Submitted:
16 March 2026
Posted:
17 March 2026
You are already at the latest version
Abstract
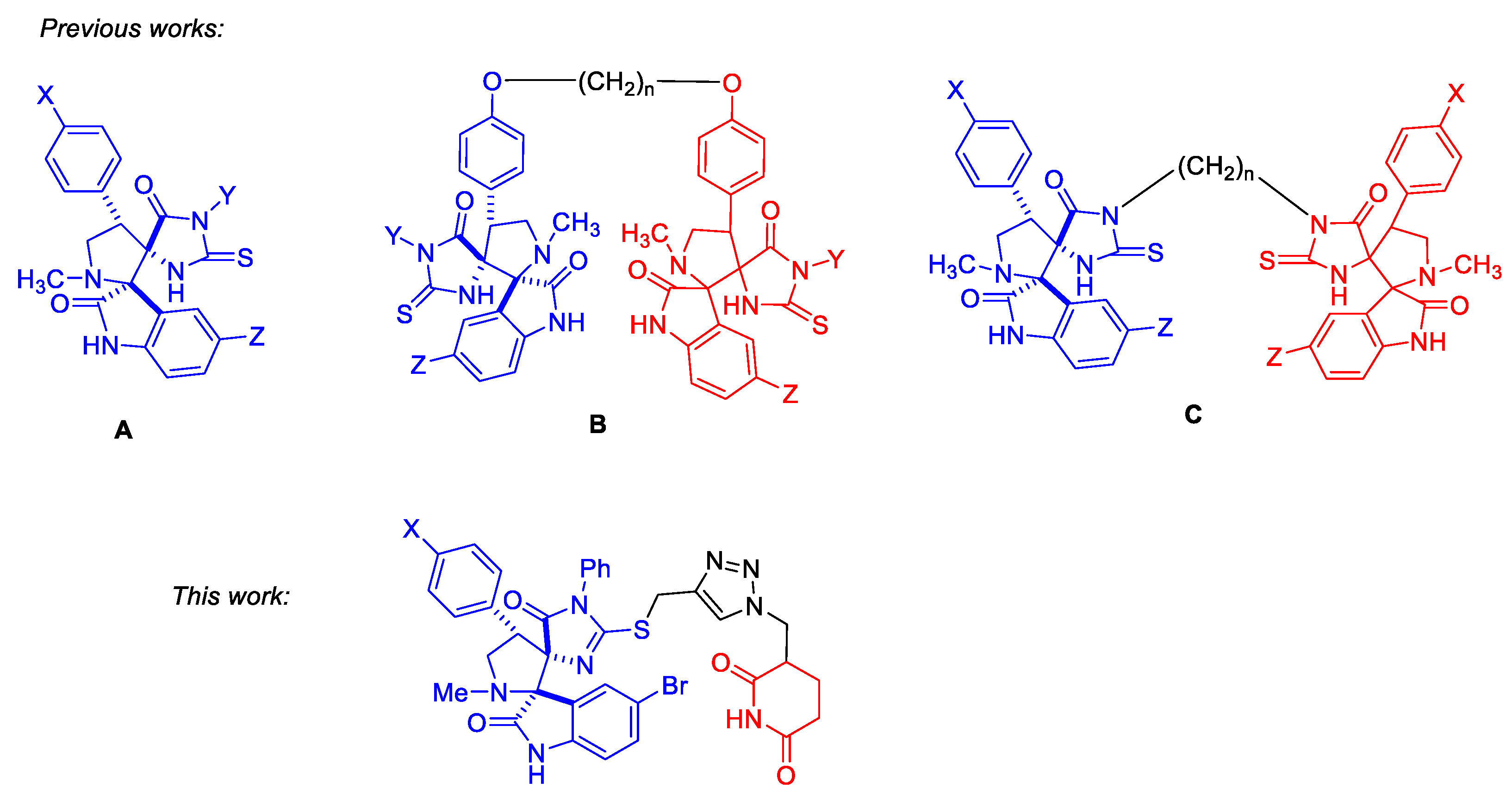
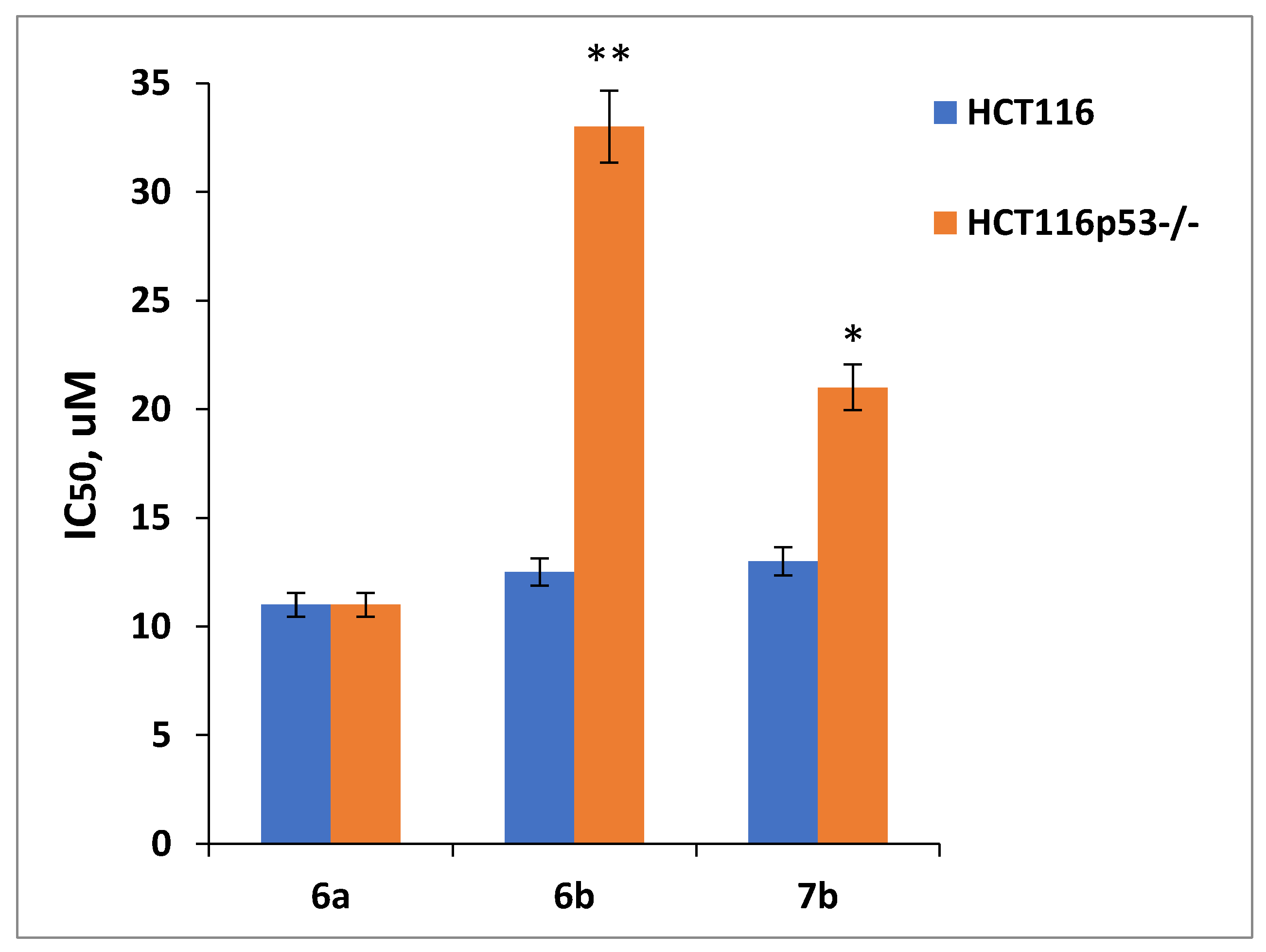
Aiming at p53-reactivating compounds, a convergent scheme for the preparation of conjugates with the dispiro-indolinone-pyrrolidine-thioimidazolone and glutarimide moieties connected via a triazole-containing linker were proposed. Target conjugates were synthesized by azide-alkyne cycloaddition reactions between propargylthio-substituted dispiro-indolinone-pyrrolidine-imidazolones and an azido-glutarimide derivative. The starting compounds were available isothiocyanates, glycine, substituted benzaldehydes, chloroacetamide, and ethyl acrylate. The key azide-alkyne cycloaddition step was carried out using TBTA as a catalyst, achieving >70% product yields. The resulting bifunctional compounds contained a fragment of dispiroindolinone (p53-MDM2 interaction inhibitor) and glutarimide, an ubiquitin ligase ligand. The dispiroindolinone-glutarimide conjugate with 5-bromoisatine and 4-bromophenyl moieties showed a potential for p53 re-activation as determined by preferential cytotoxicity against HCT116 colon carcinoma cells (wild type53) compared to the isogenic HCT116p53-/- subline.
Keywords:
1. Introduction
2. Results and Discussion
2.1. Synthesis
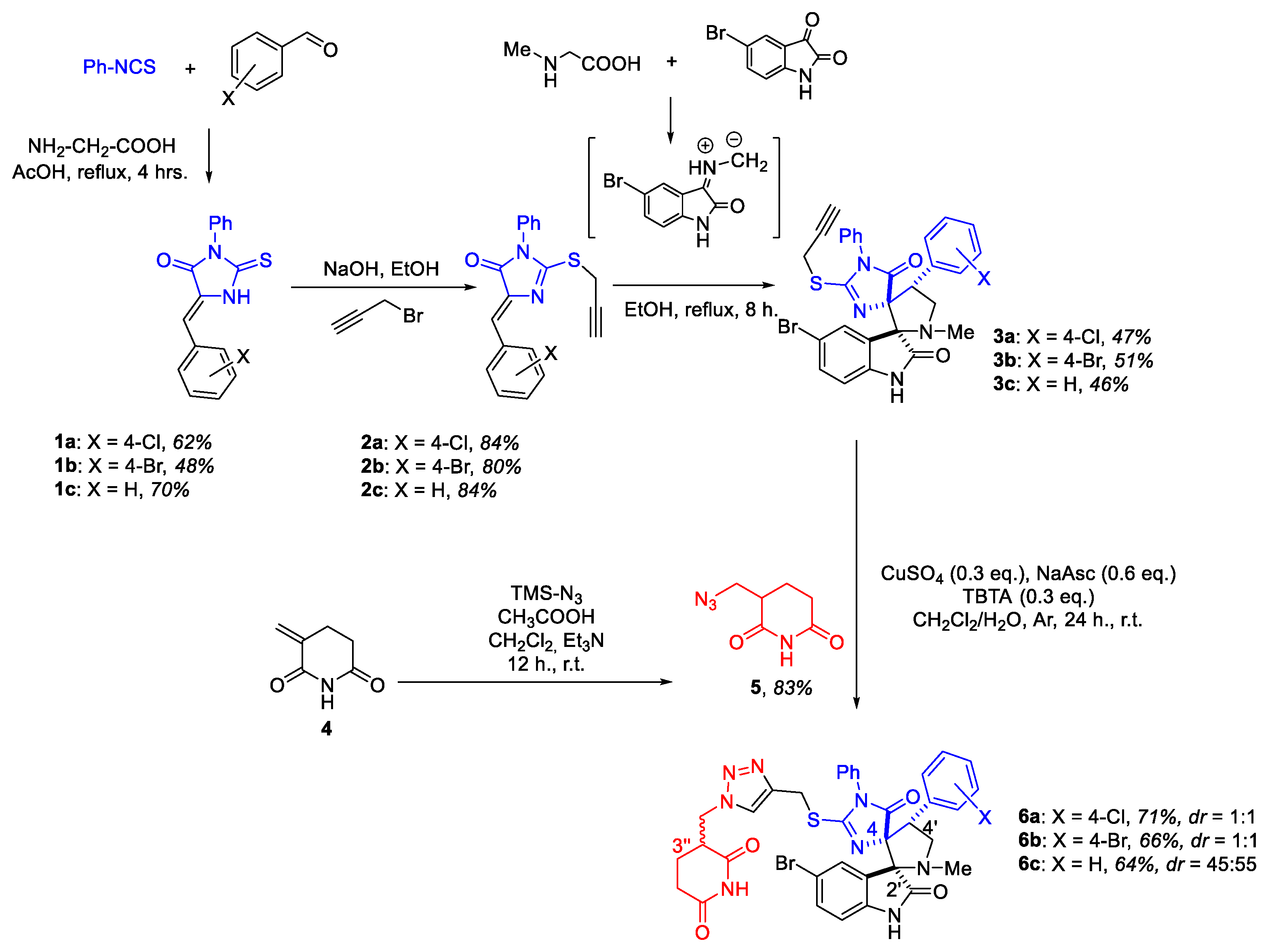
2.2. Biological Testing
3. Materials and Methods
3.1. General
3.2. Synthesis
3.3. Cell Culture and Treatment
3.3.1. Cell Culture and Cytotoxicity Assays
3.3.2. Statistics
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
References
- Levine, A.J. Targeting the P53 Protein for Cancer Therapies: The Translational Impact of P53 Research. Cancer Research 2022, 82, 362–364. [CrossRef]
- Aubrey, B.J.; Kelly, G.L.; Janic, A.; Herold, M.J.; Strasser, A. How does p53 induce apoptosis and how does this relate to p53-mediated tumour suppression? Cell Death & Differentiation 2018, 25, 104–113. [CrossRef]
- Wu, D.; Prives, C. Relevance of the p53–MDM2 axis to aging. Cell Death & Differentiation 2018, 25, 169–179. [CrossRef]
- Zhu, H.; Gao, H.; Ji, Y.; Zhou, Q.; Du, Z.; Tian, L.; Jiang, Y.; Yao, K.; Zhou, Z. Targeting p53–MDM2 interaction by small-molecule inhibitors: learning from MDM2 inhibitors in clinical trials. J Hematol Oncol. 2022, 15, 91. [CrossRef]
- Beloglazkina, A.; Zyk, N.; Majouga, A.; Beloglazkina E. Recent small-molecule inhibitors of the p53–mdm2 protein–protein interaction. Molecules 2020, 25, 2011–2027. [CrossRef]
- Synoradzki, K.J.; Bartnik, E.; Czarnecka, A.M.; Fiedorowicz, M.; Firlej, W.; Brodziak, A.; Stasinska, A.; Rutkowski, P.; Grieb, P. TP53 in Biology and Treatment of Osteosarcoma. Cancers 2021, 13, 4284. [CrossRef]
- Konopleva, M.; Martinelli, G.; Daver, N.; Papayannidis, C.; Wei, A.; Higgins, B.; Ott, M.; Mascarenhas J.; Andreeff, M. MDM2 inhibition: an important step forward in cancer therapy. Leukemia 2020, 34, 2858–2874. [CrossRef]
- Patel, K.R.; Patel, H.D. p53: An Attractive Therapeutic Target for Cancer. Curr Med Chem 2020, 27, 3706–3734. [CrossRef]
- Ivanenkov, Y.A.; Vasilevski, S.V.; Beloglazkina, E.K.; Kukushkin, M.E.; Machulkin, A.E.; Veselov, M.S.; Chufarova, N.V.; Chernyagina, E.S.; Vanzcool, A.S.; Zyk, N.V.; Skvortsov, D.A.; Khutornenko, A.A.; Rusanov, A.L.; Tonevitsky, A.G.; Dontsova, O.A.; Majouga, A.G. Design, synthesis and biological evaluation of novel potent MDM2/p53 small-molecule inhibitors. Bioorg Med Chem Lett 2015, 404–409. [CrossRef]
- Bora, D.; Kaushal, A., Shankaraiah, N. Anticancer potential of spirocompounds in medicinal chemistry: A pentennial expedition. Eur J Med Chem 2021, 215, 113263. [CrossRef]
- Arnhold, V.; Schmelz, K.; Proba. J.; Winkler, A.; Wünschel, J.; Toedling, J.; Deubzer, H.E.; Künkele, A.; Eggert, A.; Schulte, J.H.; Hundsdoerfer, P. Reactivating TP53 signaling by the novel MDM2 inhibitor DS-3032b as a therapeutic option for high-risk neuroblastoma. Oncotarget 2017, 2, 2304-2319. [CrossRef]
- Gomez-Monterrey, I.; Bertamino, A.; Porta, A.; Carotenuto, A.; Musella, S.; Aquino, C.; Granata, I.; Sala, M.; Brancaccio, D.; Picone, D.; Ercole, C.; Stiuso, P.; Campiglia, P.; Grieco, P.; Ianelli, P.; Maresca, B.; Novellino, E. Identification of the Spiro(oxindole-3,3′-thiazolidine)-Based Derivatives as Potential p53 Activity Modulators. J Med Chem 2010, 53, 8319–8329. [CrossRef]
- Wang, C.; Zhang, Y.; Chen, W.; Wu, Y.; Xing D. New-generation advanced PROTACs as potential therapeutic agents in cancer therapy. Mol Cancer 2024, 23, 110. [CrossRef]
- Yang, G.; Zhong, H.; Xia, X.; Qb, Z.; Wang, C.; Li, S. Potential application of proteolysis targeting chimera (PROTAC) modification technology in natural products for their targeted protein degradation. Food Sci Human Wellness 2022, 11, 199–207. [CrossRef]
- He, S.; Ma, J.; Fang, Y.; Liu, Y.; Wu, S.; Dong, G.; Wang, W.; Sheng, C. Homo-PROTAC mediated suicide of MDM2 to treat non-small cell lung cancer. Acta Pharma Sinica B 2021, 11, 1617–1628. [CrossRef]
- Powell, C.E.; Du, G.; Bushman, J.W.; He, Z.; Zhang, T.; Fischer, E.S.; Gray, N.S. Selective degradation-inducing probes for studying cereblon (CRBN) biology. RSC Med Chem 2021, 12, 1381–1390. [CrossRef]
- Alabi, S.B.; Crews, C.M. Major advances in targeted protein degradation: PROTACs, LYTACs, and MADTACs. J Biol Chem. 2021, 296, 100647. [CrossRef]
- Li, Y.; Yang, J.; Aguilar, A.; McEachern, D.; Przybranowski, S.; Liu, L.; Yang, C.-Y.; Wang, M.; Han, X.; Wang, S. Discovery of MD-224 as a First-in-Class, Highly Potent, and Efficacious Proteolysis Targeting Chimera Murine Double Minute 2 Degrader Capable of Achieving Complete and Durable Tumor Regression. J Med Chem 2019, 62, 448–466. [CrossRef]
- Anifowose, A.; Agbowuro, A.A.; Yang, X.; Wang, B. Anticancer strategies by upregulating p53 through inhibition of its ubiquitination by MDM2. Med Chem Res 2020, 29, 1105–1121. [CrossRef]
- Wang, B.; Wu, S.; Liu, J.; Yang, K.; Xie, H.; Tang W. Development of selective small molecule MDM2 degraders based on nutlin. Eur J Med Chem 2019,176, 476–491. [CrossRef]
- Wang, B.; Liu, J.; Tandon, I.; Wu, S.; Teng, P.; Liao, J.; Tang, W. Development of MDM2 degraders based on ligands derived from Ugi reactions: Lessons and discoveries. Eur J Med Chem 2021, 219, 113425. [CrossRef]
- Ito, T.; Ando, H.; Suzuki, T.; Ogura, T.; Hotta, K.; Imamura, Y.; Yamaguchi, Y.; Handa, H. Identification of a Primary Target of Thalidomide Teratogenicity. Science 2010, 327, 1345–1350. [CrossRef]
- Angers, S.; Li, T.; Yi, X.; MacCoss, M.J.; Moon, R.T.; Zheng, N. Molecular architecture and assembly of the DDB1–CUL4A ubiquitin ligase machinery. Nature 2006, 443, 590–593. [CrossRef]
- Ottis, P.; Toure, M.; Cromm, P.M.; Ko, E.; Gustafson, J.L.; Crews, C.M. Assessing Different E3 Ligases for Small Molecule Induced Protein Ubiquitination and Degradation. ACS Chem Biol 2017, 12, 2570–2578. [CrossRef]
- Petzold, G.; Fischer, E.S.; Thomä, N.H. Structural basis of lenalidomide-induced CK1α degradation by the CRL4CRBN ubiquitin ligase. Nature 2016, 532, 127–130. [CrossRef]
- Matyskiela, M.E.; Lu, G.; Ito, T.; Pagarigan, B.; Lu, C.-C.; Miller, K.; Fang, W.; Wang, N.-Y.; Nguyen, D.; Houston, J.; Carmel, G.; Tran, T.; Riley, M.; Nosaka, L.; Lander, G.C.; Gaidarova, S.; Xu, S.; Ruchelman, A.L.; Handa, H.; Carmichael, J.; Daniel, T.O.; Cathers, B.E.; Lopez-Girona A.; Chamberlain P.P. A novel cereblon modulator recruits GSPT1 to the CRL4CRBN ubiquitin ligase. Nature 2016, 535, 252–257. [CrossRef]
- Winter, G.E.; Buckley, D.L.; Paulk, J.; Roberts, J.M.; Souza, A.; Dhe-Paganon, S.; Bradner, J.E. Phthalimide conjugation as a strategy for in vivo target protein degradation. Science 2015, 348, 1376–1381. [CrossRef]
- Lu, J.; Qian, Y.; Altieri, M.; Dong, H.; Wang, J.; Raina, K.; Hines, J.; Winkler, J.D.; Crew, A.P.; Coleman, K.; Crews, C.M. Hijacking the E3 Ubiquitin Ligase Cereblon to Efficiently Target BRD4. Chem Biol 2015, 22, 755–763. [CrossRef]
- Khuzhakhmetova, L.R.; Ananeva, A.A.; Kantin, G.P.; Dar’in, D.V.; Bunev, A.S.; Ebeling, S.; Herrmann, A.; Hartmann, M.D.; Kalinin, S.A.; Bakulina, O.Yu. Synthesis of α-(azidomethyl)glutarimide and its applicationin construction of potential Cereblon ligands via the CuAAC reaction. Mendeleev Commun 2025, 35, 69–72. [CrossRef]
- Kantin, G.; Golubev, P.; Sapegin, A.; Bunev, A.; Dar’in, D. N-Boc-α-diazo glutarimide as efficient reagent for assembling N-heterocycle-glutarimide diads via Rh(II)-catalyzed N–H insertion reaction. Beilstein J Org Chem 2023, 19, 1841–1848. [CrossRef]
- Neklesa, T.; Snyder, L.B.; Willard, R.R.; Vitale, N.; Pizzano, J.; Gordon, D.A.; Bookbinder, M.; Macaluso, J.; Dong, H.; Ferraro, C.; Wang, G.; Wang, J.; Crews, C.M.; Houston, J.; Crew, A.P.; Taylor, I. ARV-110: An oral androgen receptor PROTAC degrader for prostate cancer. J Clin Oncol 2019, 37(7_suppl), 259-259. [CrossRef]
- Gough, S.M.; Flanagan, J.J.; The, J.; Andreoli, M.; Rousseau, E.; Pannone, M.; Bookbinder, M.; Willard, R.; Davenport, K.; Bortolon, E.; Cadelina, G.; Gordon, D.; Pizzano, J.; Macaluso, J.; Soto, L.; Corradi, J.; Digianantonio, K.; Drulyte, I.; Morgan, A.; Quinn, C.; Békés, M.; Ferraro, C.; Chen, X.; Wang, G.; Dong, H.; Wang, J.; Langley, D.R.; Houston, J.; Gedrich, R.; Taylor, I.C. Oral Estrogen Receptor PROTAC Vepdegestrant (ARV-471) Is Highly Efficacious as Monotherapy and in Combination with CDK4/6 or PI3K/mTOR Pathway Inhibitors in Preclinical ER+ Breast Cancer Models. Clin Cancer Res 2024, 30, 3549–3563. [CrossRef]
- Bricelj, A.; Steinebach, C.; Kuchta, R.; Gütschow, M.; Sosič, I. E3 ligase ligands in successful PROTACs: an overview of syntheses and linker attachment points. Front Chem 2021, 9, 707317. [CrossRef]
- Majouga, A.G.; Beloglazkina, E.K.; Beloglazkina, A.A.; Kukushkin, M.E.; Ivanenkov, Ya.A.; Veselov, M.S. New dispiro-indolinones, inhibitors of mdm2/p53 interaction, method of production and use. Patent #RU2629750C2, 9 Apryl 2015.
- Ivanenkov, Y.A.; Kukushkin, M.E.; Beloglazkina, A.A.; Shafikov, R.R.; Barashkin, A.A.; Ayginin, A.A.; Serebryakova, M.V.; Majouga, A.G.; Skvortsov, D.A.; Tafeenko, V.A.; Beloglazkina, E.K. Synthesis and biological evaluation of novel dispiro-indolinones with anticancer activity. Molecules 2023, 28, 1325. [CrossRef]
- Beloglazkina, A.A.; Karpov, N.A.; Mefedova S.R.; Polyakov, V.S.; Skvortsov, D.A.; Kalinina, M.A.; Tafeenko, V.A.; Majouga, A.G.; Zyk N.V.; Beloglazkina E.K. Synthesis of dispirooxindoles containing N-unsubstituted heterocyclic moieties and study of their anticancer activity. Russ Chem Bull 2019, 68, 1006–1013. [CrossRef]
- Kukushkin, M.E.; Skvortsov, D.A.; Kalinina, M.A.; Tafeenko, V.A.; Burmistrov, V.V.; Butov, G.M.; Zyk, N.V.; Majouga, A.G.; Beloglazkina, E.K. Synthesis and cytotoxicity of oxindoles dispiro derivatives with thiohydantoin and adamantane fragments. Phosphorus Sulfur Silicon 2020, 7, 544–555. [CrossRef]
- Polyakov, V.S.; Pervakova, E.V.; Zyk, N.V.; Beloglazkina, E.K. Synthesis of bis(dispiro[indolinone-pyrrolidine-imidazolones]). Russ Chem Bull 2025, 74, 2092–2101. [CrossRef]
- Polyakov, V.S.; Pervakova, E.V.; Grishin, Yu.K.; Beloglazkina, E.K. Bis-dispiro-indolinone-pyrrolidine-imidazolones with polymethylene linkers between imidazolone fragments. Tetrahedron Lett 2025, 162, 155589. [CrossRef]
- Kuznetsova, O.Y.; Antipin, R.L.; Udina, A.V.; Krasnovskaya, O.O.; Beloglazkina, E.K.; Terenin, V.I.; Koteliansky, V E.; Zyk, N.V.; Majouga, A.G. An improved protocol for synthesis of 3-substituted 5-arylidene-2-thiohydantoins: Two-step procedure alternative to classical methods. J Heterocyclic Chem 2016, 53, 1570–1577. [CrossRef]
- Majouga, A.G.; Beloglazkina, E.K.; Vatsadze, S.Z.; Frolova, N.A.; Zyk, N.V. Synthesis of isomeric 3-phenyl-5-(pyridylmethylene)-2-thiohydantoins and their S-methylated derivatives. Molecular and crystal structures of (5Z)-3-phenyl-5-(pyridin-2-ylmethylene)-2-thiohydantoin and (5Z)-2-methylthio-3-phenyl-5-(pyridin-2-ylmethylene)-3,5-dihydro-4H-imidazol-4-one. Russ Chem Bull 2004, 53, 2850–2855. [CrossRef]
- Wanner, M.J.; Koomen, G.J. 2-Substituted Glutarimides via Preformed Wittig Reagents. Synthesis 1988, 4, 325-327. [CrossRef]
- Krasavin, M.; Adamchik, M.; Bubyrev, A.; Heim, C.; Maiwald, S.; Zhukovsky, D.; Zhmurov, P.; Bunev, A.; Hartmann, M.D. Synthesis of novel glutarimide ligands for the E3 ligase substrate receptor Cereblon (CRBN): Investigation of their binding mode and antiproliferative effects against myeloma cell lines. Eur. J. Med. Chem., 2023, 246, 114990. [CrossRef]
- V.S. Polyakov, A.A. Barashkin, Y.K. Grishin, E.A. Tarasenko, E.K. Beloglazkina. Synthesis of chiral dispiro-indolinone-pyrrolidine-imidazolones using (r) or(s)-1-(2,4-dimethoxyphenyl)ethyl amine. ChemistrySelect 2025, 10, e01276. [CrossRef]
- Dai, H., Liu, G., Zhang, X., Yan, H., Lu, C. Pyrrolylmethyl Functionalized o-Carborane Derivatives. Organometallics, 2016, 35(10), 1488–1496. [CrossRef]
- Post, E. A. J., Fletcher, S. P. Controlling the Kinetics of Self-Reproducing Micelles by Catalyst Compartmentalization in a Biphasic System. The Journal of Organic Chemistry, 2019, 84(5), 2741–2755. [CrossRef]
- S. Jaiswal, I. R. Siddiqui & Subhadra Rajput. A novel α-haloacid based clay-catalysed expeditious synthesis of pyrazoloimidazole-2-thione-N-nucleosides, International Journal of Scientific & Engineering Research, 2013, 4(9), 2093.
- Ding, M. W., Sun, Y., & Liu, Z. J. An Efficient Synthesis of 2-Alkylthio- 5-phenylmethylidene-4 H -imidazolin-4-ones. Synthetic Communications, 2003, 33(8), 1267–1274. [CrossRef]
- Sirivolu, V. R. S.; Vernekar, K. V.; Marchand, C.; Naumova, A.; Chergui, A.; Renaud, A.; Stephen, A. G. Chen, F.; Sham, Y. Y.; Pommier, Y.; Wang. Z. 5-Arylidenethioxothiazolidinones as inhibitors of tyrosyl–DNA phosphodiesterase I. Journal of Medicinal Chemistry, 2012, 55(20), 8671-8684. [CrossRef]
- Korohoda, M.J. Introduction of selenium into heterocyclic compounds. i: synthesis of 3-aryl-2-selenohydantoins with double bond at c-5. Polish J. Chem., 1980, 54(4), 6830692.
- Guk, D. A.; Krasnovskaya, O. O.; Zyk, N. V.; Beloglazkina, E. K. Convenient synthesis of 2-thioimidazolone/menadione conjugates via a two-step sequence starting with direct amination of menadione. SynOpen, 2020, 04(02), 38–43. [CrossRef]
- Tsymbal, S.A.; Moiseeva, A.A.; Agadzhanian, N.A.; Efimova, S.S.; Markova, A.A.; Guk, D.A.; Krasnovskaya, O.O.; Alpatova, V.M.; Zaitsev, A.V.; Shibaeva, A.V.; et al. Copper-containing nanoparticles and organic complexes: Metal reduction triggers rapid cell death via oxidative burst. Int. J. Mol. Sci. 2021, 22, 11065. [CrossRef]
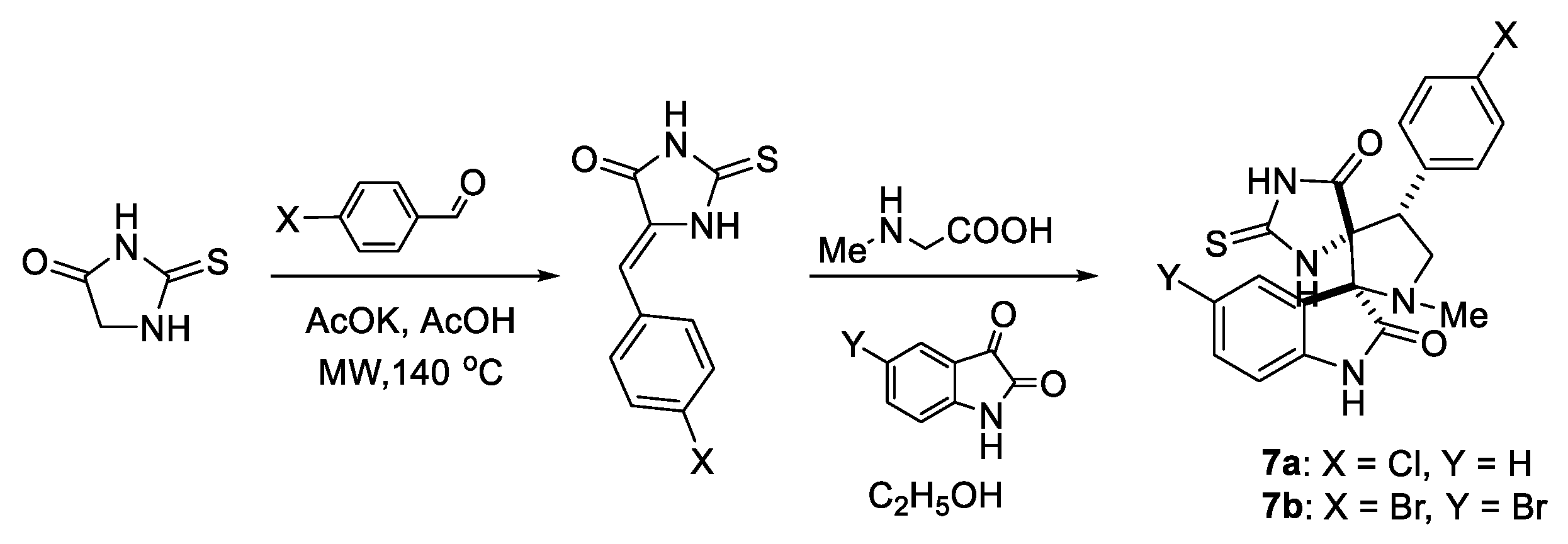
| Compound | Mean IC50, μM |
|---|---|
| 6a | 11.2+2.3 |
| 6b | 12.5+3.4 |
| 6c | 22.5+3.3 |
| 7a | 44.3+3.2 |
| 7b | 13.0+2.3 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).