Submitted:
17 March 2026
Posted:
17 March 2026
You are already at the latest version
Abstract
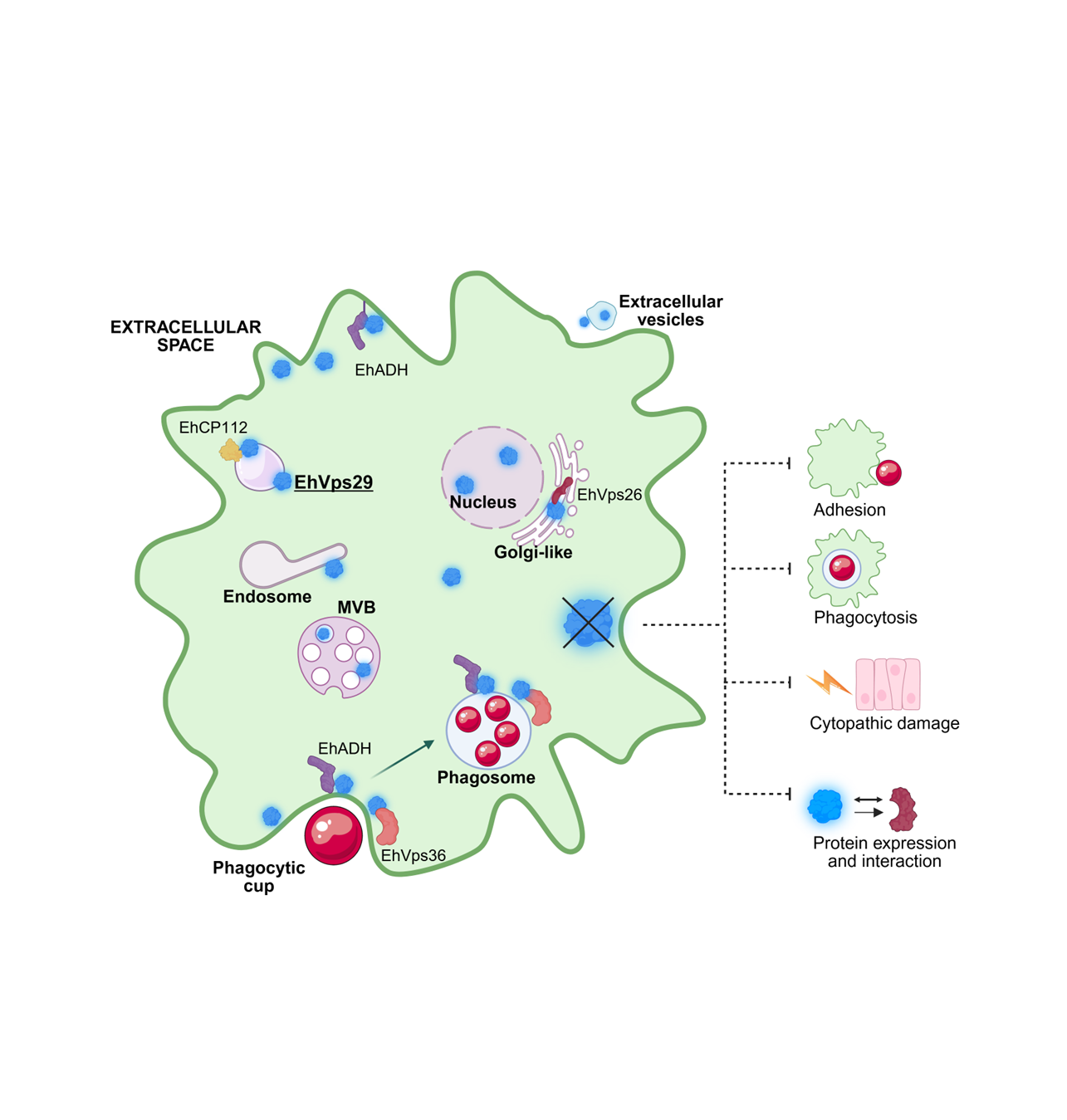
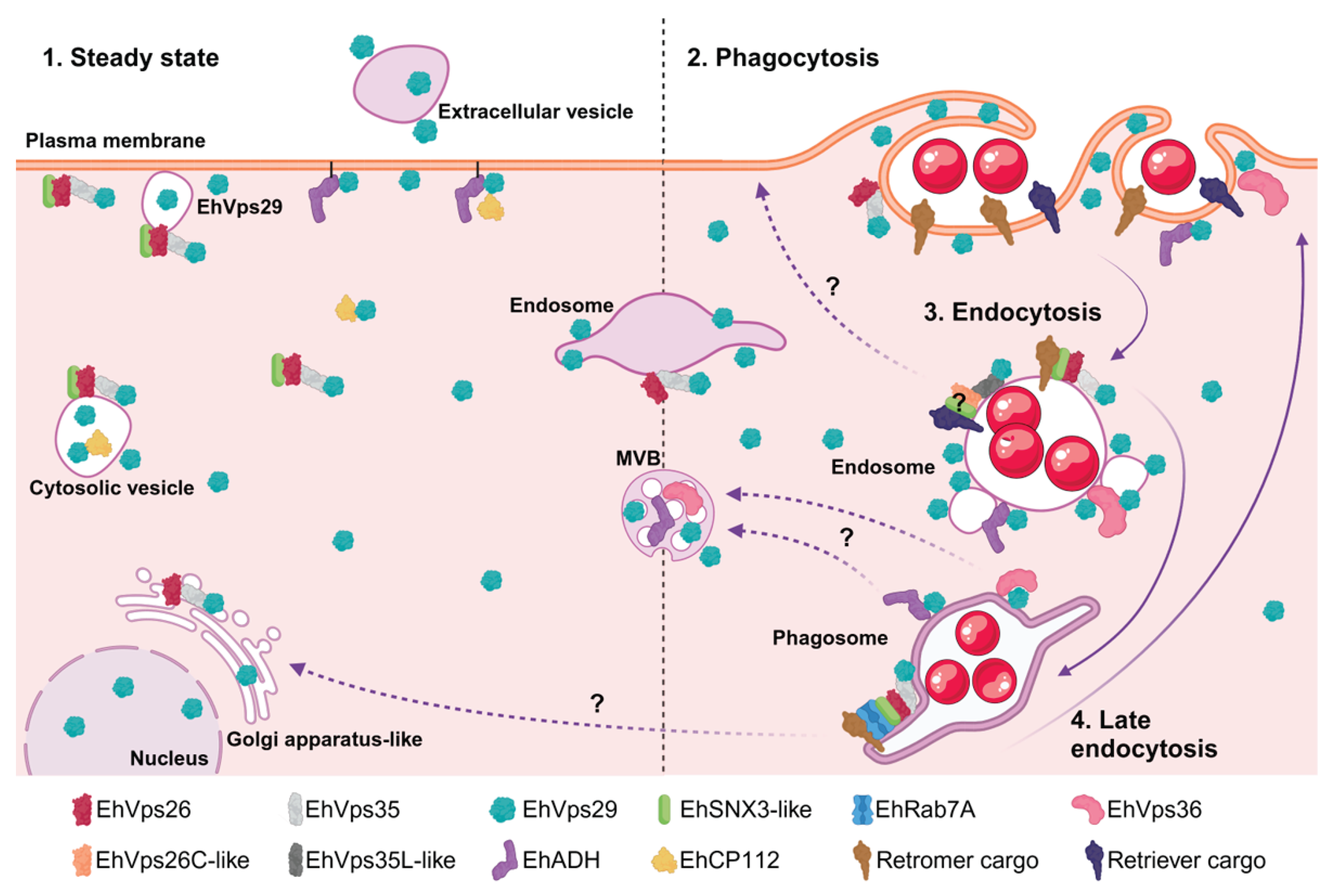
The retromer is a highly conserved complex that mediates the trafficking of cargo proteins to plasma membrane or trans-Golgi network. In pathogenic microorganisms, retromer-dependent transport contributes to the delivery of virulence factors and promotes infection. The retromer consists of a sorting nexin dimer (SNX) and a cargo-selection complex (CSC), formed by Vps26, Vps35, and Vps29. In Entamoeba histolytica, the parasite causative of human amoebiasis, the retromer functions as a Rab7A GTPase effector and participates in phagocytosis and cytotoxicity. Although we previously characterized the roles of EhVps26 and EhVps35, the function of EhVps29 remained unclear. In this study, we analyzed the subcellular localization and functional role of EhVps29 in adhesion, phagocytosis, and cytopathic effect. EhVps29 localized to the plasma membrane, cytosol, vesicles, tubules, Golgi-like structures, MVBs and, for the first time, in the nucleus. Immunofluorescence and western blot assays demonstrated that EhVps29 modulates the localization of the EhVps26, EhADH adhesin and EhCP112 cysteine protease. The Ehvps29 gene silencing and overexpression confirmed its involvement in virulence-associated processes. Immunoprecipitation and confocal microscopy results showed the interaction among EhVps29, EhVps36 and EhADH ESCRT machinery members. Our results indicate that EhVps29 is involved in parasite virulence and protein trafficking through recycling or degradation pathways.
Keywords:
1. Introduction
2. Materials and Methods
2.1. Plasmids Construction and Recombinant Protein Obtaining
2.2. Institutional Review Board Statement
2.3. α-EhVps29 Polyclonal Antibodies Generation
2.4. E. Histolytica Cultures
2.5. Western Blot Assays
2.6. Confocal Microscopy
2.7. Transmission Electron Microscopy (TEM)
2.8. Ehvps29 Knock Down Trophozoites
2.9. Overexpression of the Ehvps29 Gene in Trophozoites
2.10. Adhesion Assays
2.11. Phagocytosis Assays
2.12. Cytopathic Effect of Trophozoites on Cell Monolayers
2.13. Immunoprecipitation
2.14. Data Analysis and Statistical Methods
2.15. Tridimensional Models
2.16. Molecular Docking
3. Results
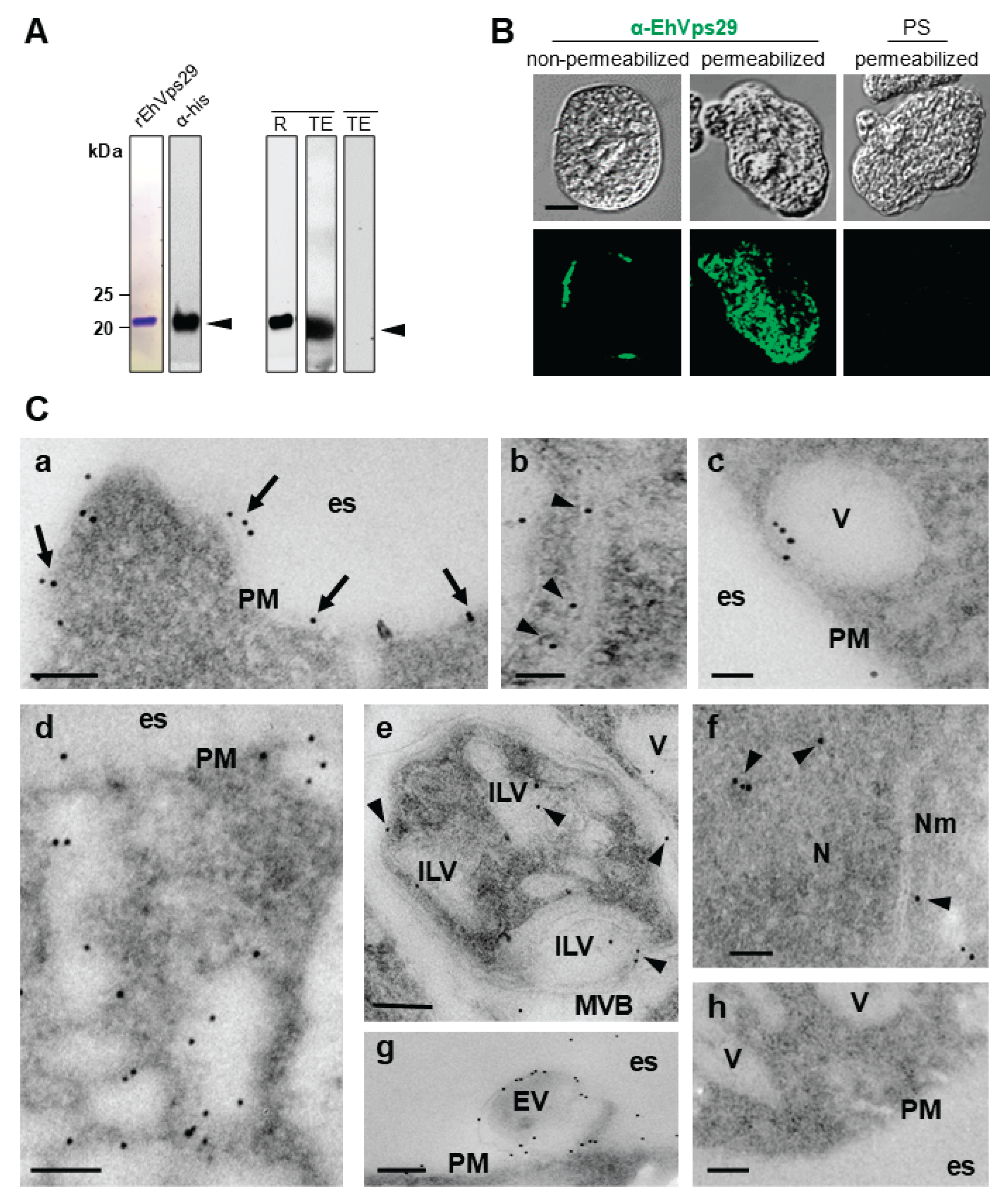
3.1. EhVps29 Is Present at Trophozoites Plasma Membrane, Cytosol, Vesicles, Tubular Structures, Extracellular Vesicles and Nucleus
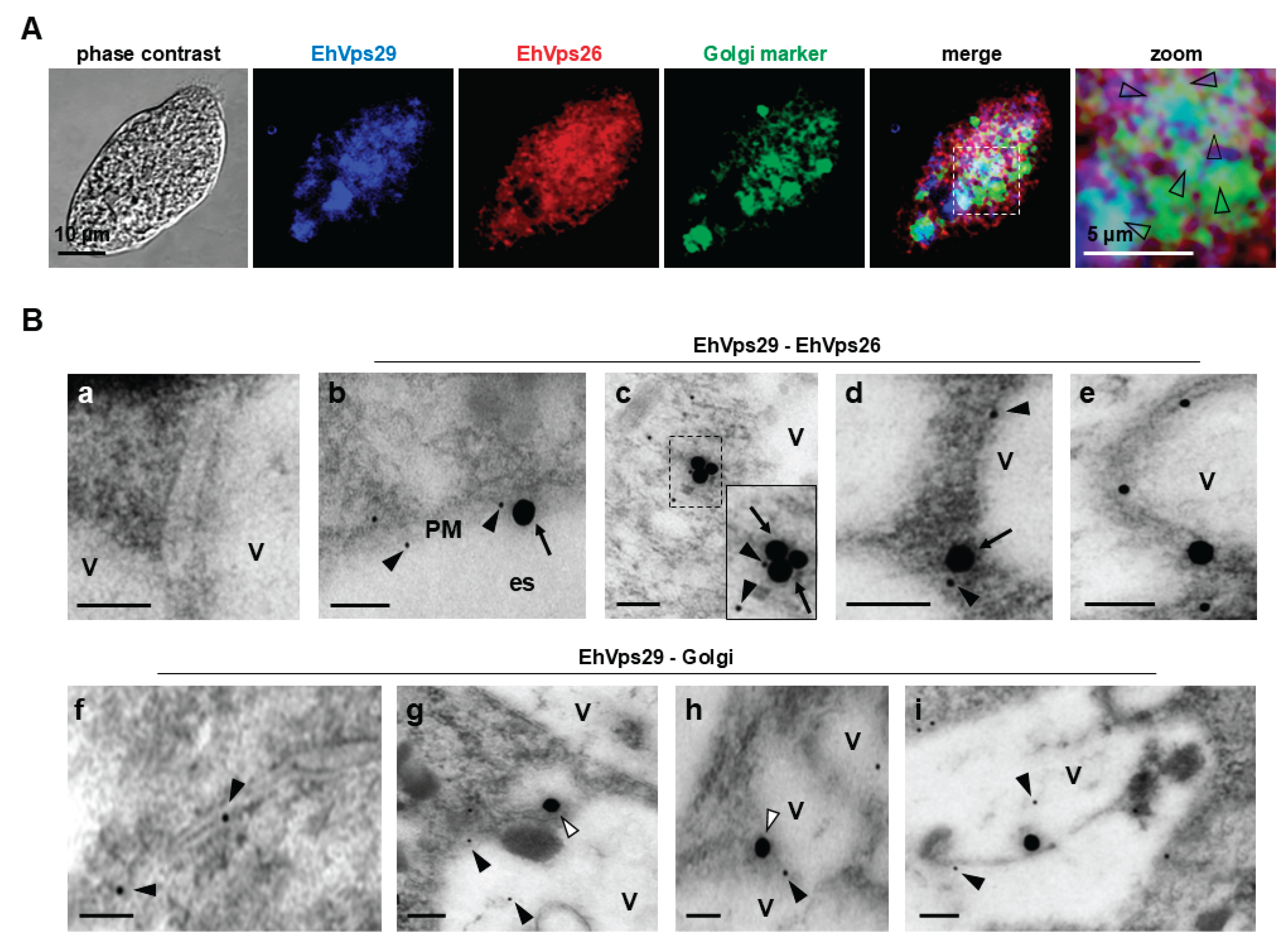
3.2. EhVps29 is in Golgi-like Structures
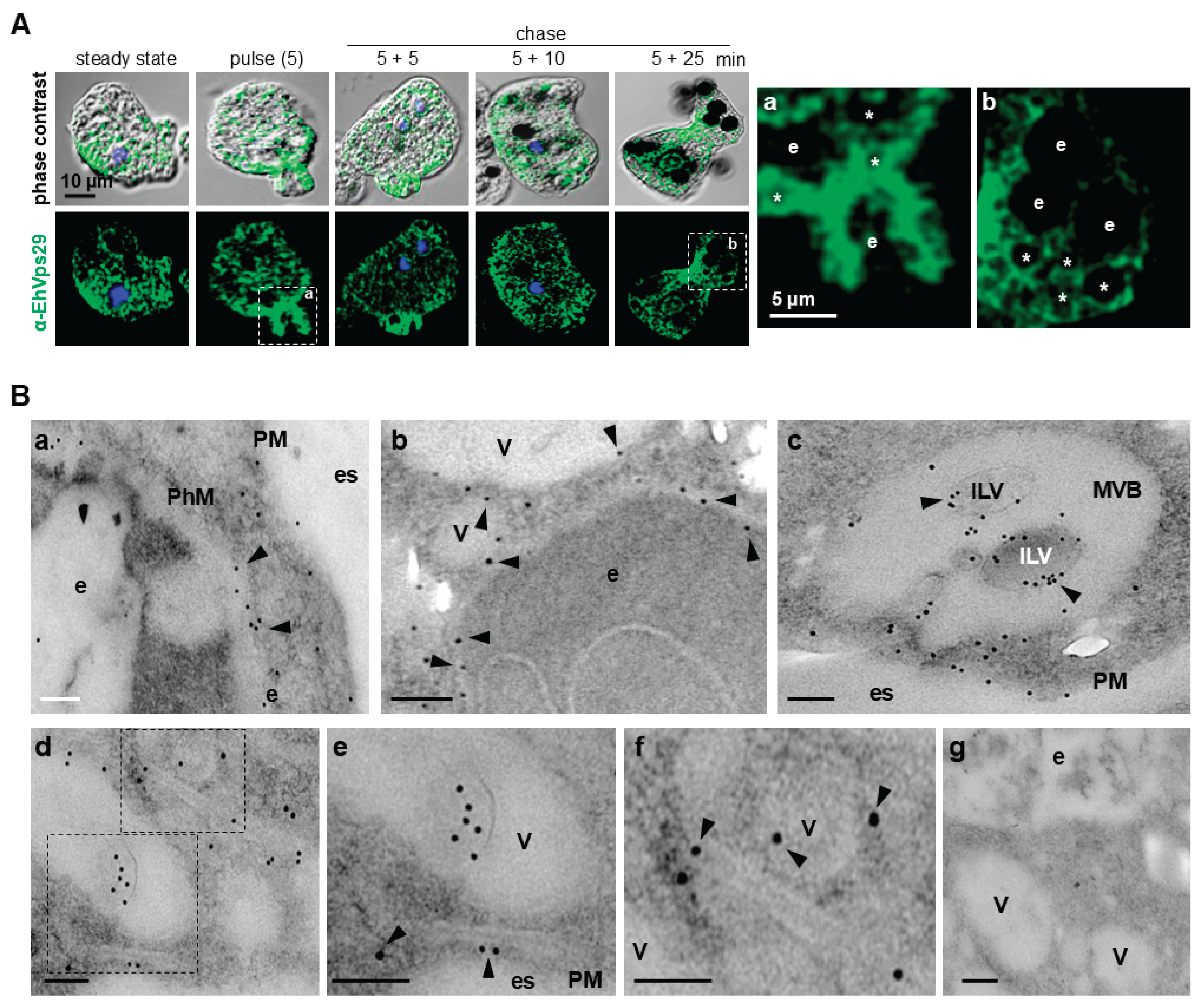
3.3. During Phagocytosis, EhVps29 Is Mobilized to Phagocytic Cups, Phagosomal Membranes, and Double-Membrane Tubular Structures
3.4. EhVps29 and ESCRT Components Interact During Phagocytosis
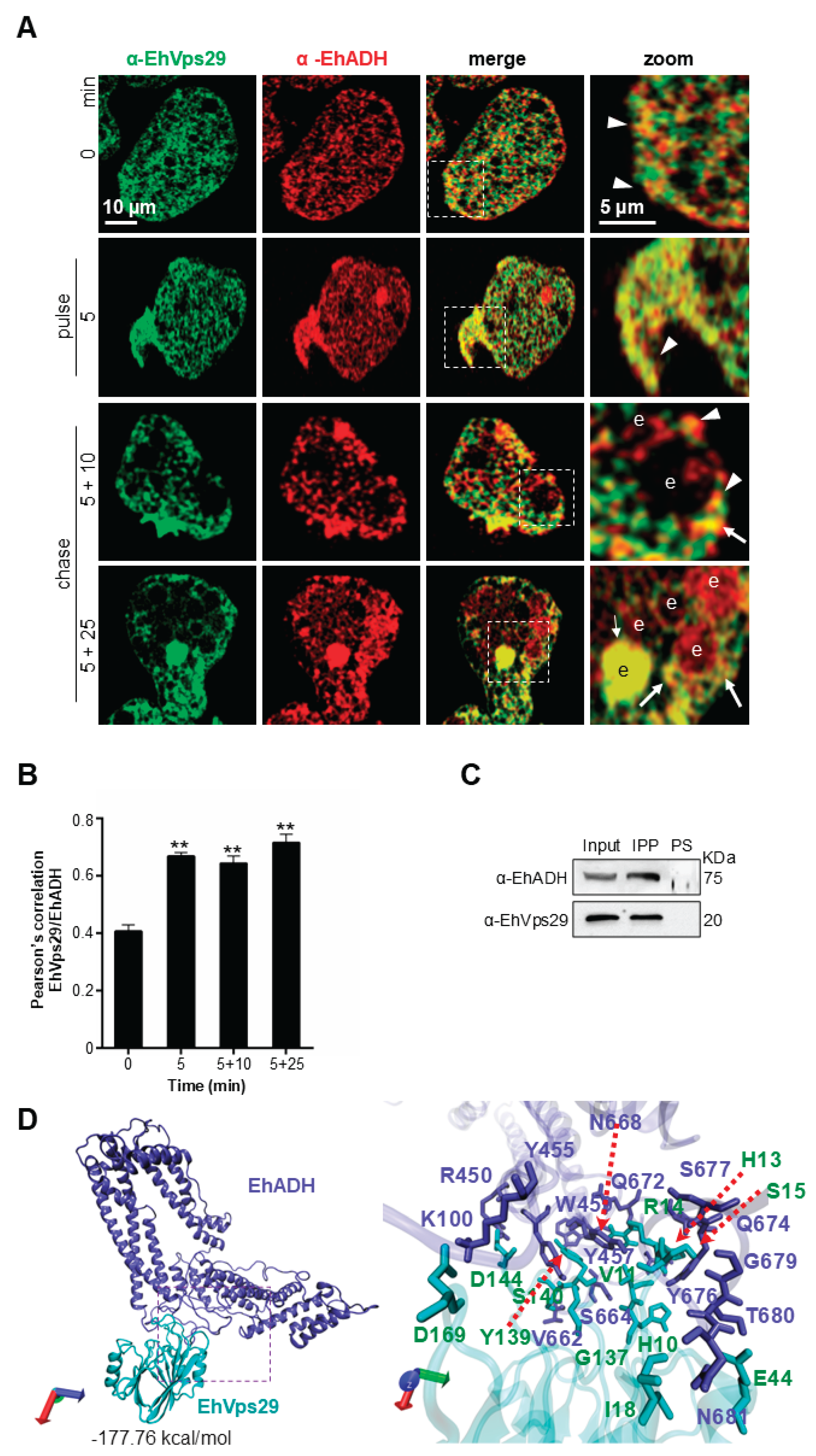
3.5. EhVps29 Interacts with the EhADH Adhesin
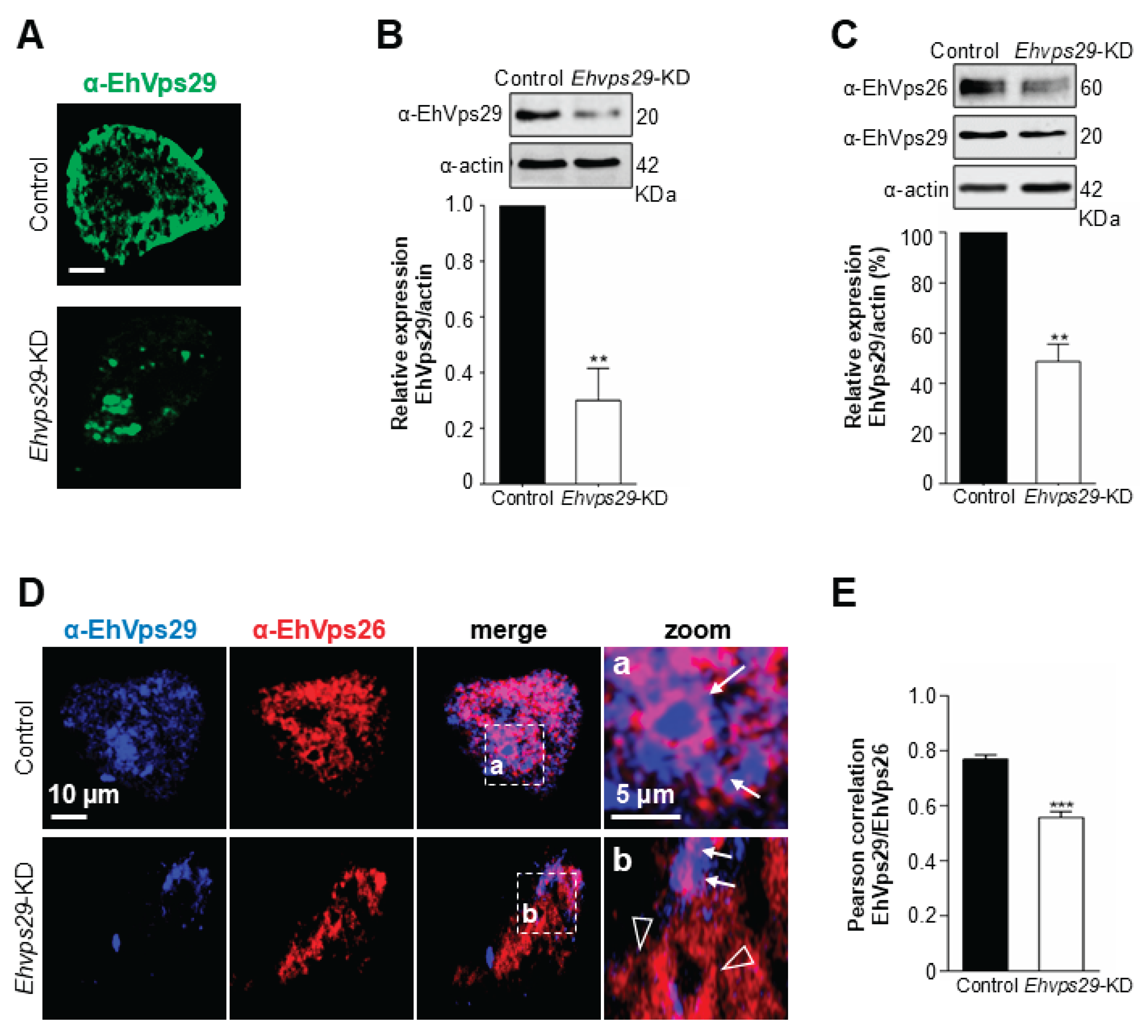
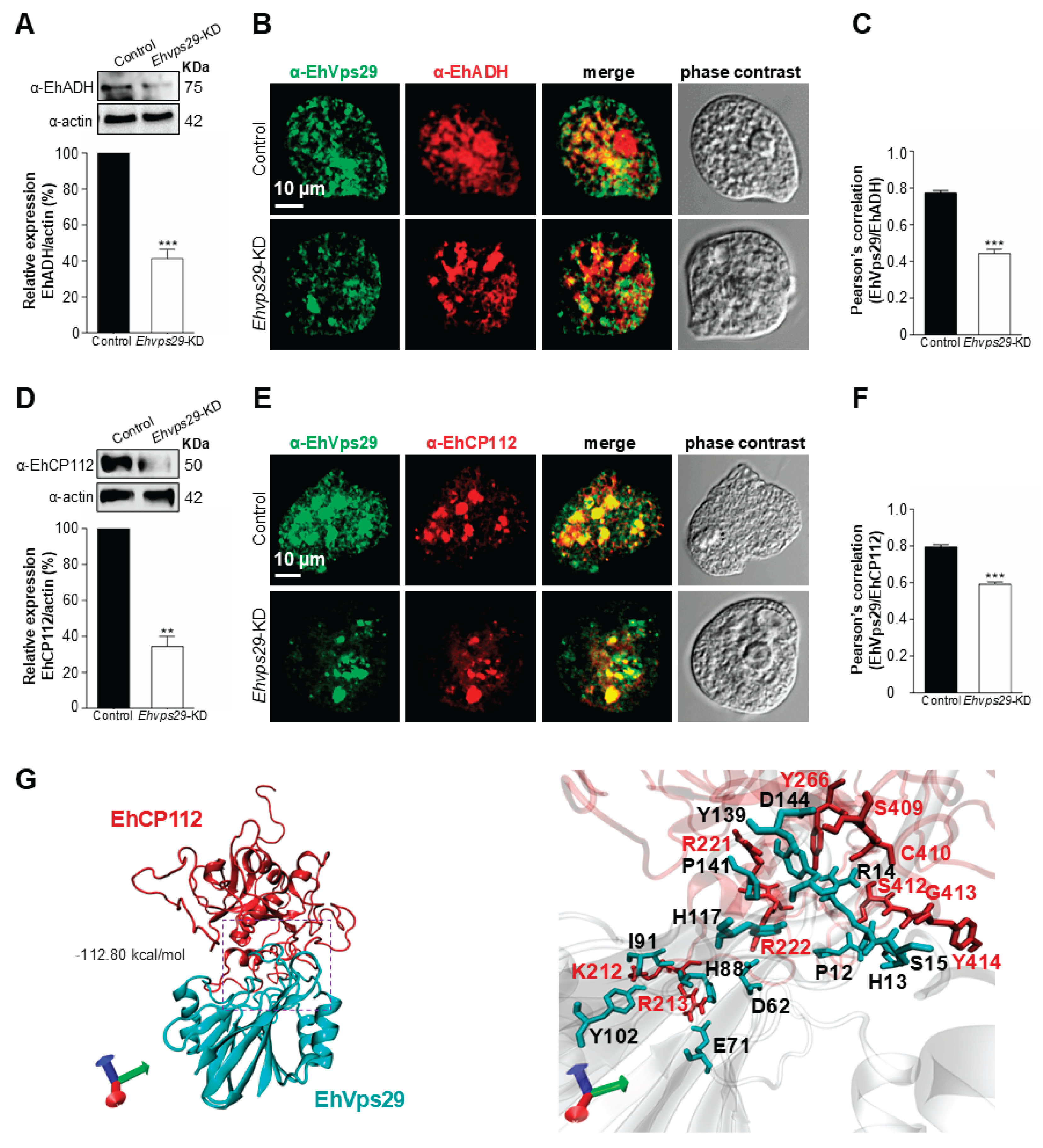
3.6. The Ehvps29 Knock Down Affects EhVps26
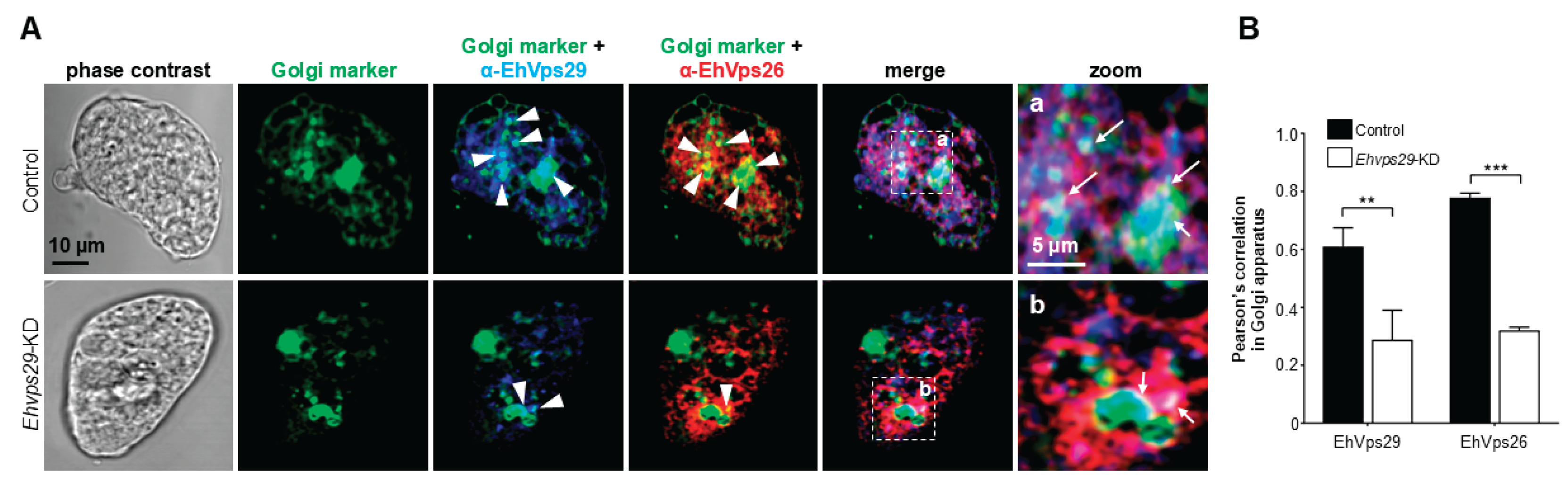
3.7. Ehvps29-KD Trophozoites Display EhVps26 and EhVps29 Mislocalization in Golgi-like Structures
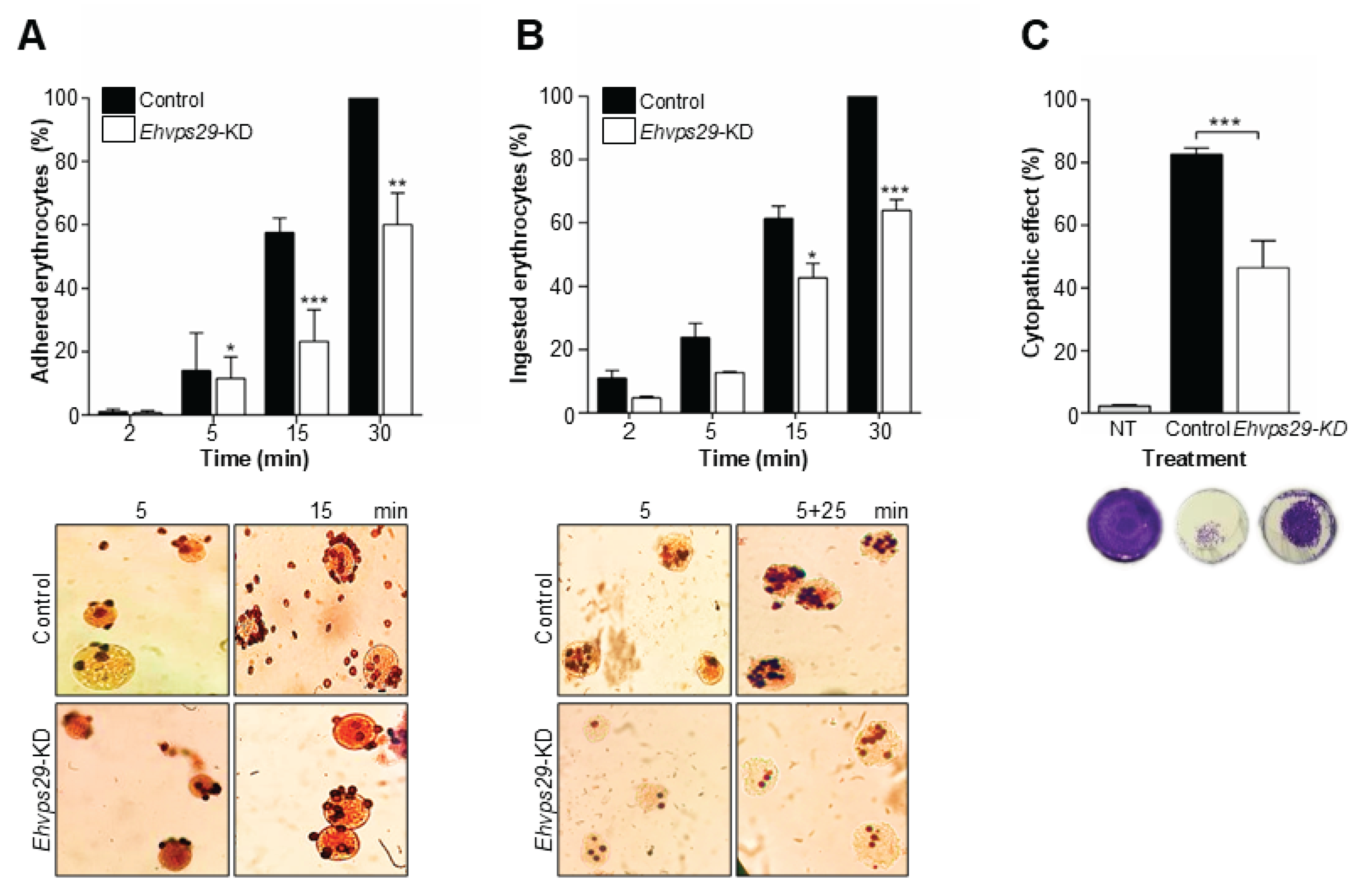
3.8. Ehvps29 Knock Down Affects Trophozoites Adhesion, Phagocytosis, and Cytopathic Effect
3.9. EhVps29 Interacts with EhCP112 Virulence Factor
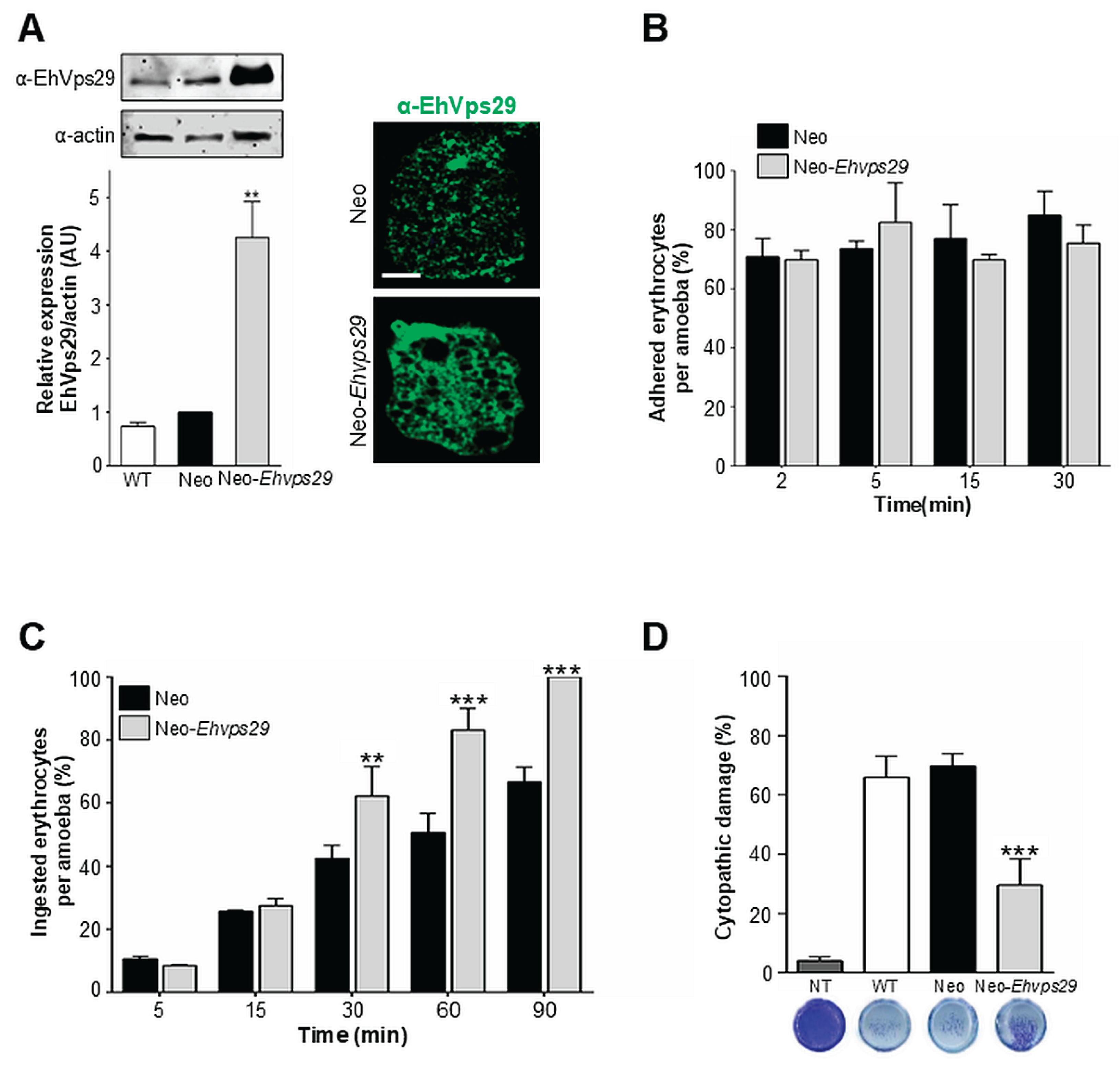
3.10. Ehvps29 Overexpression Does not Alter Rates of Adhesion, Improves Phagocytosis and Decreases the Cytopathic Effect
4. Discussion
Author Contributions
Funding
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Redpath, G. M. I.; Betzler, V. M.; Rossatti, P.; Rossy, J. Membrane Heterogeneity Controls Cellular Endocytic Trafficking. Front. Cell Dev. Biol. 2020, 8, 757. [Google Scholar] [CrossRef]
- Harterink, M.; Port, F.; Lorenowicz, M. J.; McGough, I. J.; Silhankova, M.; Betist, M. C.; Van Weering, J. R. T.; Van Heesbeen, R. G. H. P.; Middelkoop, T. C.; Basler, K.; Cullen, P. J.; Korswagen, H. C. A SNX3-Dependent Retromer Pathway Mediates Retrograde Transport of the Wnt Sorting Receptor Wntless and Is Required for Wnt Secretion. Nat Cell Biol 2011, 13(8), 914–923. [Google Scholar] [CrossRef]
- Seaman, M. N. J.; Michael McCaffery, J.; Emr, S. D. A Membrane Coat Complex Essential for Endosome-to-Golgi Retrograde Transport in Yeast. The Journal of Cell Biology 1998, 142(3), 665–681. [Google Scholar] [CrossRef]
- Fjorback, A. W.; Seaman, M.; Gustafsen, C.; Mehmedbasic, A.; Gokool, S.; Wu, C.; Militz, D.; Schmidt, V.; Madsen, P.; Nyengaard, J. R.; Willnow, T. E.; Christensen, E. I.; Mobley, W. B.; Nykjær, A.; Andersen, O. M. Retromer Binds the FANSHY Sorting Motif in SorLA to Regulate Amyloid Precursor Protein Sorting and Processing. J. Neurosci. 2012, 32(4), 1467–1480. [Google Scholar] [CrossRef] [PubMed]
- McGough, I. J.; De Groot, R. E. A.; Jellett, A. P.; Betist, M. C.; Varandas, K. C.; Danson, C. M.; Heesom, K. J.; Korswagen, H. C.; Cullen, P. J. SNX3-Retromer Requires an Evolutionary Conserved MON2:DOPEY2:ATP9A Complex to Mediate Wntless Sorting and Wnt Secretion. Nat Commun 2018, 9(1), 3737. [Google Scholar] [CrossRef] [PubMed]
- Zhang, D.; Isack, N. R.; Glodowski, D. R.; Liu, J.; Chen, C. C.-H.; Xu, X. Z. S.; Grant, B. D.; Rongo, C. RAB-6.2 and the Retromer Regulate Glutamate Receptor Recycling through a Retrograde Pathway. Journal of Cell Biology 2012, 196(1), 85–101. [Google Scholar] [CrossRef]
- Krai, P.; Dalal, S.; Klemba, M. Evidence for a Golgi-to-Endosome Protein Sorting Pathway in Plasmodium Falciparum. PLoS ONE 2014, 9(2), e89771. [Google Scholar] [CrossRef]
- Sloves, P.-J.; Delhaye, S.; Mouveaux, T.; Werkmeister, E.; Slomianny, C.; Hovasse, A.; Dilezitoko Alayi, T.; Callebaut, I.; Gaji, R. Y.; Schaeffer-Reiss, C.; Van Dorsselear, A.; Carruthers, V. B.; Tomavo, S. Toxoplasma Sortilin-like Receptor Regulates Protein Transport and Is Essential for Apical Secretory Organelle Biogenesis and Host Infection. Cell Host & Microbe 2012, 11(5), 515–527. [Google Scholar] [CrossRef]
- Tomavo, S.; Slomianny, C.; Meissner, M.; Carruthers, V. B. Protein Trafficking through the Endosomal System Prepares Intracellular Parasites for a Home Invasion. PLoS Pathog 2013, 9(10), e1003629. [Google Scholar] [CrossRef]
- Koumandou, V. L.; Klute, M. J.; Herman, E. K.; Nunez-Miguel, R.; Dacks, J. B.; Field, M. C. Evolutionary Reconstruction of the Retromer Complex and Its Function in Trypanosoma Brucei. Journal of Cell Science 2011, 124(9), 1496–1509. [Google Scholar] [CrossRef] [PubMed]
- Horazdovsky, B. F.; Davies, B. A.; Seaman, M. N.; McLaughlin, S. A.; Yoon, S.; Emr, S. D. A Sorting Nexin-1 Homologue, Vps5p, Forms a Complex with Vps17p and Is Required for Recycling the Vacuolar Protein-Sorting Receptor. MBoC 1997, 8(8), 1529–1541. [Google Scholar] [CrossRef]
- Van Weering, J. R. T.; Sessions, R. B.; Traer, C. J.; Kloer, D. P.; Bhatia, V. K.; Stamou, D.; Carlsson, S. R.; Hurley, J. H.; Cullen, P. J. Molecular Basis for SNX-BAR-Mediated Assembly of Distinct Endosomal Sorting Tubules: SNX-BAR-Mediated Formation of Distinct Sorting Tubules. The EMBO Journal 2012, 31(23), 4466–4480. [Google Scholar] [CrossRef]
- Wassmer, T.; Attar, N.; Harterink, M.; Van Weering, J. R. T.; Traer, C. J.; Oakley, J.; Goud, B.; Stephens, D. J.; Verkade, P.; Korswagen, H. C.; Cullen, P. J. The Retromer Coat Complex Coordinates Endosomal Sorting and Dynein-Mediated Transport, with Carrier Recognition by the Trans-Golgi Network. Developmental Cell 2009, 17(1), 110–122. [Google Scholar] [CrossRef]
- Carlton, J.; Bujny, M.; Peter, B. J.; Oorschot, V. M. J.; Rutherford, A.; Mellor, H.; Klumperman, J.; McMahon, H. T.; Cullen, P. J. Sorting Nexin-1 Mediates Tubular Endosome-to-TGN Transport through Coincidence Sensing of High- Curvature Membranes and 3-Phosphoinositides. Current Biology 2004, 14(20), 1791–1800. [Google Scholar] [CrossRef]
- Wassmer, T.; Attar, N.; Bujny, M. V.; Oakley, J.; Traer, C. J.; Cullen, P. J. A Loss-of-Function Screen Reveals SNX5 and SNX6 as Potential Components of the Mammalian Retromer. Journal of Cell Science 2007, 120(1), 45–54. [Google Scholar] [CrossRef]
- Griffin, C. T.; Trejo, J.; Magnuson, T. Genetic Evidence for a Mammalian Retromer Complex Containing Sorting Nexins 1 and 2. Proc. Natl. Acad. Sci. U.S.A. 2005, 102(42), 15173–15177. [Google Scholar] [CrossRef] [PubMed]
- Lucas, M.; Gershlick, D. C.; Vidaurrazaga, A.; Rojas, A. L.; Bonifacino, J. S.; Hierro, A. Structural Mechanism for Cargo Recognition by the Retromer Complex. Cell 2016, 167(6), 1623–1635.e14. [Google Scholar] [CrossRef]
- Suzuki, S. W.; Chuang, Y.-S.; Li, M.; Seaman, M. N. J.; Emr, S. D. A Bipartite Sorting Signal Ensures Specificity of Retromer Complex in Membrane Protein Recycling. Journal of Cell Biology 2019, 218(9), 2876–2886. [Google Scholar] [CrossRef] [PubMed]
- Makhoul, C.; Gosavi, P.; Duffield, R.; Delbridge, B.; Williamson, N. A.; Gleeson, P. A. Intersectin-1 Interacts with the Golgin GCC88 to Couple the Actin Network and Golgi Architecture. MBoC 2019, 30(3), 370–386. [Google Scholar] [CrossRef]
- Collins, B. M.; Skinner, C. F.; Watson, P. J.; Seaman, M. N. J.; Owen, D. J. Vps29 Has a Phosphoesterase Fold That Acts as a Protein Interaction Scaffold for Retromer Assembly. Nat Struct Mol Biol 2005, 12(7), 594–602. [Google Scholar] [CrossRef] [PubMed]
- Seaman, M. N. J.; Harbour, M. E.; Tattersall, D.; Read, E.; Bright, N. Membrane Recruitment of the Cargo-Selective Retromer Subcomplex Is Catalysed by the Small GTPase Rab7 and Inhibited by the Rab-GAP TBC1D5. Journal of Cell Science 2009, 122(14), 2371–2382. [Google Scholar] [CrossRef] [PubMed]
- Ye, H.; Ojelade, S. A.; Li-Kroeger, D.; Zuo, Z.; Wang, L.; Li, Y.; Gu, J. Y.; Tepass, U.; Rodal, A. A.; Bellen, H. J.; Shulman, J. M. Retromer Subunit, VPS29, Regulates Synaptic Transmission and Is Required for Endolysosomal Function in the Aging Brain. eLife 2020, 9, e51977. [Google Scholar] [CrossRef]
- Hesketh, G. G.; Pérez-Dorado, I.; Jackson, L. P.; Wartosch, L.; Schäfer, I. B.; Gray, S. R.; McCoy, A. J.; Zeldin, O. B.; Garman, E. F.; Harbour, M. E.; Evans, P. R.; Seaman, M. N. J.; Luzio, J. P.; Owen, D. J. VARP Is Recruited on to Endosomes by Direct Interaction with Retromer, Where Together They Function in Export to the Cell Surface. Developmental Cell 2014, 29(5), 591–606. [Google Scholar] [CrossRef]
- Zhang, X.; He, X.; Fu, X.-Y.; Chang, Z. Varp Is a Rab21 Guanine Nucleotide Exchange Factor and Regulates Endosome Dynamics. Journal of Cell Science 2006, 119(6), 1053–1062. [Google Scholar] [CrossRef]
- Kvainickas, A.; Jimenez-Orgaz, A.; Nägele, H.; Hu, Z.; Dengjel, J.; Steinberg, F. Cargo-Selective SNX-BAR Proteins Mediate Retromer Trimer Independent Retrograde Transport. Journal of Cell Biology 2017, 216(11), 3677–3693. [Google Scholar] [CrossRef]
- McNally, K. E.; Faulkner, R.; Steinberg, F.; Gallon, M.; Ghai, R.; Pim, D.; Langton, P.; Pearson, N.; Danson, C. M.; Nägele, H.; Morris, L. L.; Singla, A.; Overlee, B. L.; Heesom, K. J.; Sessions, R.; Banks, L.; Collins, B. M.; Berger, I.; Billadeau, D. D.; Burstein, E.; Cullen, P. J. Retriever Is a Multiprotein Complex for Retromer-Independent Endosomal Cargo Recycling. Nat Cell Biol 2017, 19(10), 1214–1225. [Google Scholar] [CrossRef] [PubMed]
- Nakada-Tsukui, K.; Saito-Nakano, Y.; Ali, V.; Nozaki, T. A Retromerlike Complex Is a Novel Rab7 Effector That Is Involved in the Transport of the Virulence Factor Cysteine Protease in the Enteric Protozoan Parasite Entamoeba Histolytica. MBoC 2005, 16(11), 5294–5303. [Google Scholar] [CrossRef] [PubMed]
- Srivastava, V. K.; Yadav, R.; Watanabe, N.; Tomar, P.; Mukherjee, M.; Gourinath, S.; Nakada-Tsukui, K.; Nozaki, T.; Datta, S. Structural and Thermodynamic Characterization of Metal Binding in Vps29 from Entamoeba Histolytica : Implication in Retromer Function. Molecular Microbiology 2017, 106(4), 562–581. [Google Scholar] [CrossRef]
- Martínez-Valencia, D.; Bañuelos, C.; García-Rivera, G.; Talamás-Lara, D.; Orozco, E. The Entamoeba Histolytica Vps26 (EhVps26) Retromeric Protein Is Involved in Phagocytosis: Bioinformatic and Experimental Approaches. PLoS ONE 2024, 19(8), e0304842. [Google Scholar] [CrossRef]
- Díaz-Valdez, J.; Javier-Reyna, R.; Montaño, S.; Talamás-Lara, D.; Orozco, E. EhVps35, a Retromer Component, Is Involved in the Recycling of the EhADH and Gal/GalNac Virulent Proteins of Entamoeba Histolytica. Front. Parasitol. 2024, 3, 1356601. [Google Scholar] [CrossRef]
- Hu, S.; Li, B.; Wu, F.; Zhu, D.; Zouhar, J.; Gao, C.; Shimada, T.; Rojo, E.; Hara-Nishimura, I.; Jiang, L.; Shen, J. Plant ESCRT Protein ALIX Coordinates with Retromer Complex in Regulating Receptor-Mediated Sorting of Soluble Vacuolar Proteins. Proc. Natl. Acad. Sci. U.S.A. 2022, 119(20), e2200492119. [Google Scholar] [CrossRef] [PubMed]
- Zhang, A.; He, X.; Zhang, L.; Yang, L.; Woodman, P.; Li, W. Biogenesis of Lysosome-Related Organelles Complex-1 Subunit 1 (BLOS1) Interacts with Sorting Nexin 2 and the Endosomal Sorting Complex Required for Transport-I (ESCRT-I) Component TSG101 to Mediate the Sorting of Epidermal Growth Factor Receptor into Endosomal Compartments. Journal of Biological Chemistry 2014, 289(42), 29180–29194. [Google Scholar] [CrossRef] [PubMed]
- Strochlic, T. I.; Setty, T. G.; Sitaram, A.; Burd, C. G. Grd19/Snx3p Functions as a Cargo-Specific Adapter for Retromer-Dependent Endocytic Recycling. The Journal of Cell Biology 2007, 177(1), 115–125. [Google Scholar] [CrossRef]
- Hamann, L.; Nickel, R.; Tannich, E. Transfection and Continuous Expression of Heterologous Genes in the Protozoan Parasite Entamoeba Histolytica. Proc. Natl. Acad. Sci. U.S.A. 1995, 92(19), 8975–8979. [Google Scholar] [CrossRef]
- Diamond, L. S.; Harlow, D. R.; Cunnick, C. C. A New Medium for the Axenic Cultivation of Entamoeba Histolytica and Other Entamoeba. Transactions of the Royal Society of Tropical Medicine and Hygiene 1978, 72(4), 431–432. [Google Scholar] [CrossRef]
- Zanatta, D.; Betanzos, A.; Azuara-Liceaga, E.; Montaño, S.; Orozco, E. Entamoeba Histolytica: EhADH, an Alix Protein, Participates in Several Virulence Events through Its Different Domains. IJMS 2024, 25(14), 7609. [Google Scholar] [CrossRef]
- Ocádiz-Ruiz, R.; Fonseca, W.; Martínez, M. B.; Ocádiz-Quintanar, R.; Orozco, E.; Rodríguez, M. A. Effect of the Silencing of the Ehcp112 Gene on the in Vitro Virulence of Entamoeba Histolytica. Parasites Vectors 2013, 6(1), 248. [Google Scholar] [CrossRef]
- Díaz-Hernández, M.; Javier-Reyna, R.; Martínez-Valencia, D.; Montaño, S.; Orozco, E. Dynamic Association of ESCRT-II Proteins with ESCRT-I and ESCRT-III Complexes during Phagocytosis of Entamoeba Histolytica. IJMS 2023, 24(6), 5267. [Google Scholar] [CrossRef]
- Bolte, S.; Cordelières, F. P. A Guided Tour into Subcellular Colocalization Analysis in Light Microscopy. Journal of Microscopy 2006, 224(3), 213–232. [Google Scholar] [CrossRef]
- Solis, C. F.; Santi-Rocca, J.; Perdomo, D.; Weber, C.; Guillén, N. Use of Bacterially Expressed dsRNA to Downregulate Entamoeba Histolytica Gene Expression. PLoS ONE 2009, 4(12), e8424. [Google Scholar] [CrossRef] [PubMed]
- Novikoff y Novikoff - 1972 - PEROXISOMES IN ABSORPTIVE CELLS OF MAMMALIAN SMALL.
- Montaño, S.; Orozco, E.; Correa-Basurto, J.; Bello, M.; Chávez-Munguía, B.; Betanzos, A. Heterodimerization of the Entamoeba Histolytica EhCPADH Virulence Complex through Molecular Dynamics and Protein–Protein Docking. Journal of Biomolecular Structure and Dynamics 2017, 35(3), 486–503. [Google Scholar] [CrossRef]
- Kozakov, D.; Hall, D. R.; Xia, B.; Porter, K. A.; Padhorny, D.; Yueh, C.; Beglov, D.; Vajda, S. The ClusPro Web Server for Protein–Protein Docking. Nat Protoc 2017, 12(2), 255–278. [Google Scholar] [CrossRef] [PubMed]
- Kozakov, D.; Beglov, D.; Bohnuud, T.; Mottarella, S. E.; Xia, B.; Hall, D. R.; Vajda, S. How Good Is Automated Protein Docking? Proteins 2013, 81(12), 2159–2166. [Google Scholar] [CrossRef]
- Andrusier, N.; Nussinov, R.; Wolfson, H. J. FireDock: Fast Interaction Refinement in Molecular Docking. Proteins 2007, 69(1), 139–159. [Google Scholar] [CrossRef] [PubMed]
- Mashiach, E.; Schneidman-Duhovny, D.; Andrusier, N.; Nussinov, R.; Wolfson, H. J. FireDock: A Web Server for Fast Interaction Refinement in Molecular Docking. Nucleic Acids Research 2008, 36 (Web Server), W229–W232. [Google Scholar] [CrossRef]
- Laskowski, R. A.; Jabłońska, J.; Pravda, L.; Vařeková, R. S.; Thornton, J. M. PDBsum: Structural Summaries of PDB Entries. Protein Science 2018, 27(1), 129–134. [Google Scholar] [CrossRef]
- Humphrey, W.; Dalke, A.; Schulten, K. VMD: Visual Molecular Dynamics. Journal of Molecular Graphics 1996, 14(1), 33–38. [Google Scholar] [CrossRef] [PubMed]
- Avalos-Padilla, Y.; Betanzos, A.; Javier-Reyna, R.; García-Rivera, G.; Chávez-Munguía, B.; Lagunes-Guillén, A.; Ortega, J.; Orozco, E. EhVps32 Is a Vacuole-Associated Protein Involved in Pinocytosis and Phagocytosis of Entamoeaba Histolytica. PLoS Pathog 2015, 11(7), e1005079. [Google Scholar] [CrossRef]
- Kovtun, O.; Leneva, N.; Bykov, Y. S.; Ariotti, N.; Teasdale, R. D.; Schaffer, M.; Engel, B. D.; Owen, David. J.; Briggs, J. A. G.; Collins, B. M. Structure of the Membrane-Assembled Retromer Coat Determined by Cryo-Electron Tomography. Nature 2018, 561(7724), 561–564. [Google Scholar] [CrossRef]
- Joyal, J.-S.; Nim, S.; Zhu, T.; Sitaras, N.; Rivera, J. C.; Shao, Z.; Sapieha, P.; Hamel, D.; Sanchez, M.; Zaniolo, K.; St-Louis, M.; Ouellette, J.; Montoya-Zavala, M.; Zabeida, A.; Picard, E.; Hardy, P.; Bhosle, V.; Varma, D. R.; Gobeil, F.; Beauséjour, C.; Boileau, C.; Klein, W.; Hollenberg, M.; Ribeiro-da-Silva, A.; Andelfinger, G.; Chemtob, S. Subcellular Localization of Coagulation Factor II Receptor-like 1 in Neurons Governs Angiogenesis. Nat Med 2014, 20(10), 1165–1173. [Google Scholar] [CrossRef]
- Sharma, M.; Morgado, P.; Zhang, H.; Ehrenkaufer, G.; Manna, D.; Singh, U. Characterization of Extracellular Vesicles from Entamoeba Histolytica Identifies Roles in Intercellular Communication That Regulates Parasite Growth and Development. Infect Immun 2020, 88(10), e00349-20. [Google Scholar] [CrossRef]
- Watanabe, N.; Nakada-Tsukui, K.; Nozaki, T. Two Isotypes of Phosphatidylinositol 3-phosphate - Binding Sorting Nexins Play Distinct Roles in Trogocytosis in Entamoeba Histolytica. Cellular Microbiology 2020, 22(3). [Google Scholar] [CrossRef] [PubMed]
- Liu, J.-J. Retromer-Mediated Protein Sorting and Vesicular Trafficking. Journal of Genetics and Genomics 2016, 43(4), 165–177. [Google Scholar] [CrossRef]
- Wang, J.; Fedoseienko, A.; Chen, B.; Burstein, E.; Jia, D.; Billadeau, D. D. Endosomal Receptor Trafficking: Retromer and Beyond. Traffic 2018, 19(8), 578–590. [Google Scholar] [CrossRef] [PubMed]
- Perdomo, D.; Aït-Ammar, N.; Syan, S.; Sachse, M.; Jhingan, G. D.; Guillén, N. Cellular and Proteomics Analysis of the Endomembrane System from the Unicellular Entamoeba Histolytica. Journal of Proteomics 2015, 112, 125–140. [Google Scholar] [CrossRef]
- Constantino-Jonapa, L. A.; Hernández-Ramírez, V. I.; Osorio-Trujillo, C.; Talamás-Rohana, P. EhRab21 Associates with the Golgi Apparatus in Entamoeba Histolytica. Parasitol Res 2020, 119(5), 1629–1640. [Google Scholar] [CrossRef]
- Talamás-Lara, D.; Acosta-Virgen, K.; Chávez-Munguía, B.; Lagunes-Guillén, A.; Salazar-Villatoro, L.; Espinosa-Cantellano, M.; Martínez-Palomo, A. Golgi Apparatus Components in Entamoeba Histolytica and Entamoeba Dispar after Monensin Treatment. Microsc Res Tech 2021, 84(8), 1887–1896. [Google Scholar] [CrossRef]
- Tu, Y.; Seaman, M. N. J. Navigating the Controversies of Retromer-Mediated Endosomal Protein Sorting. Front. Cell Dev. Biol. 2021, 9, 658741. [Google Scholar] [CrossRef] [PubMed]
- Chen, C.; Garcia-Santos, D.; Ishikawa, Y.; Seguin, A.; Li, L.; Fegan, K. H.; Hildick-Smith, G. J.; Shah, D. I.; Cooney, J. D.; Chen, W.; King, M. J.; Yien, Y. Y.; Schultz, I. J.; Anderson, H.; Dalton, A. J.; Freedman, M. L.; Kingsley, P. D.; Palis, J.; Hattangadi, S. M.; Lodish, H. F.; Ward, D. M.; Kaplan, J.; Maeda, T.; Ponka, P.; Paw, B. H. Snx3 Regulates Recycling of the Transferrin Receptor and Iron Assimilation. Cell Metabolism 2013, 17(3), 343–352. [Google Scholar] [CrossRef]
- Vagnozzi, A. N.; Praticò, D. Endosomal Sorting and Trafficking, the Retromer Complex and Neurodegeneration. Mol Psychiatry 2019, 24(6), 857–868. [Google Scholar] [CrossRef]
- MacDonald, E.; Savage, B.; Zech, T. Connecting the Dots: Combined Control of Endocytic Recycling and Degradation. Biochemical Society Transactions 2020, 48(6), 2377–2386. [Google Scholar] [CrossRef] [PubMed]
- Curtis, M. E.; Yu, D.; Praticò, D. Dysregulation of the Retromer Complex System in Down Syndrome. Annals of Neurology 2020, 88(1), 137–147. [Google Scholar] [CrossRef]
- Small, S. A.; Petsko, G. A. Retromer in Alzheimer Disease, Parkinson Disease and Other Neurological Disorders. Nat Rev Neurosci 2015, 16(3), 126–132. [Google Scholar] [CrossRef] [PubMed]
- Bañuelos, C.; García-Rivera, G.; López-Reyes, I.; Orozco, E. Functional Characterization of EhADH112: An Entamoeba Histolytica Bro1 Domain-Containing Protein. Experimental Parasitology 2005, 110(3), 292–297. [Google Scholar] [CrossRef]
- García-Rivera, G.; Rodríguez, M. A.; Ocádiz, R.; Martínez-López, M. C.; Arroyo, R.; González-Robles, A.; Orozco, E. Entamoeba Histolytica : A Novel Cysteine Protease and an Adhesin Form the 112 kDa Surface Protein. Molecular Microbiology 1999, 33(3), 556–568. [Google Scholar] [CrossRef]
- Ocádiz-Ruiz, R.; Fonseca, W.; Linford, A. S.; Yoshino, T. P.; Orozco, E.; Rodríguez, M. A. The Knockdown of Each Component of the Cysteine Proteinase-Adhesin Complex of Entamoeba Histolytica (EhCPADH) Affects the Expression of the Other Complex Element as Well as the in Vitro and in Vivo Virulence. Parasitology 2016, 143(1), 50–59. [Google Scholar] [CrossRef]
- Chen, K.-E.; Guo, Q.; Hill, T. A.; Cui, Y.; Kendall, A. K.; Yang, Z.; Hall, R. J.; Healy, M. D.; Sacharz, J.; Norwood, S. J.; Fonseka, S.; Xie, B.; Reid, R. C.; Leneva, N.; Parton, R. G.; Ghai, R.; Stroud, D. A.; Fairlie, D. P.; Suga, H.; Jackson, L. P.; Teasdale, R. D.; Passioura, T.; Collins, B. M. De Novo Macrocyclic Peptides for Inhibiting, Stabilizing, and Probing the Function of the Retromer Endosomal Trafficking Complex. Sci. Adv. 2021, 7(49), eabg4007. [Google Scholar] [CrossRef]
- Fagnani, E.; Bonì, F.; Seneci, P.; Gornati, D.; Muzio, L.; Mastrangelo, E.; Milani, M. Stabilization of the Retromer Complex: Analysis of Novel Binding Sites of Bis-1,3-Phenyl Guanylhydrazone 2a to the VPS29/VPS35 Interface. Computational and Structural Biotechnology Journal 2024, 23, 1088–1093. [Google Scholar] [CrossRef]
- Fuse, A.; Furuya, N.; Kakuta, S.; Inose, A.; Sato, M.; Koike, M.; Saiki, S.; Hattori, N. VPS29–VPS35 Intermediate of Retromer Is Stable and May Be Involved in the Retromer Complex Assembly Process. FEBS Letters 2015, 589(13), 1430–1436. [Google Scholar] [CrossRef]
- Franch-Marro, X.; Wendler, F.; Guidato, S.; Griffith, J.; Baena-Lopez, A.; Itasaki, N.; Maurice, M. M.; Vincent, J.-P. Wingless Secretion Requires Endosome-to-Golgi Retrieval of Wntless/Evi/Sprinter by the Retromer Complex. Nat Cell Biol 2008, 10(2), 170–177. [Google Scholar] [CrossRef] [PubMed]
- Cui, Y.; Carosi, J. M.; Yang, Z.; Ariotti, N.; Kerr, M. C.; Parton, R. G.; Sargeant, T. J.; Teasdale, R. D. Retromer Has a Selective Function in Cargo Sorting via Endosome Transport Carriers. Journal of Cell Biology 2019, 218(2), 615–631. [Google Scholar] [CrossRef]
- Pim, D.; Broniarczyk, J.; Bergant, M.; Playford, M. P.; Banks, L. A Novel PDZ Domain Interaction Mediates the Binding between Human Papillomavirus 16 L2 and Sorting Nexin 27 and Modulates Virion Trafficking. J Virol 2015, 89(20), 10145–10155. [Google Scholar] [CrossRef] [PubMed]
- Popa, A.; Zhang, W.; Harrison, M. S.; Goodner, K.; Kazakov, T.; Goodwin, E. C.; Lipovsky, A.; Burd, C. G.; DiMaio, D. Direct Binding of Retromer to Human Papillomavirus Type 16 Minor Capsid Protein L2 Mediates Endosome Exit during Viral Infection. PLoS Pathog 2015, 11(2), e1004699. [Google Scholar] [CrossRef]
- Sangaré, L. O.; Alayi, T. D.; Westermann, B.; Hovasse, A.; Sindikubwabo, F.; Callebaut, I.; Werkmeister, E.; Lafont, F.; Slomianny, C.; Hakimi, M.-A.; Van Dorsselaer, A.; Schaeffer-Reiss, C.; Tomavo, S. Unconventional Endosome-like Compartment and Retromer Complex in Toxoplasma Gondii Govern Parasite Integrity and Host Infection. Nat Commun 2016, 7(1), 11191. [Google Scholar] [CrossRef]
- Finsel, I.; Ragaz, C.; Hoffmann, C.; Harrison, C. F.; Weber, S.; van Rahden, V. A.; Johannes, L.; Hilbi, H. The Legionella Effector RidL Inhibits Retrograde Trafficking to Promote Intracellular Replication. Cell Host & Microbe 2013, 14(1), 38–50. [Google Scholar] [CrossRef]
- Arroyo, R.; Orozco, E. Localization and Identification of an Entamoeba Histolytica Adhesin. Molecular and Biochemical Parasitology 1987, 23(2), 151–158. [Google Scholar] [CrossRef]
- Bolognesi, B.; Lehner, B. Reaching the Limit. eLife 2018, 7, e39804. [Google Scholar] [CrossRef]
- Stoebel, D. M.; Dean, A. M.; Dykhuizen, D. E. The Cost of Expression of Escherichia Coli Lac Operon Proteins Is in the Process, Not in the Products. Genetics 2008, 178(3), 1653–1660. [Google Scholar] [CrossRef] [PubMed]
- Gao, C.; Cao, N.; Wang, Y. Metal Dependent Protein Phosphatase PPM Family in Cardiac Health and Diseases. Cellular Signalling 2021, 85, 110061. [Google Scholar] [CrossRef]
- Lu, J.; Xu, S.; Huo, Y.; Sun, D.; Hu, Y.; Wang, J.; Zhang, X.; Wang, P.; Li, Z.; Liang, M.; Wu, Z.; Liu, P. Sorting Nexin 3 Induces Heart Failure via Promoting Retromer-Dependent Nuclear Trafficking of STAT3. Cell Death Differ 2021, 28(10), 2871–2887. [Google Scholar] [CrossRef] [PubMed]
- Seaman, M. N. J. Cargo-Selective Endosomal Sorting for Retrieval to the Golgi Requires Retromer. The Journal of Cell Biology 2004, 165(1), 111–122. [Google Scholar] [CrossRef] [PubMed]
- Oliviusson, P.; Heinzerling, O.; Hillmer, S.; Hinz, G.; Tse, Y. C.; Jiang, L.; Robinson, D. G. Plant Retromer, Localized to the Prevacuolar Compartment and Microvesicles in Arabidopsis, May Interact with Vacuolar Sorting Receptors. The Plant Cell 2006, 18(5), 1239–1252. [Google Scholar] [CrossRef] [PubMed]
- Zelazny, E.; Santambrogio, M.; Pourcher, M.; Chambrier, P.; Berne-Dedieu, A.; Fobis-Loisy, I.; Miège, C.; Jaillais, Y.; Gaude, T. Mechanisms Governing the Endosomal Membrane Recruitment of the Core Retromer in Arabidopsis. Journal of Biological Chemistry 2013, 288(13), 8815–8825. [Google Scholar] [CrossRef] [PubMed]
- Betanzos, A.; Zanatta, D.; Bañuelos, C.; Hernández-Nava, E.; Cuellar, P.; Orozco, E. Epithelial Cells Expressing EhADH, An Entamoeba Histolytica Adhesin, Exhibit Increased Tight Junction Proteins. Front. Cell. Infect. Microbiol. 2018, 8, 340. [Google Scholar] [CrossRef]
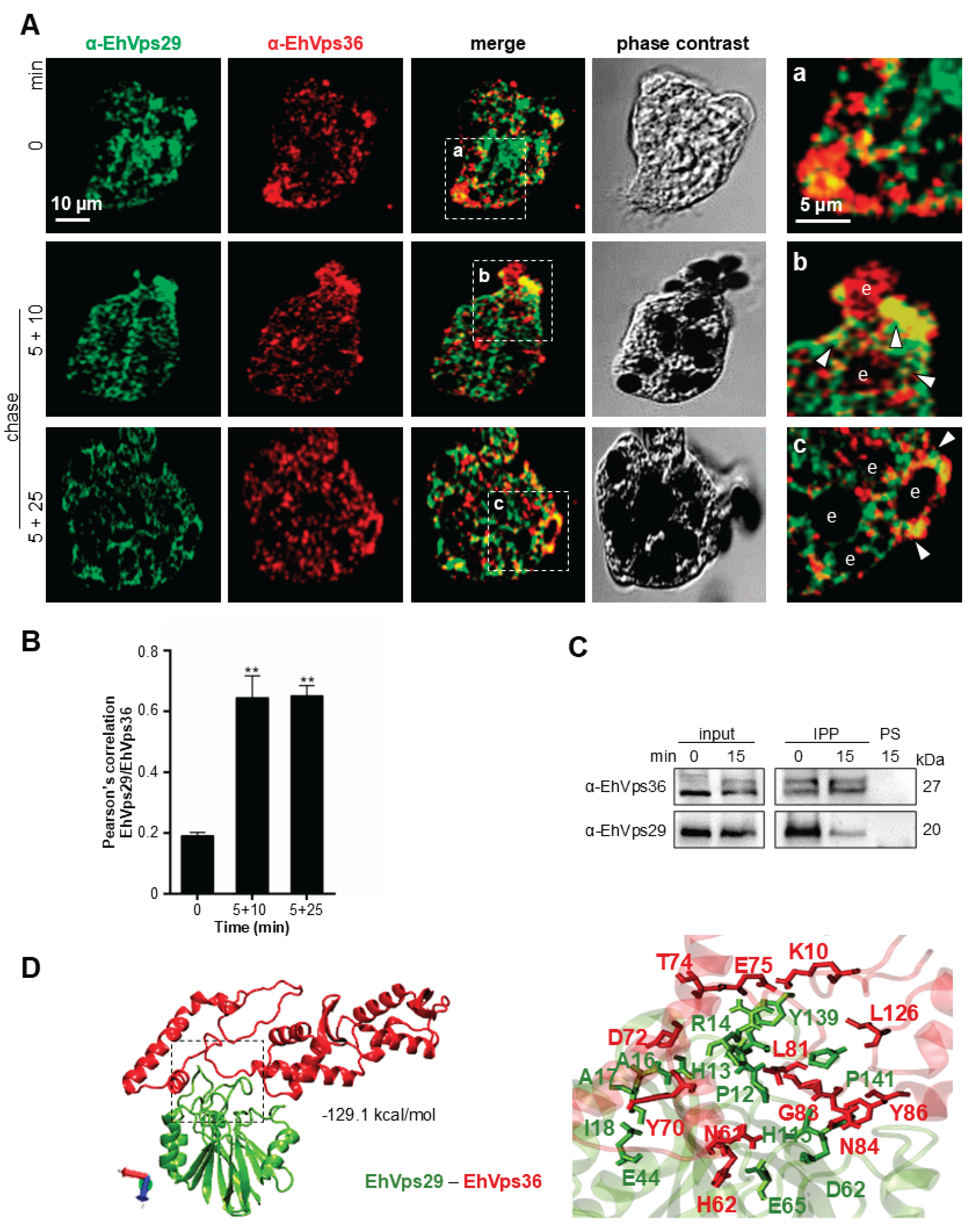
| EhVps29 | EhVps36 | Distance (Å) | EhVps29 | EhVps36 | Distance (Å) | ||
|---|---|---|---|---|---|---|---|
| 1 | H13 | R160 | 2.70 | 8 | T116 | Y233 | 2.84 |
| 2 | H13 | D161 | 2.85 | 9 | T116 | Q229 | 2.76 |
| 3 | R14 | E165 | 2.96 | 10 | T116 | Q229 | 2.93 |
| 4 | H88 | E208 | 2.93 | 11 | K118 | Y233 | 2.66 |
| 5 | W93 | T228 | 2.79 | 12 | L119 | Y235 | 2.91 |
| 6 | H115 | L231 | 2.72 | 13 | Y139 | N169 | 2.96 |
| 7 | T116 | Q232 | 2.93 | 14 | R14 | E165 | 2.71* |
| EhVps29 | EhADH | Distance (Å) | EhVps29 | EhADH | Distance (Å) | ||
|---|---|---|---|---|---|---|---|
| 1 | D169 | K100 | 2.58 | 12 | V11 | Y676 | 2.73 |
| 2 | D144 | R450 | 2.99 | 13 | H13 | Y676 | 2.90 |
| 3 | G137 | Y455 | 2.85 | 14 | S15 | S677 | 2.89 |
| 4 | S140 | Y455 | 2.93 | 15 | S15 | G679 | 2.81 |
| 5 | S15 | Y457 | 2.89 | 16 | E44 | T680 | 2.79 |
| 6 | Y139 | W459 | 2.81 | 17 | E44 | N681 | 3.04 |
| 7 | S140 | V662 | 2.75 | 18 | I18 | N681 | 3.01 |
| 8 | H10 | S664 | 3.08 | 19 | E44 | N681 | 2.97 |
| 9 | H13 | N668 | 3.19 | 20 | D169 | K100 | 2.58* |
| 10 | R14 | Q672 | 2.65 | 21 | E44 | R450 | 2.76* |
| 11 | H13 | Q674 | 2.75 | 22 | D62 | R663 | 2.75* |
| EhVps29 | EhCP112 | Distance (Å) | EhVps29 | EhCP112 | Distance (Å) | ||
|---|---|---|---|---|---|---|---|
| 1 | I91 | K212 | 2.60 | 9 | D144 | S409 | 2.89 |
| 2 | Y102 | K212 | 2.69 | 10 | R14 | C410 | 2.71 |
| 3 | E71 | R213 | 2.70 | 11 | P12 | S412 | 2.71 |
| 4 | H88 | R213 | 2.90 | 12 | H13 | S412 | 2.97 |
| 5 | Y139 | R221 | 2.73 | 13 | H13 | G413 | 2.81 |
| 6 | P141 | R221 | 3.09 | 14 | S15 | Y414 | 2.73 |
| 7 | H117 | R222 | 2.90 | 15 | E71 | R213 | 2.70* |
| 8 | R14 | Y266 | 2.68 | 16 | D62 | R222 | 2.65* |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




