Submitted:
17 March 2026
Posted:
18 March 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Experimental Details
| 1H NMR Spectroscopic Assignments | |||
| 4 (DMF-d7) | 5 (THF-d8) | 6 (THF-d8) | |
| Pt – CH3 | 2.35 ppm (s with 195Pt satellites), 2JPt-H = 73 Hz |
2.35 ppm (s with 195Pt satellites), 2JPt-H = 73 Hz |
2.58 ppm (s with 195Pt satellites), 2JPt-H = 75 Hz |
| CH3 groups bonded to bipyridyl ligand | 2.68 ppm (s) | 2.50 ppm (s) |
2.56 ppm (s) |
| Hydrogens bonded to aromatic carbons in Bipy-R2 C(5) – H C(4) – H C(3) – H C(6) – H |
7.79 ppm (d) 3JH-H = 5 Hz N/A 8.80 ppm (s) 8.90 ppm (d with 195Pt satellites) 3JH-H = 5 Hz 3JPt-H = 13 Hz |
N/A 7.94 ppm (d) 3JH-H = 8 Hz 8.38 ppm (d) 3JH-H = 8 Hz 8.77 ppm (s with 195Pt satellites) 3JPt-H = 14 Hz |
7.54 ppm (d) 3JH-H = 8 Hz 7.95 ppm (d of d) 3JH-H = 8 Hz 8.24 ppm (d) 3JH-H = 8 Hz N/A |
| 13C{1H} NMR Spectroscopic Assignments | |||
| 4 (DMF-d7) | 5 (THF-d8) | 6 (THF-d8) | |
| Pt – CH3 | – 14.8 ppm (s with 195Pt satellites) 1JPt-C = 519 Hz |
– 10.2 ppm (s with 195Pt satellites) 1JPt-C = 520 Hz |
|
| CH3 groups attached to bipyridyl ligand | 18.5 ppm | 26.2 ppm | |
| Carbons in aromatic rings of bipy-R2 (R = H or CH3) | 124.2 ppm 138.4 ppm 140.9 ppm 148.8 ppm 154.0 ppm |
122.1 ppm 128.7 ppm 139.9 ppm 160.3 ppm 163.3 ppm |
|
| 195Pt NMR Spectroscopic Assignments | |||
| 4 (DMF-d7) | 5 (THF-d8) | 6 (THF-d8) | |
| – 3691 ppm | – 3740 ppm | – 3487 ppm | |
| Compound | Infrared Absorption Frequencies |
| 4 | 3433 (m), 2961 (m), 2901 (s), 2813 (w), 1613 (vs), 1558 (m), 1483 (s), 1442 (m), 1416 (m), 1373 (w), 1301 (m), 1246 (s), 1229 (w), 1218 (m), 1136 (w), 1118 (w), 1075 (m), 1026 (s), 920 (m), 891 (w), 836 (s), 554 (m), 520 (w), 486 (w), 419 (w). |
| 5 | 3436 (s), 3039 (w), 2973 (m), 2898 (s), 2810 (w), 1630 (m), 1607 (s), 1575 (w), 1500 (w), 1480 (vs), 1392 (w), 1313 (m), 1248 (m), 1234 (m), 1163 (m), 1150 (w), 1064 (m), 1052 (m), 1002 (w), 836 (s), 727 (w), 695 (w). |
| 6 | 3437 (s), 3078 (w), 2984 (w), 2918 (m), 1631 (w), 1601 (s), 1569 (m), 1464 (m), 1443 (s), 1375 (m), 1322 (w), 1249 (w), 1244 (m), 1222 (w), 1175 (w), 1120 (m), 1104 (w), 1034 (w), 1008 (m), 899 (w), 887 (w), 815 (w), 790 (vs), 732 (w), 706 (w), 643 (w), 617 (w). |
3. Results
4. Discussion
5. Conclusions
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Escolà, A.; Crespo, M.; López, C.; Quirante, J.; Jayaraman, A.; Polat, I. H.; Badía, J.; Baldomà, L.; Cascante, M. On the Stability and Biological Behavior of Cyclometallated Pt(IV) Complexes with Halido and Aryl Ligands in the Axial Positions. Bioorg. Med. Chem. 2016, 24, 5804–5815. [Google Scholar] [CrossRef]
- Tan, M.-X.; Wang, Z.-F.; Qin, Q.-P.; Huang, X.-L.; Zou, B.-Q.; Liang, H. Complexes of Platinum(II/IV) with 2-Phenylpyridine Derivatives as a New Class of Promising Anticancer Agents. Inorg. Chem. Commun. 2019, 108, 107510. [Google Scholar] [CrossRef]
- Annunziata, A.; Amoresano, A.; Cucciolito, M. E.; Esposito, R.; Ferraro, G.; Iacobucci, I.; Imbimbo, P.; Lucignano, R.; Melchiorre, M.; Monti, M.; et al. Pt(II) Versus Pt(IV) in Carbene Glycoconjugate Antitumor Agents: Minimal Structural Variations and Great Performance Changes. Inorg. Chem. 2020, 59, 4002–4014. [Google Scholar] [CrossRef]
- Bauer, E.; Domingo, X.; Balcells, C.; Polat, I. H.; Crespo, M.; Quirante, J.; Badía, J.; Baldomà, L.; Font-Bardia, M.; Cascante, M. Synthesis, Characterization and Biological Activity of New Cyclometallated Platinum(IV) Iodido Complexes. Dalton Trans. 2017, 46, 14973–14987. [Google Scholar] [CrossRef]
- Johnston, T. C.; Suntharalingam, K.; Lippard, S. J. The Next Generation of Platinum Drugs: Targeted Pt(II) Agents, Nanoparticle Delivery, and Pt(IV) Prodrugs. Chem. Rev. 2016, 116, 3436–3486. [Google Scholar] [CrossRef] [PubMed]
- Ghezzi, A. R.; Aceto, M.; Cassino, C.; Gabano, E.; Osella, D. Uptake of Antitumor Platinum(II)-Complexes by Cancer Cells, Assayed by Inductively Coupled Plasma Mass Spectrometry (ICP-MS). J. Inorg. Biochem. 2004, 98, 73–78. [Google Scholar] [CrossRef] [PubMed]
- Crespo, M. Cyclometallated Platinum(IV) Compounds as Promising Antitumour Agents. J. Organomet. Chem. 2019, 879, 15–26. [Google Scholar] [CrossRef]
- Pouryasin, Z.; Yousefi, R.; Nabavizadeh, S. M.; Rashidi, M.; Hamidizadeh, P.; Alavianmehr, M.-M.; Moosavi-Movahedi, A. A. Anticancer and DNA Binding Activities of Platinum(IV) Complexes: Importance of Leaving Group Departure Rate. Appl. Biochem. Biotechnol. 2014, 172, 2604–2617. [Google Scholar] [CrossRef] [PubMed]
- Levenson, A. S.; Jordan, V. C. MCF-7: The First Hormone-Responsive Breast Cancer Cell Line. Cancer Res. 1997, 57, 3071–3078. [Google Scholar]
- Lee, A. V.; Oesterreich, S.; Davidson, N. E. MCF-7 Cells – Changing the Course of Breast Cancer Research and Care for 45 Years. J. Natl. Cancer Inst. 2015, 107, djv073. [Google Scholar] [CrossRef]
- Schneider, U.; Schwenk, H.-U.; Bornkamm, G. Characterization of EVB- Genome Negative “Null” and “T” Cell Lines Derived from Children with Acute Lymphoblastic Leukemia and Leukemic Transformed Non-Hodgkin Lymphoma. Int. J. Cancer 1977, 19, 621–626. [Google Scholar] [CrossRef] [PubMed]
- Schwenk, H.-U.; Schneider, U. Cell Cycle Dependency of a T Cell Marker on Lymphoblasts. Blut: Zeit. Für die Gesamte Blutforschung 1975, 31, 299–306. [Google Scholar] [CrossRef]
- Lozzio, C. B.; Lozzio, B. B. Human Chronic Myelogenous Leukemia Cell-Line with Positive Philadelphia Chromosome. Blood 1975, 45, 321–334. [Google Scholar] [CrossRef]
- Kelly, M. E.; Gómez-Ruiz, S.; Kluge, R.; Merzweiler, K.; Steinborn, D.; Wagner, C.; Schmidt, H. Studies of Mononuclear and Dinuclear Complexes of Dibromodimethylplatinum(IV): Preparation, Characterization and Crystal Structures. Inorg. Chim. Acta 2009, 362, 1323–1332. [Google Scholar] [CrossRef]
- Batsanov, S. S. Van der Waals Radii of Elements. Inorg. Materials 2001, 37, 1031–1046. [Google Scholar] [CrossRef]
- Zamora, A.; Wachter, E.; Vera, M.; Heidary, D. K.; Rodríguez, V.; Ortega, E.; Fernández-Espín, V.; Janiak, C.; Glazer, E. C.; Barone, G.; et al. Organoplatinum(II) Complexes Self-Assemble and Recognize AT-Rich Duplex DNA Sequences. Inorg. Chem. 2021, 60, 2178–2187. [Google Scholar] [CrossRef] [PubMed]
- Veclani, D.; Tolazzi, M.; Cerón-Carrasco, J. P.; Melchior, A. Intercalation Ability of Novel Monofunctional Platinum Anticancer Drugs: A Key Step in Their Biological Action. J. Chem. Inf. Model. 2021, 61, 4391–4399. [Google Scholar] [CrossRef]
- Wang, F.-Y.; Liu, R.; Huang, K.-B.; Feng, H.-W.; Liu, Y.-N.; Liang, H. New Platinum(II)-Based DNA Intercalator: Synthesis, Characterization, and Anticancer Activity. Inorg. Chem. Commun. 2019, 105, 182–187. [Google Scholar] [CrossRef]
- Pages, B. J.; Garbutcheon-Singh, K. B.; Aldrich-Wright, J. R. Platinum Intercalators of DNA as Anticancer Agents. Eur. J. Inorg. Chem. 2017, 2017, 1613–1624. [Google Scholar] [CrossRef]
- Arabi, A.; Cogley, M. O.; Fabrizio, D.; Stitz, S.; Howard, W. A.; Wheeler, K. A. Anticancer Activity of Nonpolar Pt(CH3)2I2{bipy} is Found to be Superior among Four Similar Organoplatinum(IV) Complexes. J. Mol. Struct. 2023, 1274, 134551. [Google Scholar] [CrossRef]
- Engel, L. W.; Young, N. A.; Tralka, T. S.; Lippman, M. E.; O’Brien, S. J.; Joyce, M. J. Establishment and Characterization of Three New Continuous Cell Lines Derived from Human Breast Carcinomas. Cancer Res. 1978, 38, 3352–3364. [Google Scholar]
- O’Brien, A. M.; Wheeler, K. A.; Howard, W. A. Synthesis, Structure, and Anticancer Activity of a Dinuclear Organoplatinum(IV) Complex Stabilized by Adenine. Compounds 2025, 5, 16–27. [Google Scholar] [CrossRef]
- Clark, H. C.; Manzer, L. E. Reactions of π-(1,5-Cyclooctadiene)organoplatinum(II) Compounds and the Synthesis of Perfluoroalkylplatinum Complexes. J. Organomet. Chem. 1973, 59, 411–28. [Google Scholar]
- Appleton, T. G. Encyclopedia of Spectroscopy and Spectrometry, 3rd Ed. ed; 2017. [Google Scholar]
- Sheldrick, G. M. SADABS and TWINABS – Program for Area Detector Absorption Corrections; University of Göttingen: Göttingen, Germany, 2014. [Google Scholar]
- Barbour, L. J. X-Seed 4: Updates to a Program for Small-Molecule Supramolecular Crystallography. J. Appl. Cryst. 2020, 53, 1141–1146. [Google Scholar] [CrossRef]
- Sheldrick, G. M. SHELXT-Integrated Space-Group and Crystal-Structure Determination. Acta Cryst. A 2015, 71, 3–8. [Google Scholar] [CrossRef]
- Kumar, C. V.; Asuncion, E. H. DNA Binding Studies and Site Selective Fluorescence Sensitization of an Anthryl Probe. J. Am. Chem. Soc. 1993, 115, 8547–8553. [Google Scholar] [CrossRef]
- Morris, J.; Kunkel, M. W.; White, S. L.; Wishka, D. G.; Lopez, O. D.; Bowles, L.; Brady, P. S.; Ramsey, P.; Grams, J.; Rohrer, T.; et al. Targeted Investigational Oncology Agents in the NCI60: A Phenotypic Systems-Based Resource. Mol. Cancer Ther. 2023, 22, 1270–1279. [Google Scholar] [CrossRef]
- Shoemaker, R. H. The NCI60 Human Tumor Cell Line Anticancer Drug Screen. Nat. Rev. Cancer 2006, 6, 813–823. [Google Scholar] [CrossRef]
- Monks, A.; Scudiero, D.; Skehan, P.; Shoemaker, R.; Paull, K.; Vistica, D.; Hose, C.; Langley, J.; Cronise, P.; Vaigro-Wolff, A.; et al. Feasibility of a High-Flux Anticancer Drug Screen Using a Diverse Panel of Cultured Human Tumor Cell Lines. J. Natl. Cancer Inst. 1991, 83, 757–766. [Google Scholar] [CrossRef]
- D’Abramo, M.; Castellazzi, C. L.; Orozco, M.; Amadei, A. On the Nature of DNA Hyperchromic Effect. J. Phys. Chem. B 2013, 117, 8697–8704. [Google Scholar] [CrossRef]
- Tu, B.; Chen, Z.-F.; Liu, Z.-J.; Cheng, L.-Y.; Hu, Y.-J. Interaction of Flavones with DNA in Vitro: Structure-Activity Relationships. RSC Adv. 2015, 5, 33058–33066. [Google Scholar] [CrossRef]
- Brattain, M. G.; Fine, W. D.; Khaled, F. M.; Thompson, J.; Brattain, D. E. Heterogeneity of Malignant Cells from a Human Colonic Carcinoma. Cancer Res. 1981, 41, 1751–1756. [Google Scholar]
- Semple, T. U.; Quinn, L. A.; Woods, L. K.; Moore, G. E. Tumor and Lymphoid Cell Lines from a Patient with Carcinoma of the Colon for a Cytotoxicity Model. Cancer Res. 1978, 38, 1345–1355. [Google Scholar]
- Kunkel, M. W.; Coussens, N. P.; Morris, J.; Taylor, R. C.; Dexheimer, T. S.; Jones, E. M.; Doroshow, J. H.; Teicher, B. A. HTS384 NCI60: the Next Phase of the NCI60 Screen. Cancer Res. 2024, 84, 2403–2416. [Google Scholar] [CrossRef]
- Chen, T. R.; Dorotinsky, C. S.; McGuire, L. J.; Macy, M. L.; Hay, R. J. DLD-1 and HCT-15 Cell Lines Derived Separately from Colorectal Carcinomas Have Totally Different Chromosome Changes but the Same Genetic Origin. Cancer Genet. Cytogenet. 1995, 81, 103–108. [Google Scholar] [CrossRef]
- Marshall, C. J.; Franks, L. M.; Carbonell, A. W. Markers of Neoplastic Transformation in Epithelial Cell Lines Derived from Human Carcinomas. J. Natl. Cancer Inst. 1977, 58, 1743–1751. [Google Scholar] [PubMed]
- Camps, J.; Morales, C.; Prat, E.; Ribas, M.; Capellà, G.; Egozcue, J.; Peinado, M. A.; Miró, R. Genetic Evolution in Colon Cancer KM12 Cells and Metastatic Derivates. Int. J. Cancer 2004, 110, 869–874. [Google Scholar] [CrossRef] [PubMed]
- Stragand, J. J.; Barlogie, B.; White, R. A.; Drewinko, B. Biological Properties of the Human Colonic Adenocarcinoma Cell Line SW 620 Grown as a Xenograft in the Athymic Mouse. Cancer Res. 1981, 41, 3364–3369. [Google Scholar] [PubMed]
- Boland, C. R.; Goel, A. Microsatellite Instability in Colorectal Cancer. Gastroenterology 2010, 138, 2073–2087. [Google Scholar] [CrossRef]
- Foley, G. E.; Lazarus, H.; Farber, S.; Uzman, B. G.; Boone, B. A.; McCarthy, R. E. Continuous Culture of Human Lymphoblasts from Peripheral Blood of a Child with Acute Leukemia. Cancers 1965, 18, 522–529. [Google Scholar]
- Birnie, G. D. The HL60 Cell Line: A Model System for Studying Human Myeloid Cell Differentiation. Br. J. Cancer 1988, 58 Suppl. IX, 41–45. [Google Scholar]
- Greenberg, J. M.; Gonzalez-Sarmiento, R.; Arthur, D. C.; Wilkowski, C. W.; Streifel, B. J.; Kersey, J. H. Immunophenotypic and Cytogenetic Analysis of Molt-3 and Molt-4: Human T-Lymphoid Cell Lines with Rearrangement of Chromosome 7. Blood 1988, 72, 1755–1760. [Google Scholar] [CrossRef]
- Matsuoka, Y.; Moore, G. E.; Yagi, Y.; Pressman, D. Production of Free Light Chains of Immunoglobulin by a Hematopoietic Cell Line Derived from a Patient with Multiple Myeloma. Proc. Soc. Exp. Biol. Med. 1967, 125, 1246–1250. [Google Scholar]
- Yang, S.; Damiano, M. G.; Zhang, H.; Tripathy, S.; Luthi, A. J.; Rink, J. S.; Ugolkov, A. V.; Singh, A. T. K.; Dave, S. S.; Gordon, L. I.; Thaxton, C. S. Biomimetic, Synthetic HDL Nanostructures for Lymphoma. Proc. Natl. Acad. Sci. 2013, 110, 2511–2516. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Dai, Z.; Sadee, W.; Hancock, W. S. A Pharmacoproteomics Study of the Cancer Cell Line EKVX Using Capillary-LC/MS/MS. Mol. Pharm. 2006, 3, 566–578. [Google Scholar] [CrossRef] [PubMed]
- Monks, A.; Scudiero, D.; Skehan, P.; Shoemaker, R.; Paull, K.; Vistica, D.; Hose, C.; Langley, J.; Cronise, P.; Vaigro-Wolff, A.; et al. Feasibility of a High-Flux Anticancer Drug Screen Using a Diverse Panel of Cultured Human Tumor Cell Lines. J. Natl. Cancer Inst. 1991, 83, 757–766. [Google Scholar] [PubMed]
- Zhang, H.; Li, N.; Chen, Y.; Huang, L.-Y.; Wang, Y.-C.; Fang, G.; He, D.-C.; Xiao, X.-Y. Protein Profile of Human Lung Squamous Carcinoma Cell Line NCI-H226. Biomed. Eng. Sci. 2007, 20, 24–32. [Google Scholar]
- Shi, Y.; Fu, X.; Hua, Y.; Han, Y.; Lu, Y.; Wang, J. The Side Population in Human Lung Cancer Cell Line NCI-H460 Is Enriched in Stem-Like Cancer Cells. PLoS One 2012, 7, e33358. [Google Scholar]
- Gazdar, A. F.; Minna, J. D. NCI Series of Cell Lines: An Historical Perspective. J. Cellular Biochem. Suppl. 1996, 24, 1–11. [Google Scholar]
- Rutka, J. T.; Giblin, J. R.; Høifødt, H. K.; Dougherty, D. V.; Bell, C. W.; McCulloch, J. R.; Davis, R. L.; Wilson, C. B.; Rosenblum, M. L. Establishment and Characterization of a Cell Line from a Human Gliosarcoma. Cancer Research 1986, 46, 5893–5902. [Google Scholar]
- Torsvik, A.; Stieber, D.; Enger, P. Ø.; Golebiewska, A.; Molven, A.; Svendsen, A.; Westermark, B.; Niclou, S. P.; Olsen, T. K.; Enger, M. C.; Bjerkvig, R. U-251 Revisited: Genetic Drift and Phenotypic Consequences of Long-Term Cultures of Glioblastoma Cells. Cancer Med. 2014, 3, 812–824. [Google Scholar] [CrossRef]
- Fogh, J.; Fogh, J. M.; Orfeo, T. One Hundred and Twenty-Seven Cultured Human Tumor Cell Lines Producing Tumors in Nude Mice. J. Natl. Cancer Inst. 1977, 59, 221–226. [Google Scholar] [CrossRef]
- Sustarsic, E. G.; Junnila, R. K.; Kopchick, J. J. Human Metastatic Melanoma Cell Lines Express High Levels of Growth Hormone Receptor and Respond to GH Treatment. Biochem. Biophys. Res. Commun. 2013, 441, 144–150. [Google Scholar] [CrossRef]
- Bénard, J.; Da Silva, J.; De Blois, M.-C.; Boyer, P.; Duvillard, P.; Chiric, E.; Riou, G. Characterization of a Human Ovarian Adenocarcinoma Line, IGROV1, in Tissue Culture and in Nude Mice. Cancer Res. 1985, 45, 4970–4979. [Google Scholar]
- Hamilton, T. C.; Young, R. C.; McKoy, W. M.; Grotzinger, K. R.; Green, J. A.; Chu, E. W.; Whang-Peng, J.; Rogan, A. M.; Green, W. R.; Ozols, R. F. Characterization of Human Ovarian Carcinoma Cell Line (NIH:OVCAR-3) with Androgen and Estrogen Receptors. Cancer Res. 1983, 43, 5379–5389. [Google Scholar] [PubMed]
- Nunes, M.; Silva, P. M. A.; Coelho, R.; Pinto, C.; Resende, A.; Bousbaa, H.; Almeida, G. M.; Ricardo, S. Generation of Two Paclitaxel-Resistant High-Grade Serous Carcinoma Cell Lines with Increased Expression of P-Glycoprotein. Front. Oncol. 2021, 11, 752127. [Google Scholar]
- Kaighn, M. E.; Narayan, K. S.; Ohnuki, Y.; Lechner, J. F.; Jones, L. W. Establishment and Characterization of a Human Prostatic Carcinoma Cell Line (PC-3). Invest. Urol. 1979, 17, 16–23. [Google Scholar] [PubMed]
- Neve, R. M.; Chin, K.; Fridlyand, J.; Yeh, J.; Baehner, F. L.; Fevr, T.; Clark, L.; Bayani, N.; Coppe, J.-P.; Tong, F.; Speed, T.; Spellman, P. T.; DeVries, S.; Lapuk, A.; Wang, N. J.; Kuo, W.-L.; Stilwell, J. L.; Pinkel, D.; Albertson, D. G.; Waldman, F. M.; McCormick, F.; Dickson, R. B.; Johnson, M. D.; Lippman, M.; Ethier, S.; Gazdar, A.; Gray, J. W. A Collection of Breast Cancer Cell Lines for the Study of Functionally Distinct Cancer Subtypes. Cancer Cell 2006, 10, 515–527. [Google Scholar] [CrossRef] [PubMed]
- Koi, M.; Umar, A.; Chauhan, D. P.; Cherian, S. P.; Carethers, J. M.; Kunkel, T. A.; Boland, C. R. Human Chromosome 3 Corrects Mismatch Repair Deficiency and Microsatellite Instability and Reduces N-Methyl-N’-nitro-N-nitrosoguanidine Tolerance in Colon Tumor Cells with Homozygous hMLH1 Mutation. Cancer Res. 1994, 54, 4308–4312. [Google Scholar]
- Mitra, A. K.; Davis, D. A.; Tomar, S.; Roy, L.; Gurler, H.; Xie, J.; Lantvit, D. D.; Cardenas, H.; Fang, F.; Liu, Y.; Loughran, E.; Yang, J.; Stack, M. S.; Emerson, R. E.; Cowden Dahl, K. D.; Barbolina, M. V.; Nephew, K. P.; Matei, D.; Burdette, J. E. In Vivo Tumor Growth of High-Grade Serous Ovarian Cancer Cell Lines. Gynecol. Oncol. 2015, 138, 372–377. [Google Scholar] [CrossRef]
- Vert, A.; Castro, J.; Ribó, M.; Vilanova, M.; Benito, A. Transcriptional Profiling of NCI/ADR-RES Cells Unveils a Complex Network of Signaling Pathways and Molecular Mechanisms of Drug Resistance. OncoTarg. Ther. 2018, 11, 221–237. [Google Scholar]
- Nakatsuka, E.; Tan, L.; Cunneen, B.; Foster, C.; Lei, Y. L.; McLean, K. Characterization of DNA Damage Repair Pathway Utilization in High-Grade Serous Ovarian Cancers Yields Rational Therapeutic Approaches. Transl. Oncol. 2024, 50, 102119. [Google Scholar] [CrossRef] [PubMed]
- Meijer, T. G.; Martens, J. W. M.; Prager-van der Smissen, W. J. C.; Verkaik, N. S.; Beaufort, C. M.; van Herk, S.; Robert-Finestra, T.; Hoogenboezem, R. M.; Ruigrok-Ritstier, K.; Paul, M. W.; Gribnau, J.; Bindels, E. M. J.; Kanaar, R.; Jager, A.; van Gent, D. C.; Hollestelle, A. Functional Homologous Recombination (HR) Screening Shows the Majority of BRCA1/2-Mutant Breast and Ovarian Cancer Cell Lines Are HR-Proficient. Cancers 2024, 16, 741–758. [Google Scholar] [CrossRef] [PubMed]

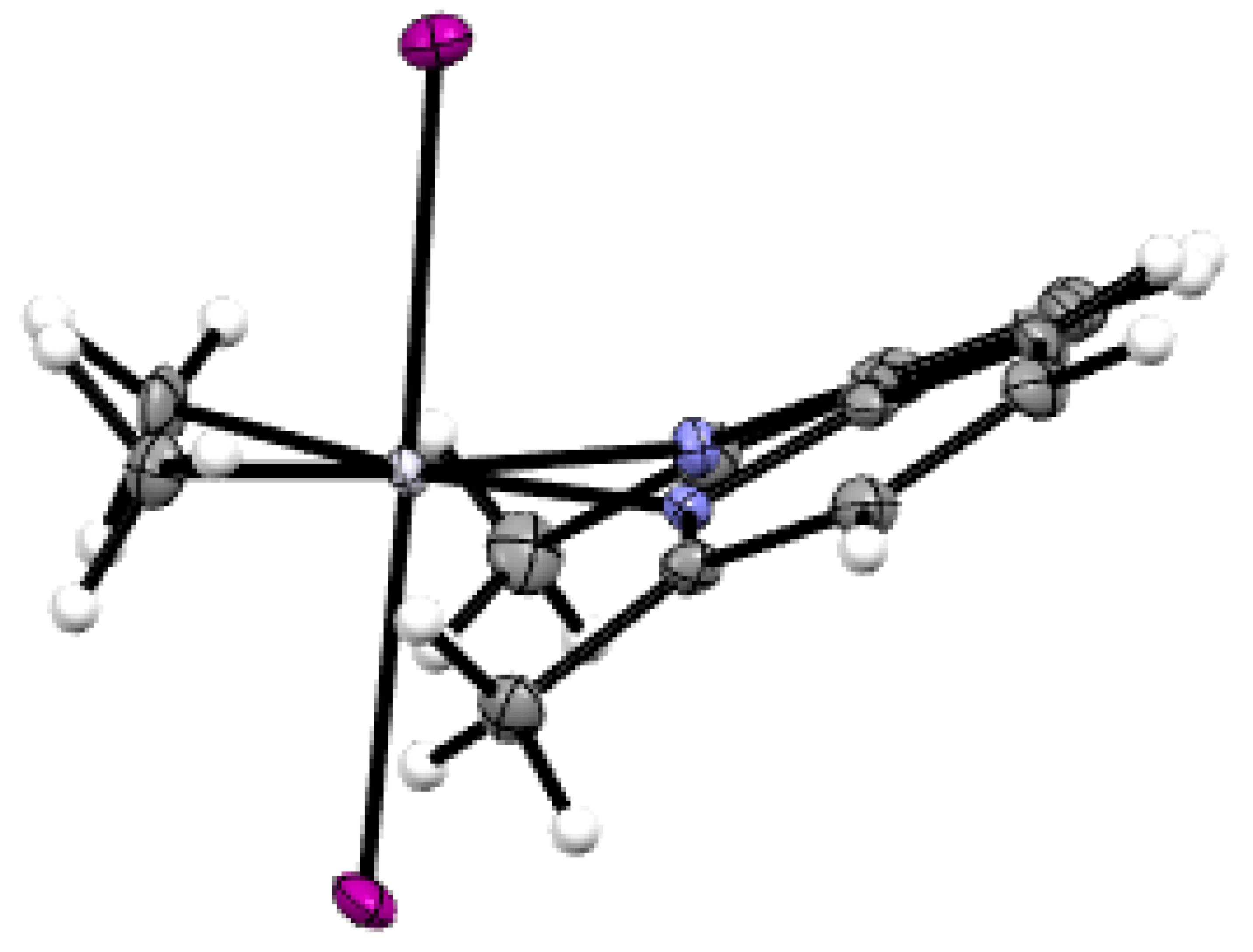
| Compound Number | 5 | 6 |
| Chemical formula | C14H18I2N2Pt | C14H18I2N2Pt |
| molecular weight, g mol-1 | 663.19 | 663.19 |
| temperature, K | 100(2) | 100(2) |
| wavelength, Å | 0.71073 | 0.71073 |
| lattice | orthorhombic | orthorhombic |
| space group | Pbca | Pbca |
| cell constants | ||
| a, Å | 12.7995(8) | 14.7706(10) |
| b, Å | 14.4289(10) | 13.6001(10) |
| c, Å | 18.0758(11) | 16.4126(12) |
| α, deg. | 90 | 90 |
| β, deg. | 90 | 90 |
| γ, deg. | 90 | 90 |
| volume, Å3 | 3338.3(4) | 3297.0(4) |
| Z | 8 | 8 |
| ρ(calc.) g cm-3 | 2.639 | 2.672 |
| absorption coefficient, mm-1 | 12.095 | 12.247 |
| F(000) | 2400 | 2400 |
| crystal size, mm3 | 0.266 x 0.138 x 0.055 | 0.231 x 0.177 x 0.134 |
| θrange | 2.253 to 30.526° | 2.384 to 30.564° |
|
index ranges |
– 18 ≤ h ≤ +18 – 20 ≤ k ≤ +20 – 23 ≤ l ≤ +25 |
– 19 ≤ h ≤ +21 – 19 ≤ k ≤ +19 – 23 ≤ l ≤ +23 |
| reflections collected | 111,168 | 60,295 |
| independent reflections | 5101 [Rint = 0.0626] | 5039 [Rint = 0.0699] |
| coverage, independent reflections | 99.9% | 99.9% |
| Absorption correction | Multi-scan | Multi-scan |
| max. & min. transmission | 0.746 and 0.375 | 0.7461 and 0.2833 |
| refinement method | Full matrix least squares on F2 | Full matrix least squares on F2 |
| data/restraints/parameters | 5101/8/186 | 5039/0/176 |
| goodness-of-fit on F2 | 1.146 | 1.117 |
| Final R indices [I > 2σ(I)] | R1 = 0.0208 wR2 = 0.0475 | R1 = 0.0363 wR2 = 0.0917 |
| Rindices (all data) | R1 = 0.0245 wR2 = 0.0486 | R1 = 0.0398 wR2 = 0.0942 |
| largest difference peak and hole | 1.252 & – 1.723 e Å-3 | 2.542 & – 4.484 e Å-3 |
| Compound Number | 5 | 6 |
| Bond Lengths (Å) | ||
| Pt – I(1) | 2.6500(3) | 2.6453(4) |
| Pt – I(2) | 2.6429(3) | 2.6634(4) |
| Pt – I(3) | 2.539(8) | |
| Pt – C(13) | 2.064(3) | 2.063(5) |
| Pt – C(14) | 2.065(5) | 2.058(5) |
| Pt – C(14A) | 2.065(5) | |
| Pt – N(1) | 2.154(3) | 2.200(4) |
| Pt – N(2) | 2.163(2) | 2.218(4) |
| Bond Angles (deg.) | ||
| I(1) – Pt – I(2) | 178.905(8) | 177.727(13) |
| C(13) – Pt – N(2) | 174.38(12) | 173.95(19) |
| C(14) – Pt – N(1) | 176.15(16) | 173.93(19) |
| I(1) – Pt – C(13) | 90.38(11) | 86.60(16) |
| I(1) – Pt – C(14) | 89.26(15) | 87.90(15) |
| I(1) – Pt – N(1) | 90.29(7) | 87.77(11) |
| I(1) – Pt – N(2) | 91.81(7) | 88.36(10) |
| C(13) – Pt – C(14) | 85.83(17) | 83.8(2) |
| C(13) – Pt – N(1) | 98.00(11) | 100.2(2) |
| C(14) – Pt – N(2) | 99.37(16) | 99.34(18) |
| N(1) – Pt – N(2) | 76.82(10) | 76.29(15) |
| Pt – N(1) – C(5) | 115.33(19) | 110.7(3) |
| Dihedral Angle (deg.) | ||
| N(1) – C(5) – C(6) – N(2) | – 2.5(4) | 5.8(6) |
| Pt – N(1) – C(1) – C(2) | – 178.6(2) | 162.0(4) |
| Cell Line, Reference | Type of Cancer | GI50, μM | TGI, μM | LC50, μM |
| CCRF-CEM,[42] | Leukemia | 5.42 (1.47) | 18.0 (> 100) | 42.5 (> 100) |
| HL-60(TB),[43] | Leukemia | 3.98 (4.12) | 16.0 (75.1) | 40.5 (> 100) |
| K-562,13 | Leukemia | 4.62 | 19.3 | 60.2 |
| MOLT-4,[44] | Leukemia | 5.53 (2.75) | 18.0 (> 100) | 42.8 (> 100) |
| RPMI-8226,[45] | Leukemia | 2.50 (6.44) | 11.2 (> 100) | 33.6 (> 100) |
| SR,[46] | Leukemia | 4.19 (0.496) | 16.2 (16.0) | 40.5 (> 100) |
| EKVX,[47] | Non-Small Cell Lung | 1.41 (6.54) | 4.52 (62.9) | 79.5 (> 100) |
| HOP-62,[48] | Non-Small Cell Lung | 3.40 (1.54) | 18.4 (13.2) | 82.2 (> 100) |
| NCI-H226,[49] | Non-Small Cell Lung | 10.1 (4.42) | 29.0 (25.6) | 83.1 (> 100) |
| NCI-H460,[50] | Non-Small Cell Lung | 8.75 (0.455) | 22.7 (78.2) | 53.4 (> 100) |
| NCI-H522,[51] | Non-Small Cell Lung | 3.05 | 16.5 | 41.2 |
| HCT-116,34 | Colon Cancer | 1.79 (9.24) | 3.83 (> 100) | 8.17 (> 100) |
| SF-539,[52] | Central Nervous System | 7.93 (0.600) | 23.6 (7.67) | 58.9 (> 100) |
| U251,[53] | Central Nervous System | 11.6 (1.57) | 26.0 (24.7) | 58.5 (> 100) |
| SK-MEL-2,[54] | Melanoma | 2.54 | 11.6 | 50.9 |
| UACC-62,[55] | Melanoma | 6.77 (1.34) | 22.9 (9.39) | 60.1 (37.2) |
| IGROV-1,[56] | Ovarian Cancer | 2.07 (1.70) | 5.10 (7.05) | 42.6 (> 100) |
| OVCAR-3,[57] | Ovarian Cancer | 1.69 (1.93) | 3.52 (4.27) | 7.34 |
| OVCAR-8,[58] | Ovarian Cancer | 12.3 (4.09) | 29.1 (> 100) | 68.7 (> 100) |
| PC-3,[59] | Prostate Cancer | 2.53 (4.08) | 7.82 (> 100) | 73.0 (> 100) |
| MCF-7,9,10 | Breast Cancer | 4.24 (2.66) | 19.5 (79.3) | 91.5 (> 100) |
| BT-549,[60] | Breast Cancer | 13.7 (3.36) | 32.5 (44.9) | 77.0 (> 100) |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




