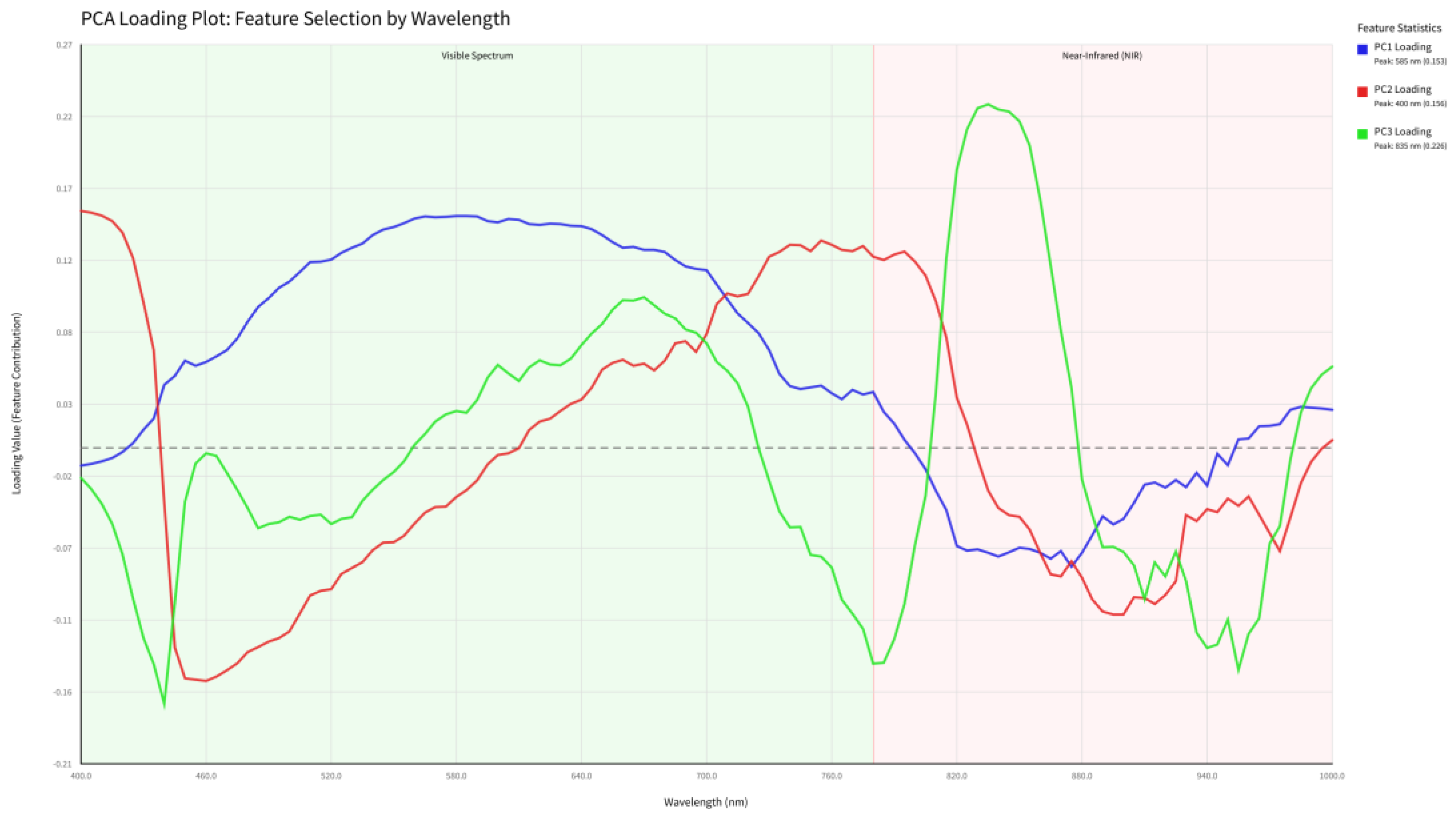
3.5. Interpretation of PCA Loading Features
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| Micro-HSI | Micro-Hyperspectral Imaging |
| VNIR | Visible and Near-Infrared |
| NIR | Near-Infrared |
| UV | Ultraviolet |
| PCA | Principal Component Analysis |
| LDA | Linear Discriminant Analysis |
| PC | Principal Component |
| LD | Linear Discriminant |
| CV | Coefficient of Variation |
| SD | Standard Deviation |
| ROI | Region of Interest |
| S–G | Savitzky–Golay |
| NDT | Non-Destructive Testing |
| JIS | Japanese Industrial Standard |
References
- Area, M.C.; Cheradame, H. Paper aging and degradation: Recent findings and research methods. BioResources 2011, 6, 5307–5337. [Google Scholar] [CrossRef]
- Avataneo, C.; Sablier, M. New criteria for the characterization of traditional East Asian papers. Environmental science and pollution research international 2017, 24, 2166–2181. [Google Scholar] [CrossRef] [PubMed]
- Japanese Industrial Standards. JIS P 8120:1998; Paper, board and pulps—Fiber analysis. Japanese Standards Association: Tokyo, Japan, 1998.
- Lukesova, H.; Holst, B. Identifying plant fibers in cultural heritage with optical and electron microscopy: how to present results and avoid pitfalls. Heritage Science 2024, 12, 12. [Google Scholar] [CrossRef]
- Garside, P.; Wyeth, P. Identification of Cellulosic Fibres by FTIR Spectroscopy - Thread and Single Fibre Analysis by Attenuated Total Reflectance. Studies in Conservation 2003, 48, 269–275. [Google Scholar] [CrossRef]
- Okuyama, M.; Sato, M.; Akada, M. The Study on Excavated Bast Fibres Using Synchrotron Polarized FT-IR Micro-Spectroscopy. Sen’i Gakkaishi 2012, 68, 55–58. [Google Scholar] [CrossRef]
- Wu, Y.; Wang, B.; Chen, J.; Huang, X.; Xu, J.; Wei, W.; Chen, K. Non-destructive prediction and pixel-level visualization of polysaccharide-based properties in ancient paper using SWNIR hyperspectral imaging and machine learning. Carbohydrate Polymers 2025, 352, 123198. [Google Scholar] [CrossRef] [PubMed]
- Yagi, C.; Yoshimura, N.; Takayanagi, M.; Kikuchi, R.; Yasunaga, T.; Hayakawa, N. Discrimination of traditional plant fibers used in Japanese cultural artifacts by infrared spectroscopy. Vibrational Spectroscopy 2022, 123, 103466. [Google Scholar] [CrossRef]
- Kamiya, N.; Ashino, K.; Sakai, Y.; Zhou, Y.; Ohyanagi, Y.; Shibazaki, K. Non-Destructive Estimation of Paper Fiber Using Macro Images: A Comparative Evaluation of Network Architectures and Patch Sizes for Patch-Based Classification. NDT 2024, 2, 487–503. [Google Scholar] [CrossRef]
- Schwanninger, M.; Rodrigues, J.C.; Fackler, K. A review of band assignments in near infrared spectra of wood and wood components. J. Near Infrared Spectrosc. 2011, 19, 287–308. [Google Scholar] [CrossRef]
- Sadeghifar, H.; Ragauskas, A. Lignin as a UV light blocker—A review. Polymers 2020, 12, 1134. [Google Scholar] [CrossRef] [PubMed]
- Rinnan, A.; van den Berg, F.; Engelsen, S.B. Review of the most common pre-processing techniques for near-infrared spectra. TrAC Trends in Analytical Chemistry 2009, 28, 1201–1222. [Google Scholar] [CrossRef]
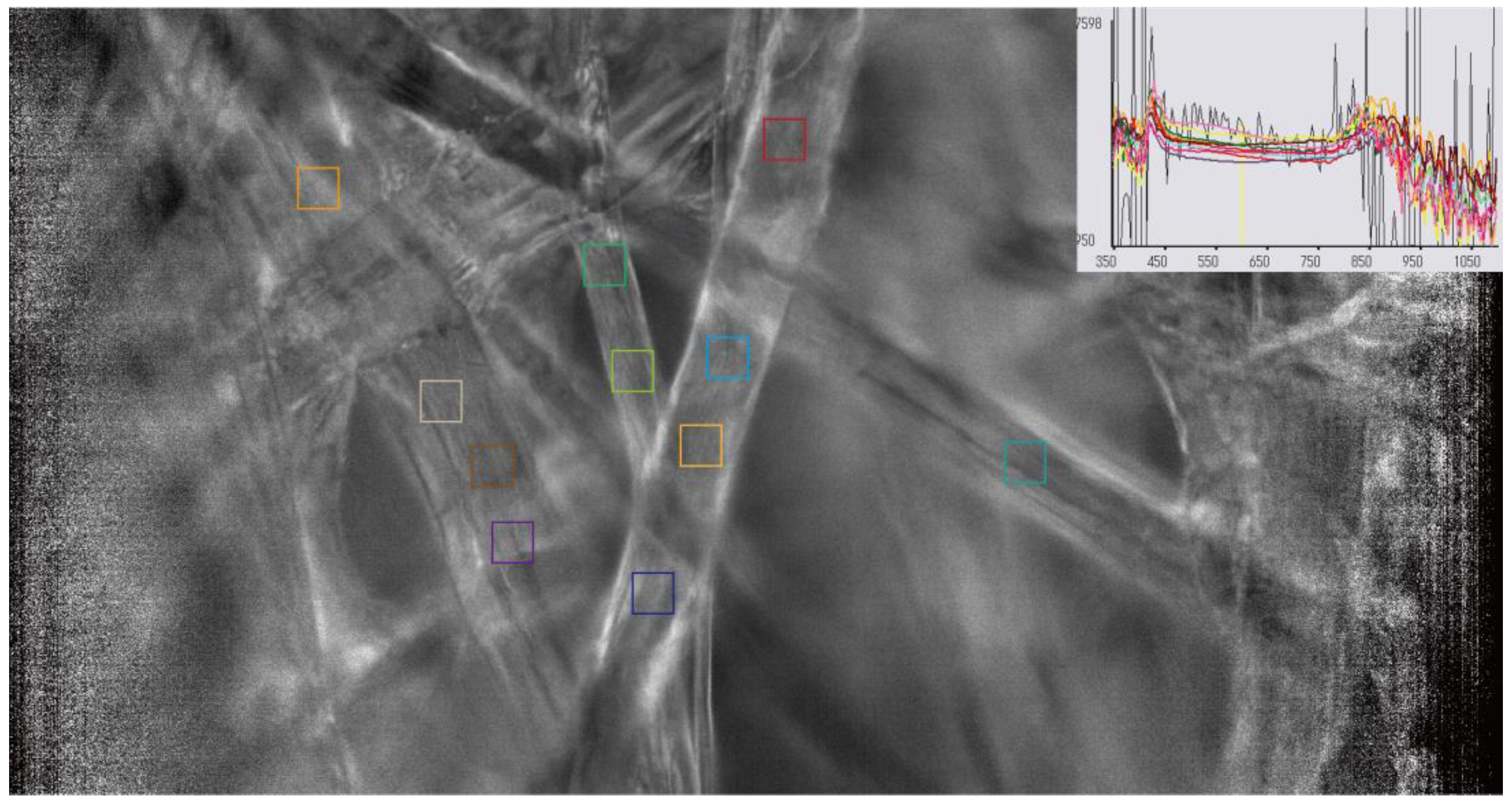

| Parameter | Specification |
| Camera type | Push-broom HSI |
| Spectral range | 350 nm–1100 nm (covering UV-Vis-NIR) |
| Spectral resolution | 5 nm |
| Detector resolution | 2048 (H) × 1080 (V) pixels |
| Microscope platform | Nikon ECLIPSE LV100ND |
| Objective lens | 50× (NA 0.8) |
| Illumination | 12 V-50 W halogen lamp |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




