Submitted:
29 March 2026
Posted:
31 March 2026
Read the latest preprint version here
Abstract
Keywords:
1. Introduction
2. Molecular Basis of Antibody-Dependent Enhancement
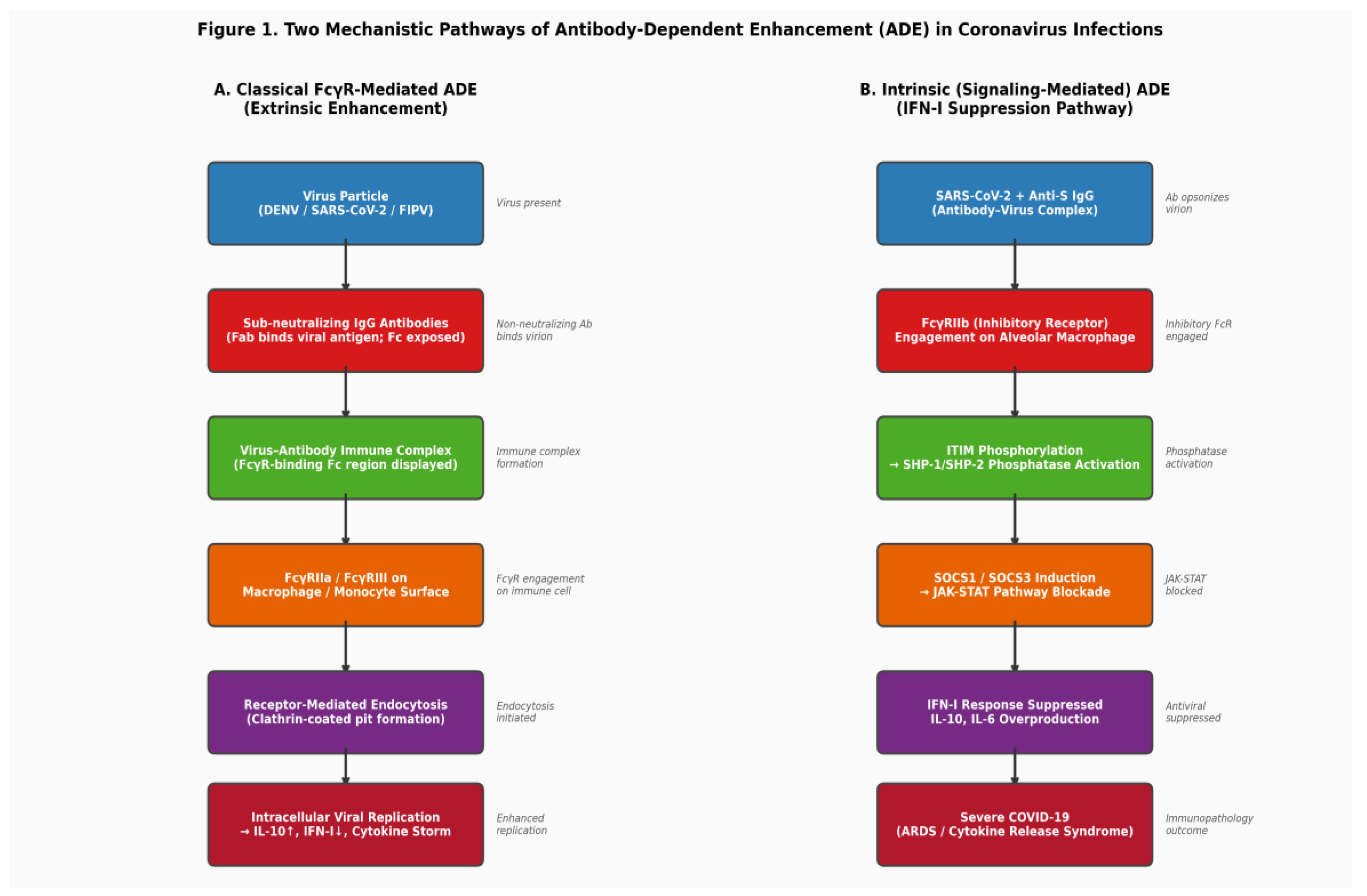
2.1. Classical Extrinsic ADE: FcγR-Mediated Viral Entry
2.2. Intrinsic ADE: FcγRIIb-Mediated IFN-I Suppression
2.3. Antibody Glycosylation and ADE Risk
3. Fc Gamma Receptor Biology in Immune Cells
3.1. Classification, Structure, and Expression
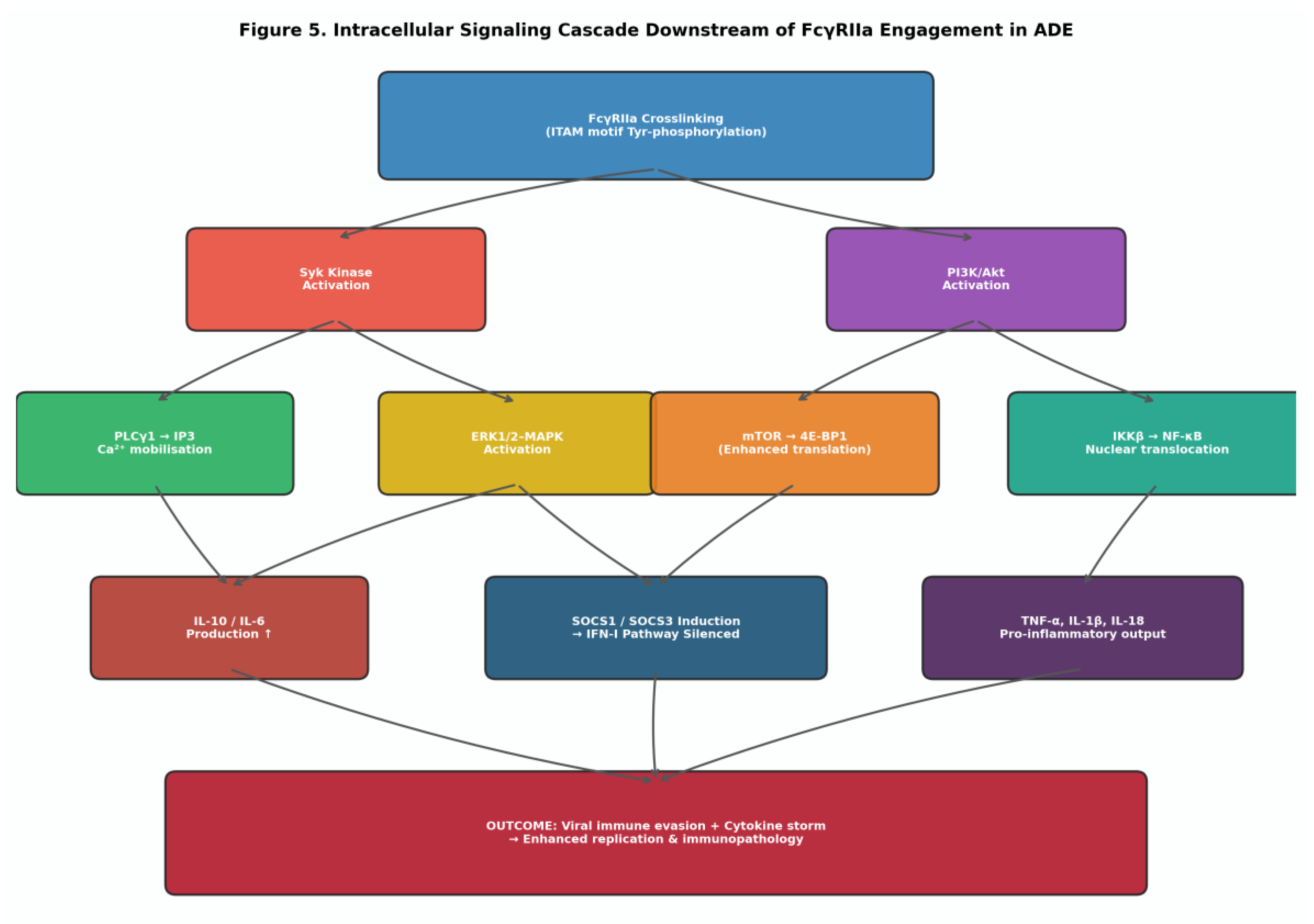
3.2. Intracellular Signalling Cascades in ADE
3.3. FcγR Polymorphisms and Individual ADE Susceptibility
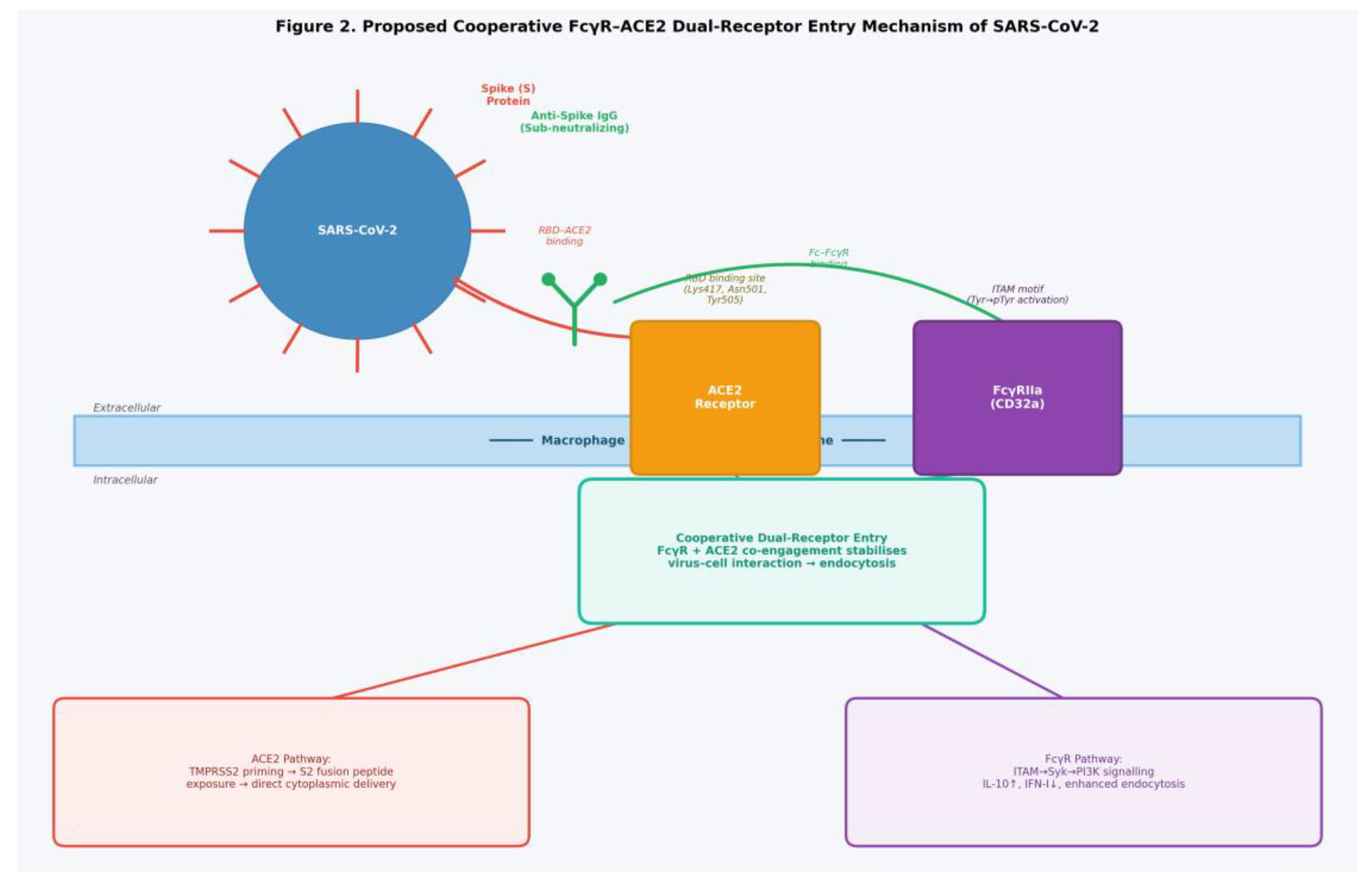
5. The Cooperative FcγR–ACE2 Entry Model
5.1. Rationale, Structural Feasibility, and Experimental Evidence
5.2. Implications for Antibody-Based Therapeutics
6. Intracellular Trafficking and Viral Fate After FcγR-Mediated Entry
8. Nano-Engineered Immunomodulatory Platforms for ADE-Safe Vaccination
8.1. Antigen Engineering Principles to Minimise ADE Risk
8.2. Lipid Nanoparticle mRNA Vaccines
8.3. Virus-Like Particle Platforms
8.4. Biodegradable Polymeric Nanoparticles (PLGA/Chitosan)
9. ADE in Comparative Perspective: Evidence Across Viral Families
11. Future Research Priorities
- • Structural biology: Cryo-electron tomography of SARS-CoV-2–IgG–macrophage ternary complexes to directly visualise cooperative FcγR–ACE2 engagement geometry and endosomal sorting.
- • Single-cell transcriptomics: sc-RNA-seq/ATAC-seq profiling of macrophages infected via ADE vs standard entry to map ADE-specific transcriptional signatures and identify targetable molecular checkpoints.
- • FcγR-humanised animal models: Development of mice expressing human FcγRIIa (H131/R131 alleles) and FcγRIIb to enable in vivo ADE mechanistic studies not possible in standard murine models.
- • FIPV vaccine re-engineering: Systematic evaluation of ADE-safe mucosal FIPV vaccine platforms (intranasal LNP-mRNA, VLP) as direct translational templates for ADE-proof human coronavirus vaccines.
- • Nanomedicine platform comparison: Head-to-head comparison of LNP-mRNA, VLP, and PLGA nanoparticle vaccines in ADE-capable macaque and FIPV models, correlating antibody glycosylation profiles and nAb:non-nAb ratios with ADE protection.
- • Fc engineering: Clinical evaluation of LALAPG and GASDALIE Fc-modified therapeutic mAbs to establish the therapeutic window between ADE risk and beneficial ADCC effector function.
- • Polyherbal-nano interface: Investigation of plant-derived bioactive compounds (andrographolide from Andrographis paniculata; nimbin from Azadirachta indica; baeyer from Aegle marmelos) as nano-formulated adjunct immunomodulators capable of downregulating FcγR expression on macrophages to reduce ADE susceptibility while maintaining phagocytic function.
- • Variant surveillance: Continuous monitoring of antibody cross-reactivity and neutralisation profiles against emerging SARS-CoV-2 variants of concern to identify ADE-permissive antibody configurations arising from immune imprinting or antigenic sin.
12. Conclusions
References
- Periasamy S, Tang J, Olaoba OT, Spearman P, Damania BA, de Silva AM. SARS-CoV-2 antibody–dependent enhancement of infection depends on antibody binding to both ACE2 and Fc receptors. JCI Insight. 2026;11(4):e175432. [CrossRef]
- Halstead SB. Dengue antibody-dependent enhancement: knowns and unknowns. Microbiol Spectr. 2014;2(6):AID-0022-2014.
- Takada A, Kawaoka Y. Antibody-dependent enhancement of viral infection: molecular mechanisms and in vivo implications. Rev Med Virol. 2003;13(6):387-398.
- Wan Y, et al. Molecular mechanism for antibody-dependent enhancement of coronavirus entry. J Virol. 2020;94(5):e02015-19.
- Liu L, et al. Anti-spike IgG causes severe acute lung injury by skewing macrophage responses during acute SARS-CoV infection. JCI Insight. 2019;4(4):e123158.
- Zhao J, et al. Antibody responses to SARS-CoV-2 in patients with COVID-19. Clin Infect Dis. 2020;71(16):2027-2034.
- Lee WS, et al. Antibody-dependent enhancement and SARS-CoV-2 vaccines and therapies. Nat Microbiol. 2020;5(10):1185-1191.
- Katzelnick LC, et al. Antibody-dependent enhancement of severe dengue disease in humans. Science. 2017;358(6365):929-932.
- Olsen CW, et al. Monoclonal antibodies to FIPV spike protein mediate antibody-dependent enhancement of infection in macrophages. J Virol. 1992;66(2):956-965.
- Vennema H, et al. Early death after FIPV challenge due to recombinant vaccinia virus immunization. J Virol. 1990;64(3):1407-1409.
- Peiris JS, Porterfield JS. Antibody-mediated enhancement of Flavivirus replication in macrophage-like cell lines. Nature. 1979;282(5738):509-511.
- Hoffmann M, et al. SARS-CoV-2 cell entry depends on ACE2 and TMPRSS2. Cell. 2020;181(2):271-280.
- Walls AC, et al. Structure, function, and antigenicity of the SARS-CoV-2 spike glycoprotein. Cell. 2020;181(2):281-292.
- Wrapp D, et al. Cryo-EM structure of the 2019-nCoV spike in the prefusion conformation. Science. 2020;367(6483):1260-1263.
- Shang J, et al. Cell entry mechanisms of SARS-CoV-2. Proc Natl Acad Sci USA. 2020;117(21):11727-11734.
- Millet JK, Whittaker GR. Physiological and molecular triggers for SARS-CoV membrane fusion. Virology. 2018;517:3-8.
- Poland GA, et al. Vaccinomics and personalized vaccinology. PLoS Pathog. 2011;7(12):e1002344.
- Dinnon KH, et al. A mouse-adapted model of SARS-CoV-2. Nature. 2020;586(7830):560-566.
- Zohar T, Alter G. Dissecting antibody-mediated protection against SARS-CoV-2. Nat Rev Immunol. 2020;20(7):392-394.
- Nimmerjahn F, Ravetch JV. Fcγ receptors as regulators of immune responses. Nat Rev Immunol. 2008;8(1):34-47.
- Bournazos S, et al. The role of IgG Fc domain in antibody-mediated protection against bacteria. J Exp Med. 2015;212(9):1387-1392.
- Ferrara C, et al. Carbohydrate interactions required for high affinity FcγRIII–antibody binding. Proc Natl Acad Sci USA. 2011;108(31):12669-12674.
- Larsen MD, et al. Afucosylated IgG characterizes enveloped viral infections and COVID-19 severity. Science. 2021;371(6532):eabc8378.
- Cheung CY, et al. Cytokine responses in SARS-CoV-infected macrophages in vitro. J Virol. 2005;79(12):7819-7826.
- Pedersen NC. An update on feline infectious peritonitis: diagnostics and therapeutics. Vet J. 2014;201(2):133-141.
- Murphy BG, et al. GS-441524 strongly inhibits FIPV in tissue culture and experimental cat infection. Vet Microbiol. 2018;219:226-233.
- Polack FP, et al. Safety and efficacy of the BNT162b2 mRNA COVID-19 vaccine. N Engl J Med. 2020;383(27):2603-2615.
- Baden LR, et al. Efficacy and safety of mRNA-1273 SARS-CoV-2 vaccine. N Engl J Med. 2021;384(5):403-416.
- Vogel AB, et al. BNT162b vaccines protect rhesus macaques from SARS-CoV-2. Nature. 2021;592(7853):283-289.
- Keech C, et al. Phase 1-2 trial of a SARS-CoV-2 recombinant spike nanoparticle vaccine. N Engl J Med. 2020;383(24):2320-2332.
- Coleman CM, et al. Innate immune responses to MERS-CoV. mBio. 2017;8(4):e00099-17.
- de Alwis R, et al. Impact of immune enhancement on COVID-19 polyclonal hyperimmune globulin therapy. EBioMedicine. 2020;55:102768.
- Ricke DO. Two ADE risks for SARS-CoV-2 antibodies. Front Immunol. 2021;12:443.
- Arvin AM, et al. Potential antibody-dependent enhancement of SARS-CoV-2—a perspective. Nature. 2020;584(7821):353-363.
- Bolles M, et al. A double-inactivated SARS vaccine provides incomplete protection in mice. J Virol. 2011;85(23):12201-12215.
- Pinto D, et al. Cross-neutralization of SARS-CoV-2 by a human monoclonal SARS-CoV antibody. Nature. 2020;583(7815):290-295.
- Rogers TF, et al. Isolation of potent SARS-CoV-2 neutralizing antibodies. Science. 2020;369(6506):956-963.
- Suthar MS, et al. Rapid generation of neutralizing antibody responses in COVID-19 patients. Cell Rep Med. 2020;1(3):100040.
- Brouwer PJM, et al. Potent neutralizing antibodies from COVID-19 patients. Science. 2020;369(6504):643-650.
- Guzman MG, et al. Dengue: a continuing global threat. Nat Rev Microbiol. 2010;8(12 Suppl):S7-16.
- Kiyotani K, et al. Coronavirus immunology: lessons from SARS-CoV-1. Ann NY Acad Sci. 2021;1503(1):18-29.
- Corti D, Lanzavecchia A. Broadly neutralizing antiviral antibodies. Annu Rev Immunol. 2013;31:705-742.
- Bhatt S, et al. The global distribution and burden of dengue. Nature. 2013;496(7446):504-507.
- Ou X, et al. Characterization of spike glycoprotein of SARS-CoV-2 on virus entry. Nat Commun. 2020;11(1):1620.
- Zhou P, et al. A pneumonia outbreak with a new coronavirus of probable bat origin. Nature. 2020;579(7798):270-273.
- Pedersen NC. A review of feline infectious peritonitis virus infection: 1963-2008. J Feline Med Surg. 2009;11(4):225-258.
- Legendre AM, Bartges JW. Effect of Polyprenyl Immunostimulant on FIP survival times. J Feline Med Surg. 2009;11(8):624-626.
- Mair-Jenkins J, et al. Effectiveness of convalescent plasma for severe acute respiratory infections. J Infect Dis. 2015;211(1):80-90.
- Ravichandran S, et al. Longitudinal antibody repertoire in convalescent COVID-19 patients. JCI Insight. 2020;5(24):e143380.
- Dijkman R, et al. Human coronavirus NL63 and 229E: disease associations and receptor usage. J Clin Virol. 2012;55(2):111-116.
- Wrobel AG, et al. SARS-CoV-2 and bat coronavirus RaTG13 spike structure. Nat Commun. 2020;11(1):5173.
| FcγR Type | CD Name | IgG Affinity | Cell Expression | Signal Motif | ADE Relevance |
| FcγRI | CD64 | High (Kd ~10−9 M) | Monocytes, Macrophages, DCs | ITAM (γ-chain) | Primary mediator; binds monomeric IgG; high-avidity ADE |
| FcγRIIa | CD32a | Intermediate | Macrophages, Neutrophils, Platelets | ITAM (intrinsic) | Activating; central coronavirus ADE receptor; H131 polymorphism |
| FcγRIIb | CD32b | Intermediate | B cells, Macrophages, DCs | ITIM (inhibitory) | Intrinsic ADE; IFN-I suppression via SHP-1/2; SOCS induction |
| FcγRIIIa | CD16a | Low–Intermediate | NK cells, Macrophages, Monocytes | ITAM (γ/ζ-chain) | ADCC; ADE in monocytes; V158 polymorphism affects affinity |
| FcγRIIIb | CD16b | Low | Neutrophils only | GPI-linked (no signalling) | Phagocytosis; minimal direct ADE role |
| Parameter | SARS-CoV-2 (Human) | FIPV (Feline) |
| Coronavirus lineage | Betacoronavirus (lineage B) | Alphacoronavirus (FCoV type I/II) |
| Primary entry receptor | ACE2 (+ TMPRSS2 co-factor) | APN/CD13; FcγR in ADE |
| ADE mechanism | Extrinsic (FcγRIIa + ACE2 cooperative) + Intrinsic (FcγRIIb-SOCS) | Classical FcγR-mediated macrophage entry; antibody accelerates entry |
| Primary ADE target cell | Alveolar macrophages, monocytes | Peritoneal/pleural macrophages (tissue-resident) |
| Key pathogenic cytokines | IL-6, IL-10, TNF-α; IFN-I suppressed | IL-6, TNF-α, IL-1β; IL-12 reduced |
| Tissue pathology | Diffuse alveolar damage, ARDS | Pyogranulomatous vasculitis, exudative effusions |
| Mortality in severe cases | ~15–40% (critically ill); overall ~1–3% | Near 100% without antiviral therapy |
| Vaccine-enhanced disease | Not observed with mRNA vaccines; theoretical risk exists | Documented with whole-virus and recombinant spike vaccines |
| Therapeutic antiviral | Remdesivir, Paxlovid, molnupiravir | GS-441524 (remdesivir parent compound) |
| Gold-standard ADE model? | Human—limited by ethical constraints on controlled challenge | YES—natural cat model; controlled challenge studies possible |
| Platform | Antigen Type | Size Range | Adjuvant? | ADE Risk Mitigation | Stage/Examples |
| LNP-mRNA | Encoded prefusion spike | 80–200 nm | No (self-adj) | Selective nAb1 induction; core-fucosylated IgG Fc; brief Ag expression | Approved: BNT162b2, mRNA-1273 |
| Virus-Like Particle | RBD/spike multimer | 20–100 nm | Often co-formulated | No Fc-activating material; high nAb density; no Fc-triggering genome | HPV/HepB approved; CoV pre-clinical |
| PLGA NP | Protein/peptide subunit | 100–500 nm | MPL, CpG | Sustained release prevents sub-nAb surge window; Th1 bias | Pre-clinical CoV; FDA-approved platforms |
| Chitosan NP | Mucosal protein Ag | 100–600 nm | Intrinsic | Mucosal sIgA induction; sIgA does NOT engage FcγR → no ADE | Pre-clinical; intranasal route |
| Self-assembling NP | Ferritin-RBD, I3-01 scaffold | 10–50 nm | Alum/AS01B | Precise epitope control; avoids immunodominance shift to non-nAb regions | Pre-clinical (HexaPro, SpFN, I53-50) |
| Virus | Family | ADE Mechanism | Target Cell | Key Receptor(s) | Clinical Impact |
| DENV (Dengue) | Flaviviridae | Classical FcγR | Monocytes/Macrophages | FcγRIIa (CD32a) | DHF/DSS; ~25,000 deaths/yr; strongest clinical ADE evidence |
| SARS-CoV-2 | Coronaviridae | Classical + Intrinsic | Alveolar Macrophages, Monocytes | FcγRIIa + ACE2 (cooperative) | COVID-19 cytokine storm; >6M deaths; both pathways implicated |
| SARS-CoV-1 | Coronaviridae | Classical FcγR | Macrophages, DCs | FcγRI, FcγRII | In vitro ADE; vaccine-enhanced disease in animal models |
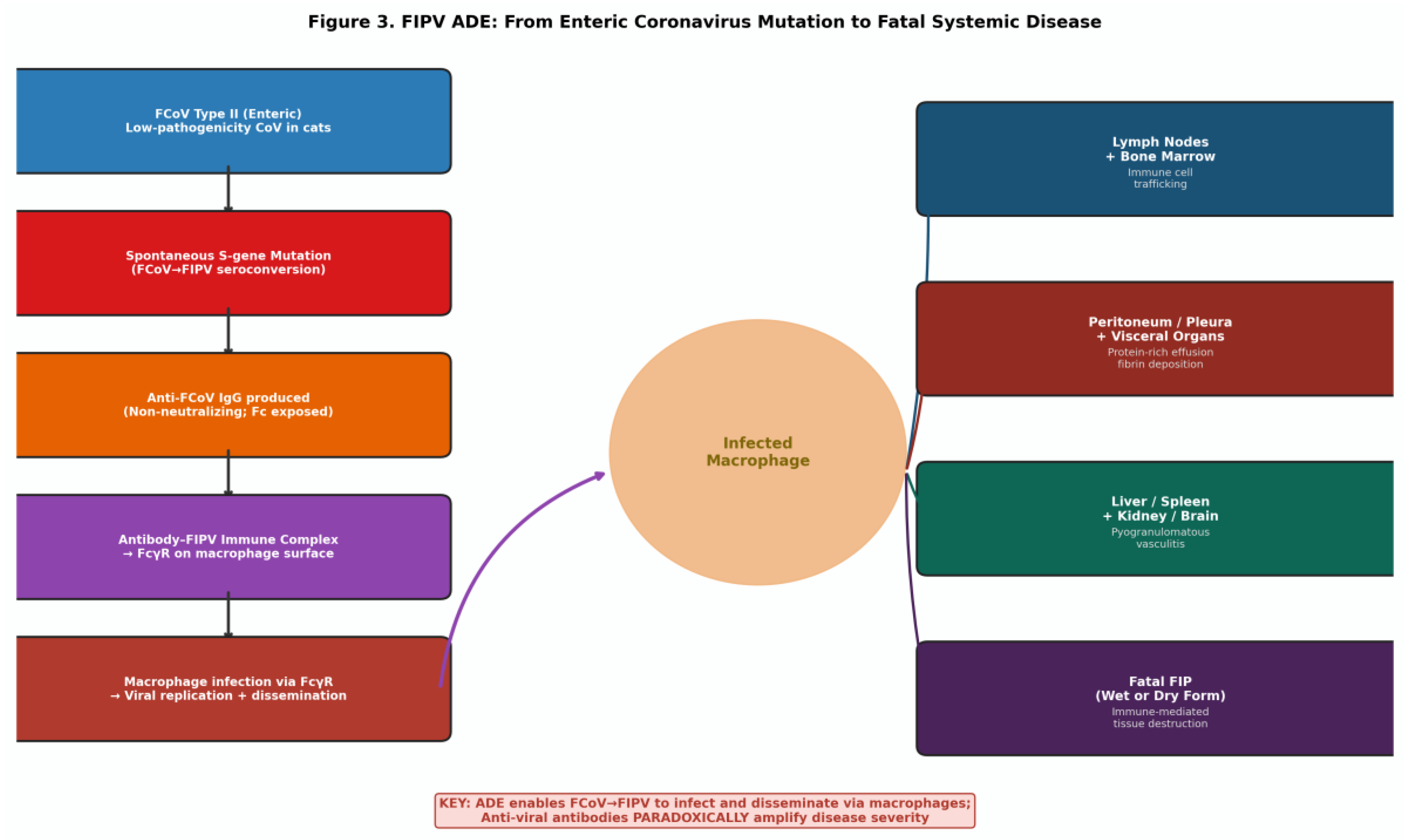
| FIPV (FCoV) | Coronaviridae | Classical FcγR | Peritoneal Macrophages | Feline FcγR | Fatal FIP; ADE is central, documented, and unavoidable mechanism |
| MERS-CoV | Coronaviridae | Putative Classical | Macrophages, T cells | DPP4 + FcγR (hypothetical) | In vitro evidence; uncertain in vivo significance |
| HIV-1 | Retroviridae | Classical + Complement | CD4+ T cells, Macrophages | FcγRIII, CR3 | Established in vitro; clinical relevance debated |
| Zika Virus | Flaviviridae | Classical FcγR | Placental macrophages, DCs | FcγRIIIa | Congenital Zika risk; cross-reactive DENV Ab implicated |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




