Submitted:
20 March 2026
Posted:
23 March 2026
You are already at the latest version
Abstract
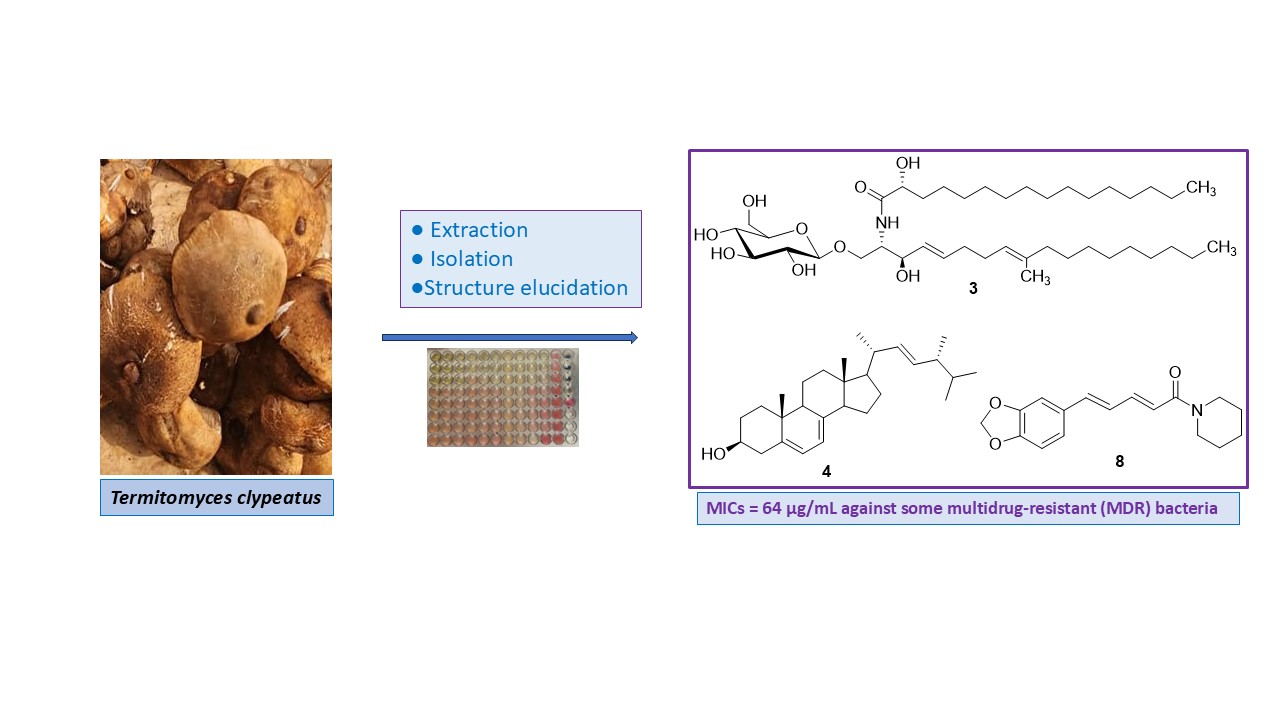
Keywords:
1. Introduction
2. Results
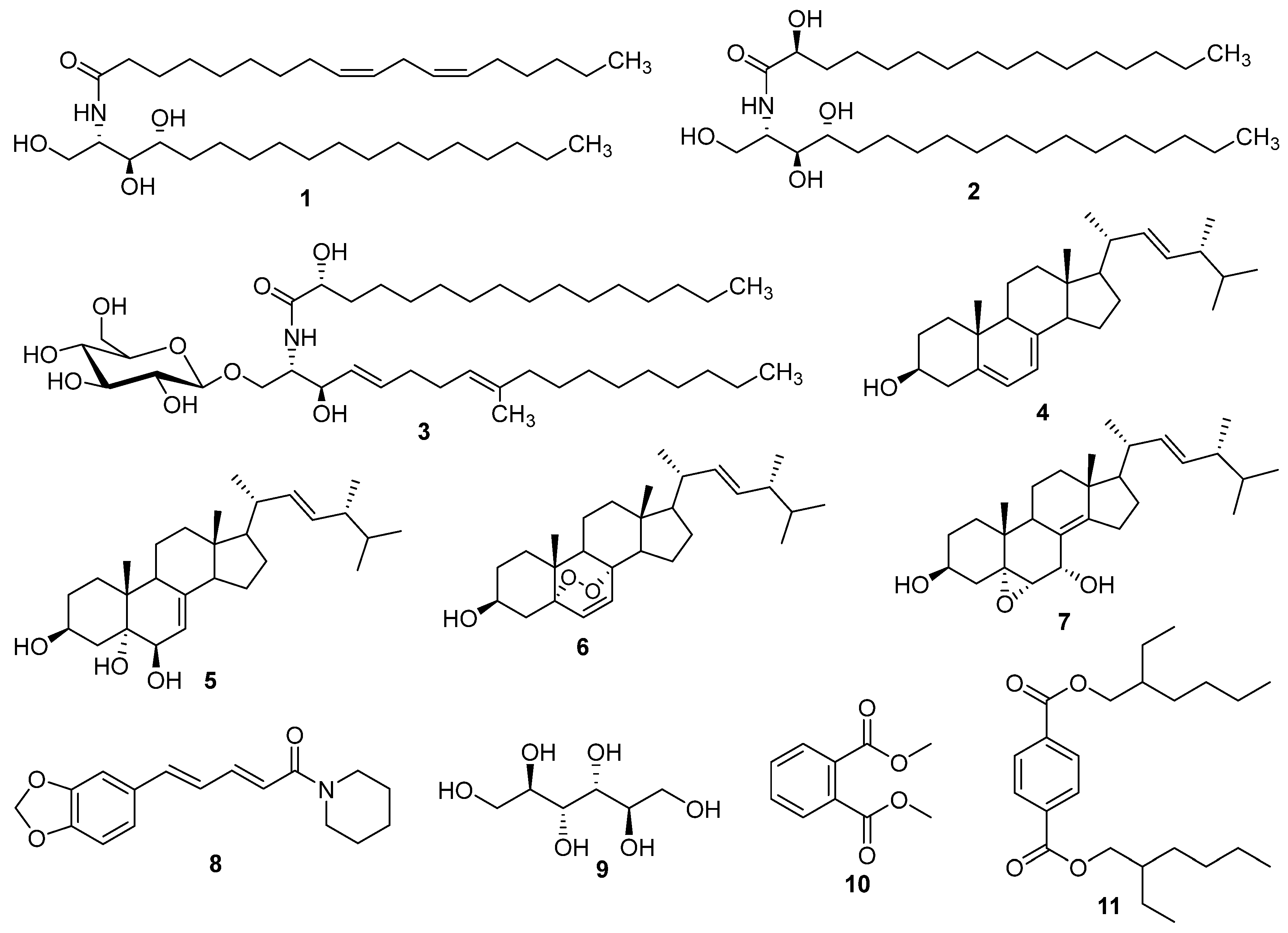
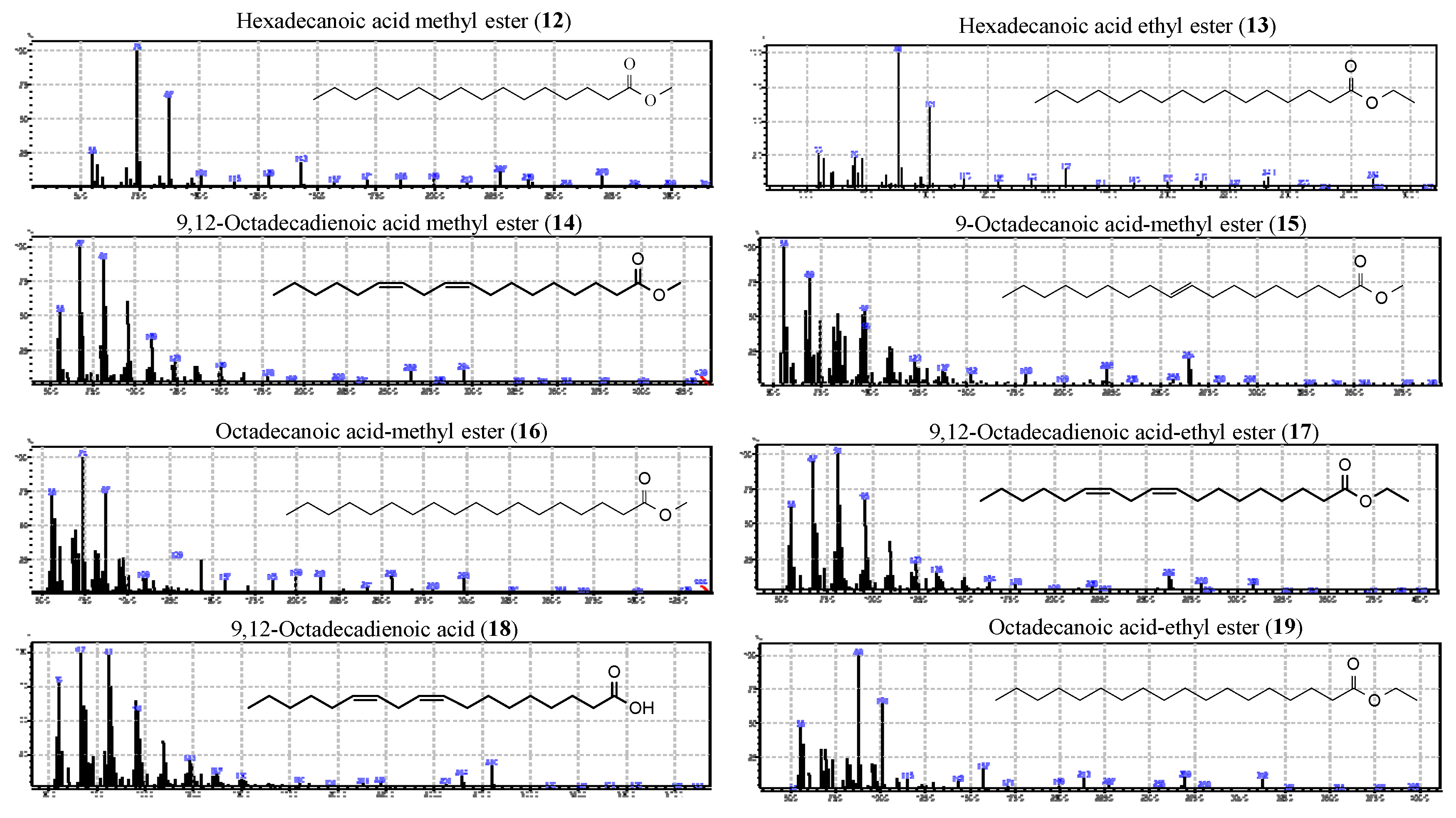
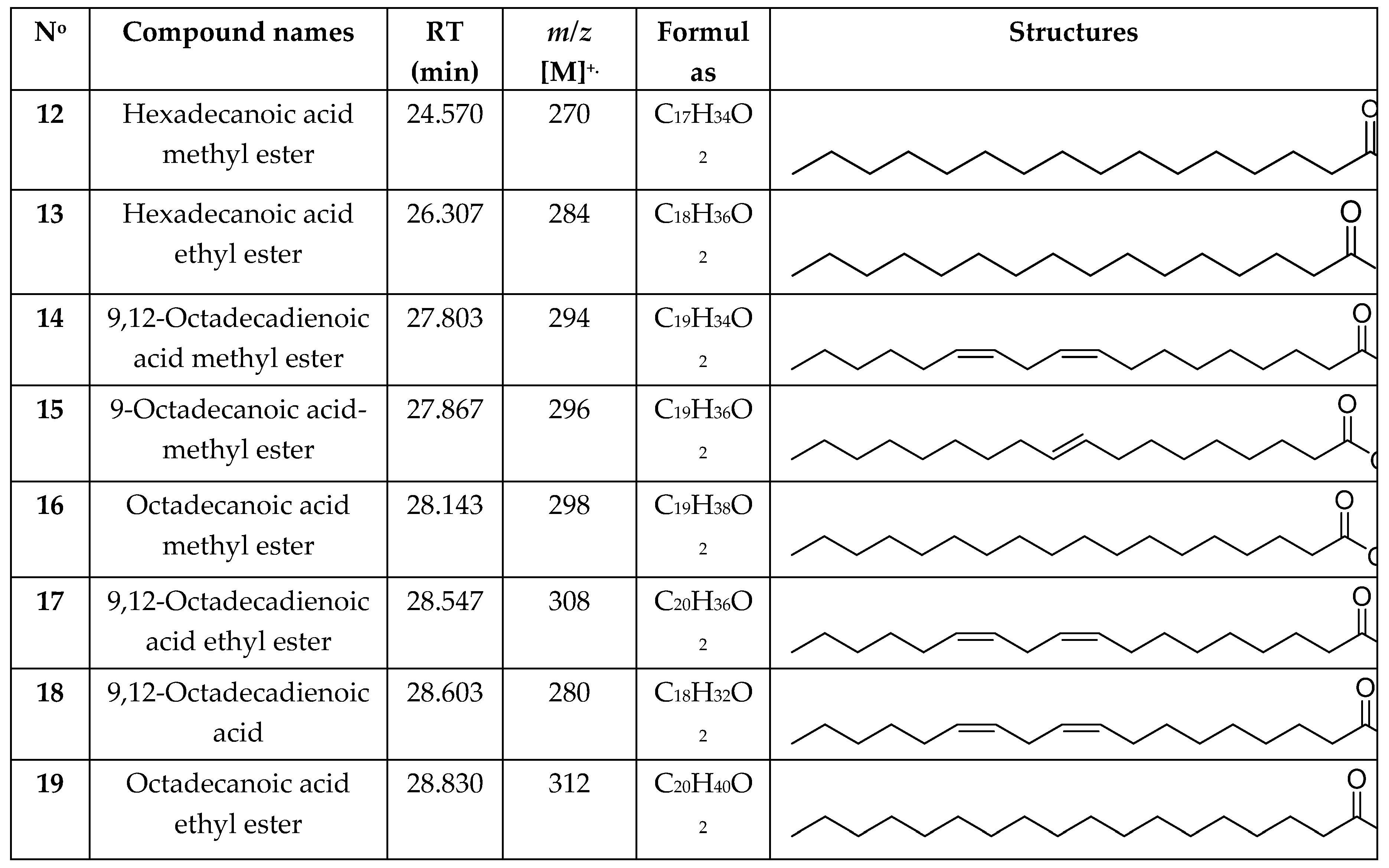
2.1. Isolation and Identification of Compounds
2.2. Antibacterial Activity of Extracts and Compounds
3.3. Effect of Samples in Presence of Efflux Pumps Inhibitors
3.4. Determination of the Antibiotic-Potentiating Effects of the Samples
3. Discussion
4. Materials and Methods
4.1. General Experimental Procedures
4.2. Fungal Material
4.3. Extraction and Isolation
4.4. Antibacterial Activity
4.4.1. Culture Media and Chemicals
4.4.2. Microorganisms
4.4.3. Minimal Inhibitory and Bactericidal Concentrations
4.4.4. Effect of Efflux Pumps on the Antibacterial Activity of the Samples
4.4.5. Antibiotic Potentiating Effect
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Falagas, M.E.; Bliziotis, I.A.; Tam, V.H. Intraventricular or Intrathecal Use of Polymyxins in Patients with Gram-Negative Meningitis: A Systematic Review of the Available Evidence. Int J Antimicrob Agents 2007, 29, 9–25. [CrossRef]
- Rice, L.B. Antimicrobial Resistance in Gram-Positive Bacteria. Am J Infect Control 2006, 34, S11-19; discussion S64-73. [CrossRef]
- Ferraz, M.P. Antimicrobial Resistance: The Impact from and on Society According to One Health Approach. Societies 2024, 14, 187. [CrossRef]
- Ferraz, M.P. Antimicrobial Resistance: The Impact from and on Society According to One Health Approach. Societies 2024, 14, 187. [CrossRef]
- Ikuta, K.S.; Swetschinski, L.R.; Robles Aguilar, G.; Sharara, F.; Mestrovic, T.; Gray, A.P.; Davis Weaver, N.; Wool, E.E.; Han, C.; Gershberg Hayoon, A.; et al. Global Mortality Associated with 33 Bacterial Pathogens in 2019: A Systematic Analysis for the Global Burden of Disease Study 2019. The Lancet 2022, 400, 2221–2248. [CrossRef]
- Ahmed, S.K.; Hussein, S.; Qurbani, K.; Ibrahim, R.H.; Fareeq, A.; Mahmood, K.A.; Mohamed, M.G. Antimicrobial Resistance: Impacts, Challenges, and Future Prospects. Journal of Medicine, Surgery, and Public Health 2024, 2, 100081. [CrossRef]
- Sharaf, O.M.; Al-Gamal, M.S.; Ibrahim, G.A.; Dabiza, N.M.; Salem, S.S.; El-ssayad, M.F.; Youssef, A.M. Evaluation and Characterization of Some Protective Culture Metabolites in Free and Nano-Chitosan-Loaded Forms against Common Contaminants of Egyptian Cheese. Carbohydrate Polymers 2019, 223, 115094. [CrossRef]
- Hari, P.A.; Usha, B. Ethnomycology of Termitomyces R. Heim in Nepal. J. Yeast Fungal Res. 2016, 7, 28–38. [CrossRef]
- Mahamat, O.; André-Ledoux, N.; Chrisopher, T.; Mbifu, A.A.; Albert, K. Assessment of Antimicrobial and Immunomodulatory Activities of Termite Associated Fungi, Termitomyces Clypeatus R. Heim (Lyophyllaceae, Basidiomycota). Clin Phytosci 2018, 4, 28. [CrossRef]
- Hussein, J.M.; Tibuhwa, D.D.; Tibell, S. Phylogenetic Position and Taxonomy of Kusaghiporia Usambarensis Gen. et Sp. Nov. (Polyporales). Mycology 2018, 9, 136–144. [CrossRef]
- Teponno, R.B.; Noumeur, S.R.; Helaly, S.E.; Hüttel, S.; Harzallah, D.; Stadler, M. Furanones and Anthranilic Acid Derivatives from the Endophytic Fungus Dendrothyrium Variisporum. Molecules 2017, 22, 1674. [CrossRef]
- Noumeur, S.R.; Teponno, R.B.; Helaly, S.E.; Wang, X.-W.; Harzallah, D.; Houbraken, J.; Crous, P.W.; Stadler, M. Diketopiperazines from Batnamyces Globulariicola, Gen. & Sp. Nov. (Chaetomiaceae), a Fungus Associated with Roots of the Medicinal Plant Globularia Alypum in Algeria. Mycol Progress 2020, 19, 589–603. [CrossRef]
- Kuete, J.R.N.; Fadeyi, O.G.; Olou, B.A.; Kenmoe, F.W.T.; Tsakem, B.; Teponno, R.B.; Tapondjou, L.A.; Kouam, S.F.; Yorou, N.S. Two Hispidin Dimers from Piptoporellus Baudonii (Fomitopsidaceae) with Anticancer Activity. Phytochem. Lett. 2025, 65, 19–23. [CrossRef]
- Fadeyi, O.G.; Olou, B.A.; Tchuenguem, R.T.; Tsakem, B.; Sinda, P.V.K.; Teponno, R.B.; Tapondjou, L.A.; Dzoyem, J.P.; Yorou, N.S. Antimicrobial and Antioxidant Activities of a Brown Wood Rotting Mushroom Piptoporellus bAudonii from Benin (Tropical Africa). ABC 2024, 14, 145–170. [CrossRef]
- Tsakem, B.; Tchamgoue, J.; Kinge, R.T.; Tiani, G.L.M.; Teponno, R.B.; Kouam, S.F. Diversity of African Fungi, Chemical Constituents and Biological Activities. Fitoterapia 2024, 178, 106154. [CrossRef]
- Gao, J.M.; Yang, X.; Wang, C.Y.; Liu, J.K. Armillaramide, a New Sphingolipid from the Fungus Armillaria Mellea. Fitoterapia 2001, 72, 858–864. [CrossRef]
- Sitrin, R.D.; Chan, G.; Dingerdissen, J.; DeBrosse, C.; Mehta, R.; Roberts, G.; Rottschaefer, S.; Staiger, D.; Valenta, J.; Snader, K.M. Isolation and Structure Determination of Pachybasium Cerebrosides Which Potentiate the Antifungal Activity of Aculeacin. J Antibiot (Tokyo) 1988, 41, 469–480. [CrossRef]
- Alexandre, T.R.; Lima, M.L.; Galuppo, M.K.; Mesquita, J.T.; Do Nascimento, M.A.; Dos Santos, A.L.; Sartorelli, P.; Pimenta, D.C.; Tempone, A.G. Ergosterol Isolated from the Basidiomycete Pleurotus Salmoneostramineus Affects Trypanosoma Cruzi Plasma Membrane and Mitochondria. J Venom Anim Toxins Incl Trop Dis 2017, 23, 30. [CrossRef]
- Piccialli, V.; Sica, D. Four New Trihydroxylated Sterols from the Sponge Spongionella Gracilis. J. Nat. Prod. 1987, 50, 915–920. [CrossRef]
- Huong, L.M.; Nghi, D.H.; Ha, T.H.; Hang, T.N.; Chi, H.K.; Toan, M.N.; Hofrichter, M. Cytotoxic and Antibacterial Compounds from Fermented Broth of Phellinus Adamantinus. J. Chem. 2010, 48, 544–548.
- Luo, X.; Li, F.; Shinde, P.B.; Hong, J.; Lee, C.-O.; Im, K.S.; Jung, J.H. 26,27-Cyclosterols and Other Polyoxygenated Sterols from a Marine Sponge Topsentia Sp. J Nat Prod 2006, 69, 1760–1768. [CrossRef]
- Nikitin, S. Structure Elucidation and Complete Assignment of 1H and 13C NMR Data of Piperine. 2020. [CrossRef]
- Nyemb, J.N.; Magnibou, L.M.; Talla, E.; Tchinda, A.T.; Tchuenguem, R.T.; Henoumont, C.; Laurent, S.; Mbafor, J.T. Lipids Constituents from Gardenia Aqualla Stapf & Hutch. Open Chemistry 2018, 16, 371–376. [CrossRef]
- Garg, B.; Bisht, T.; Ling, Y.-C. Sulfonated graphene as highly efficient and reusable acid carbocatalyst for the synthesis of ester plasticizers. RSC Adv. 2014, 4, 57297-57307.
- Grymel, A.; Latos, P.; Matuszek, K.; Erfurt, K.; Barteczko, N.; Pankalla, E.; Chrobok, A. Sustainable Method for the Synthesis of Alternative Bis(2-Ethylhexyl) Terephthalate Plasticizer in the Presence of Protic Ionic Liquids. Catalysts 2020, 10, 457. [CrossRef]
- Zhang, X.; Ma, Y.; Ouyang, B.; Wang, Y.; Wang, F. Efficient Lipase-Catalyzed Synthesis of Ceramide III Series Compounds in an Eco-Friendly Solvent. Molecular Catalysis 2024, 558, 114006. [CrossRef]
- Huang, Y.; Yang, J.; Shen, J.; Chen, F.F.; Yu, Y. Sphingolipids Are Involved in N-Methyl-N’-Nitro-N-Nitrosoguanidine-Induced Epidermal Growth Factor Receptor Clustering. Biochem Biophys Res Commun 2005, 330, 430–438. [CrossRef]
- Paloi, S.; Kumla, J.; Paloi, B.P.; Srinuanpan, S.; Hoijang, S.; Karunarathna, S.C.; Acharya, K.; Suwannarach, N.; Lumyong, S. Termite Mushrooms (Termitomyces), a Potential Source of Nutrients and Bioactive Compounds Exhibiting Human Health Benefits: A Review. JoF 2023, 9, 112. [CrossRef]
- Qi, J.; Ojika, M.; Sakagami, Y. Neuritogenic Cerebrosides from an Edible Chinese Mushroom. Part 2: Structures of Two Additional Termitomycesphins and Activity Enhancement of an Inactive Cerebroside by Hydroxylation. Bioorg Med Chem 2001, 9, 2171–2177. [CrossRef]
- Xing, L. Biosynthesis of Cerebroside by Termitomyces clypeatusUsing Serine as a Precursor. Nutr. Food Sci. 2019, 2.
- Njue, A.W.; Omolo, J.O.; Cheplogoi, P.K.; Langat, M.K.; Mulholland, D.A. Cytotoxic Ergostane Derivatives from the Edible Mushroom Termitomyces Microcarpus (Lyophyllaceae). Biochem. Syst. Ecol. 2018, 76, 12–14. [CrossRef]
- Ray, R.; Saha, S.; Paul, S. Two Novel Compounds, Ergosterol and Ergosta-5,8-Dien-3-Ol, from Termitomyces Heimii Natarajan Demonstrate Promising Anti-Hepatocarcinoma Activity. J. Trad. Chin. Med. Sci. 2022, 9, 443–453. [CrossRef]
- Ray, R.; Saha, S.; Pyne, N.; Pal, A.; Paul, S. Neoergosterol-Rich Mushroom Extract, Resourced from Wild Edible Mushroom Termitomyces Heimii Natarajan, Induces Robust Apoptosis against Lung Cancer. S. Afr. J. Bot. 2024, 170, 229–249. [CrossRef]
- Bauer, A.; Nam, J.-H.; Maulide, N. A Short, Efficient, and Stereoselective Synthesis of Piperine and Its Analogues. Synlett 2019, 30, 413–416. [CrossRef]
- Chithra, S.; Jasim, B.; Sachidanandan, P.; Jyothis, M.; Radhakrishnan, E.K. Piperine Production by Endophytic Fungus Colletotrichum Gloeosporioides Isolated from Piper Nigrum. Phytomedicine 2014, 21, 534–540. [CrossRef]
- Verma, V.C.; Lobkovsky, E.; Gange, A.C.; Singh, S.K.; Prakash, S. Piperine Production by Endophytic Fungus Periconia Sp. Isolated from Piper Longum L. J Antibiot (Tokyo) 2011, 64, 427–431. [CrossRef]
- Thiemann, T. Isolation of Phthalates and Terephthalates from Plant Material—Natural Products or Contaminants? CHEM 2021, 8, 1–36. [CrossRef]
- Tian, C.; Ni, J.; Chang, F.; Liu, S.; Xu, N.; Sun, W.; Xie, Y.; Guo, Y.; Ma, Y.; Yang, Z.; et al. Bio-Source of Di-n-Butyl Phthalate Production by Filamentous Fungi. Sci Rep 2016, 6, 19791. [CrossRef]
- Dahlin, J.L.; Nissink, J.W.M.; Strasser, J.M.; Francis, S.; Higgins, L.; Zhou, H.; Zhang, Z.; Walters, M.A. PAINS in the Assay: Chemical Mechanisms of Assay Interference and Promiscuous Enzymatic Inhibition Observed during a Sulfhydryl-Scavenging HTS. J. Med. Chem. 2015, 58, 2091–2113. [CrossRef]
- Caesar, L.K.; Cech, N.B. Synergy and Antagonism in Natural Product Extracts: When 1 + 1 Does Not Equal 2. Nat. Prod. Rep. 2019, 36, 869–888. [CrossRef]
- Anifowose, S.O.; Salih, A.M.; Oladejo, M.K.; Rady, A.; Al Mosallam, M.S.; Aljohi, H.A.; Almansour, M.I.; Alkahtani, S.H.; Alanazi, I.O.; Al-Dahmash, B.A. Bioactivity-Guided Fractionation and Mechanistic Insights into Aristolochia Ringens Root Extract-Induced G1 Phase Arrest and Mitochondria-Mediated Apoptosis in Human Colon Adenocarcinoma Cells. Pharmaceuticals 2025, 18, 1250. [CrossRef]
- Bao, N.; Yin, Y.; Wang, P. An Antimicrobial Cerebroside from the Liposoluble Constituent of Cervus Nippon Antler Velvet Layer. PJZ 2018, 50. [CrossRef]
- Plésiat, P.; Nikaido, H. Outer Membranes of Gram-negative Bacteria Are Permeable to Steroid Probes. Molecular Microbiology 1992, 6, 1323–1333. [CrossRef]
- Murase, L.S.; Perez De Souza, J.V.; Meneguello, J.E.; Palomo, C.T.; Fernandes Herculano Ramos Milaré, Á.C.; Negri, M.; Dias Siqueira, V.L.; Demarchi, I.G.; Vieira Teixeira, J.J.; Cardoso, R.F. Antibacterial and Immunological Properties of Piperine Evidenced by Preclinical Studies: A Systematic Review. Future Microbiol. 2023, 18, 1279–1299. [CrossRef]
- Price, K.E.; Orazi, G.; Ruoff, K.L.; Hebert, W.P.; O’Toole, G.A.; Mastoridis, P. Mannitol Does Not Enhance Tobramycin Killing of Pseudomonas Aeruginosa in a Cystic Fibrosis Model System of Biofilm Formation. PLoS ONE 2015, 10, e0141192. [CrossRef]
- Li, X.-Z.; Plésiat, P.; Nikaido, H. The Challenge of Efflux-Mediated Antibiotic Resistance in Gram-Negative Bacteria. Clin Microbiol Rev 2015, 28, 337–418. [CrossRef]
- Jang, S. AcrAB-TolC, a Major Efflux Pump in Gram Negative Bacteria: Toward Understanding Its Operation Mechanism. BMB Rep 2023, 56, 326–334. [CrossRef]
- Andrade, J.C.; Morais Braga, M.F.B.; Guedes, G.M.M.; Tintino, S.R.; Freitas, M.A.; Quintans, L.J.; Jr.; Menezes, I.R.A.; Coutinho, H.D.M. Cholecalciferol, Ergosterol, and Cholesterol Enhance the Antibiotic Activity of Drugs. International Journal for Vitamin and Nutrition Research 2018, 88, 244–250. [CrossRef]
- Lamers, R.P.; Cavallari, J.F.; Burrows, L.L. The Efflux Inhibitor Phenylalanine-Arginine Beta-Naphthylamide (PAβN) Permeabilizes the Outer Membrane of Gram-Negative Bacteria. PLoS ONE 2013, 8, e60666. [CrossRef]
- Ngakam, R.; Matieta, V.Y.; Kengne Fonkou, G.; Mapie Tiwa, S.; Megaptche, J.F.; Nayim, P.; Mbaveng, A.T.; Kuete, V. Antibacterial Potential and Modes of Action of Methanol Extracts of Flowers and Leaves of Vernonia Glabra (Steetz) Vatke (Asteraceae) against Multidrug-Resistant Gram-Negative Bacteria Overexpressing Efflux Pumps. Invest. Med. Chem. Pharmacol. 2024, 7, 1–10. [CrossRef]
- Mbaveng, A.T.; Sandjo, L.P.; Tankeo, S.B.; Ndifor, A.R.; Pantaleon, A.; Nagdjui, B.T.; Kuete, V. Antibacterial Activity of Nineteen Selected Natural Products against Multi-Drug Resistant Gram-Negative Phenotypes. SpringerPlus 2015, 4, 823. [CrossRef]
- Kuete, V.; Alibert-Franco, S.; Eyong, K.O.; Ngameni, B.; Folefoc, G.N.; Nguemeving, J.R.; Tangmouo, J.G.; Fotso, G.W.; Komguem, J.; Ouahouo, B.M.W.; et al. Antibacterial Activity of Some Natural Products against Bacteria Expressing a Multidrug-Resistant Phenotype. Int J Antimicrob Agents 2011, 37, 156–161. [CrossRef]
- Kova, J.; Gavari, N.; Bucar, F.; Moina, S.S. Antimicrobial and Resistance Modulatory Activity of Alpinia Katsumadai Seed Phenolic Extract, Essential Oil and Post-Distillation Extract. 2014.
 |
| Bacteria and strains | EtOH extract | EtOAc fraction | n-BuOH fraction | Sub-fraction A | |||||||||
| MIC | MBC | R | MIC | MBC | R | MIC | MBC | R | MIC | MBC | R | ||
|
Pseudomonas aeruginosa |
PA01 | >2048 | - | - | 1024 | >2048 | - | 2048 | >2048 | - | 128 | >256 | - |
| PA124 | >2048 | - | - | 512 | >2048 | - | >2048 | - | - | 128 | >256 | - | |
|
Klebsiella pneumoniae |
KP55 | >2048 | - | - | >2048 | - | - | >2048 | - | - | 64 | >256 | - |
| ATCC11296 | >2048 | - | - | >2048 | - | - | >2048 | - | - | 256 | 256 | 1 | |
|
Escherichia coli |
AG100 | 2048 | 2048 | 1 | 1024 | >2048 | - | 2048 | >2048 | - | >256 | - | - |
| ATCC10536 | >2048 | - | - | >2048 | - | - | >2048 | - | - | >256 | - | - | |
| Providencia stuartii | PS2636 | 2048 | >2048 | - | 1024 | >2048 | - | 2048 | >2048 | - | 128 | >256 | - |
| NEA16 | >2048 | - | - | >2048 | - | - | >2048 | - | - | >256 | - | - | |
| Enterobacter aerogenes | EA3 | >2048 | - | - | >2048 | - | - | >2048 | - | - | >256 | - | - |
| EA27 | 2048 | >2048 | - | 2048 | 2048 | - | >2048 | - | - | >256 | - | - | |
| Bacteria and strains | 3 | 4 | 5 | 6 | |||||||||
| MIC | MBC | R | MIC | MBC | R | MIC | MBC | R | MIC | MBC | R | ||
|
Pseudomonas aeruginosa |
PA01 | 256 | 256 | 1 | 128 | 256 | 2 | >256 | - | - | >256 | - | - |
| PA124 | 128 | 256 | 2 | 128 | - | - | >256 | - | - | >256 | - | - | |
|
Klebsiella pneumoniae |
KP55 | >256 | - | >256 | - | - | >256 | - | - | >256 | - | - | |
| ATCC11296 | 256 | >256 | - | 256 | >256 | - | 128 | 256 | 2 | >256 | - | - | |
| Escherichia coli | AG100 | 128 | >256 | - | 256 | 256 | 1 | 256 | >256 | - | >256 | - | - |
| ATCC10536 | 64 | 64 | 1 | 64 | >256 | - | 32 | 128 | 4 | 128 | 256 | 2 | |
| Providencia stuartii | PS2636 | 128 | - | - | 128 | 128 | 1 | >256 | - | - | 256 | 256 | 1 |
| NEA16 | 256 | >256 | - | 256 | >256 | - | >256 | - | - | >256 | - | - | |
| Enterobacter aerogenes | EA3 | 256 | >256 | - | 128 | >256 | - | 256 | 256 | 1 | >256 | - | - |
| EA27 | >256 | - | - | 256 | >256 | - | >256 | - | - | >256 | - | - | |
| Bacteria and strains | 7 | 8 | 9 | Chloramphenicol | |||||||||
| MIC | MBC | R | MIC | MBC | R | MIC | MBC | R | MIC | MBC | R | ||
|
Pseudomonas aeruginosa |
PA01 | >256 | - | - | >256 | >256 | - | >256 | - | - | 32 | 64 | 2 |
| PA124 | >256 | - | - | 256 | >256 | - | 256 | >256 | - | 8 | 128 | 32 | |
|
Klebsiella pneumoniae |
KP55 | >256 | - | - | >256 | >256 | - | >256 | - | - | 64 | 128 | 2 |
| ATCC11296 | >256 | - | - | 256 | 256 | 1 | >256 | - | - | 16 | 32 | 2 | |
| Escherichia coli | AG100 | 256 | 256 | 1 | 256 | 256 | 1 | 256 | >256 | - | 2 | 16 | 8 |
| ATCC10536 | 256 | >256 | - | >256 | >256 | - | 256 | 256 | 1 | 32 | 32 | 1 | |
| Providencia stuartii | PS2636 | >256 | - | - | 128 | 256 | 2 | >256 | - | - | 16 | >128 | - |
| NEA16 | >256 | - | - | >256 | >256 | - | >256 | - | - | 64 | >128 | - | |
| Enterobacter aerogenes | EA3 | >256 | - | - | 64 | 128 | 2 | >256 | - | - | 32 | 128 | 4 |
| EA27 | >256 | - | - | >256 | >256 | - | >256 | - | - | 32 | >128 | - | |
| Bacteria and strains | A | 3 | 4 | 8 | Chloramphenicol | ||||||||||
|
MIC alone |
MIC +PAβN | R |
MIC alone |
MIC +PAβN | R |
MIC alone |
MIC +PAβN | R |
MIC alone |
MIC +PAβN | R |
MIC alone |
MIC +PAβN | R | |
| PA01 | 1024 | >2048 | / | 256 | / | / | 128 | 32 | 4 | >256 | / | / | 32 | 32 | 1 |
| PA124 | 512 | >2048 | / | 128 | 512 | 4 | 128 | 64 | 2 | 256 | 128 | 2 | 64 | 64 | 1 |
| KP55 | >2048 | / | / | >256 | / | / | >256 | 32 | >8 | >256 | 256 | 1 | 128 | 16 | 8 |
| ATCC11296 | 256 | >2048 | / | 256 | 32 | 8 | 256 | 64 | 4 | 256 | 256 | 1 | 64 | 16 | 4 |
| AG100 | 1024 | >2048 | / | 128 | 32 | 4 | 256 | 256 | 1 | 256 | 32 | 8 | 128 | 64 | 2 |
| ATCC10536 | >2048 | 16 | >128 | 64 | 4 | 16 | 64 | 32 | 2 | >256 | 8 | >32 | 256 | 256 | 1 |
| PS2636 | 1024 | 256 | 4 | 128 | 4 | 32 | 128 | 128 | 1 | 128 | 4 | 32 | 128 | 8 | 16 |
| NEA16 | >2048 | / | / | 256 | / | / | 256 | 128 | 2 | >256 | / | / | 128 | 64 | 2 |
| EA3 | >2048 | / | / | 256 | 4 | >64 | 128 | 32 | 4 | 64 | 4 | 16 | 64 | 4 | 16 |
| EA27 | >2048 | / | / | >256 | 128 | >2 | 256 | 32 | 8 | >256 | / | / | 32 | 16 | 2 |
| ATBs | Sub-fractions | Concentrations | MIC of antibiotics in the presence of extract (AMF) | PSP (%) | ||||
| Kp55 | AG100 | PS2636 | PA124 | EA3 | ||||
| CIP | 0 | ˃512 | ˃512 | 256 | 512 | ˃512 | ||
| A | MIC/2 | 512 (-) | ˃512 (-) | 128 (2) | 512 (1) | 256 (~2) | 40 | |
| MIC/4 | ˃512 (-) | ˃512 (-) | 128 (2) | 512 (1) | 512 (~1) | 20 | ||
| B | MIC/2 | ˂4 (128) | ˂4 (128) | ˂4 (256) | ˂4 (128) | ˂4 (128) | 100 | |
| MIC/4 | ˂4 (128) | ˂4 (128) | ˂4 (256) | ˂4 (128) | ˂4 (128) | 100 | ||
| C | MIC/2 | ˂4 (128) | ˂4 (128) | ˂4 (256) | ˂4 (128) | ˂4 (128) | 100 | |
| MIC/4 | ˂4 (128) | ˂4 (128) | ˂4 (256) | ˂4 (128) | ˂4 (128) | 100 | ||
| D | MIC/2 | ˂4 (128) | ˂4 (128) | ˂4 (256) | ˂4 (128) | ˂4 (128) | 100 | |
| MIC/4 | ˂4 (128) | ˂4 (128) | ˂4 (256) | ˂4 (128) | ˂4 (128) | 100 | ||
| DOX | 0 | 16 | ˂4 | 32 | 64 | 64 | ||
| A | MIC/2 | ˂4 (4) | ˂4 (1) | 32 (1) | 128 (0.5) | 8 (8) | 40 | |
| MIC/4 | 8 (2) | ˂4 (1) | ˂4 (8) | 64 (1) | 8 (8) | 60 | ||
| B | MIC/2 | ˂4 (4) | ˂4 (1) | ˂4 (8) | ˂4 (16) | ˂4 (16) | 80 | |
| MIC/4 | ˂4 (4) | ˂4 (1) | ˂4 (8) | ˂4 (16) | ˂4 (16) | 80 | ||
| C | MIC/2 | ˂4 (4) | ˂4 (1) | ˂4 (8) | ˂4 (16) | ˂4 (16) | 0 | |
| MIC/4 | ˂4 (4) | ˂4 (1) | ˂4 (8) | ˂4 (16) | ˂4 (16) | 20 | ||
| D | MIC/2 | ˂4 (4) | ˂4 (1) | ˂4 (8) | ˂4 (16) | ˂4 (16) | 40 | |
| MIC/4 | ˂4 (4) | ˂4 (1) | ˂4 (8) | ˂4 (16) | ˂4 (16) | 20 | ||
| AMK | 0 | ˂4 | 16 | ˂4 | 32 | ˂4 | ||
| A | MIC/2 | ˂4 (1) | 16 (1) | ˂4 (1) | 8 (4) | ˂4 (1) | 20 | |
| MIC/4 | ˂4 (1) | 16 (1) | ˂4 (1) | 64 (0.5) | ˂4 (1) | 0 | ||
| B | MIC/2 | 8 (0.5) | ˂4 (4) | 32 (0.125) | ˂4 (8) | ˂4 (1) | 40 | |
| MIC/4 | 16 (0.25) | ˂4 (4) | 32 (0.125) | ˂4 (8) | 8 (0.5) | 40 | ||
| C | MIC/2 | ˂4 (1) | 32 (0.5) | ˂4 (1) | ˂4 (8) | 16 (0.25) | 20 | |
| MIC/4 | ˂4 (1) | 32 (0.5) | 32 (0.125) | ˂4 (8) | 16 (0.25) | 20 | ||
| D | MIC/2 | 8 (0.5) | 16 (1) | 16 (0.25) | ˂4 (8) | 16 (0.25) | 20 | |
| MIC/4 | 16 (0.25) | 64 (0.25) | 16 (0.25) | ˂4 (8) | 16 (0.25) | 20 | ||
| ATBs | Compounds | Concentration | MIC of antibiotics in the presence of extract (AMF) | PSP (%) | ||||
| Kp55 | AG100 | PS2636 | PA124 | EA3 | ||||
| CIP | 0 | ˃512 | ˃512 | 256 | 512 | ˃512 | ||
| 3 | MIC/2 | ˃512(-) | ˃512(-) | 512(0.5) | 512(1) | 256(~2) | 20 | |
| MIC/4 | ˃512(-) | ˃512(-) | 512(-2) | 512(1) | 256(~2) | 20 | ||
| 4 | MIC/2 | 256(2) | ˃512(-) | 128(2) | 512(1) | 512(~1) | 40 | |
| MIC/4 | ˃512(-) | ˃512(-) | 128(2) | 512(1) | 512(~1) | 20 | ||
| 8 | MIC/2 | ˃512(-) | ˃512(-) | 256(1) | 512(1) | 256(~2) | 20 | |
| MIC/4 | ˃512(-) | ˃512(-) | 256(1) | 512(1) | 512(~1) | 0 | ||
| DOX | 0 | 16 | ˂4 | 32 | 64 | 64 | ||
| 3 | MIC/2 | 64(0.25) | 32(0.125) | 64(0.5) | 64(1) | 64(1) | 0 | |
| MIC/4 | 32(0.5) | 64(0.06) | 64(0.5) | 64(1) | 8(8) | 20 | ||
| 4 | MIC/2 | ˂4(4) | ˂4(1) | 32(1) | 64(1) | 64(1) | 20 | |
| MIC/4 | ˂4(4) | 8(0.5) | 32(1) | 64(1) | 16(4) | 40 | ||
| 8 | MIC/2 | 32(0.5) | 16(0.25) | 16(2) | 64(1) | ˂4(16) | 40 | |
| MIC/4 | 32(-2) | 32(-8) | 16(2) | 64(1) | 64(1) | 20 | ||
| AMK | 0 | ˂4 | 16 | ˂4 | 32 | ˂4 | ||
| 3 | MIC/2 | 8(0.5) | 32(0.5) | ˂4(1) | 64(0.5) | ˂4(1) | 0 | |
| MIC/4 | 8(0.5) | 32(0.5) | ˂4(1) | 64(0.5) | ˂4(1) | 0 | ||
| 4 | MIC/2 | ˂4(1) | 32(0.5) | ˂4(1) | 16(2) | ˂4(1) | 20 | |
| MIC/4 | ˂4(1) | 32(0.5) | ˂4(1) | 16(2) | ˂4(1) | 20 | ||
| 8 | MIC/2 | 16(0.25) | 32(0.5) | ˂4(1) | 64(0.5) | ˂4(1) | 0 | |
| MIC/4 | 8(0.5) | 32(0.5) | ˂4(1) | 32(1) | ˂4(1) | 0 | ||
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




