Submitted:
17 March 2026
Posted:
18 March 2026
You are already at the latest version
Abstract
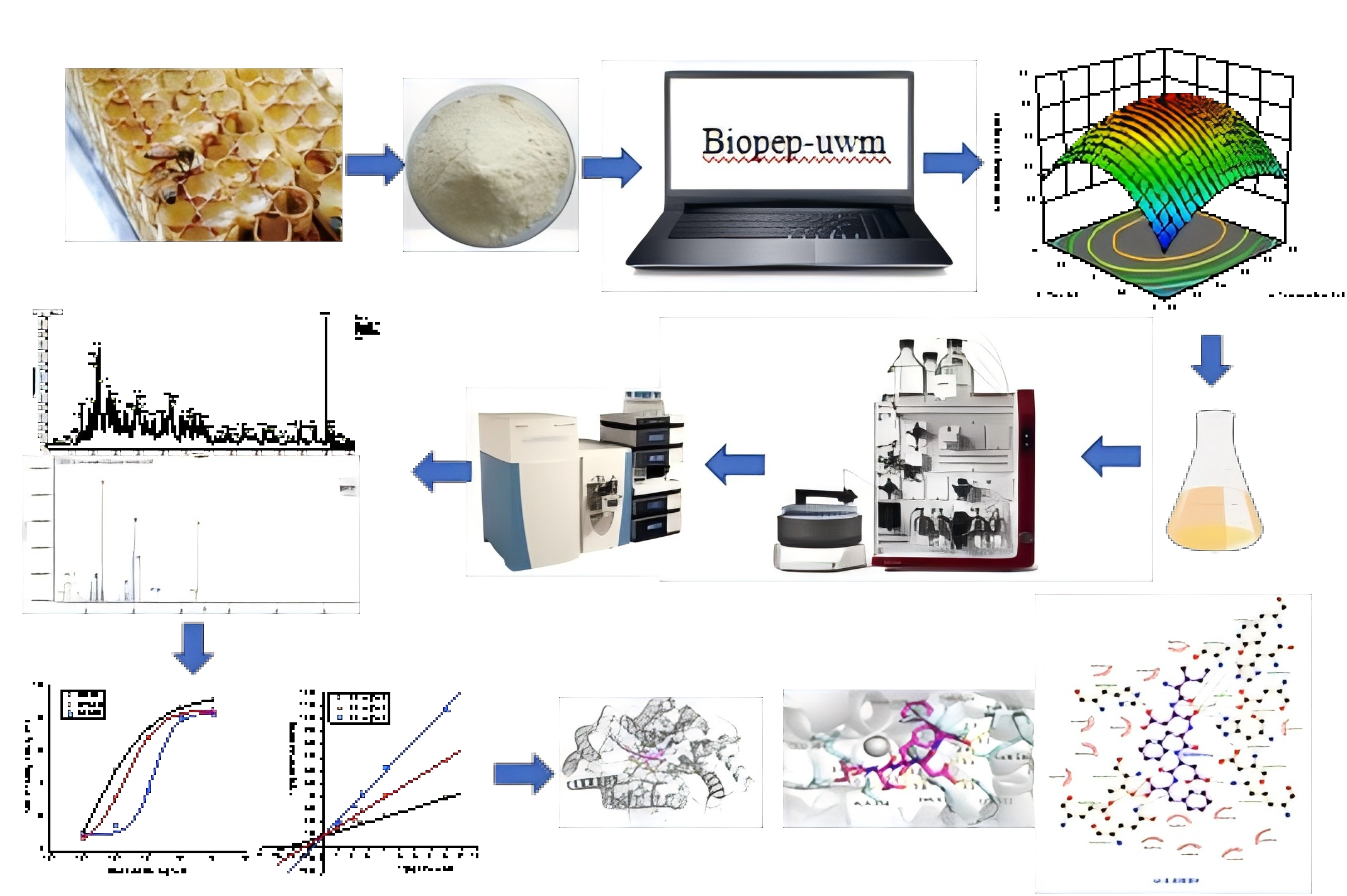
Keywords:
1. Introduction
2. Materials and Methods
2.1. Materials and Reagents
2.2. Evaluation of RJ Proteins as a Source of Bioactive Peptides
2.3. In Silico Analysis of ACE Inhibitory Peptides Released from MRJPs
2.4. Measurement of ACE Inhibitory Activity
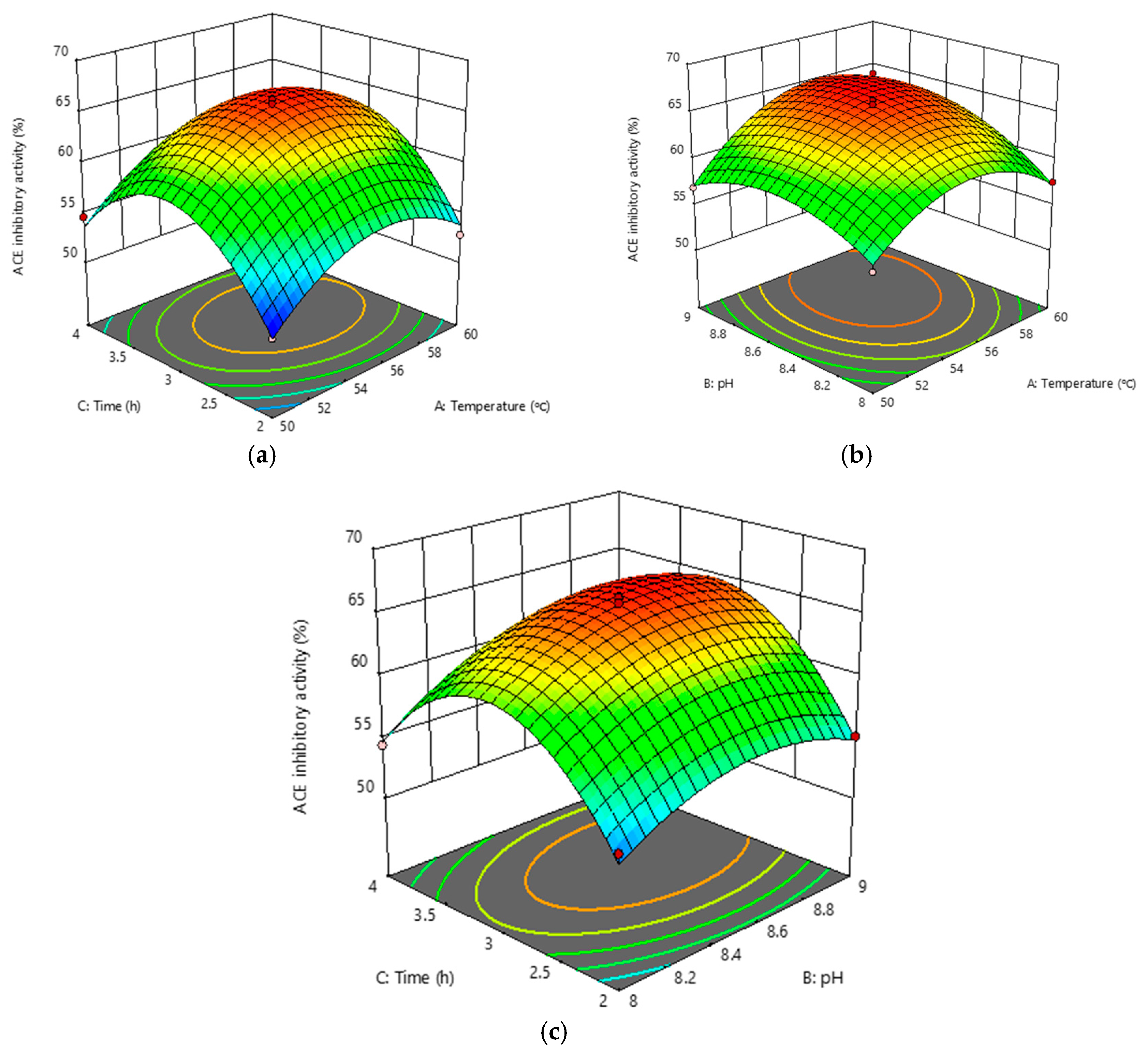
2.5. Optimization of RJ Protease Hydrolysis Process Using Response Surface Methodology
2.6. Isolation and Purification of ACE Inhibitory peptides
2.7. Peptide Sequence Analysis by LC-MS/MS
2.8. Scoring Method for de Novo Results
2.9. Synthesis of Peptides
2.10. ACE inhibition Kinetics
2.11. Molecular Docking
2.12. Statistical Analysis
3. Results
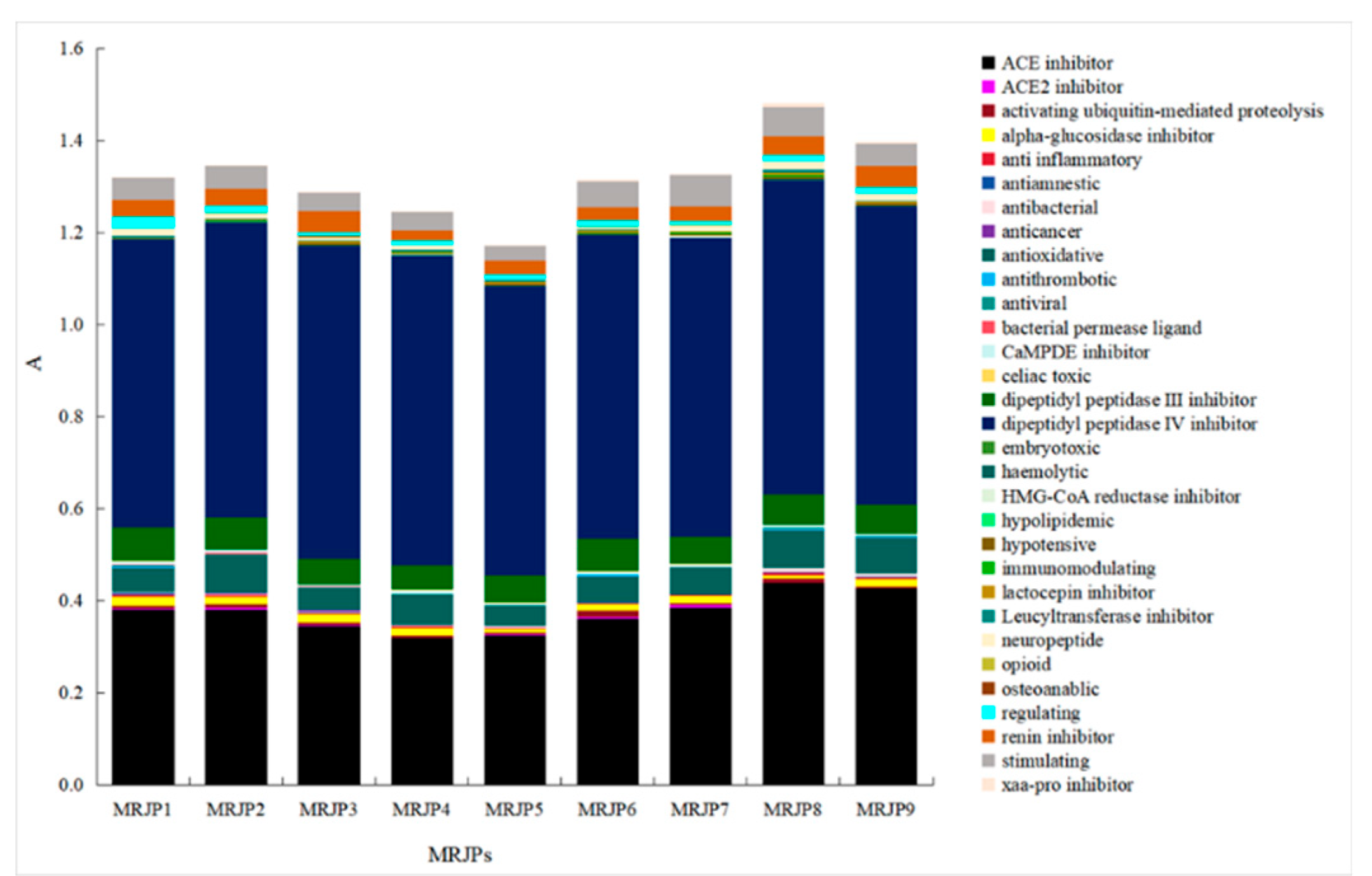
3.1. In Silico Analysis of Bioactive Peptides Encrypted in MRJPs
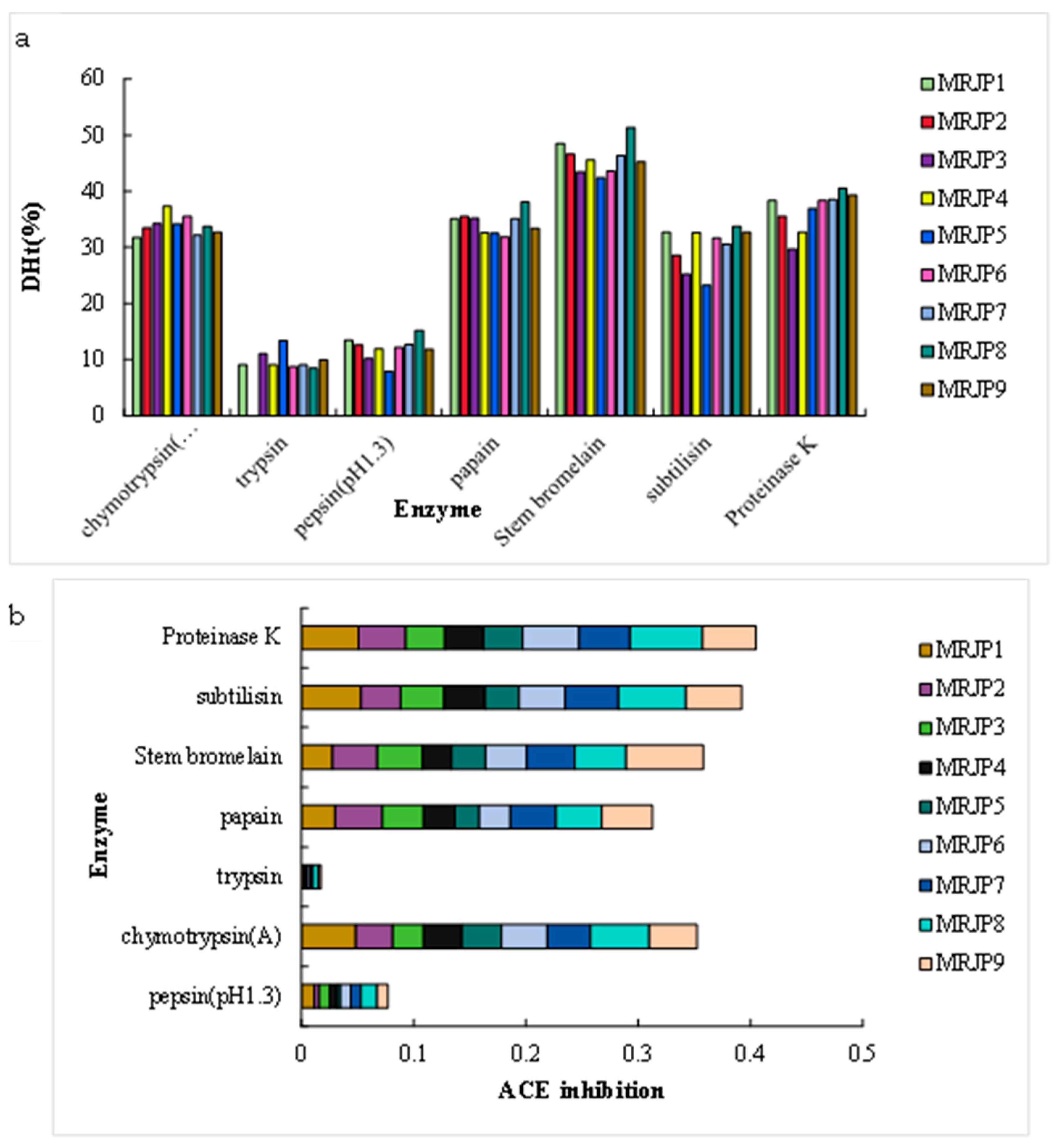
3.2. In Silico Proteolysis of MRJPs for the Production of ACE Inhibitory Peptides
3.3. In Vitro Proteolysis of RJ Proteins for the Production of ACE Inhibitory Peptides by RSM
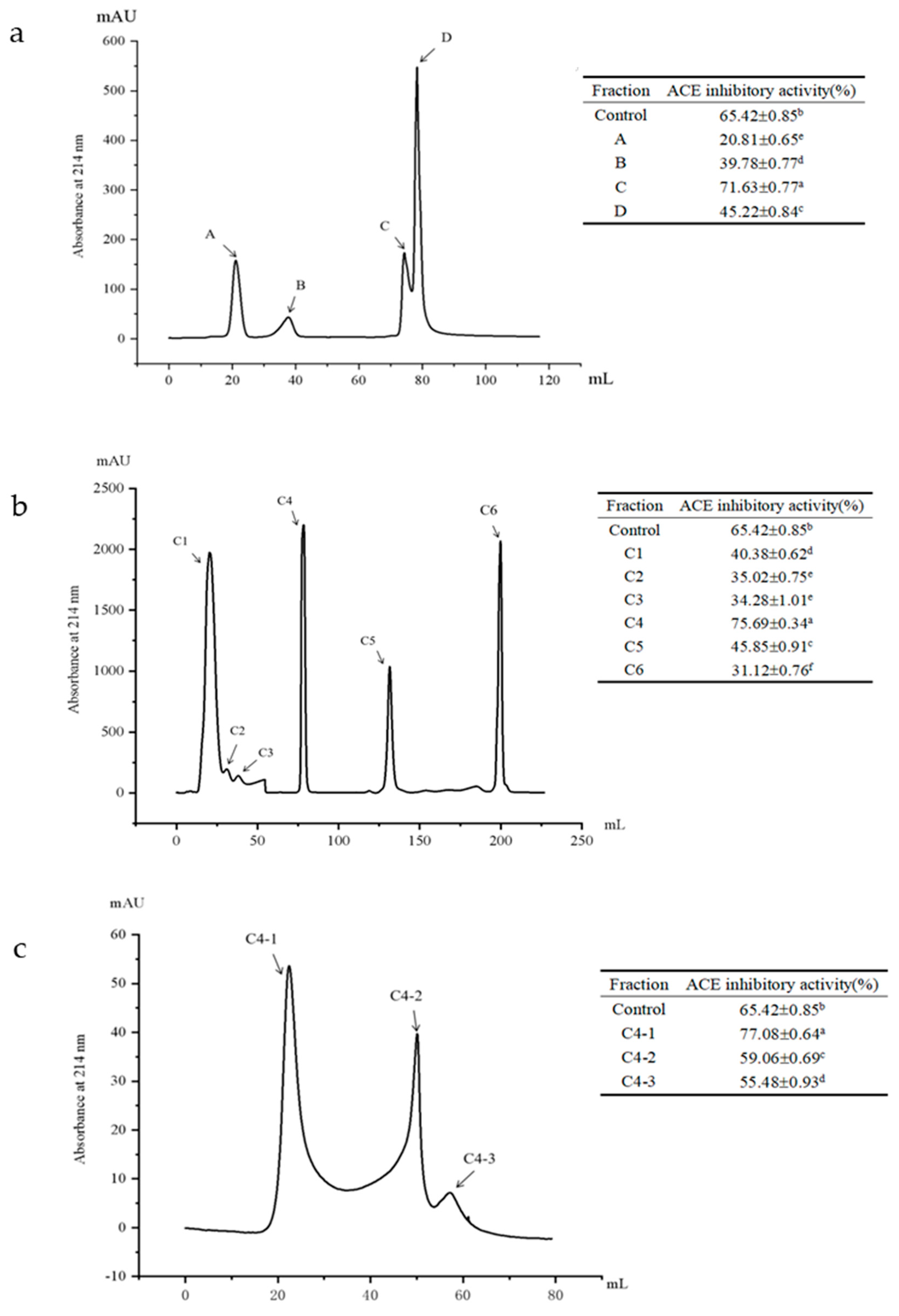
3.4. Isolation and Purification of ACE Inhibitory Peptides
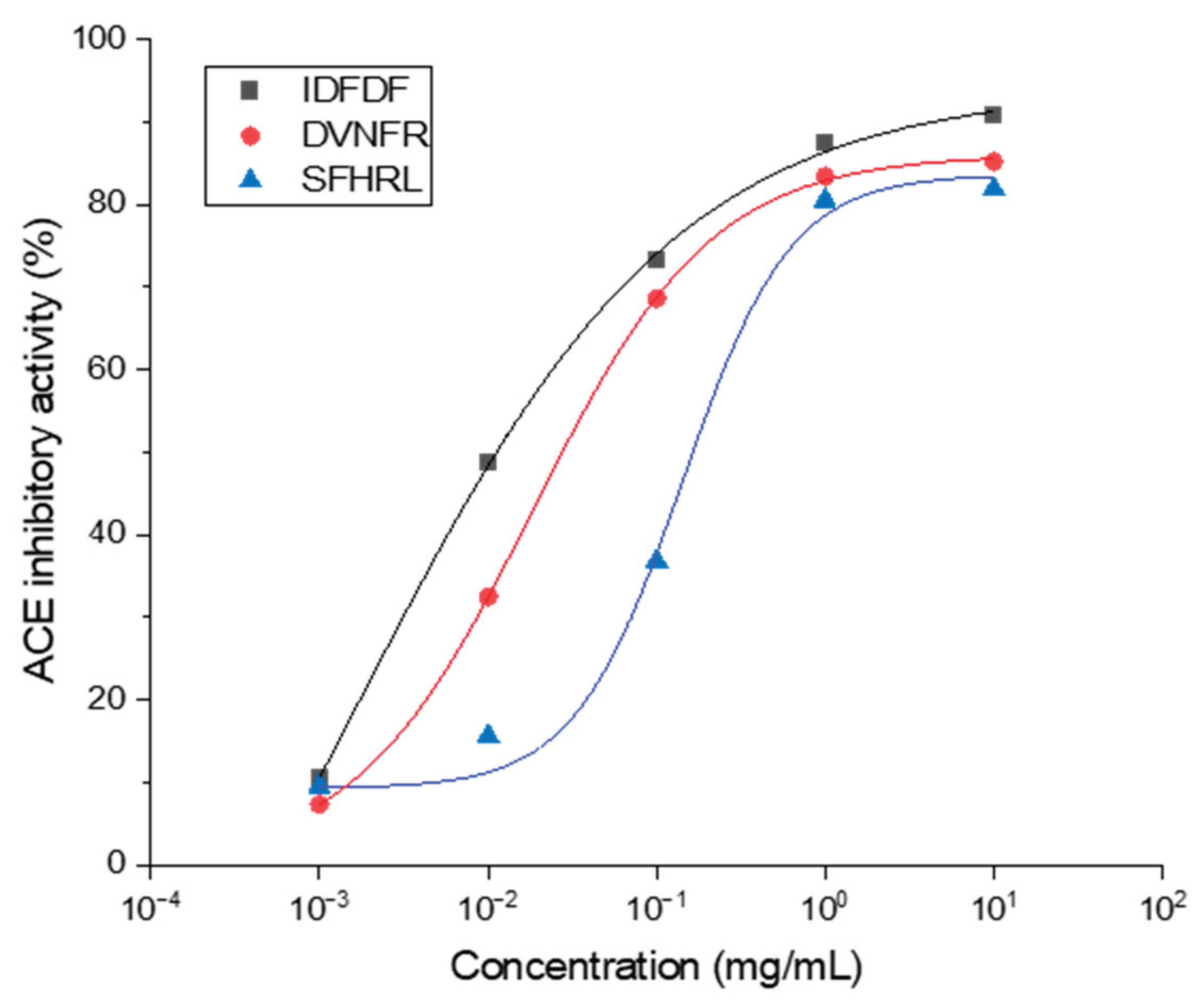
3.5. Identification, Screening and Activities of the ACE Inhibitory Peptides
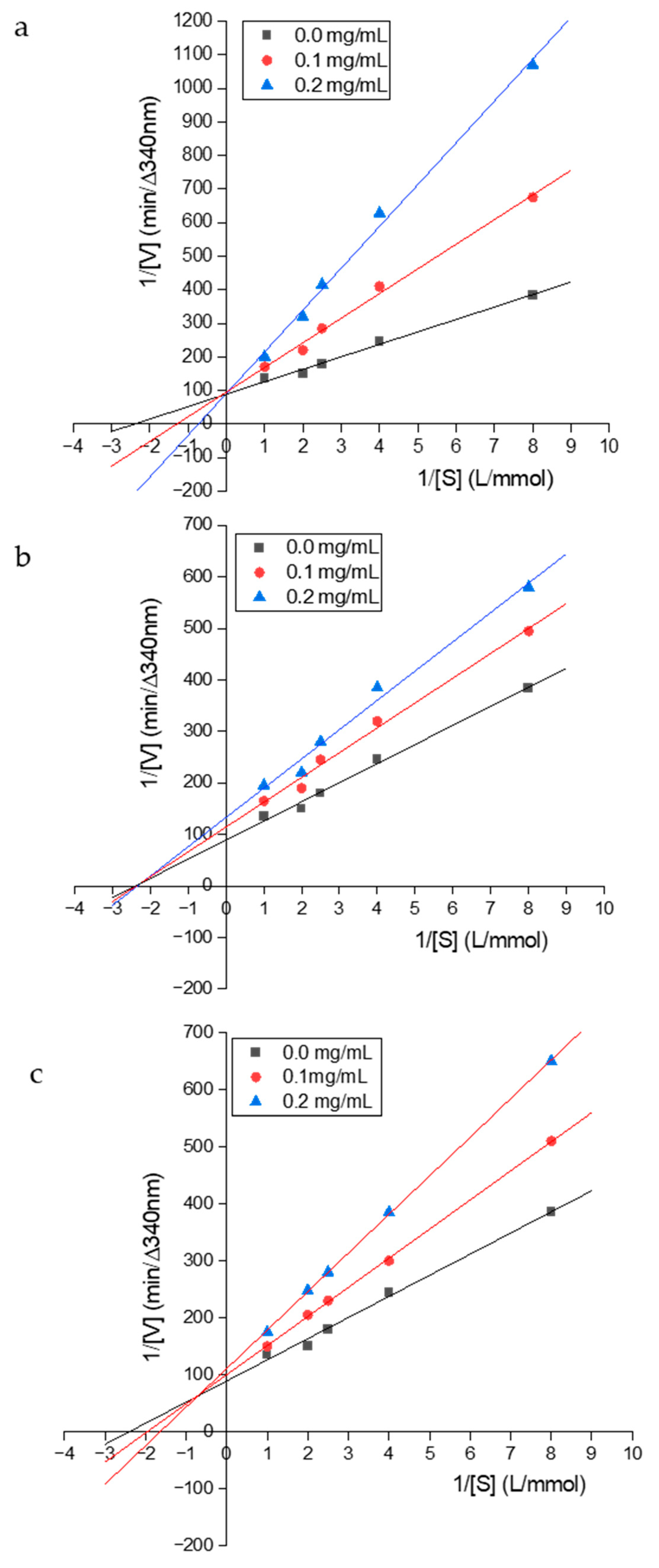
3.6. Determination of the ACE Inhibition Pattern of the Purified Peptides
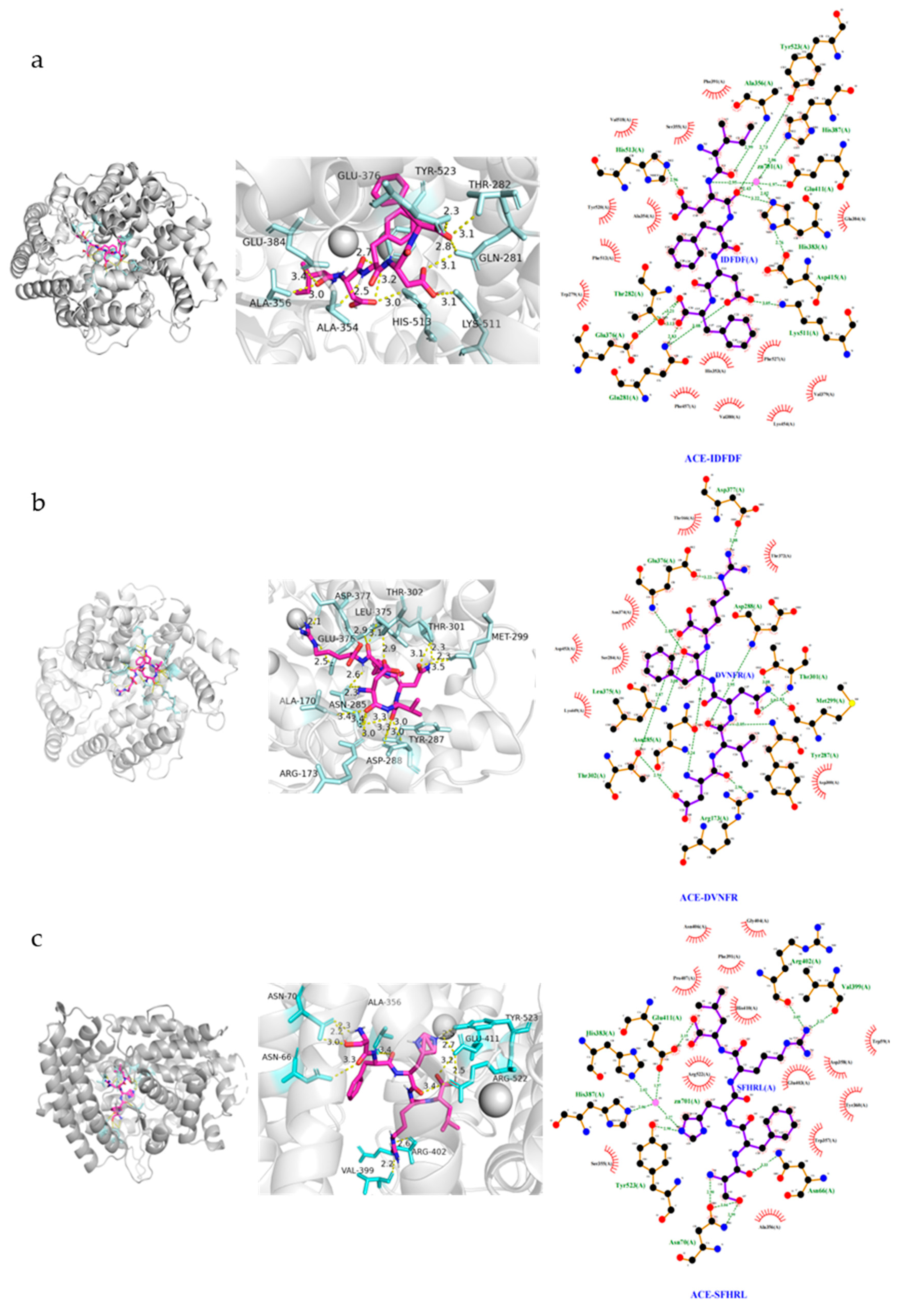
3.7. Molecular Docking
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Simuth, J. Some properties of the main protein of honeybee (Apis mellifera) royal jelly. Apidologie 2001, 32, 69–80. [CrossRef]
- Tian, W.L.; Li, M.; Guo, H.Y.; Peng, W.J.; Xue, X.F.; Hu, Y.F.; Liu, Y.; Zhao, Y.Z.; Fang, X.M.; Wang, K.; Li, X.T.; Tong, Y.F.; Conlon, M.A.; Wu, W.; Ren, F.Z.; Chen, Z.Z. Architecture of the native major royal jelly protein 1 oligomer. Nat. Commun. 2018, 9, 3373. [CrossRef]
- Buttstedt, A.; Ihling, C. H.; Pietzsch, M.; Moritz, R. F. Royalactin is not a royal making of a queen. Nature 2016, 537, E10–E12. [CrossRef]
- Kucharski, R.; Maleszka, J.; Foret, S.; Maleszka, R. Nutritional control of reproductive status in honeybees via DNA methylation. Science 2008, 319, 1827–1830. [CrossRef]
- Fratini, F.; Cilia, G.; Mancini,S.; Felicioli, A. Royal Jelly: An Ancient remedy with remarkable antibacterial properties. Microbiol. Res. 2016, 192, 130–141. [CrossRef]
- Guo, H. M.; Richel, A.; Hao, Y. Q.; Fan, X.; Everaert, N.; Yang, X. S.; Ren, G. X. Novel dipeptidyl peptidase-IV and angiotensin-I-converting enzyme inhibitory peptides released from quinoa protein by in silico proteolysis. Food Sci. Nutr. 2020, 8, 1415–1422. [CrossRef]
- Ahmad, S.; Campos, M.G.; Fratini, F.; Altaye, S.Z.; Li, J. New insights into the biological and pharmaceutical properties of royal jelly. Int. J. Mol. Sci. 2020, 21, 382. [CrossRef]
- Lv, G. D.; Kang, W.P.; Wei, Q.H.; Gao, H.N.; Hu, H. Effect of royal jelly on kidney of D-galactose induced aging mice. J. Food Sci. Technol. 2023, 5, 45-57.
- Rosmilah, M.; Shahnaz, M.; Patel, G.; Lock, J.; Rahman, D.; Masita, A.; Noormalin, A. Characterization of major allergens of royal jelly Apis mellifera. Trop. Biomed. 2008, 25, 243–251.
- Altaye, S.Z.; Meng, L.F.; Lu, Y.; Li, J.K. The Emerging Proteomic Research Facilitates in-Depth Understanding of the Biology of Honeybees. Int. J. Mol. Sci. 2019, 20, 4252. [CrossRef]
- Feng, M.; Fang,Y.; Han, B.; Xu, X.; Fan, P.; Hao, Y.; Qi, Y.; Hu, H.; Huo, X.; Meng, L. In-depth N-glycosylation reveals species-specific modifications and functions of the royal jelly protein from Western (Apis mellifera) and Eastern Honeybees (Apis cerana). J. Proteome Res. 2015,14, 5327–5340. [CrossRef]
- Abdelhedi, O.; Nasri, R.; Mora, L.; Jridi, M.; Toldra, F.; Nasri, M. In silico analysis and molecular docking study of angiotensin I-converting enzyme inhibitory peptides from smooth-hound viscera protein hydrolysates fractionated by ultrafiltration. Food Chem. 2018, 239, 453–463. [CrossRef]
- Guo, H.; Kouzuma, Y.; Yonekura, M. Structures and properties of antioxidative peptides derived from royal jelly protein. Food Chem. 2009, 113, 238–245. [CrossRef]
- Matsui, T.; Yukiyoshi, A.; Doi, S.; Sugimoto, H.; Yamada, H.; Matsumoto, K. Gastrointestinal enzyme production of bioactivepeptides from royal jelly protein and their antihypertensive abilityin SHR. J Nutr. Biochem. 2002, 13, 80–86.
- Tokunaga, K.; Yoshida, C.; Suzuki, K.; Maruyama, H.; Futamura, Y.; Araki, Y.; Mishima, S. Antihypertensive effect of peptide from royal jelly in spontaneously hypertensive rats. Biol. Pharm. Bull. 2004, 27, 189–192. [CrossRef]
- Sultana, A.; Nabi, A. H.; Nasir, U. M.; Maruyama, H.; Suzuki, K. M.; Mishima, S.; Suzuki, F. A dipeptide YY derived from royaljelly proteins inhibits renin activity. Int. J. Mol. Med. 2008, 21, 677–681. [CrossRef]
- Chen, J.; Yu, X.; Chen, Q.; Wu, Q.; He, Q. Screening and mechanisms of novel angiotensin-I-converting enzyme inhibitory peptides from rabbit meat proteins: A combined in silico and in vitro study. Food Chem. 2022, 370, 131070. [CrossRef]
- Liu, C.; Fang, L.; Min, W.; Liu, J.; Li, H. Exploration of the molecular interactions between angiotensin-I-converting enzyme (ACE) and the inhibitory peptides derived from hazelnut (Corylus heterophylla Fisch.). Food Chem. 2018, 245, 471−480. [CrossRef]
- Minkiewicz, P.; Iwaniak, A.; Darewicz, M. BIOPEP-UWM database of bioactive peptides: Current opportunities. Int. J. Mol. Sci. 2019, 20, 5978. [CrossRef]
- Cao, S.; Wang, Y.; Hao, Y.; Zhang, W.; Zhou, G. Antihypertensive effects in vitro and in vivo of novel angiotensin-converting enzyme inhibitory peptides from bovine bone gelatin hydrolysate. J. Agric. Food Chem. 2020, 68, 759–768. [CrossRef]
- Lowry, O. H.; Rosebrough, N. J.; Farr, A. L.; Randall, R. J. Protein measurement with the folin phenol reagent. J. Biol. Chem. 1951, 193, 265–275. [CrossRef]
- Tahir, R.A.; Bashir, A.; Yousaf, M.N.; Ahmed, A.; Dali, Y.; Khan, S.; Sehgal, S.A. In Silico identification of angiotensin-converting enzyme inhibitory peptides from MRJP1. PLoS ONE 2020, 15, e0228265. [CrossRef]
- Udenigwe, C. C. Bioinformatics approaches, prospects and challenges of food bioactive peptide research. Trends. Food Sci. Tech. 2014, 36, 137–143. [CrossRef]
- Ligné, T.; Pauthe, E.; Monti, J.P.; Gacel, G.; Larreta-Garde, V. Additional data about thermolysin specificity in buffer-and glycerol-containing media. Biochim. Biophys. Acta (BBA)-Protein Structure and Molecular Enzymology, 1997, 1337, 143–148. [CrossRef]
- Giampieri, F.; Quiles, J. L.; Cianciosi, D.; Forbes-Hernández, T. Y.; Orantes-Bermejo, F. J.; Alvarez-Suarez, J. M.; Battino, M. Bee Products: An Emblematic Example of Underutilized Sources of Bioactive Compounds. J. Agric. Food Chem. 2022, 70, 6833–6848. [CrossRef]
- Duan, X. J.; Dong, Y. F.; Zhang, Min.; Li, Z. H.; Bu, G. H.; Chen, F. S. Identification and molecular interactions of novel ACE inhibitory peptides from rapeseed protein. Food Chem. 2023, 422, 136085. [CrossRef]
- Sheng, J. Y.; Yang, X.Y.; Chen, J. T.; Peng, T. H.; Yin, X. Q.; Liu, W.; Liang, M.; Wan, J. L.; Yang, X. L. Antioxidative effects and mechanism study of bioactive peptides from defatted walnut (Juglans regia L.) meal hydrolysate. J. Agric. Food Chem. 2019, 67, 3305–3312.
- Zhou, H.; Safdar, B.; Li, H.; Yang, L.; Ying, Z.; Liu, X. Identification of a novel α-amylase inhibitory activity peptide from quinoa protein hydrolysate. Food Chem. 2023, 403, 134434. [CrossRef]
- Panyayai, T.; Sangsawad, P.; Pacharawongsakda, E.; Sawatdichaikul, O.; Tongsima, S.; Choowongkomon, K. The potential peptides against angiotensin-I converting enzyme through a virtual tripeptide-constructing library. Comput. Biol. Chem. 2018, 77, 207–213. [CrossRef]
| Coded Level | Independent Variable | ||
|---|---|---|---|
| A:temperature (℃) | B: pH | C: time (h) | |
| -1 | 45 | 8 | 2 |
| 0 | 50 | 8.5 | 3 |
| 1 | 55 | 9 | 4 |
| number | A:Temperature (℃) | B: pH | C:Time (h) | ACE inhibitory activity (%) |
|---|---|---|---|---|
| 1 | 50 | 8 | 3 | 55.62±0.88 |
| 2 | 60 | 8 | 3 | 57.54±0.79 |
| 3 | 50 | 9 | 3 | 56.93±1.20 |
| 4 | 60 | 9 | 3 | 63.89±0.95 |
| 5 | 50 | 8.5 | 2 | 50.80±1.10 |
| 6 | 60 | 8.5 | 2 | 52.78±0.87 |
| 7 | 50 | 8.5 | 4 | 54.60±0.89 |
| 8 | 60 | 8.5 | 4 | 57.51±0.69 |
| 9 | 55 | 8 | 2 | 53.71±0.93 |
| 10 | 55 | 9 | 2 | 55.10±0.73 |
| 11 | 55 | 8 | 4 | 54.45±0.88 |
| 12 | 55 | 9 | 4 | 58.82±1.13 |
| 13 | 55 | 8.5 | 3 | 64.89±1.04 |
| 14 | 55 | 8.5 | 3 | 66.30±0.85 |
| 15 | 55 | 8.5 | 3 | 65.41±0.96 |
| 16 | 55 | 8.5 | 3 | 66.20±1.22 |
| 17 | 55 | 8.5 | 3 | 65.89±1.51 |
| source | Sum of squares | df | mean square | F-value | P-value Prob>F |
|---|---|---|---|---|---|
| model | 453.42 | 9 | 50.38 | 58.08 | <0.0001a |
| A-Temp | 23.81 | 1 | 23.81 | 27.44 | 0.0012a |
| B-pH | 22.78 | 1 | 22.78 | 26.26 | 0.0014a |
| C-TM | 20.8 | 1 | 20.80 | 23.98 | 0.0018a |
| AB | 6.5 | 1 | 6.5 | 7.5 | 0.0290a |
| AC | 0.2025 | 1 | 0.2025 | 0.2334 | 0.6437 |
| BC | 2.25 | 1 | 2.25 | 2.59 | 0.1513 |
| A2 | 81.52 | 1 | 81.52 | 93.97 | <0.0001a |
| B2 | 33.6 | 1 | 33.6 | 38.74 | 0.0004a |
| C2 | 229.01 | 1 | 229.01 | 263.99 | <0.0001a |
| residual | 6.07 | 7 | 0.8675 | ||
| Lack of Fit | 4.55 | 3 | 1.52 | 3.99 | 0.1072 |
| Pure error | 1.52 | 4 | 0.38 | ||
| Cor Totol | 459.50 | 16 | |||
| Std.Dev. | 0.93 | R-Squared | 0.9868 | ||
| Mean | 73.83 | Adj R-Squared | 0.9698 | ||
| C.V.% | 1.26 | Pred R-Squared | 0.8363 | ||
| PRESS | 75.22 | Adeq Precision | 20.841 |
| Peptide sequence | Abundance | Score A | Confidence | Score B | Binding energy | Score C | Total score | ACE inhibitory activity (%) |
|---|---|---|---|---|---|---|---|---|
| KNYPF | 1.58×1010 | 95.05 | 418.0 | 99.09 | -8.4 | 88.42 | 94.27 | 78.56±1.24 |
| VEIPH | 1.41×1010 | 84.53 | 420.6 | 99.72 | -8.6 | 90.53 | 90.89 | 75.78±1.02 |
| KPYPDWS | 1.61×1010 | 96.26 | 420.2 | 99.62 | -7.7 | 81.05 | 92.71 | 71.55±1.91 |
| IDFDF | 1.50×1010 | 90.02 | 416.2 | 98.67 | -9.5 | 100.00 | 95.61 | 87.04±1.12 |
| FDYDFG | 1.60×1010 | 95.82 | 412.5 | 97.80 | -6.8 | 71.58 | 89.14 | 61.15±1.89 |
| SFHRL | 1.67×1010 | 100.00 | 421.8 | 100.0 | -8.8 | 92.63 | 97.79 | 81.43±1.25 |
| DVNFR | 1.56×1010 | 93.41 | 419.8 | 99.53 | -8.9 | 93.68 | 95.32 | 83.26±1.37 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




