Submitted:
18 March 2026
Posted:
20 March 2026
You are already at the latest version
Abstract
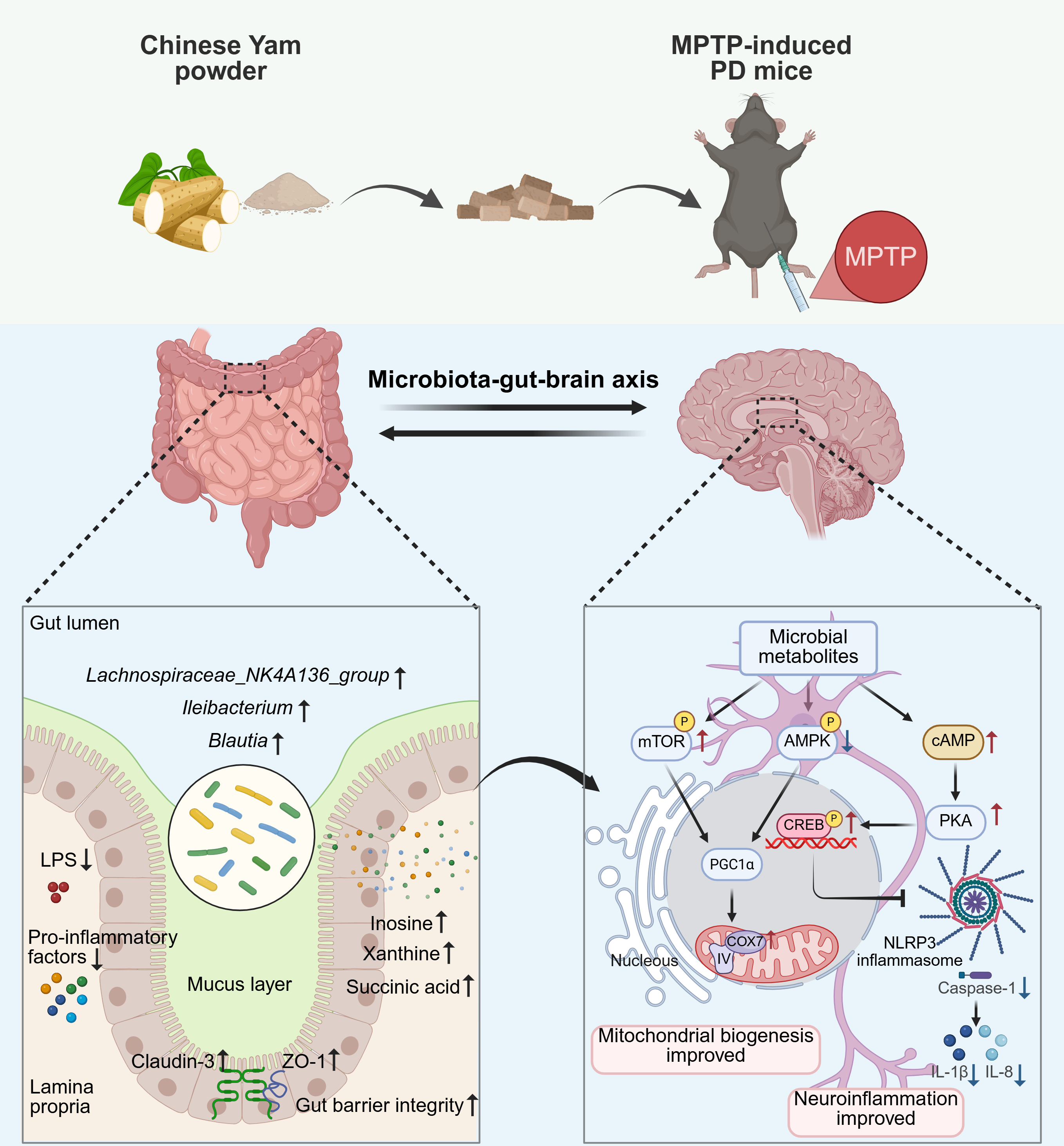
Keywords:
1. Introduction
2. Material and Methods
2.1. Animals and Experimental Design
2.2. Behavioral Tests
2.2.1. Pole Test
2.2.2. Rotarod Test
2.2.3. Inverted Grid Test
2.2.4. Hindlimb Scoring
2.3. Enzyme-Linked Immunosorbent Assay
2.4. Hematoxylin and Eosin (H&E) Staining and Alcian Blue - Periodic acid Schiff (AB-PAS) Staining
2.5. Immunofluorescence and Immunohistochemical Staining
2.6. Western Blot
2.7. Transcriptome Sequencing Analysis
2.8. 16S rRNA Sequencing Analysis
2.9. Untargeted Metabolomics Sequencing Analysis
2.10. Integrated Multi-Omics Analysis
2.11. Statistical Analysis
3. Results
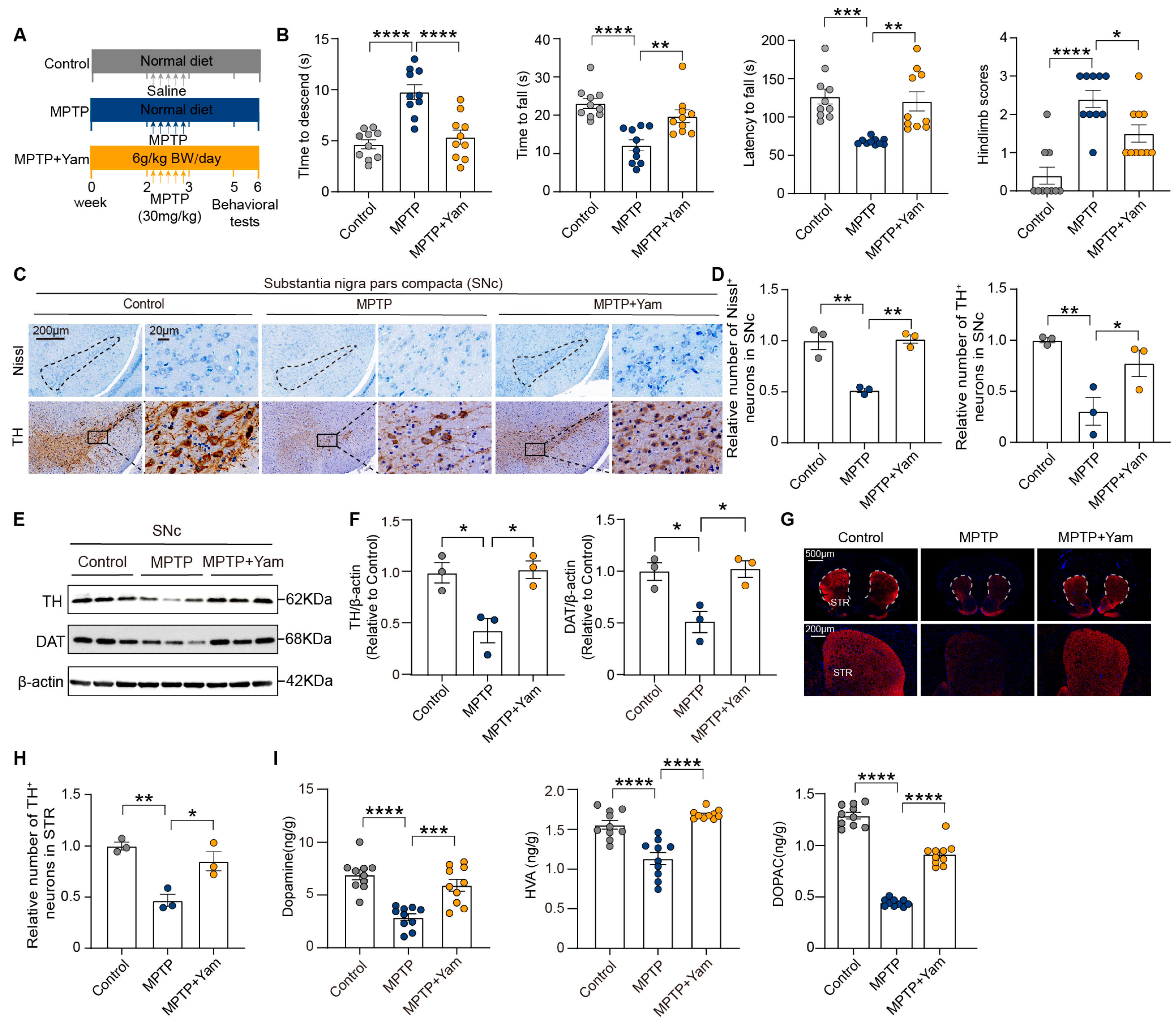
3.1. Yam Protects Against Motor Deficits and Loss of Dopaminergic Neurons in MPTP-Induced PD Mice
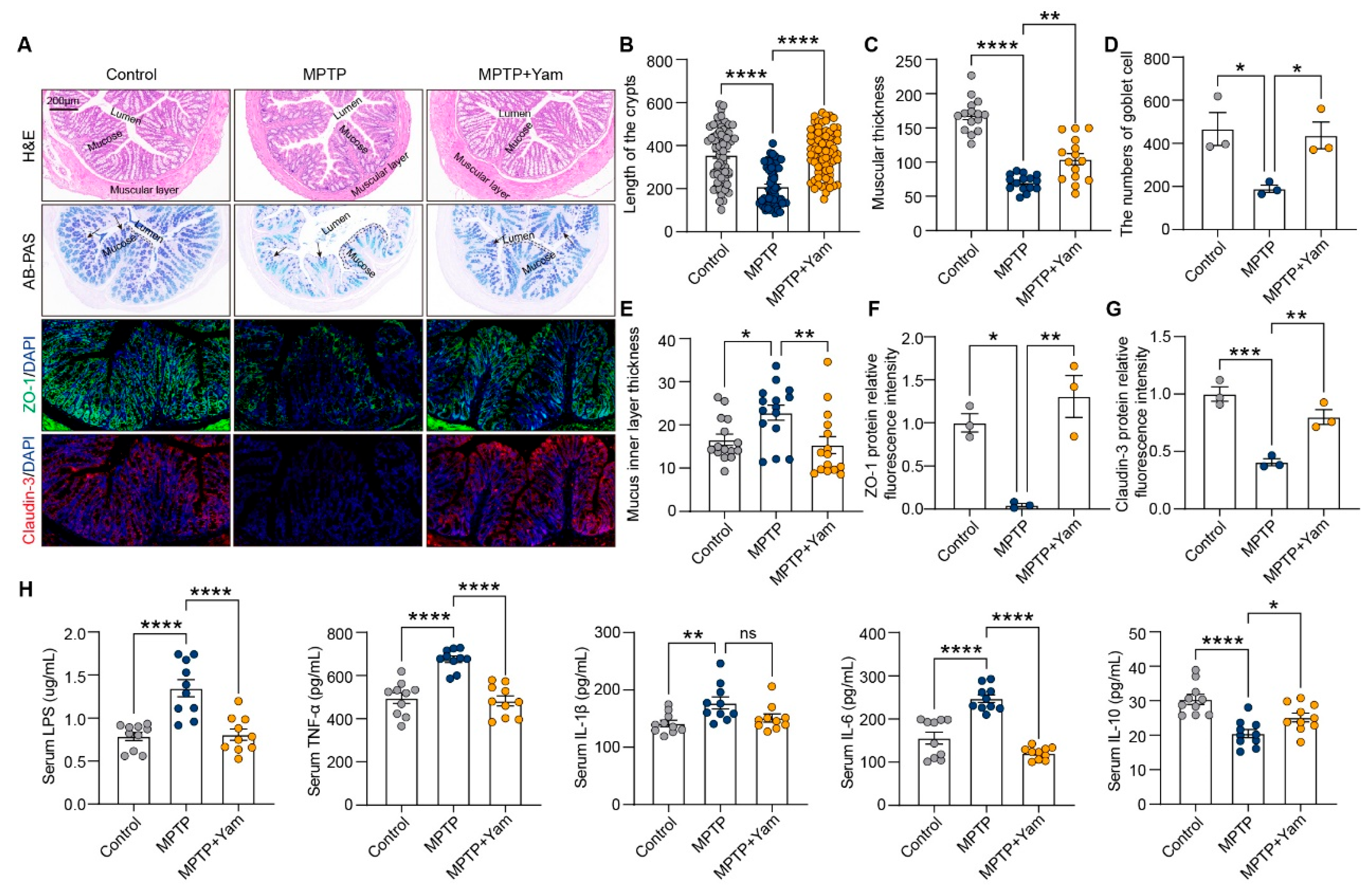
3.2. Yam Ameliorates Intestinal Barrier Dysfunction in MPTP-Induced PD Mice
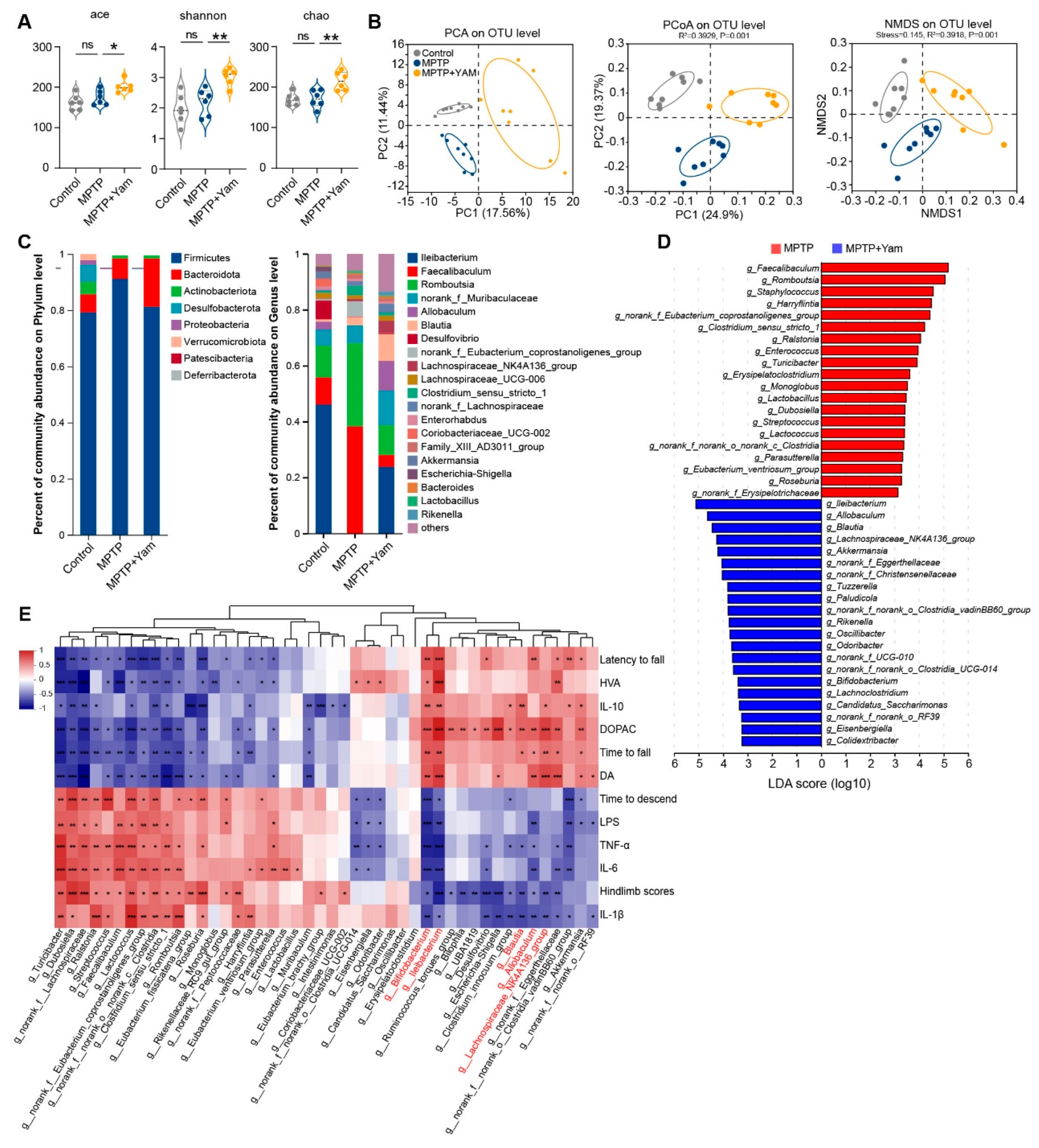
3.3. Yam Reshapes the Diversity and Composition of the Gut Microbiota in MPTP-Induced PD Mice
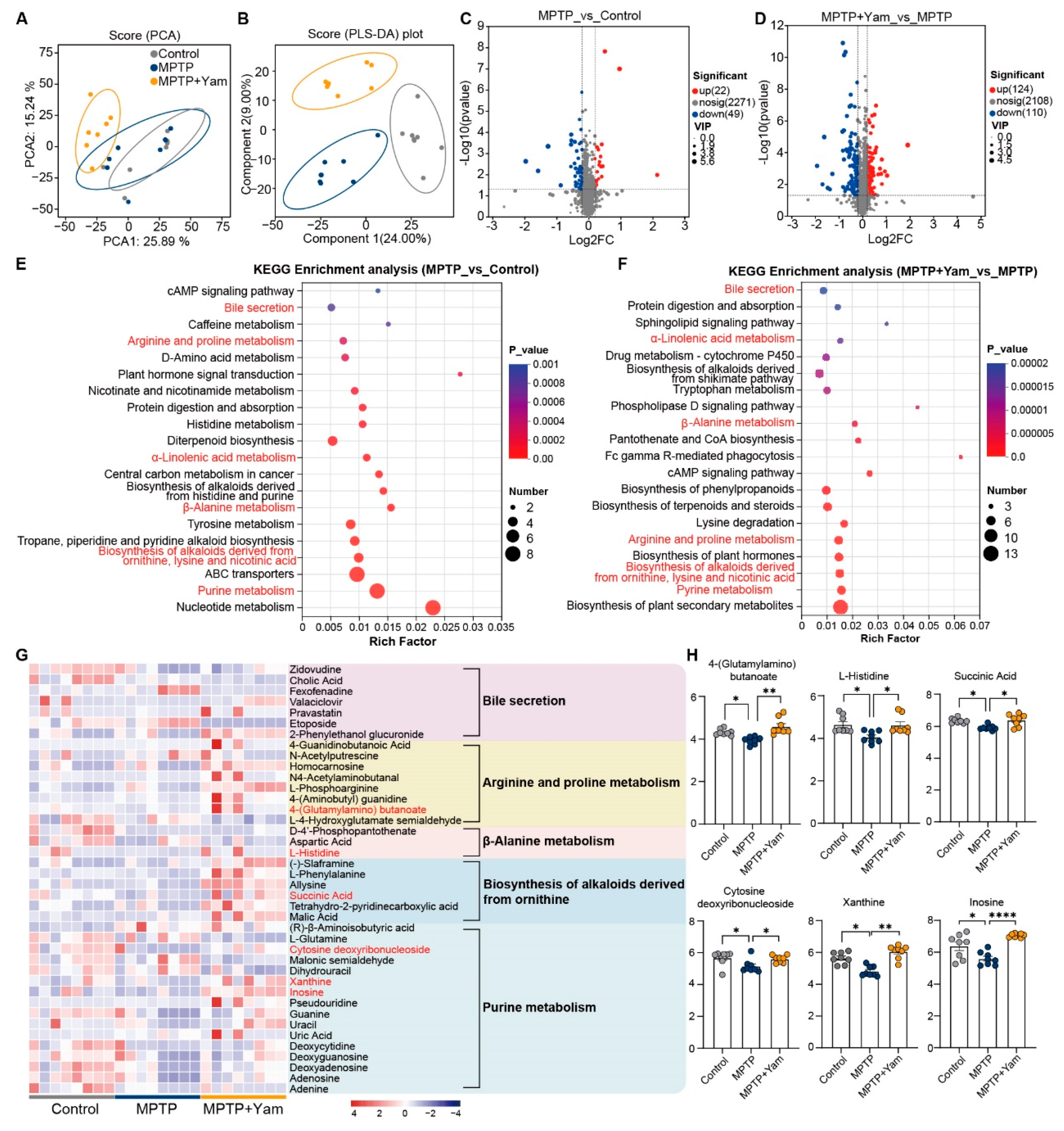
3.4. Yam Changed the Fecal Metabolite Profiles in MPTP-Induced PD Mice
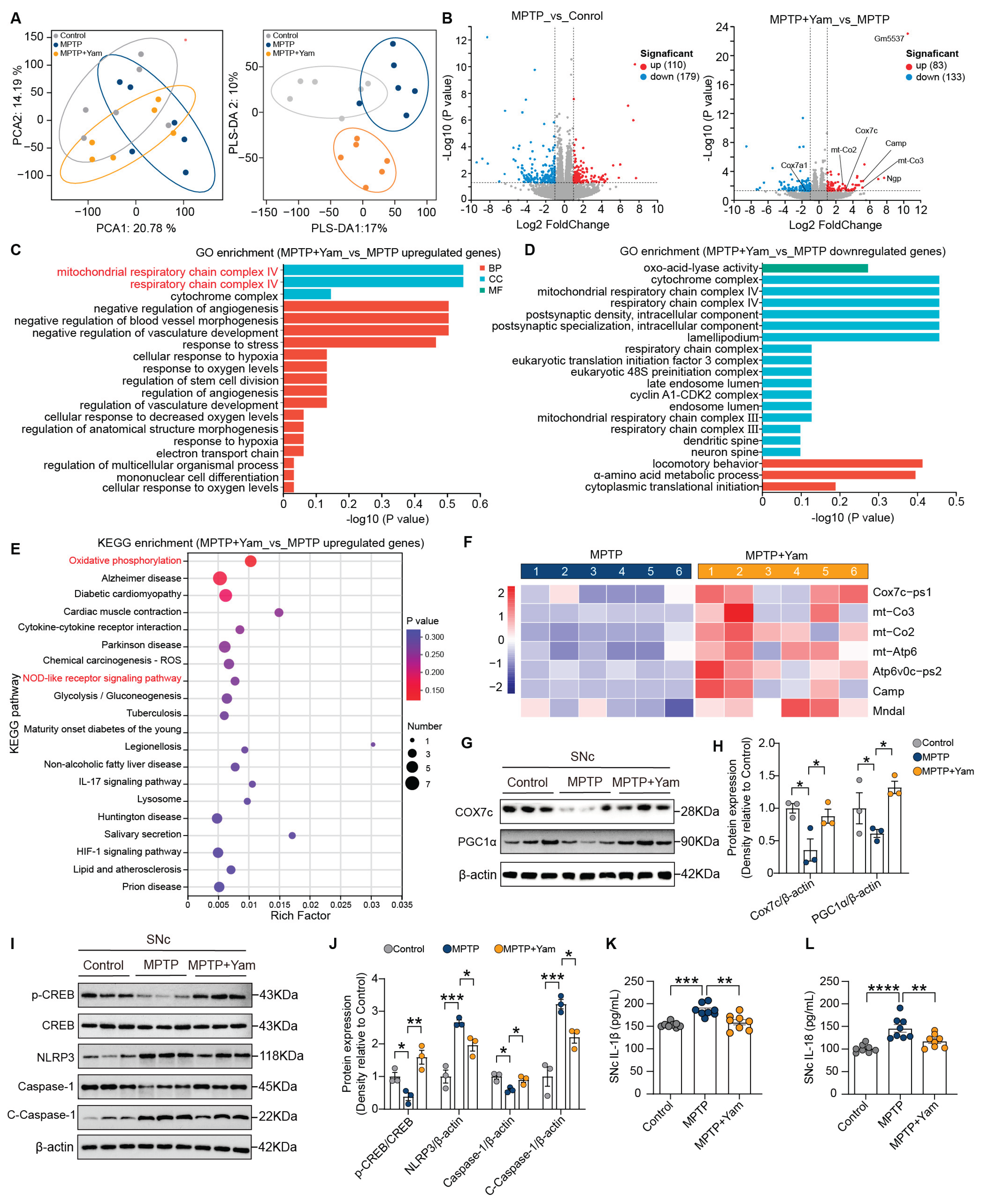
3.5. Yam Improves Mitochondrial Biogenesis and Inflammation in SNc of MPTP-Induced PD Mice
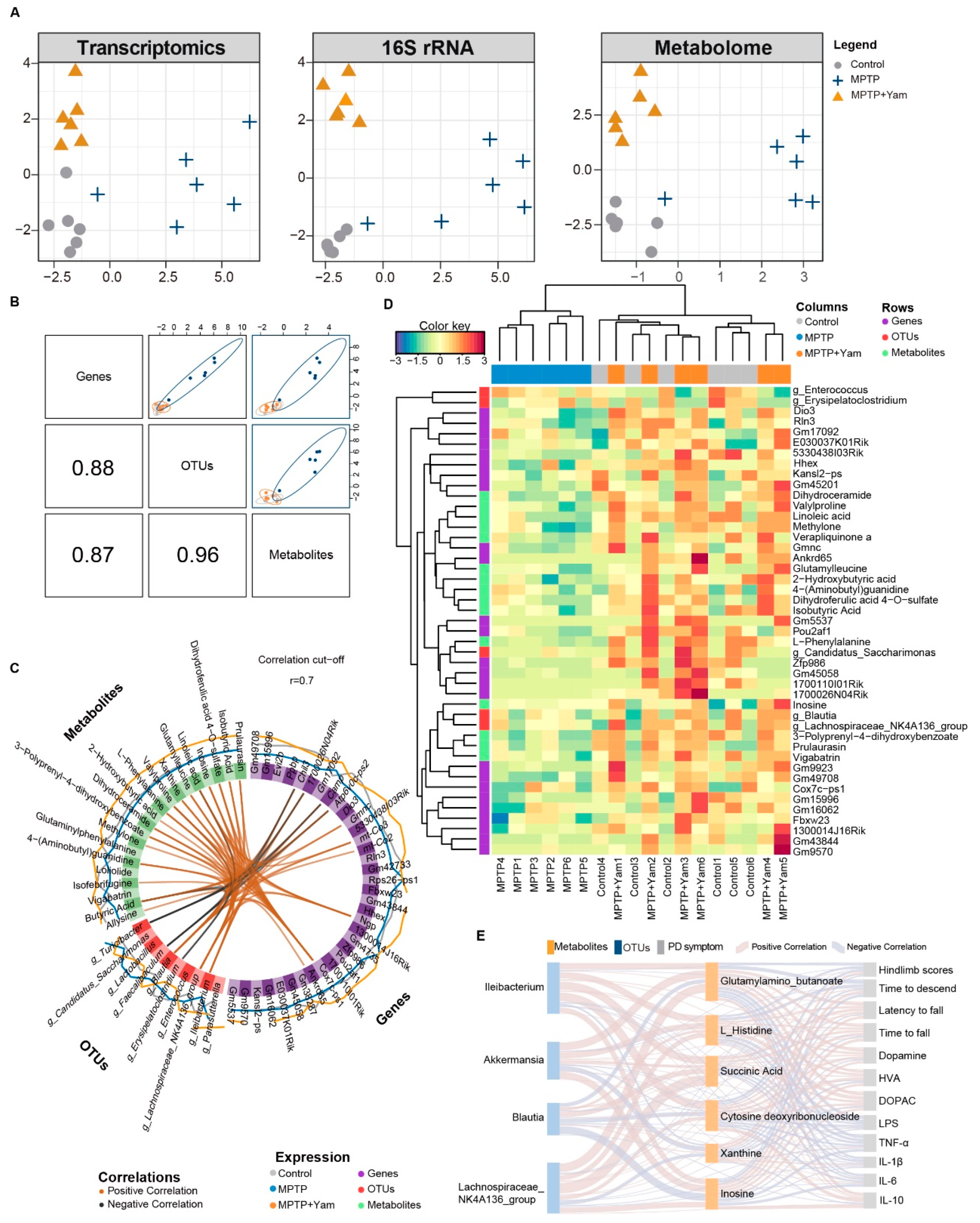
3.6. Integrated Multi-Omics Analysis on Dietary Yam Intervention
4. Discussion
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Data Availability Statement
Conflicts of Interest
References
- W. Poewe, K. Seppi, C. M. Tanner, G. M. Halliday, P. Brundin, J. Volkmann, A.-E. Schrag, A. E. Lang. Parkinson disease. Nat Rev Dis Primers. 2017, 3, 17013. [CrossRef]
- J. Jankovic. Parkinson’s disease: clinical features and diagnosis. Journal of Neurology, Neurosurgery & Psychiatry. 2008, 79, 368. [CrossRef]
- P. Calabresi, A. Castrioto, M. Di Filippo, B. Picconi. New experimental and clinical links between the hippocampus and the dopaminergic system in Parkinson’s disease. Lancet Neurol. 2013, 12, 811-821. [CrossRef]
- T. Zhou, G. Zu, X. Zhang, X. Wang, S. Li, X. Gong, Z. Liang, J. Zhao. Neuroprotective effects of ginsenoside Rg1 through the Wnt/β-catenin signaling pathway in both in vivo and in vitro models of Parkinson’s disease. Neuropharmacology. 2016, 101, 480-489. [CrossRef]
- H. Ullah, S. Arbab, Y. Tian, C.-q. Liu, Y. Chen, L. Qijie, M. I. U. Khan, I. U. Hassan, K. Li. The gut microbiota–brain axis in neurological disorder. Front Neurosci. 2023, 17 - 2023. [CrossRef]
- V. T. E. Aho, P. A. B. Pereira, S. Voutilainen, L. Paulin, E. Pekkonen, P. Auvinen, F. Scheperjans. Gut microbiota in Parkinson’s disease: Temporal stability and relations to disease progression. EBioMedicine. 2019, 44, 691-707. [CrossRef]
- E. M. Hill-Burns, J. W. Debelius, J. T. Morton, W. T. Wissemann, M. R. Lewis, Z. D. Wallen, S. D. Peddada, S. A. Factor, E. Molho, C. P. Zabetian, R. Knight, H. Payami. Parkinson’s disease and Parkinson’s disease medications have distinct signatures of the gut microbiome. Mov Disord. 2017, 32, 739-749. [CrossRef]
- P. Perez-Pardo, H. B. Dodiya, P. A. Engen, C. B. Forsyth, A. M. Huschens, M. Shaikh, R. M. Voigt, A. Naqib, S. J. Green, J. H. Kordower, K. M. Shannon, J. Garssen, A. D. Kraneveld, A. Keshavarzian. Role of TLR4 in the gut-brain axis in Parkinson’s disease: a translational study from men to mice. Gut. 2019, 68, 829. [CrossRef]
- T. R. Sampson, J. W. Debelius, T. Thron, S. Janssen, G. G. Shastri, Z. E. Ilhan, C. Challis, C. E. Schretter, S. Rocha, V. Gradinaru, M.-F. Chesselet, A. Keshavarzian, K. M. Shannon, R. Krajmalnik-Brown, P. Wittung-Stafshede, R. Knight, S. K. Mazmanian. Gut Microbiota Regulate Motor Deficits and Neuroinflammation in a Model of Parkinson Disease. Cell. 2016, 167, 1469-1480.e1412. [CrossRef]
- Y.-f. Hou, C. Shan, S.-y. Zhuang, Q.-q. Zhuang, A. Ghosh, K.-c. Zhu, X.-k. Kong, S.-m. Wang, Y.-l. Gong, Y.-y. Yang, B. Tao, L.-h. Sun, H.-Y. Zhao, X.-z. Guo, W.-q. Wang, G. Ning, Y.-y. Gu, S.-t. Li, J.-m. Liu. Gut microbiota-derived propionate mediates the neuroprotective effect of osteocalcin in a mouse model of Parkinson’s disease. Microbiome. 2021, 9, 34. [CrossRef]
- Y. Li, S. Ji, T. Xu, Y. Zhong, M. Xu, Y. Liu, M. Li, B. Fan, F. Wang, J. Xiao, B. Lu. Chinese yam (Dioscorea): Nutritional value, beneficial effects, and food and pharmaceutical applications. Trends Food Sci Technol. 2023, 134, 29-40. [CrossRef]
- Y. Bai, Y. Zhou, R. Zhang, Y. Chen, F. Wang, M. Zhang. Gut microbial fermentation promotes the intestinal anti-inflammatory activity of Chinese yam polysaccharides. Food Chem. 2023, 402, 134003. [CrossRef]
- G. Zhang, J. Pan, X. Xu, S. Nie, L. Lu, Y. Jing, F. Yang, G. Ji, H. Xu. Chinese yam polysaccharide enhances anti-PD-1 immunotherapy in colorectal cancer through alterations in the gut microbiota and metabolites. Int J Biol Macromol. 2025, 310, 143323. [CrossRef]
- J. Lu, H. Qin, L. Liang, J. Fang, K. Hao, Y. Song, T. Sun, G. Hui, Y. Xie, Y. Zhao. Yam protein ameliorates cyclophosphamide-induced intestinal immunosuppression by regulating gut microbiota and its metabolites. Int J Biol Macromol. 2024, 279, 135415. [CrossRef]
- J.-J. Cui, Z.-Y. Huang, Y.-H. Xie, J.-B. Wu, G.-H. Xu, C.-F. Li, M.-M. Zhang, L.-T. Yi. Gut microbiota mediated inflammation, neuroendocrine and neurotrophic functions involved in the antidepressant-like effects of diosgenin in chronic restraint stress. J Affect Disord. 2023, 321, 242-252. [CrossRef]
- L. Guan, Z. Mao, S. Yang, G. Wu, Y. Chen, L. Yin, Y. Qi, L. Han, L. Xu. Dioscin alleviates Alzheimer’s disease through regulating RAGE/NOX4 mediated oxidative stress and inflammation. Biomed Pharmacother. 2022, 152, 113248. [CrossRef]
- S. Som, J. Antony, S. Dhanabal, S. Ponnusankar. Neuroprotective role of Diosgenin, a NGF stimulator, against Aβ (1–42) induced neurotoxicity in animal model of Alzheimer’s disease. Metab Brain Dis. 2022, 37, 359-372. [CrossRef]
- Y. Cui, Y. Zhou, Y. Li, J. Wang, D. Li, F. Chen. Chinese Yam and Its Active Components Regulate the Structure of Gut Microbiota and Indole-like Metabolites in Anaerobic Fermentation In Vitro. Nutrients. 2023, 15, 5112. [CrossRef]
- N. Zhang, T. Liang, Q. Jin, C. Shen, Y. Zhang, P. Jing. Chinese yam (Dioscorea opposita Thunb.) alleviates antibiotic-associated diarrhea, modifies intestinal microbiota, and increases the level of short-chain fatty acids in mice. Food Res Int. 2019, 122, 191-198. [CrossRef]
- C. c. Hsu, Y.-C. Huang, M.-c. Yin, S. J. Lin. Effect of Yam (Dioscorea alata Compared to Dioscorea japonica) on Gastrointestinal Function and Antioxidant Activity in Mice. J Food Sci. 2006, 71, 27.
- M. Zhao, B. Wang, C. Zhang, Z. Su, B. Guo, Y. Zhao, R. Zheng. The DJ1-Nrf2-STING axis mediates the neuroprotective effects of Withaferin A in Parkinson’s disease. Cell Death Differ. 2021, 28, 2517-2535. [CrossRef]
- H. Shi, Y. Yu, D. Lin, P. Zheng, P. Zhang, M. Hu, Q. Wang, W. Pan, X. Yang, T. Hu, Q. Li, R. Tang, F. Zhou, K. Zheng, X.-F. Huang. β-glucan attenuates cognitive impairment via the gut-brain axis in diet-induced obese mice. Microbiome. 2020, 8, 143. [CrossRef]
- H. Weng, L. Deng, T. Wang, H. Xu, J. Wu, Q. Zhou, L. Yu, B. Chen, L. a. Huang, Y. Qu, L. Zhou, X. Chen. Humid heat environment causes anxiety-like disorder via impairing gut microbiota and bile acid metabolism in mice. Nat Commun. 2024, 15, 5697. [CrossRef]
- Z. Liu, X. Dai, H. Zhang, R. Shi, Y. Hui, X. Jin, W. Zhang, L. Wang, Q. Wang, D. Wang, J. Wang, X. Tan, B. Ren, X. Liu, T. Zhao, J. Wang, J. Pan, T. Yuan, C. Chu, L. Lan, F. Yin, E. Cadenas, L. Shi, S. Zhao, X. Liu. Gut microbiota mediates intermittent-fasting alleviation of diabetes-induced cognitive impairment. Nat Commun. 2020, 11, 855. [CrossRef]
- Z. Zhao, J. Ning, X.-q. Bao, M. Shang, J. Ma, G. Li, D. Zhang. Fecal microbiota transplantation protects rotenone-induced Parkinson’s disease mice via suppressing inflammation mediated by the lipopolysaccharide-TLR4 signaling pathway through the microbiota-gut-brain axis. Microbiome. 2021, 9, 226. [CrossRef]
- W. Hao, Q. Ma, L. Wang, N. Yuan, H. Gan, L. He, X. Li, J. Huang, J. Chen. Gut dysbiosis induces the development of depression-like behavior through abnormal synapse pruning in microglia-mediated by complement C3. Microbiome. 2024, 12, 34. [CrossRef]
- F. Rohart, B. Gautier, A. Singh, L. C. KA. mixOmics: An R package for ‘omics feature selection and multiple data integration. PLoS Comput Biol. 2017, 13, e1005752. [CrossRef]
- B. R. Bloem, M. S. Okun, C. Klein. Parkinson’s disease. Lancet. 2021, 397, 2284-2303. [CrossRef]
- C. Pellegrini, M. Fornai, V. D’Antongiovanni, L. Antonioli, N. Bernardini, P. Derkinderen. The intestinal barrier in disorders of the central nervous system. Lancet Gastroenterol Hepatol. 2023, 8, 66-80. [CrossRef]
- P. Derkinderen, F. Cossais, K. Kulcsárová, M. Škorvánek, L. Sellier-Montaigne, E. Coron, L. Leclair-Visonneau, S. Cerri, C. Pellegrini, M. Rolli-Derkinderen. How leaky is the gut in Parkinson’s disease? EBioMedicine. 2025, 117. [CrossRef]
- M. E. V. Johansson, J. K. Gustafsson, J. Holmén-Larsson, K. S. Jabbar, L. Xia, H. Xu, F. K. Ghishan, F. A. Carvalho, A. T. Gewirtz, H. Sjövall, G. C. Hansson. Bacteria penetrate the normally impenetrable inner colon mucus layer in both murine colitis models and patients with ulcerative colitis. Gut. 2014, 63, 281. [CrossRef]
- M.-F. Sun, Y.-Q. Shen. Dysbiosis of gut microbiota and microbial metabolites in Parkinson’s Disease. Ageing Res Rev. 2018, 45, 53-61. [CrossRef]
- C. Chu, T. Li, L. Yu, Y. Li, M. Li, M. Guo, J. Zhao, Q. Zhai, F. Tian, W. Chen. A Low-Protein, High-Carbohydrate Diet Exerts a Neuroprotective Effect on Mice with 1-Methyl-4-phenyl-1,2,3,6-tetrahydropyridine-Induced Parkinson’s Disease by Regulating the Microbiota-Metabolite–Brain Axis and Fibroblast Growth Factor 21. J Agric Food Chem. 2023, 71, 8877-8893. [CrossRef]
- Y. Liu, Y. Hou, G. Wang, X. Zheng, H. Hao. Gut Microbial Metabolites of Aromatic Amino Acids as Signals in Host-Microbe Interplay. Trends in Endocrinology & Metabolism. 2020, 31, 818-834. [CrossRef]
- J. L. Cooper, T. M. Jeitner. Central Role of Glutamate Metabolism in the Maintenance of Nitrogen Homeostasis in Normal and Hyperammonemic Brain. Biomolecules. 2016, 6, 16. [CrossRef]
- S. Vascellari, V. Palmas, M. Melis, S. Pisanu, R. Cusano, P. Uva, D. Perra, V. Madau, M. Sarchioto, V. Oppo, N. Simola, M. Morelli, L. Santoru Maria, L. Atzori, M. Melis, G. Cossu, A. Manzin. Gut Microbiota and Metabolome Alterations Associated with Parkinson’s Disease. mSystems. 2020, 5. [CrossRef]
- R. M. Voigt, Z. Wang, J. M. Brown, P. A. Engen, A. Naqib, C. G. Goetz, D. A. Hall, L. V. Metman, M. Shaikh, C. B. Forsyth, A. Keshavarzian. Gut microbial metabolites in Parkinson’s disease: Association with lifestyle, disease characteristics, and treatment status. Neurobiology of Disease. 2022, 170, 105780. [CrossRef]
- M. Von Seggern, C. Szarowicz, M. Swanson, S. Cavotta, S. T. Pike, J. T. Lamberts. Purine molecules in Parkinson’s disease: Analytical techniques and clinical implications. Neurochemistry International. 2020, 139, 104793. [CrossRef]
- S. Cipriani, R. Bakshi, M. A. Schwarzschild. Protection by inosine in a cellular model of Parkinson’s disease. Neuroscience. 2014, 274, 242-249. [CrossRef]
- Z. Wang, Z. Liu, S. Feng, X. Song, D. Liu, N. Ma, X. Zhang, W. Liu, D. O. Wang, X. Liu, T. Ikejima. High glucose induces hippocampal neuron impairment through the SKP1/COX7C pathway: A potential mechanism for perimenopausal depression. Acta Pharm Sin B. 2025, 15, 5832-5853. [CrossRef]
- Bruckbauer, M. B. Zemel. Effects of dairy consumption on SIRT1 and mitochondrial biogenesis in adipocytes and muscle cells. Nutr Metab (Lond). 2011, 8, 91. [CrossRef]
- S.-L. Lee, S.-C. Tu, M.-Y. Hsu, T.-Y. Chin. Diosgenin Prevents Microglial Activation and Protects Dopaminergic Neurons from Lipopolysaccharide-Induced Neural Damage In Vitro and In Vivo. Int J Mol Sci. 2021, 22, 10361. [CrossRef]
- B. Li, P. Xu, S. Wu, Z. Jiang, Z. Huang, Q. Li, D. Chen. Diosgenin Attenuates Lipopolysaccharide-Induced Parkinson’s Disease by Inhibiting the TLR/NF-κB Pathway. Journal of Alzheimer’s Disease. 2018, 64, 943-955. [CrossRef]
- Z. Mao, H. Hui, X. Zhao, L. Xu, Y. Qi, L. Yin, L. Qu, L. Han, J. Peng. Protective effects of dioscin against Parkinson’s disease via regulating bile acid metabolism through remodeling gut microbiome/GLP-1 signaling. J Pharm Anal. 2023, 13, 1153-1167. [CrossRef]
- F. Piancone, M. Saresella, F. La Rosa, I. Marventano, M. Meloni, J. Navarro, M. Clerici. Inflammatory Responses to Monomeric and Aggregated α-Synuclein in Peripheral Blood of Parkinson Disease Patients. Front Neurosci. 2021, 15 - 2021. [CrossRef]
- Z. Yan, F. Yang, L. Sun, J. Yu, L. Sun, Y. Si, L. Yao. Role of gut microbiota-derived branched-chain amino acids in the pathogenesis of Parkinson’s disease: An animal study. Brain, Behavior, and Immunity. 2022, 106, 307-321. [CrossRef]
- J. Liu, X. Lv, T. Ye, M. Zhao, Z. Chen, Y. Zhang, W. Yang, H. Xie, L. Zhan, L. Chen, W.-C. Liu, K.-P. Su, J. Sun. Microbiota-microglia crosstalk between Blautia producta and neuroinflammation of Parkinson’s disease: A bench-to-bedside translational approach. Brain, Behavior, and Immunity. 2024, 117, 270-282. [CrossRef]
- P. Angoorani, H.-S. Ejtahed, S. Hasani-Ranjbar, S. D. Siadat, A. R. Soroush, B. Larijani. Gut microbiota modulation as a possible mediating mechanism for fasting-induced alleviation of metabolic complications: a systematic review. Nutr Metab (Lond). 2021, 18, 105. [CrossRef]
- M.-M. Liu, N. Zhou, N. Jiang, K.-M. Lu, C.-F. Wu, J.-K. Bao. Neuroprotective Effects of Oligosaccharides From Periplaneta Americana on Parkinson’s Disease Models In Vitro and In Vivo. Front Pharmacol. 2022, 13 - 2022. [CrossRef]
- P. LeWitt, L. Schultz, P. Auinger, M. Lu. CSF xanthine, homovanillic acid, and their ratio as biomarkers of Parkinson’s disease. Brain Res. 2011, 1408, 88-97. [CrossRef]
- Z. Khan, S. A. Ali. Oxidative stress-related biomarkers in Parkinson’s disease: A systematic review and meta-analysis. Iran J Neurol. 2018, 17, 137-144.
- S.-P. D. I. The Parkinson Study Group. Effect of Urate-Elevating Inosine on Early Parkinson Disease Progression: The SURE-PD3 Randomized Clinical Trial. JAMA. 2021, 326, 926-939. [CrossRef]
- S.-P. D. I. The Parkinson Study Group. Inosine to Increase Serum and Cerebrospinal Fluid Urate in Parkinson Disease: A Randomized Clinical Trial. JAMA Neurol. 2014, 71, 141-150. [CrossRef]
- P. M. Abou-Sleiman, M. M. K. Muqit, N. W. Wood. Expanding insights of mitochondrial dysfunction in Parkinson’s disease. Nat Rev Neurosci. 2006, 7, 207-219. [CrossRef]
- K. Čunátová, D. P. Reguera, J. Houštěk, T. Mráček, P. Pecina. Role of cytochrome c oxidase nuclear-encoded subunits in health and disease. Physiol Res. 2020, 69, 947-965. [CrossRef]
- X. Wang, Z. Wu, S. Wei, X. Zhao, J. Lin, F. Zhao, X. Liu. Integrated transcriptomic and single-cell RNA sequencing identifies lysosomal ion channel genes as potential biomarkers for Alzheimer’s disease. Front Genet. 2025, 16 - 2025. [CrossRef]
- B. Zheng, Z. Liao, J. J. Locascio, K. A. Lesniak, S. S. Roderick, M. L. Watt, A. C. Eklund, Y. Zhang-James, P. D. Kim, M. A. Hauser, E. Grünblatt, L. B. Moran, S. A. Mandel, P. Riederer, R. M. Miller, H. J. Federoff, U. Wüllner, S. Papapetropoulos, M. B. Youdim, I. Cantuti-Castelvetri, A. B. Young, J. M. Vance, R. L. Davis, J. C. Hedreen, C. H. Adler, T. G. Beach, M. B. Graeber, F. A. Middleton, J.-C. Rochet, C. R. Scherzer, P. D. G. E. C. the Global. PGC-1α, A Potential Therapeutic Target for Early Intervention in Parkinson’s Disease. Sci Transl Med. 2010, 2, 52ra73. [CrossRef]
- K.-J. Yang, H. Park, Y.-K. Chang, C. W. Park, S. Y. Kim, Y. A. Hong. Xanthine oxidoreductase inhibition ameliorates high glucose-induced glomerular endothelial injury by activating AMPK through the purine salvage pathway. Scientific Reports. 2024, 14, 11167. [CrossRef]
- S. Khanal, E.-J. Shin, C. J. Yoo, J. Kim, D.-Y. Choi. Inosine exerts dopaminergic neuroprotective effects via mitigation of NLRP3 inflammasome activation. Neuropharmacology. 2025, 266, 110278. [CrossRef]
- W. Xu, Y. Huang, R. Zhou. NLRP3 inflammasome in neuroinflammation and central nervous system diseases. Cell Mol Immunol. 2025, 22, 341-355. [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).



