Submitted:
15 March 2026
Posted:
16 March 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Evolutionary History
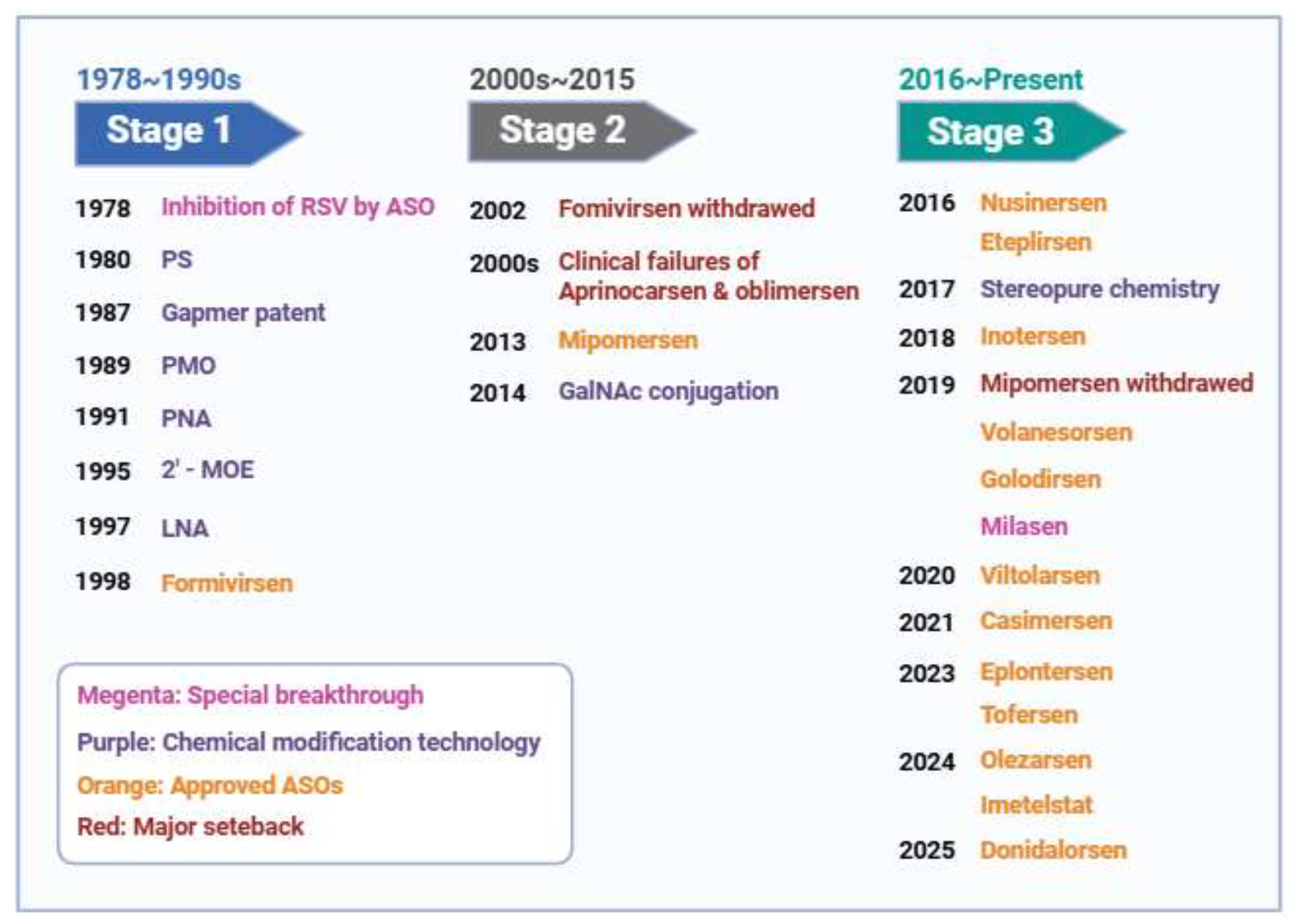
2.1. Inception and Preliminary Validation (1978–1999)
2.2. Setbacks and Technological Reshaping (2000–2015)
2.3. Clinical Explosion and the Golden Age (2016–Present)
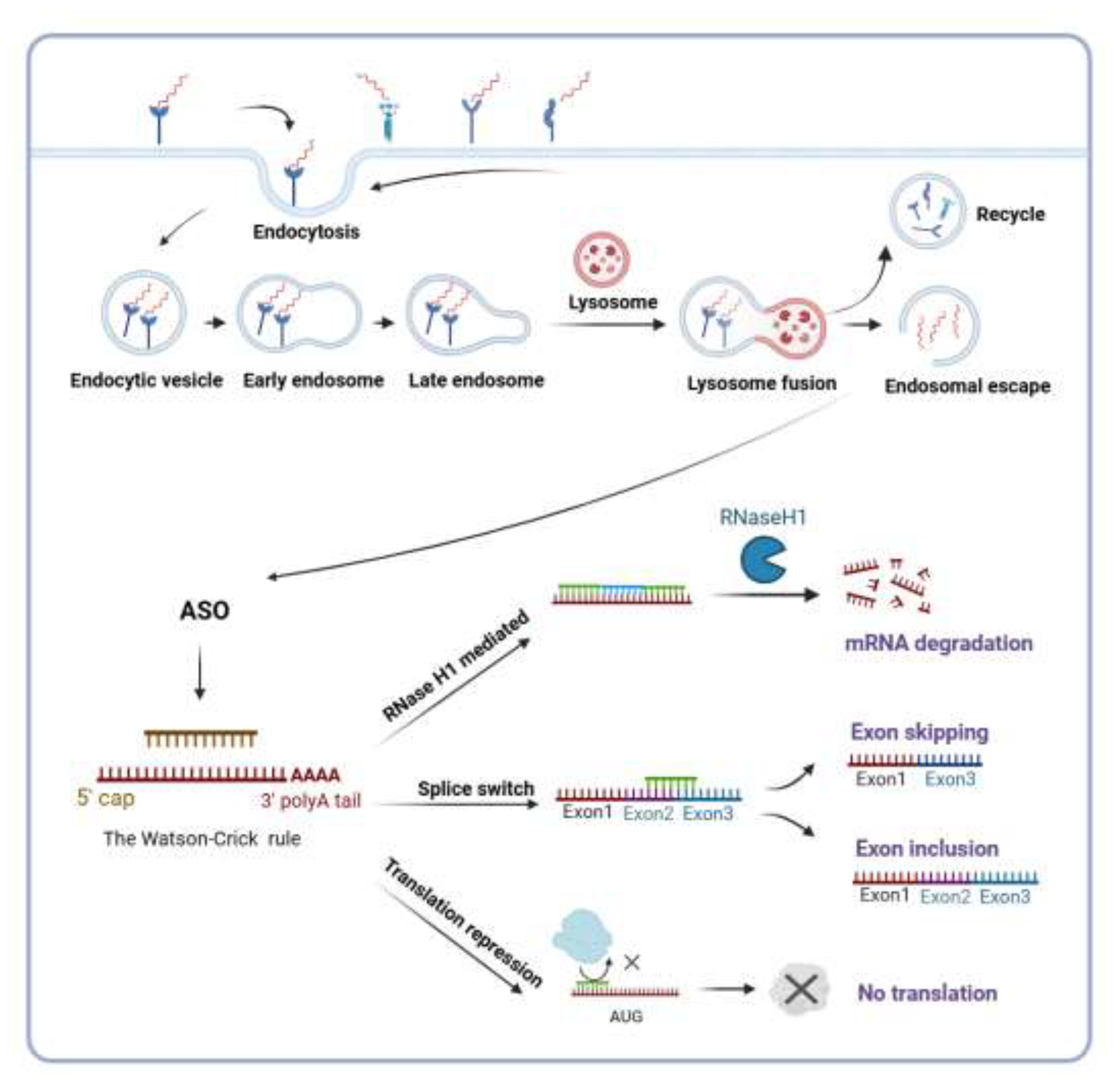
3. Mechanisms of Action
3.1. Pre-hybridization: Cellular Uptake and Trafficking
3.2. Hybridization: Target RNA Recognition
3.3. Post-hybridization: Functional Modulation of Target RNA
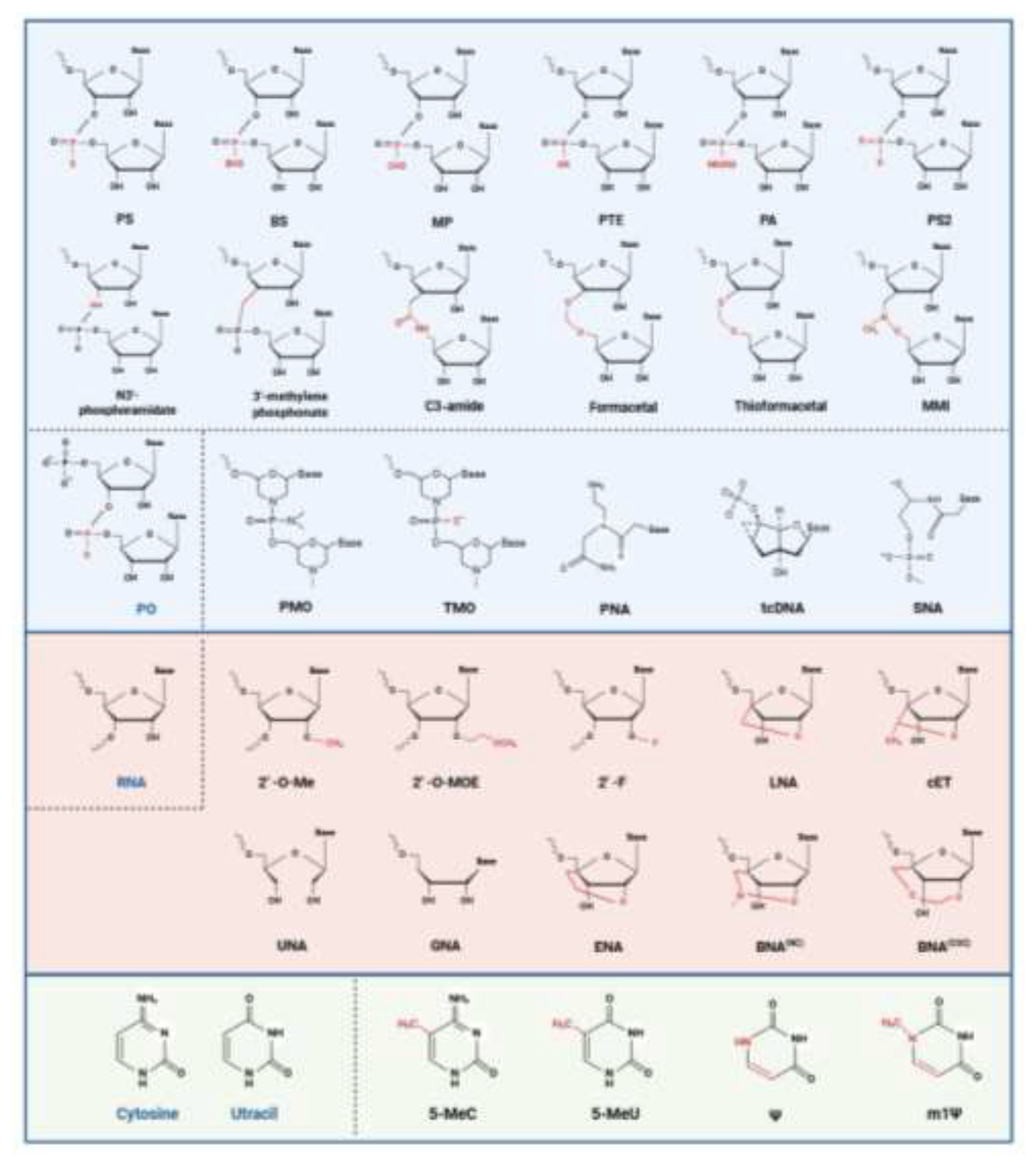
4. Chemical Modifications
4.1. Backbone Modifications
4.2. Sugar Modifications
4.3. Nucleobase Modification
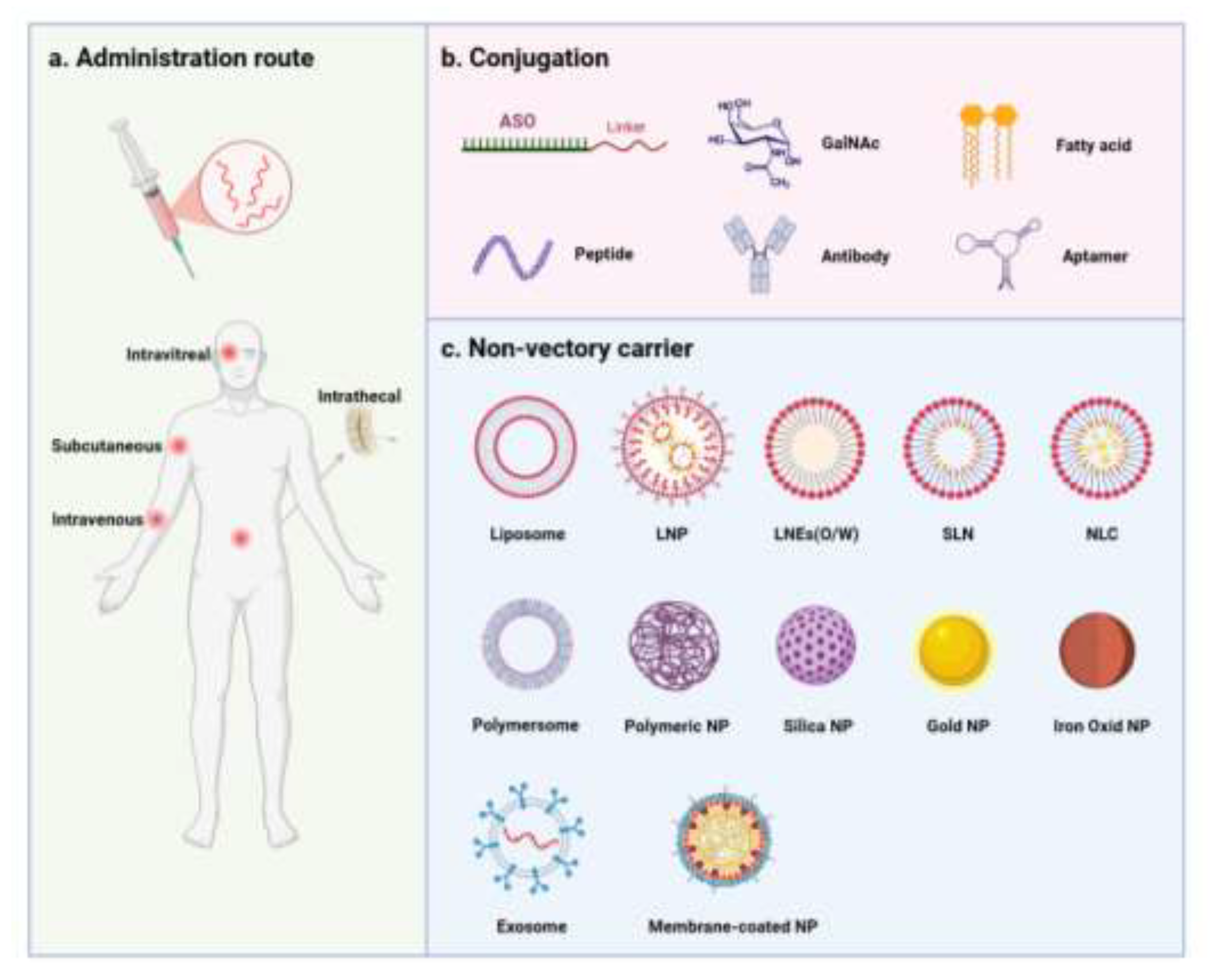
5. Delivery Strategies
5.1. Naked ASOs
5.2. Conjugate-Based Delivery
5.3. Carrier-Based Delivery
6. Clinical Translation Landscape
6.1 Insights from Approved Drugs
6.2. Lessons from Failed Attempts
6.3. Trends in the Clinical Pipeline
7. Challenges and Perspectives
7.1 Challenges
7.2. Future Directions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
Abbreviations
| ASOs | Antisense oligonucleotides |
| mRNA | messenger RNA |
| ncRNA | non-coding RNA |
| siRNAs | small interfering RNAs |
| miRNAs | microRNAs |
| saRNAs | small activating RNAs |
| PS | phosphorothioate |
| FDA | Food and Drug Administration |
| 2′-MOE | 2′-O-methoxyethyl |
| GalNAc | N-acetylgalactosamine |
References
- K.D. Warner, C.E. Hajdin, K.M. Weeks, Principles for targeting RNA with drug-like small molecules, Nat Rev Drug Discov 17(8) (2018) 547-558. [CrossRef]
- Z. Cai, H. Ma, F. Ye, D. Lei, Z. Deng, Y. Li, R. Gu, H. Wen, Discovery of RNA-Targeting Small Molecules: Challenges and Future Directions, MedComm (2020) 6(9) (2025) e70342. [CrossRef]
- M. Liu, Y. Wang, Y. Zhang, D. Hu, L. Tang, B. Zhou, L. Yang, Landscape of small nucleic acid therapeutics: moving from the bench to the clinic as next-generation medicines, Signal Transduct Target Ther 10(1) (2025) 73. [CrossRef]
- C.M. Ballantyne, S. Vasas, M. Azizad, P. Clifton, R.S. Rosenson, T. Chang, S. Melquist, R. Zhou, M. Mushin, N.J. Leeper, J. Hellawell, D. Gaudet, Plozasiran, an RNA Interference Agent Targeting APOC3, for Mixed Hyperlipidemia, N Engl J Med 391(10) (2024) 899-912. [CrossRef]
- A. Mullard, FDA approves anti-prekallikrein drug for hereditary angioedema, Nat Rev Drug Discov 24(10) (2025) 732. [CrossRef]
- C.F. Bennett, Therapeutic Antisense Oligonucleotides Are Coming of Age, Annu Rev Med 70 (2019) 307-321. [CrossRef]
- P.C. Zamecnik, M.L. Stephenson, Inhibition of Rous sarcoma virus replication and cell transformation by a specific oligodeoxynucleotide, Proc Natl Acad Sci U S A 75(1) (1978) 280-4. [CrossRef]
- M.L. Stephenson, P.C. Zamecnik, Inhibition of Rous sarcoma viral RNA translation by a specific oligodeoxyribonucleotide, Proc Natl Acad Sci U S A 75(1) (1978) 285-8. [CrossRef]
- S.T. Crooke, T.A. Vickers, X.H. Liang, Phosphorothioate modified oligonucleotide-protein interactions, Nucleic Acids Res 48(10) (2020) 5235-5253. [CrossRef]
- M.D. de Smet, C.J. Meenken, G.J. van den Horn, Fomivirsen - a phosphorothioate oligonucleotide for the treatment of CMV retinitis, Ocul Immunol Inflamm 7(3-4) (1999) 189-98. [CrossRef]
- R.S. Geary, S.P. Henry, L.R. Grillone, Fomivirsen: clinical pharmacology and potential drug interactions, Clin Pharmacokinet 41(4) (2002) 255-60.
- P.H. Hagedorn, B.R. Hansen, T. Koch, M. Lindow, Managing the sequence-specificity of antisense oligonucleotides in drug discovery, Nucleic Acids Res 45(5) (2017) 2262-2282. [CrossRef]
- M. Egli, M. Manoharan, Chemistry, structure and function of approved oligonucleotide therapeutics, Nucleic Acids Res 51(6) (2023) 2529-2573. [CrossRef]
- P. Hair, F. Cameron, K. McKeage, Mipomersen sodium: first global approval, Drugs 73(5) (2013) 487-93. [CrossRef]
- D.J. Blom, F.J. Raal, R.D. Santos, A.D. Marais, Lomitapide and Mipomersen-Inhibiting Microsomal Triglyceride Transfer Protein (MTP) and apoB100 Synthesis, Curr Atheroscler Rep 21(12) (2019) 48. [CrossRef]
- K. Paunovska, D. Loughrey, J.E. Dahlman, Drug delivery systems for RNA therapeutics, Nat Rev Genet 23(5) (2022) 265-280. [CrossRef]
- R. Obexer, M. Nassir, E.R. Moody, P.S. Baran, S.L. Lovelock, Modern approaches to therapeutic oligonucleotide manufacturing, Science 384(6692) (2024) eadl4015. [CrossRef]
- F. Alhamadani, K. Zhang, R. Parikh, H. Wu, T.P. Rasmussen, R. Bahal, X.B. Zhong, J.E. Manautou, Adverse Drug Reactions and Toxicity of the Food and Drug Administration-Approved Antisense Oligonucleotide Drugs, Drug Metab Dispos 50(6) (2022) 879-887. [CrossRef]
- E. Mercuri, B.T. Darras, C.A. Chiriboga, J.W. Day, C. Campbell, A.M. Connolly, S.T. Iannaccone, J. Kirschner, N.L. Kuntz, K. Saito, P.B. Shieh, M. Tulinius, E.S. Mazzone, J. Montes, K.M. Bishop, Q. Yang, R. Foster, S. Gheuens, C.F. Bennett, W. Farwell, E. Schneider, D.C. De Vivo, R.S. Finkel, Nusinersen versus Sham Control in Later-Onset Spinal Muscular Atrophy, N Engl J Med 378(7) (2018) 625-635. [CrossRef]
- E.S.G. Stroes, V.J. Alexander, E. Karwatowska-Prokopczuk, R.A. Hegele, M. Arca, C.M. Ballantyne, H. Soran, T.A. Prohaska, S. Xia, H.N. Ginsberg, J.L. Witztum, S. Tsimikas, Olezarsen, Acute Pancreatitis, and Familial Chylomicronemia Syndrome, N Engl J Med 390(19) (2024) 1781-1792.
- T. Coelho, W. Marques, Jr., N.R. Dasgupta, C.C. Chao, Y. Parman, M.C. França, Jr., Y.C. Guo, J. Wixner, L.S. Ro, C.R. Calandra, P.A. Kowacs, J.L. Berk, L. Obici, F.A. Barroso, M. Weiler, I. Conceição, S.W. Jung, G. Buchele, M. Brambatti, J. Chen, S.G. Hughes, E. Schneider, N.J. Viney, A. Masri, M.R. Gertz, Y. Ando, J.D. Gillmore, S. Khella, P.J.B. Dyck, M. Waddington Cruz, Eplontersen for Hereditary Transthyretin Amyloidosis With Polyneuropathy, Jama 330(15) (2023) 1448-1458. [CrossRef]
- R.L. Juliano, The delivery of therapeutic oligonucleotides, Nucleic Acids Res 44(14) (2016) 6518-48.
- V. Kumar, W.B. Turnbull, Targeted delivery of oligonucleotides using multivalent protein-carbohydrate interactions, Chem Soc Rev 52(4) (2023) 1273-1287. [CrossRef]
- C.M. Miller, A.J. Donner, E.E. Blank, A.W. Egger, B.M. Kellar, M.E. Østergaard, P.P. Seth, E.N. Harris, Stabilin-1 and Stabilin-2 are specific receptors for the cellular internalization of phosphorothioate-modified antisense oligonucleotides (ASOs) in the liver, Nucleic Acids Res 44(6) (2016) 2782-94. [CrossRef]
- A.J. Debacker, J. Voutila, M. Catley, D. Blakey, N. Habib, Delivery of Oligonucleotides to the Liver with GalNAc: From Research to Registered Therapeutic Drug, Mol Ther 28(8) (2020) 1759-1771. [CrossRef]
- S.F. Dowdy, Endosomal escape of RNA therapeutics: How do we solve this rate-limiting problem?, Rna 29(4) (2023) 396-401. [CrossRef]
- X.H. Liang, H. Sun, W. Shen, S.T. Crooke, Identification and characterization of intracellular proteins that bind oligonucleotides with phosphorothioate linkages, Nucleic Acids Res 43(5) (2015) 2927-45. [CrossRef]
- E. Bäckström, A. Bonetti, P. Johnsson, S. Öhlin, A. Dahlén, P. Andersson, S. Andersson, P. Gennemark, Tissue pharmacokinetics of antisense oligonucleotides, Mol Ther Nucleic Acids 35(1) (2024) 102133. [CrossRef]
- B.T. Le, S. Chen, R.N. Veedu, Rational Design of Chimeric Antisense Oligonucleotides on a Mixed PO-PS Backbone for Splice-Switching Applications, Biomolecules 14(7) (2024). [CrossRef]
- N. Iwamoto, D.C.D. Butler, N. Svrzikapa, S. Mohapatra, I. Zlatev, D.W.Y. Sah, Meena, S.M. Standley, G. Lu, L.H. Apponi, M. Frank-Kamenetsky, J.J. Zhang, C. Vargeese, G.L. Verdine, Control of phosphorothioate stereochemistry substantially increases the efficacy of antisense oligonucleotides, Nat Biotechnol 35(9) (2017) 845-851. [CrossRef]
- R.S. Geary, D. Norris, R. Yu, C.F. Bennett, Pharmacokinetics, biodistribution and cell uptake of antisense oligonucleotides, Adv Drug Deliv Rev 87 (2015) 46-51. [CrossRef]
- T.A. Vickers, J.R. Wyatt, S.M. Freier, Effects of RNA secondary structure on cellular antisense activity, Nucleic Acids Res 28(6) (2000) 1340-7. [CrossRef]
- M.E. Rogalska, E. Mancini, S. Bonnal, A. Gohr, B.M. Dunyak, N. Arecco, P.G. Smith, F.H. Vaillancourt, J. Valcárcel, Transcriptome-wide splicing network reveals specialized regulatory functions of the core spliceosome, Science 386(6721) (2024) 551-560. [CrossRef]
- C. Terada, K. Oh, R. Tsubaki, B. Chan, N. Aibara, K. Ohyama, M.A. Shibata, T. Wada, M. Harada-Shiba, A. Yamayoshi, T. Yamamoto, Dynamic and static control of the off-target interactions of antisense oligonucleotides using toehold chemistry, Nat Commun 14(1) (2023) 7972. [CrossRef]
- X. Shen, D.R. Corey, Chemistry, mechanism and clinical status of antisense oligonucleotides and duplex RNAs, Nucleic Acids Res 46(4) (2018) 1584-1600. [CrossRef]
- X.H. Liang, H. Sun, J.G. Nichols, S.T. Crooke, RNase H1-Dependent Antisense Oligonucleotides Are Robustly Active in Directing RNA Cleavage in Both the Cytoplasm and the Nucleus, Mol Ther 25(9) (2017) 2075-2092. [CrossRef]
- J. Scharner, I. Aznarez, Clinical Applications of Single-Stranded Oligonucleotides: Current Landscape of Approved and In-Development Therapeutics, Mol Ther 29(2) (2021) 540-554. [CrossRef]
- L. Torres-Masjoan, S. Aguti, H. Zhou, F. Muntoni, Clinical applications of exon-skipping antisense oligonucleotides in neuromuscular diseases, Mol Ther 33(6) (2025) 2689-2704. [CrossRef]
- X.H. Liang, W. Shen, H. Sun, M.T. Migawa, T.A. Vickers, S.T. Crooke, Translation efficiency of mRNAs is increased by antisense oligonucleotides targeting upstream open reading frames, Nat Biotechnol 34(8) (2016) 875-80. [CrossRef]
- T. Merkle, S. Merz, P. Reautschnig, A. Blaha, Q. Li, P. Vogel, J. Wettengel, J.B. Li, T. Stafforst, Precise RNA editing by recruiting endogenous ADARs with antisense oligonucleotides, Nat Biotechnol 37(2) (2019) 133-138. [CrossRef]
- V. Genna, G. Portella, A. Sala, M. Terrazas, I. Serrano-Chacón, J. González, N. Villegas, L. Mateo, C. Castellazzi, M. Labrador, A. Aviño, A. Hospital, A. Gandioso, P. Aloy, I. Brun-Heath, C. Gonzalez, R. Eritja, M. Orozco, Systematic study of hybrid triplex topology and stability suggests a general triplex-mediated regulatory mechanism, Nucleic Acids Res 53(5) (2025). [CrossRef]
- A.M. Vanderplow, G.E. Dodis, Y. Rhee, J.J. Cikowski, S. Gonzalez, M.L. Smith, R.G. Gogliotti, Site-blocking antisense oligonucleotides as a mechanism to fine-tune MeCP2 expression, Rna 30(12) (2024) 1554-1571. [CrossRef]
- J. Bhamra, M. Krishna, G. Samaan, S. Pattanayak, S. Mukhopadhyay, Toxicity of Antisense Oligonucleotides is Determined by the Synergistic Interplay of Chemical Modifications and Nucleotide Sequences, Not by Either Factor Alone, Chembiochem 26(20) (2025) e202500584. [CrossRef]
- C.R. Hofman, D.R. Corey, Targeting RNA with synthetic oligonucleotides: Clinical success invites new challenges, Cell Chem Biol 31(1) (2024) 125-138. [CrossRef]
- P. Kandasamy, G. McClorey, M. Shimizu, N. Kothari, R. Alam, N. Iwamoto, J. Kumarasamy, G.R. Bommineni, A. Bezigian, O. Chivatakarn, D.C.D. Butler, M. Byrne, K. Chwalenia, K.E. Davies, J. Desai, J.D. Shelke, A.F. Durbin, R. Ellerington, B. Edwards, J. Godfrey, A. Hoss, F. Liu, K. Longo, G. Lu, S. Marappan, J. Oieni, I.H. Paik, E.P. Estabrook, C. Shivalila, M. Tischbein, T. Kawamoto, C. Rinaldi, J. Rajão-Saraiva, S. Tripathi, H. Yang, Y. Yin, X. Zhao, C. Zhou, J. Zhang, L. Apponi, M.J.A. Wood, C. Vargeese, Control of backbone chemistry and chirality boost oligonucleotide splice switching activity, Nucleic Acids Res 50(10) (2022) 5443-5466. [CrossRef]
- Y. Takahashi, K. Sato, T. Wada, Solid-Phase Synthesis of Boranophosphate/Phosphorothioate/Phosphate Chimeric Oligonucleotides and Their Potential as Antisense Oligonucleotides, J Org Chem 87(6) (2022) 3895-3909. [CrossRef]
- Patutina, S.K. Gaponova Miroshnichenko, A.V. Sen'kova, I.A. Savin, D.V. Gladkikh, E.A. Burakova, A.A. Fokina, M.A. Maslov, E.V. Shmendel, M.J.A. Wood, V.V. Vlassov, S. Altman, D.A. Stetsenko, M.A. Zenkova, Mesyl phosphoramidate backbone modified antisense oligonucleotides targeting miR-21 with enhanced in vivo therapeutic potency, Proc Natl Acad Sci U S A 117(51) (2020) 32370-32379. [CrossRef]
- Y.Y. Syed, Eteplirsen: First Global Approval, Drugs 76(17) (2016) 1699-1704. [CrossRef]
- B.T. Le, S. Paul, K. Jastrzebska, H. Langer, M.H. Caruthers, R.N. Veedu, Thiomorpholino oligonucleotides as a robust class of next generation platforms for alternate mRNA splicing, Proc Natl Acad Sci U S A 119(36) (2022) e2207956119. [CrossRef]
- V. MacLelland, M. Kravitz, A. Gupta, Therapeutic and diagnostic applications of antisense peptide nucleic acids, Mol Ther Nucleic Acids 35(1) (2024) 102086. [CrossRef]
- H. Asanuma, Y. Kamiya, H. Kashida, K. Murayama, Xeno nucleic acids (XNAs) having non-ribose scaffolds with unique supramolecular properties, Chem Commun (Camb) 58(25) (2022) 3993-4004. [CrossRef]
- K. Murayama, Y. Yamano, H. Asanuma, 8-Pyrenylvinyl Adenine Controls Reversible Duplex Formation between Serinol Nucleic Acid and RNA by [2 + 2] Photocycloaddition, J Am Chem Soc 141(24) (2019) 9485-9489. [CrossRef]
- T. Yoshida, T. Hagihara, Y. Uchida, Y. Horiuchi, K. Sasaki, T. Yamamoto, T. Yamashita, Y. Goda, Y. Saito, T. Yamaguchi, S. Obika, S. Yamamoto, T. Inoue, Introduction of sugar-modified nucleotides into CpG-containing antisense oligonucleotides inhibits TLR9 activation, Sci Rep 14(1) (2024) 11540. [CrossRef]
- S.T. Crooke, B.F. Baker, R.M. Crooke, X.H. Liang, Antisense technology: an overview and prospectus, Nat Rev Drug Discov 20(6) (2021) 427-453. [CrossRef]
- S.T. Crooke, B.F. Baker, J.L. Witztum, T.J. Kwoh, N.C. Pham, N. Salgado, B.W. McEvoy, W. Cheng, S.G. Hughes, S. Bhanot, R.S. Geary, The Effects of 2'-O-Methoxyethyl Containing Antisense Oligonucleotides on Platelets in Human Clinical Trials, Nucleic Acid Ther 27(3) (2017) 121-129. [CrossRef]
- H. Abou Assi, A.K. Rangadurai, H. Shi, B. Liu, M.C. Clay, K. Erharter, C. Kreutz, C.L. Holley, H.M. Al-Hashimi, 2'-O-Methylation can increase the abundance and lifetime of alternative RNA conformational states, Nucleic Acids Res 48(21) (2020) 12365-12379. [CrossRef]
- L. Sheng, F. Rigo, C.F. Bennett, A.R. Krainer, Y. Hua, Comparison of the efficacy of MOE and PMO modifications of systemic antisense oligonucleotides in a severe SMA mouse model, Nucleic Acids Res 48(6) (2020) 2853-2865. [CrossRef]
- Y. Masaki, Y. Iriyama, H. Nakajima, Y. Kuroda, T. Kanaki, S. Furukawa, M. Sekine, K. Seio, Application of 2'-O-(2-N-Methylcarbamoylethyl) Nucleotides in RNase H-Dependent Antisense Oligonucleotides, Nucleic Acid Ther 28(5) (2018) 307-311. [CrossRef]
- T. Yamaguchi, H. Komine, T. Sugiura, R. Kumagai, T. Yoshida, K. Sasaki, T. Nakayama, H. Kamada, T. Inoue, S. Obika, Cycloalkane Incorporation Into the 2',4'-Bridge of Locked Nucleic Acid: Enhancing Nuclease Stability, Reducing Phosphorothioate Modifications, and Lowering Hepatotoxicity in Antisense Oligonucleotides, JACS Au 5(10) (2025) 5111-5120. [CrossRef]
- P.H. Hagedorn, R. Persson, E.D. Funder, N. Albæk, S.L. Diemer, D.J. Hansen, M.R. Møller, N. Papargyri, H. Christiansen, B.R. Hansen, H.F. Hansen, M.A. Jensen, T. Koch, Locked nucleic acid: modality, diversity, and drug discovery, Drug Discov Today 23(1) (2018) 101-114. [CrossRef]
- N. Papargyri, M. Pontoppidan, M.R. Andersen, T. Koch, P.H. Hagedorn, Chemical Diversity of Locked Nucleic Acid-Modified Antisense Oligonucleotides Allows Optimization of Pharmaceutical Properties, Mol Ther Nucleic Acids 19 (2020) 706-717. [CrossRef]
- T. Matsubayashi, K. Yoshioka, S.S. Lei Mon, M. Katsuyama, C. Jia, T. Yamaguchi, R.I. Hara, T. Nagata, O. Nakagawa, S. Obika, T. Yokota, Favorable efficacy and reduced acute neurotoxicity by antisense oligonucleotides with 2',4'-BNA/LNA with 9-(aminoethoxy)phenoxazine, Mol Ther Nucleic Acids 35(2) (2024) 102161.
- U.S. Haque, T. Yokota, Enhancing Antisense Oligonucleotide-Based Therapeutic Delivery with DG9, a Versatile Cell-Penetrating Peptide, Cells 12(19) (2023). [CrossRef]
- Y. Nan, Y.J. Zhang, Antisense Phosphorodiamidate Morpholino Oligomers as Novel Antiviral Compounds, Front Microbiol 9 (2018) 750. [CrossRef]
- X. Sun, S. Setrerrahmane, C. Li, J. Hu, H. Xu, Nucleic acid drugs: recent progress and future perspectives, Signal Transduct Target Ther 9(1) (2024) 316. [CrossRef]
- A.J. Pollak, L. Zhao, T.A. Vickers, I.J. Huggins, X.H. Liang, S.T. Crooke, Insights into innate immune activation via PS-ASO-protein-TLR9 interactions, Nucleic Acids Res 50(14) (2022) 8107-8126. [CrossRef]
- J. He, F. Seela, Propynyl groups in duplex DNA: stability of base pairs incorporating 7-substituted 8-aza-7-deazapurines or 5-substituted pyrimidines, Nucleic Acids Res 30(24) (2002) 5485-96. [CrossRef]
- J.I. Gyi, D. Gao, G.L. Conn, J.O. Trent, T. Brown, A.N. Lane, The solution structure of a DNA*RNA duplex containing 5-propynyl U and C; comparison with 5-Me modifications, Nucleic Acids Res 31(10) (2003) 2683-93.
- A. Das, A. Ghosh, J. Kundu, M. Egli, M. Manoharan, S. Sinha, Synthesis and Biophysical Studies of High-Affinity Morpholino Oligomers Containing G-Clamp Analogs, J Org Chem 88(21) (2023) 15168-15175. [CrossRef]
- G. Fracchioni, S. Vailati, M. Grazioli, V. Pirota, Structural Unfolding of G-Quadruplexes: From Small Molecules to Antisense Strategies, Molecules 29(15) (2024). [CrossRef]
- L.P. Aiello, A.J. Brucker, S. Chang, E.T. Cunningham, Jr., D.J. D'Amico, H.W. Flynn, Jr., L.R. Grillone, S. Hutcherson, J.M. Liebmann, T.P. O'Brien, I.U. Scott, R.F. Spaide, C. Ta, M.T. Trese, Evolving guidelines for intravitreous injections, Retina 24(5 Suppl) (2004) S3-19. [CrossRef]
- J.M. Migliorati, S. Liu, A. Liu, A. Gogate, S. Nair, R. Bahal, T.P. Rasmussen, J.E. Manautou, X.B. Zhong, Absorption, Distribution, Metabolism, and Excretion of US Food and Drug Administration-Approved Antisense Oligonucleotide Drugs, Drug Metab Dispos 50(6) (2022) 888-897.
- D. Wu, Q. Chen, X. Chen, F. Han, Z. Chen, Y. Wang, The blood-brain barrier: structure, regulation, and drug delivery, Signal Transduct Target Ther 8(1) (2023) 217. [CrossRef]
- M. Monine, D. Norris, Y. Wang, I. Nestorov, A physiologically-based pharmacokinetic model to describe antisense oligonucleotide distribution after intrathecal administration, J Pharmacokinet Pharmacodyn 48(5) (2021) 639-654. [CrossRef]
- C.A. Chiriboga, K.J. Swoboda, B.T. Darras, S.T. Iannaccone, J. Montes, D.C. De Vivo, D.A. Norris, C.F. Bennett, K.M. Bishop, Results from a phase 1 study of nusinersen (ISIS-SMN(Rx)) in children with spinal muscular atrophy, Neurology 86(10) (2016) 890-7. [CrossRef]
- P. Anand, Y. Zhang, S. Patil, K. Kaur, Metabolic Stability and Targeted Delivery of Oligonucleotides: Advancing RNA Therapeutics Beyond The Liver, J Med Chem 68(7) (2025) 6870-6896. [CrossRef]
- M.F. Yuen, S.G. Lim, R. Plesniak, K. Tsuji, H.L.A. Janssen, C. Pojoga, A. Gadano, C.P. Popescu, T. Stepanova, T. Asselah, G. Diaconescu, H.J. Yim, J. Heo, E. Janczewska, A. Wong, N. Idriz, M. Imamura, G. Rizzardini, K. Takaguchi, P. Andreone, M. Arbune, J. Hou, S.J. Park, A. Vata, J. Cremer, R. Elston, T. Lukić, G. Quinn, L. Maynard, S. Kendrick, H. Plein, F. Campbell, M. Paff, D. Theodore, Efficacy and Safety of Bepirovirsen in Chronic Hepatitis B Infection, N Engl J Med 387(21) (2022) 1957-1968. [CrossRef]
- M.F. Yuen, J. Heo, J.W. Jang, J.H. Yoon, Y.O. Kweon, S.J. Park, Y. Tami, S. You, P. Yates, Y. Tao, J. Cremer, F. Campbell, R. Elston, D. Theodore, M. Paff, C.F. Bennett, T.J. Kwoh, Safety, tolerability and antiviral activity of the antisense oligonucleotide bepirovirsen in patients with chronic hepatitis B: a phase 2 randomized controlled trial, Nat Med 27(10) (2021) 1725-1734. [CrossRef]
- A. Vaillant, Bepirovirsen/GSK3389404: Antisense or TLR9 agonists?, J Hepatol 78(3) (2023) e107-e108. [CrossRef]
- M.F. Yuen, J. Heo, H. Kumada, F. Suzuki, Y. Suzuki, Q. Xie, J. Jia, Y. Karino, J. Hou, K. Chayama, M. Imamura, J.Y. Lao-Tan, S.G. Lim, Y. Tanaka, W. Xie, J.H. Yoon, Z. Duan, M. Kurosaki, S.J. Park, M.E. Labio, R. Kumar, Y.O. Kweon, H.J. Yim, Y. Tao, J. Cremer, R. Elston, M. Davies, S. Baptiste-Brown, K. Han, F.M. Campbell, M. Paff, D. Theodore, Phase IIa, randomised, double-blind study of GSK3389404 in patients with chronic hepatitis B on stable nucleos(t)ide therapy, J Hepatol 77(4) (2022) 967-977. [CrossRef]
- R.W. Hui, L.Y. Mak, J. Fung, W.K. Seto, M.F. Yuen, Prospect of emerging treatments for hepatitis B virus functional cure, Clin Mol Hepatol 31(Suppl) (2025) S165-181. [CrossRef]
- R.A. Goodnow, Reality check: lipid-oligonucleotide conjugates for therapeutic applications, Expert Opin Drug Discov 18(2) (2023) 129-134. [CrossRef]
- S. Shahnoor, I.F. Raza, M. Rashid, H. Panhwar, I. Fatima, S. Gul, K. Nadeem, S. Khan, M. Mahmmoud Fadelallah Eljack, FDA approval of imetelstat: a new era in the treatment of lower-risk myelodysplastic syndrome, Ann Med Surg (Lond) 87(12) (2025) 8385-8390. [CrossRef]
- U. Platzbecker, V. Santini, P. Fenaux, M.A. Sekeres, M.R. Savona, Y.F. Madanat, M. Díez-Campelo, D. Valcárcel, T. Illmer, A. Jonášová, P. Bělohlávková, L.J. Sherman, T. Berry, S. Dougherty, S. Shah, Q. Xia, L. Sun, Y. Wan, F. Huang, A. Ikin, S. Navada, F. Feller, R.S. Komrokji, A.M. Zeidan, Imetelstat in patients with lower-risk myelodysplastic syndromes who have relapsed or are refractory to erythropoiesis-stimulating agents (IMerge): a multinational, randomised, double-blind, placebo-controlled, phase 3 trial, Lancet 403(10423) (2024) 249-260. [CrossRef]
- T.P. Prakash, A.E. Mullick, R.G. Lee, J. Yu, S.T. Yeh, A. Low, A.E. Chappell, M.E. Østergaard, S. Murray, H.J. Gaus, E.E. Swayze, P.P. Seth, Fatty acid conjugation enhances potency of antisense oligonucleotides in muscle, Nucleic Acids Res 47(12) (2019) 6029-6044. [CrossRef]
- A.A. Balachandran, B.H. Poudel, K. Rahimizadeh, A. Chikkanna, R.N. Veedu, Enhancing the intracellular delivery of antisense oligonucleotides (ASO) : a comparative study of aptamer, vitamin E, and cholesterol ASO conjugates, RSC Adv 15(51) (2025) 43727-43736. [CrossRef]
- P. Mangla, Q. Vicentini, A. Biscans, Therapeutic Oligonucleotides: An Outlook on Chemical Strategies to Improve Endosomal Trafficking, Cells 12(18) (2023). [CrossRef]
- B. Malecova, R.S. Burke, M. Cochran, M.D. Hood, R. Johns, P.R. Kovach, V.R. Doppalapudi, G. Erdogan, J.D. Arias, B. Darimont, C.D. Miller, H. Huang, A. Geall, H.S. Younis, A.A. Levin, Targeted tissue delivery of RNA therapeutics using antibody-oligonucleotide conjugates (AOCs), Nucleic Acids Res 51(12) (2023) 5901-5910. [CrossRef]
- M. Cochran, D. Arias, R. Burke, D. Chu, G. Erdogan, M. Hood, P. Kovach, H.W. Kwon, Y. Chen, M. Moon, C.D. Miller, H. Huang, A. Levin, V.R. Doppalapudi, Structure-Activity Relationship of Antibody-Oligonucleotide Conjugates: Evaluating Bioconjugation Strategies for Antibody-siRNA Conjugates for Drug Development, J Med Chem 67(17) (2024) 14852-14867. [CrossRef]
- L.V. Gushchina, T.A. Vetter, E.C. Frair, A.J. Bradley, K.M. Grounds, J.W. Lay, N. Huang, A. Suhaiba, F.J. Schnell, G. Hanson, T.R. Simmons, N. Wein, K.M. Flanigan, Systemic PPMO-mediated dystrophin expression in the Dup2 mouse model of Duchenne muscular dystrophy, Mol Ther Nucleic Acids 30 (2022) 479-492. [CrossRef]
- W. Yang, C. Ran, X. Lian, Z. Wang, Z. Du, T. Bing, Y. Zhang, W. Tan, Aptamer-based targeted drug delivery and disease therapy in preclinical and clinical applications, Adv Drug Deliv Rev 226 (2025) 115680. [CrossRef]
- F. Millozzi, P. Milán-Rois, A. Sett, G. Delli Carpini, M. De Bardi, M. Gisbert-Garzarán, M. Sandonà, C. Rodríguez-Díaz, M. Martínez-Mingo, I. Pardo, F. Esposito, M.T. Viscomi, M. Bouché, O. Parolini, V. Saccone, J.J. Toulmé, Á. Somoza, D. Palacios, Aptamer-conjugated gold nanoparticles enable oligonucleotide delivery into muscle stem cells to promote regeneration of dystrophic muscles, Nat Commun 16(1) (2025) 577. [CrossRef]
- M. Mehta, T.A. Bui, X. Yang, Y. Aksoy, E.M. Goldys, W. Deng, Lipid-Based Nanoparticles for Drug/Gene Delivery: An Overview of the Production Techniques and Difficulties Encountered in Their Industrial Development, ACS Mater Au 3(6) (2023) 600-619. [CrossRef]
- P.R. Cullis, P.L. Felgner, The 60-year evolution of lipid nanoparticles for nucleic acid delivery, Nat Rev Drug Discov 23(9) (2024) 709-722. [CrossRef]
- R. Jeitler, C. Glader, G. König, J. Kaplan, C. Tetyczka, J. Remmelgas, M. Mußbacher, E. Fröhlich, E. Roblegg, On the Structure, Stability, and Cell Uptake of Nanostructured Lipid Carriers for Drug Delivery, Mol Pharm 21(7) (2024) 3674-3683. [CrossRef]
- M.L. Borrajo, A. Quijano, P. Lapuhs, A.I. Rodriguez-Perez, S. Anthiya, J.L. Labandeira-Garcia, R. Valenzuela, M.J. Alonso, Ionizable nanoemulsions for RNA delivery into the central nervous system - importance of diffusivity, J Control Release 372 (2024) 295-303. [CrossRef]
- X. Cai, R. Dou, C. Guo, J. Tang, X. Li, J. Chen, J. Zhang, Cationic Polymers as Transfection Reagents for Nucleic Acid Delivery, Pharmaceutics 15(5) (2023). [CrossRef]
- H. Takakusa, N. Iwazaki, M. Nishikawa, T. Yoshida, S. Obika, T. Inoue, Drug Metabolism and Pharmacokinetics of Antisense Oligonucleotide Therapeutics: Typical Profiles, Evaluation Approaches, and Points to Consider Compared with Small Molecule Drugs, Nucleic Acid Therapeutics 33(2) (2023) 83-94. [CrossRef]
- D.C. Luther, R. Huang, T. Jeon, X. Zhang, Y.W. Lee, H. Nagaraj, V.M. Rotello, Delivery of drugs, proteins, and nucleic acids using inorganic nanoparticles, Adv Drug Deliv Rev 156 (2020) 188-213. [CrossRef]
- Y. Liu, M. Zhao, M. Zhang, B. Yang, Y.K. Qi, Q. Fu, Mesoporous silica nanoparticle-based nanomedicine: Preparation, functional modification, and theranostic applications, Mater Today Bio 34 (2025) 102223. [CrossRef]
- A.R. Sharma, Y.H. Lee, A. Bat-Ulzii, M. Bhattacharya, C. Chakraborty, S.S. Lee, Recent advances of metal-based nanoparticles in nucleic acid delivery for therapeutic applications, J Nanobiotechnology 20(1) (2022) 501. [CrossRef]
- C.R. Garza-Cardenas, A. Leon-Buitimea, A.A. Siller-Ceniceros, J.R. Morones-Ramirez, Antisense Oligonucleotide-Capped Gold Nanoparticles as a Potential Strategy for Tackling Antimicrobial Resistance, Microbiology Research 16(3) (2025) 70. [CrossRef]
- R. Lapusan, R. Borlan, M. Focsan, Advancing MRI with magnetic nanoparticles: a comprehensive review of translational research and clinical trials, Nanoscale Adv 6(9) (2024) 2234-2259. [CrossRef]
- M. Liu, B. Chu, R. Sun, J. Ding, H. Ye, Y. Yang, Y. Wu, H. Shi, B. Song, Y. He, H. Wang, J. Hong, Antisense Oligonucleotides Selectively Enter Human-Derived Antibiotic-Resistant Bacteria through Bacterial-Specific ATP-Binding Cassette Sugar Transporter, Adv Mater 35(28) (2023) e2300477. [CrossRef]
- G. Xu, J. Jin, Z. Fu, G. Wang, X. Lei, J. Xu, J. Wang, Extracellular vesicle-based drug overview: research landscape, quality control and nonclinical evaluation strategies, Signal Transduct Target Ther 10(1) (2025) 255. [CrossRef]
- M. Zhang, S. Hu, L. Liu, P. Dang, Y. Liu, Z. Sun, B. Qiao, C. Wang, Engineered exosomes from different sources for cancer-targeted therapy, Signal Transduct Target Ther 8(1) (2023) 124. [CrossRef]
- S. Kamerkar, C. Leng, O. Burenkova, S.C. Jang, C. McCoy, K. Zhang, K. Dooley, S. Kasera, T. Zi, S. Sisó, W. Dahlberg, C.L. Sia, S. Patel, K. Schmidt, K. Economides, T. Soos, D. Burzyn, S. Sathyanarayanan, Exosome-mediated genetic reprogramming of tumor-associated macrophages by exoASO-STAT6 leads to potent monotherapy antitumor activity, Sci Adv 8(7) (2022) eabj7002. [CrossRef]
- L. Liu, D. Pan, S. Chen, M.V. Martikainen, A. Kårlund, J. Ke, H. Pulkkinen, H. Ruhanen, M. Roponen, R. Käkelä, W. Xu, J. Wang, V.P. Lehto, Systematic design of cell membrane coating to improve tumor targeting of nanoparticles, Nat Commun 13(1) (2022) 6181. [CrossRef]
- A. Gogate, J. Belcourt, M. Shah, A.Z. Wang, A. Frankel, H. Kolmel, M. Chalon, P. Stephen, A. Kolli, S.M. Tawfik, J. Jin, R. Bahal, T.P. Rasmussen, J.E. Manautou, X.B. Zhong, Targeting the Liver with Nucleic Acid Therapeutics for the Treatment of Systemic Diseases of Liver Origin, Pharmacol Rev 76(1) (2023) 49-89. [CrossRef]
- J.L. Witztum, D. Gaudet, S.D. Freedman, V.J. Alexander, A. Digenio, K.R. Williams, Q. Yang, S.G. Hughes, R.S. Geary, M. Arca, E.S.G. Stroes, J. Bergeron, H. Soran, F. Civeira, L. Hemphill, S. Tsimikas, D.J. Blom, L. O'Dea, E. Bruckert, Volanesorsen and Triglyceride Levels in Familial Chylomicronemia Syndrome, N Engl J Med 381(6) (2019) 531-542. [CrossRef]
- L. De Michieli, A. Lupi, G. Sinigiani, A. Tietto, A. Salvalaggio, A. Branca, S. Da Pozzo, S. Rizzo, D. Cecchin, M. Perazzolo Marra, T. Berno, D. Corrado, C. Briani, A. Cipriani, Pharmacological Management of Transthyretin Amyloid Cardiomyopathy: Where We Are and Where We Are Going, J Clin Med 14(10) (2025). [CrossRef]
- L. Xu, I. Irony, W.W. Bryan, B. Dunn, Development of gene therapies-lessons from nusinersen, Gene Ther 24(9) (2017) 527-528. [CrossRef]
- A. McGuigan, H.A. Blair, Tofersen: A Review in Amyotrophic Lateral Sclerosis Associated with SOD1 Mutations, CNS Drugs 39(9) (2025) 903-912. [CrossRef]
- S.J. Keam, Imetelstat: First Approval, Drugs 84(9) (2024) 1149-1155. [CrossRef]
- Y. Abaza, A.E. DeZern, Imetelstat: a new addition to the therapeutic landscape of lower-risk MDS, Blood 145(5) (2025) 469-474. [CrossRef]
- M.A. Riedl, R. Tachdjian, W.R. Lumry, T. Craig, G. Karakaya, A. Gelincik, M. Stobiecki, J.S. Jacobs, N.M. Gokmen, A. Reshef, M.M. Gompels, M.E. Manning, L. Bordone, K.B. Newman, S. Treadwell, S. Wang, A. Yarlas, D.M. Cohn, Efficacy and Safety of Donidalorsen for Hereditary Angioedema, N Engl J Med 391(1) (2024) 21-31. [CrossRef]
- B.A. Bergmark, N.A. Marston, C.R. Bramson, M. Curto, V. Ramos, A. Jevne, J.F. Kuder, J.G. Park, S.A. Murphy, S. Verma, W. Wojakowski, S.G. Terra, M.S. Sabatine, S.D. Wiviott, Effect of Vupanorsen on Non-High-Density Lipoprotein Cholesterol Levels in Statin-Treated Patients With Elevated Cholesterol: TRANSLATE-TIMI 70, Circulation 145(18) (2022) 1377-1386. [CrossRef]
- K. Chwalenia, M.J.A. Wood, T.C. Roberts, Progress and prospects in antisense oligonucleotide-mediated exon skipping therapies for Duchenne muscular dystrophy, J Muscle Res Cell Motil 46(4) (2025) 293-300. [CrossRef]
- Food and Drug Administration (FDA), Clinical Pharmacology Considerations for the Development of Oligonucleotide Therapeutics, 2026-01-30. https://www.fda.gov/regulatory-information/search-fda-guidance-documents/clinical-pharmacology-considerations-development-oligonucleotide-therapeutics.
- T.B. Emran, A. Shahriar, A.R. Mahmud, T. Rahman, M.H. Abir, M.F. Siddiquee, H. Ahmed, N. Rahman, F. Nainu, E. Wahyudin, S. Mitra, K. Dhama, M.M. Habiballah, S. Haque, A. Islam, M.M. Hassan, Multidrug Resistance in Cancer: Understanding Molecular Mechanisms, Immunoprevention and Therapeutic Approaches, Front Oncol 12 (2022) 891652. [CrossRef]
- M. Nikanjam, S. Kato, T. Allen, J.K. Sicklick, R. Kurzrock, Novel clinical trial designs emerging from the molecular reclassification of cancer, CA Cancer J Clin 75(3) (2025) 243-267. [CrossRef]
- K.N. Chi, C.S. Higano, B. Blumenstein, J.M. Ferrero, J. Reeves, S. Feyerabend, G. Gravis, A.S. Merseburger, A. Stenzl, A.M. Bergman, S.D. Mukherjee, P. Zalewski, F. Saad, C. Jacobs, M. Gleave, J.S. de Bono, Custirsen in combination with docetaxel and prednisone for patients with metastatic castration-resistant prostate cancer (SYNERGY trial): a phase 3, multicentre, open-label, randomised trial, Lancet Oncol 18(4) (2017) 473-485. [CrossRef]
- T.M. Beer, S.J. Hotte, F. Saad, B. Alekseev, V. Matveev, A. Fléchon, G. Gravis, F. Joly, K.N. Chi, Z. Malik, B. Blumenstein, P.S. Stewart, C.A. Jacobs, K. Fizazi, Custirsen (OGX-011) combined with cabazitaxel and prednisone versus cabazitaxel and prednisone alone in patients with metastatic castration-resistant prostate cancer previously treated with docetaxel (AFFINITY): a randomised, open-label, international, phase 3 trial, Lancet Oncol 18(11) (2017) 1532-1542. [CrossRef]
- Clinicaltrials.gov, Study of SRP-4045 (Casimersen) and SRP-4053 (Golodirsen) in Participants With Duchenne Muscular Dystrophy (DMD) (ESSENCE), 2025-11-18. https://clinicaltrials.gov/study/NCT02500381.
- Parent Project Muscular Dystrophy, Sarepta Announces Completion of ESSENCE Trial for Exon 45 & 53 Skipping Therapies, 2026-01-30. https://www.parentprojectmd.org/sarepta-announces-completion-of-essence-trial-for-exon-45-53-skipping-therapies.
- Clinicaltrials.gov, A Study to Evaluate Efficacy, Safety, Tolerability and Exposure After a Repeat-dose of Sepofarsen (QR-110) in LCA10 (ILLUMINATE) (ILLUMINATE), 2022-03-17. https://clinicaltrials.gov/study/NCT03913143.
- S.R. Russell, A.V. Drack, A.V. Cideciyan, S.G. Jacobson, B.P. Leroy, C. Van Cauwenbergh, A.C. Ho, A.V. Dumitrescu, I.C. Han, M. Martin, W.L. Pfeifer, E.H. Sohn, J. Walshire, A.V. Garafalo, A.K. Krishnan, C.A. Powers, A. Sumaroka, A.J. Roman, E. Vanhonsebrouck, E. Jones, F. Nerinckx, J. De Zaeytijd, R.W.J. Collin, C. Hoyng, P. Adamson, M.E. Cheetham, M.R. Schwartz, W. den Hollander, F. Asmus, G. Platenburg, D. Rodman, A. Girach, Intravitreal antisense oligonucleotide sepofarsen in Leber congenital amaurosis type 10: a phase 1b/2 trial, Nat Med 28(5) (2022) 1014-1021. [CrossRef]
- J. Saade, T.A. Mestre, Huntington's Disease: Latest Frontiers in Therapeutics, Curr Neurol Neurosci Rep 24(8) (2024) 255-264. [CrossRef]
- P. McColgan, A. Thobhani, L. Boak, S.A. Schobel, A. Nicotra, G. Palermo, D. Trundell, J. Zhou, V. Schlegel, P. Sanwald Ducray, D.J. Hawellek, J. Dorn, C. Simillion, M. Lindemann, V. Wheelock, A. Durr, K.E. Anderson, J.D. Long, E.J. Wild, G.B. Landwehrmeyer, B.R. Leavitt, S.J. Tabrizi, R. Doody, Tominersen in Adults with Manifest Huntington's Disease, N Engl J Med 389(23) (2023) 2203-2205. [CrossRef]
- J.Y. Yao, T. Liu, X.R. Hu, H. Sheng, Z.H. Chen, H.Y. Zhao, X.J. Li, Y. Wang, L. Hao, An insight into allele-selective approaches to lowering mutant huntingtin protein for Huntington's disease treatment, Biomed Pharmacother 180 (2024) 117557. [CrossRef]
- European Medicines Agency(EMA), Vitravene, 2002-07-08. https://www.ema.europa.eu/en/medicines/human/EPAR/vitravene.
- Clinicaltrials.gov, Assessing the Impact of Lipoprotein (a) Lowering With Pelacarsen (TQJ230) on Major Cardiovascular Events in Patients With CVD (Lp(a)HORIZON), 2025-12-31. https://clinicaltrials.gov/study/NCT04023552.
- ClinicalTrials.gov, Phase 3 Efficacy and Safety Study of GTX-102 in Pediatric Subjects With Angelman Syndrome (AS) (Aspire), 2025-11-06. https://clinicaltrials.gov/study/NCT06617429.
- ClinicalTrials.gov, Study of Bepirovirsen in Nucleos(t)Ide Analogue-treated Participants With Chronic Hepatitis B (B-Well 2) (B-Well 2), 2025-12-12. https://clinicaltrials.gov/study/NCT05630820.
- F. Jaschinski, T. Rothhammer, P. Jachimczak, C. Seitz, A. Schneider, K.H. Schlingensiepen, The antisense oligonucleotide trabedersen (AP 12009) for the targeted inhibition of TGF-β2, Curr Pharm Biotechnol 12(12) (2011) 2203-13. [CrossRef]
- S. Mariathasan, S.J. Turley, D. Nickles, A. Castiglioni, K. Yuen, Y. Wang, E.E. Kadel, III, H. Koeppen, J.L. Astarita, R. Cubas, S. Jhunjhunwala, R. Banchereau, Y. Yang, Y. Guan, C. Chalouni, J. Ziai, Y. Şenbabaoğlu, S. Santoro, D. Sheinson, J. Hung, J.M. Giltnane, A.A. Pierce, K. Mesh, S. Lianoglou, J. Riegler, R.A.D. Carano, P. Eriksson, M. Höglund, L. Somarriba, D.L. Halligan, M.S. van der Heijden, Y. Loriot, J.E. Rosenberg, L. Fong, I. Mellman, D.S. Chen, M. Green, C. Derleth, G.D. Fine, P.S. Hegde, R. Bourgon, T. Powles, TGFβ attenuates tumour response to PD-L1 blockade by contributing to exclusion of T cells, Nature 554(7693) (2018) 544-548. [CrossRef]
- ClinicalTrials.gov,, Trabedersen (OT-101) With Pembrolizumab for Newly Diagnosed Advanced NSCLC and Positive PD-L1, 2025-07-24. https://clinicaltrials.gov/study/NCT06579196.
- M. Li, H. An, J. Zhang, W. Li, C. Yu, L. Wang, Advances in the pharmaceutical development of antibody-oligonucleotide conjugates, Eur J Pharm Sci 215 (2025) 107292. [CrossRef]
- O. Kovecses, F.E. Mercier, M. McKeague, Nucleic acid therapeutics as differentiation agents for myeloid leukemias, Leukemia 38(7) (2024) 1441-1454. [CrossRef]
- V. Bhati, S. Prasad, A. Kabra, RNA-based therapies for neurodegenerative disease: Targeting molecular mechanisms for disease modification, Mol Cell Neurosci 133 (2025) 104010. [CrossRef]
- A. Bruch, A.A. Kelani, M.G. Blango, RNA-based therapeutics to treat human fungal infections, Trends Microbiol 30(5) (2022) 411-420. [CrossRef]
- D. Araújo, D. Mil-Homens, M. Henriques, S. Silva, Anti-EFG1 2'-OMethylRNA oligomer inhibits Candida albicans filamentation and attenuates the candidiasis in Galleria mellonella, Mol Ther Nucleic Acids 27 (2022) 517-523. [CrossRef]
- A. Barbosa, D. Araújo, M. Henriques, S. Silva, The combined application of the anti-RAS1 and anti-RIM101 2'-OMethylRNA oligomers enhances Candida albicans filamentation control, Med Mycol 59(10) (2021) 1024-1031.
- J.Y. Chung, Y.K. Hong, E. Jeon, S. Yang, A. Park, R. Weissleder, Y.P. Chong, H.J. Chung, Effective treatment of systemic candidiasis by synergistic targeting of cell wall synthesis, Nat Commun 16(1) (2025) 5532. [CrossRef]
- P.R. Carstens, T. Yokota, From Genomic Diagnosis to Personalized RNA Medicine: Advances in Next-Generation Sequencing and N-of-1 Antisense Oligonucleotide Therapies for Rare Genetic Diseases, Preprints, Preprints, 2025. [CrossRef]
- H. Wilton-Clark, E. Yan, T. Yokota, Milasen: The Emerging Era of Patient-Customized N-of-1 Antisense Oligonucleotides as Therapeutic Agents for Genetic Diseases, Methods Mol Biol 2964 (2025) 85-93.
- A. Aartsma-Rus, A. Garanto, W. van Roon-Mom, E.M. McConnell, V. Suslovitch, W.X. Yan, J.K. Watts, T.W. Yu, Consensus Guidelines for the Design and In Vitro Preclinical Efficacy Testing N-of-1 Exon Skipping Antisense Oligonucleotides, Nucleic Acid Ther 33(1) (2023) 17-25. [CrossRef]
- Y.F. Wu, J.A. Chen, Y.J. Jong, Treating neuromuscular diseases: unveiling gene therapy breakthroughs and pioneering future applications, J Biomed Sci 32(1) (2025) 30. [CrossRef]
- K.D. Kernohan, K.M. Boycott, The expanding diagnostic toolbox for rare genetic diseases, Nature Reviews Genetics 25(6) (2024) 401-415. [CrossRef]
- B.J. Booth, S. Nourreddine, D. Katrekar, Y. Savva, D. Bose, T.J. Long, D.J. Huss, P. Mali, RNA editing: Expanding the potential of RNA therapeutics, Mol Ther 31(6) (2023) 1533-1549. [CrossRef]
| Drug Name | Trade Name | First Approval | Company | Target | Indication | Mechanism | Modification | Delivery | Delivery route | Status |
|---|---|---|---|---|---|---|---|---|---|---|
| Fomivirsen | Vitravene | 1998 | Ionis & Novartis | CMV mRNA | CMV retinitis | RNase H mediated | PS | naked | IVT | Withdrawn |
| Mipomersen | Kynamro | 2013 | Ionis & Genzyme | ApoB-100 | HoFH | RNase H mediated | 2'-MOE Gapmer | naked | SC | Withdrawn |
| Eteplirsen | Exondys 51 | 2016 | Sarepta | Dys Exon 51 | DMD | Steric blocking | PMO | naked | IV | Marketed |
| Nusinersen | Spinraza | 2016 | Ionis & Biogen | SMN2 | SMA | Steric blocking | 2'-MOE, PS | naked | IT | Marketed |
| Inotersen | Tegsedi | 2018 | Ionis & Sobi | TTR | hATTR Amyloidosis | RNase H mediated | 2'-MOE Gapmer | naked | SC | Marketed |
| Volanesorsen | Waylivra | 2019 | Ionis & Sobi | APOC3 | FCS | RNase H mediated | 2'-MOE Gapmer | naked | SC | Marketed |
| Golodirsen | Vyondys 53 | 2019 | Sarepta | Dys Exon 53 | DMD | Steric blocking | PMO | naked | IV | Marketed |
| Viltolarsen | Viltepso | 2020 | Nippon Shinyaku | Dys Exon 53 | DMD | Steric blocking | PMO | naked | IV | Marketed |
| Casimersen | Amondys 45 | 2021 | Sarepta | Dys Exon 45 | DMD | Steric blocking | PMO | naked | IV | Marketed |
| Tofersen | Qalsody | 2023 | Ionis & Biogen | SOD1 | ALS | RNase H mediated | 2'-MOE Gapmer | naked | IT | Marketed |
| Eplontersen | Wainua | 2023 | AstraZeneca & Ionis | TTR | hATTR Amyloidosis |
RNase H mediated | 2'-MOE Gapmer | GalNAc | SC | Marketed |
| Imetelstat | Rytelo | 2024 | Geron Corporation | Telomerase hTR | MDS | Telomerase inhibition | N3'-P5' Thio | Lipid | IV | Marketed |
| Olezarsen | Tryngolza | 2024 | Ionis | APOC3 | FCS | RNase H mediated | 2'-MOE Gapmer | GalNAc | SC | Marketed |
| Phase | Drug Name / Code | Clinical Trial ID | Target | Indication | Disease Category | Mechanism | Modification | Delivery | Delivery route |
|---|---|---|---|---|---|---|---|---|---|
| III | Trabedersen (AP 12009) |
NCT00761280 NCT00431561 NCT00844064 NCT05935774 |
TGF-β2 | Glioma | Oncology & Hematology | RNase H mediated | PS | Intratumoral Perfusion | naked |
| III | Aprinocarsen (ISIS 3521/LY900003) |
NCT00017407 NCT00034268 NCT00003989 |
PKC-α | Multiple Solid Tumors | Oncology & Hematology | RNase H mediated | PS | IV | naked |
| III | Custirsen (OGX-011) |
NCT01188187 NCT01578655 |
Clusterin | CRPC | Oncology & Hematology | RNase H mediated | 2'-MOE Gapmer | IV | naked |
| III | Oblimersen (G3139/Genasense) |
NCT00024440 NCT00518895 NCT00021749 |
BCL2 | Bcl-2 Positive Malignancies | Oncology & Hematology | RNase H mediated | PS | IV | naked |
| III | Drisapersen (PRO051/GSK2402968) |
NCT01254019 NCT01153932 NCT01462292 |
DMD Exon 51 | DMD | Neuromuscular Diseases | Steric blocking | 2'-O-Me PS | SC | naked |
| III | Tominersen (RG6042/IONIS-HTTRx) |
NCT03761849 NCT03842969 NCT02519036 |
HTT | HD | Neurological Diseases | RNase H mediated | 2'-MOE Gapmer | IT | naked |
| III | Sepofarsen (QR-110) |
NCT03913143 NCT03140969 |
CEP290 | LCA10 | Ophthalmic Diseases | Steric blocking | 2'-O-Me PS | IVT | naked |
| III | Alicaforsen (ISIS 2302) |
NCT02525523 NCT00063830 NCT00048113 |
ICAM-1 | Crohn's Disease | Immunological Diseases | RNase H mediated | PS | IV/Enema | naked |
| III | Mongersen (GED-0301) |
NCT02596893 NCT02367183 |
SMAD7 | Crohn's Disease | Immunological Diseases | RNase H mediated | PS | PO | pH-dependent Coating |
| II | IONIS-DGAT2Rx | NCT03334214 | DGAT2 | Hepatic Steatosis | Cardiovascular & Metabolic Diseases | RNase H mediated | 2'-MOE Gapmer | SC | naked |
| II | ISIS-GCGRRx | NCT02824003 NCT01885260 NCT02583919 |
GCGR | T2DM | Cardiovascular & Metabolic Diseases | RNase H mediated | 2'-MOE Gapmer | SC | naked |
| II | ISIS-GCCRRx | NCT01968265 | GCCR | T2DM | Cardiovascular & Metabolic Diseases | RNase H mediated | 2'-MOE Gapmer | SC | naked |
| II | ISIS-FGFR4Rx | NCT02476019 | FGFR4 | Obesity | Cardiovascular & Metabolic Diseases | RNase H mediated | 2'-MOE Gapmer | SC | naked |
| II | CIVI-007 | NCT04164888 NCT03427710 |
PCSK9 | Hypercholesterolemia | Cardiovascular & Metabolic Diseases | RNase H mediated | LNA Gapmer | SC | GalNAc |
| II | IONIS-GHR-LRx | NCT04522180 NCT03967249 NCT03548415 |
GHR | Acromegaly | Cardiovascular & Metabolic Diseases | RNase H mediated | 2'-MOE / cEt Gapmer | SC | GalNAc |
| II | ISIS-PTP1BRx | NCT01918865 | PTP1B | T2DM | Cardiovascular & Metabolic Diseases | RNase H mediated | 2'-MOE Gapmer | SC | naked |
| II | IONIS-PTP1BRx (ISIS-404173) |
NCT01918865 | PTP1B | T2DM | Cardiovascular & Metabolic Diseases | RNase H mediated | 2'-MOE Gapmer | SC | naked |
| II | Vupanorsen (ISIS 703802) |
NCT04516291 NCT03514420 NCT03360747 |
ANGPTL3 | Severe Hypertriglyceridemia / CV Risk Reduction | Cardiovascular & Metabolic Diseases | RNase H mediated | 2'-MOE Gapmer | SC | GalNAc |
| II | Atesidorsen (ATL1103) |
EUCTR2012-003147-30 ACTRN12615000289516 |
GHR | Acromegaly | Cardiovascular & Metabolic Diseases | RNase H mediated | 2'-MOE Gapmer | SC | naked |
| II | Miravirs (SPC3649) |
NCT01200420 | miR-122 | HCV | Infectious Diseases | Anti-miR | LNA anti-miR | SC | naked |
| II | RG-101 | EudraCT:2015-001535-21 EudraCT:2015-004702-42 EudraCT:2016-002069-77 EudraCT:2013-002978-49 |
miR-122 | HCV | Infectious Diseases | Anti-miR | LNA anti-miR | SC | GalNAc |
| II | GSK3389404 (GalNAc-bepirovirsen) |
NCT03020745 | All HBV RNAs | HBV | Infectious Diseases | RNase H mediated | 2'-MOE Gapmer | SC | GalNAc |
| II | Donidalorsen (ISIS 721744) |
NCT04549922 | ASKCOV | COIVD-19 | Infectious Diseases | RNase H mediated | 2'-MOE Gapmer | SC | GalNAc |
| II | OGX-427 | NCT01829113 NCT01120470 NCT01681433 |
Hsp27 | Multiple Solid Tumors | Oncology & Hematology | RNase H mediated | 2'-MOE Gapmer | IV | naked |
| II | Danvatirsen (AZD9150) |
NCT02983578 NCT02417753 NCT03334617 NCT03794544 NCT01839604 NCT02546661 NCT02499328 NCT03527147 NCT02549651 NCT03421353 |
STAT3 | Multiple Solid Tumors | Oncology & Hematology | RNase H mediated | 2'-cEt Gapmer | IV | naked |
| II | ISIS 5132 (CGP69846A) |
NCT00002587 NCT00002588 NCT00002589 |
C-RAF-1 | Advanced Solid Tumors | Oncology & Hematology | RNase H mediated | PS | IV | naked |
| II | ISIS 2503 | NCT00004193 NCT00005594 NCT00006467 |
HRAS | Pancreatic Cancer | Oncology & Hematology | RNase H mediated | PS | IV | naked |
| II | G4460 (LR-3001) |
NCT00002592 | c-myb | CLL | Oncology & Hematology | RNase H mediated | PS | IV | naked |
| II | AEG35156 (GEM640) |
NCT00882869 | XIAP Mrna | Hepatocellular Carcinoma | Oncology & Hematology | RNase H mediated | 2'-MOE Gapmer | IV | naked |
| II | Gataparsen (LY2181308/ISIS-23722) |
NCT01107444 NCT00620321 NCT00642018 |
BIRC5 (Survivin) |
Second-line NSCLC | Oncology & Hematology | RNase H mediated | 2'-MOE Gapmer | IV | naked |
| II | Apatorsen (OGX-427) |
NCT00487786 NCT01829113 NCT02423590 NCT01454089 NCT01844817 |
HSPB1 (Hsp27) |
Multiple Solid Tumors | Oncology & Hematology | RNase H mediated | 2'-MOE Gapmer | IV | naked |
| II | QR-421a (Sepofarsen) |
NCT03780257 NCT05158296 |
USH2A exon 13 | arRP | Ophthalmic Diseases | Steric blocking | 2'-O-Me PS | IVT | naked |
| II | QR-1123 (IONIS-RHO-2.5Rx) |
NCT04123626 | RHO P23H | adRP | Ophthalmic Diseases | RNase H mediated | 2'-cEt Gapmer | IVT | naked |
| II | PGN-EDO51 | NCT06079736 | DMD Exon 51 | DMD | Neuromuscular Diseases | Steric blocking | PPMO | IV | CPP |
| II | Avicursen (ATL1102) |
ACTRN12618000936203 | CD49d | DMD | Neuromuscular Diseases | RNase H mediated | 2'-MOE Gapmer | SC | naked |
| II | SRP-5051 (Vesleteplirsen) |
NCT04004065 | DMD Exon 51 | DMD | Neuromuscular Diseases | Steric blocking | PPMO | IV | CPP |
| II | IONIS-PKKRx | NCT03254362 | PKK | Chronic Migraine | Neurological Diseases | RNase H mediated | 2'-MOE Gapmer | SC | naked |
| II | WVE-120101 | NCT03225833 NCT04617847 |
mHTT SNP1 | HD | Neurological Diseases | RNase H mediated | PN Chemistry | IT | naked |
| II | WVE-120102 | NCT03225846 NCT04617860 |
mHTT SNP1 | HD | Neurological Diseases | RNase H mediated | PN Chemistry | IT | naked |
| II | ION-827359 (IONIS-ENaC-2.5Rx) |
NCT03647228 | SCNN1A/B/G | Cystic Fibrosis | Respiratory Diseases | RNase H mediated | 2'-cEt Gapmer | INH | naked |
| II | Fesomersen (ISIS 416858) |
NCT03358030 NCT02553889 |
Factor XI | Thromboprophylaxis / Anticoagulation | Hematological Diseases | RNase H mediated | 2'-MOE Gapmer | SC | naked |
| II | QR-313 (WNG-313) |
NCT03605069 | COL7A1 Exon73 | RDEB | Genodermatoses | Steric blocking | 2'-O-Me PS | TOP | naked |
| I/II | RG125 (AZD4076) |
NCT02826525 NCT02612662 |
miR-103/107 | T2DM with NAFLD / NASH | Cardiovascular & Metabolic Diseases | Anti-miR | LNA anti-miR | SC | GalNAc |
| I/II | Cavrotolimod (AST-008) |
NCT03684785 NCT03086278 |
TLR9 | PD-1 Resistant Tumors | Oncology & Hematology | Immune Activation | SNA | SC | naked |
| I/II | AZD5312 | NCT02144051 NCT03300505 |
AR | CRPC | Oncology & Hematology | RNase H mediated | 2'-cEt Gapmer | IV | naked |
| I/II | BIIB105 (ION541) |
NCT04494256 | ATXN2 | ALS(ATXN2) | Neurological Diseases | RNase H mediated | 2'-MOE Gapmer | IT | naked |
| I | EZN-2968 | NCT00466583 NCT01120288 |
HIF-1α | Solid Tumors or Lymphoma | Oncology & Hematology | RNase H mediated | LNA Gapmer | IV | naked |
| I | RO7070179 | NCT02564614 | HIF1A | HCC | Oncology & Hematology | RNase H mediated | Unknown | IV | naked |
| I | CDK-004 | NCT05375604 | STAT6 | HCC | Oncology & Hematology | RNase H mediated | Unknown | IV | exosome |
| I | Radavirsen (AVI-7100) |
NCT01747148 | M1/M2 | Influenza A Virus | Infectious Diseases | Steric blocking | PMO | IV | PMOplus |
| I | RO7062931 | NCT03038113 NCT03505190 |
All HBV RNAs | HBV | Infectious Diseases | RNase H mediated | LNA Gapmer | SC | GalNAc |
| I | ALG-020572 | NCT05001022 | All HBV RNAs | HBV | Infectious Diseases | RNase H mediated | BNA Gapmer | SC | GalNAc |
| I | BIIB078 (IONIS-C9Rx) |
NCT03626012 NCT04288856 |
C9orf72 | ALS/FTD | Neurological Diseases | RNase H mediated | 2'-MOE Gapmer | IT | naked |
| I | WVE-004 | NCT04931862 NCT05683860 |
C9orf72 | ALS/FTD | Neurological Diseases | RNase H mediated | PN Chemistry | IT | naked |
| I | NIO752 | NCT04539041 | TAU | PSP | Neurological Diseases | RNase H mediated | 2'-MOE Gapmer | IT | naked |
| I | ISIS 388626 | NCT00836225 | SGLT2 | T2DM & Obesity | Cardiovascular & Metabolic Diseases | RNase H mediated | 2'-MOE Gapmer | SC | naked |
| Phase | Drug Name / Code | Clinical Trial ID | Target | Indication | Disease Category | Mechanism | Modification | Delivery | Delivery route |
|---|---|---|---|---|---|---|---|---|---|
| III | Zilganersen (ION373) |
NCT04849741 | GFAP | Alexander Syndrome | Neurological Diseases | RNase H mediated | 2'-MOE Gapmer | IT | naked |
| III | ION582 (BIIB121) |
NCT06914609 NCT05127226 |
UBE3A-ATS | Angelman Syndrome | Neurological Diseases | RNase H mediated | 2'-MOE Gapmer | IT | naked |
| III | GTX-102 (Apazunersen) |
NCT06617429 NCT07157254 NCT04259281 |
UBE3A-ATS | Angelman Syndrome | Neurological Diseases | RNase H mediated | 2'-MOE Gapmer | IT | naked |
| III | ION363 (Jacifusen,Ulefnersen) |
NCT04768972 | FUS | ALS (FUS) | Neurological Diseases | RNase H mediated | 2'-MOE Gapmer | IT | naked |
| III | Zorevunersen (STK-001) |
NCT06872125 NCT04740476 NCT04442295 |
SCN1A | Dravet Syndrome | Neurological Diseases | TANGO | 2'-MOE ODN | IT | naked |
| III | Eteplirsen (approved LTE) |
NCT02420379 NCT02286947 |
DMD Exon 51 | DMD | Neuromuscular Diseases | Steric blocking | PMO | IV | naked |
| III | Bepirovirsen (GSK3228836) |
NCT05630820 NCT05630807 NCT04449029 NCT04954859 NCT04676724 NCT04544956 NCT02981602 |
All HBV RNAs | HBV;CHB | Infectious Diseases | RNase H mediated Immune Activation |
2'-MOE Gapmer | SC | naked |
| III | AHB-137 | NCT07246889 NCT07146100 NCT05717686 NCT06550128 NCT07069569 NCT06115993 |
All HBV RNAs | HBV | Infectious Diseases | RNase H mediated Immune Activation |
Med-Oligo™ | SC | naked |
| III | NEXAGON (Lufepirsen) |
NCT05966493 NCT04081103 NCT01165450 |
Connexin 43 | PCED | Ophthalmic Diseases | Steric blocking | ODN | Eye Gel | naked |
| III | Sefaxersen (IONIS-FB-LRx, RO7434656) |
NCT05797610 NCT03815825 NCT04014335 |
Complement Factor B | IgA Nephropathy (IgAN) | Immunological Diseases | RNase H mediated | 2'-MOE Gapmer | SC | GalNAc |
| III | Pelacarsen (TQJ230) |
NCT04023552 NCT06875973 NCT05305664 NCT05900141 NCT06267560 NCT06813911 NCT05646381 NCT03070782 |
LPA | Lp(a), CVD | Cardiovascular & Metabolic Diseases | RNase H mediated | 2'-MOE Gapmer | SC | GalNAc |
| III | Olezarsen | NCT05079919 NCT05552326 NCT05681351 |
APOC3 | sHTG– CORE | Cardiovascular & Metabolic Diseases | RNase H mediated | 2'-MOE Gapmer | SC | GalNAc |
| III | Olezarsen | NCT05355402 NCT05610280 NCT03385239 NCT02900027 |
APOC3 | Hypertriglyceridemia w/ ASCVD or High CV Risk | Cardiovascular & Metabolic Diseases | RNase H mediated | 2'-MOE Gapmer | SC | GalNAc |
| III | Eplontersen | NCT04136171 | TTR | ATTR-CM | Cardiovascular & Metabolic Diseases | RNase H mediated | 2'-MOE Gapmer | SC | GalNAc |
| III | Olezarsen (approved LTE) |
NCT05185843 | APOC3 | FCS | Cardiovascular & Metabolic Diseases | RNase H mediated | 2'-MOE Gapmer | SC | GalNAc |
| III | Donidalorsen (approved LTE) |
NCT05139810 | PKK | HAE | Immunological Diseases | RNase H mediated | 2'-MOE Gapmer | SC | GalNAc |
| II | AZD2693 (ION839) |
NCT05809934 (CTR20232127) |
PNPLA3 | MASH | Cardiovascular & Metabolic Diseases | RNase H mediated | 2'-MOE Gapmer | SC | GalNAc |
| II | IONIS-AGT-LRx | NCT03714776 NCT04083222 |
AGT | Resistant Hypertension | Cardiovascular & Metabolic Diseases | RNase H mediated | 2'-MOE Gapmer | SC | GalNAc |
| II | ION224 (IONIS-DGAT2Rx) |
NCT03334214 NCT04932512 |
DGAT2 | MASH with Fibrosis | Cardiovascular & Metabolic Diseases | RNase H mediated | 2'-MOE Gapmer | SC | GalNAc |
| II | QR-421a (Ultevursen) |
NCT06627179 | USH2A exon 13 | arRP | Ophthalmic Diseases | Steric blocking | 2'-O-Me PS | IVT | naked |
| II | Fesomersen (BAY2976217) |
NCT04534114 | Factor XI | Thromboprophylaxis / Anticoagulation | Hematological Diseases | RNase H mediated | 2'-MOE Gapmer | SC | GalNAc |
| II | AZD2373 (Opemalirsen) |
NCT06824987 | APOL1 | AMKD | Kidney Disease | RNase H mediated | 2'-cEt Gapmer | SC | naked |
| II | OT-101 | NCT06079346 NCT05425576 |
TGF-β2 | PDAC; MPM | Oncology & Hematology | RNase H mediated | PS | Intratumoral Perfusion | naked |
| II | BP1001 (Prexigebersen) |
NCT02781883 | Grb-2 | AML、ALL、CML-BP、MDS | Oncology & Hematology | RNase H mediated | P-ethoxy-DNA | IV | Liposome |
| II | Danvatirsen (AZD9150) |
NCT05814666 | STAT3 | HNSCC | Oncology & Hematology | RNase H mediated | 2'-cEt Gapmer | IV | naked |
| II | BIIB080 (IONIS-MAPT Rx) |
NCT05399888 NCT03186989 |
MAPT | AD | Neurological Diseases | RNase H mediated | 2'-MOE Gapmer | IT | naked |
| II | WVE-003 | NCT05032196 | mHTT SNP3 | HD | Neurological Diseases | RNase H mediated | PN Chemistry (Stereopure) | IT | naked |
| II | WVE-N531 | NCT04906460 | Dystrophin Exon 53 | DMD | Neuromuscular Diseases | Steric blocking | PN Chemistry (Stereopure) | IV | naked |
| I/II | Elsunersen (PRAX-222) |
NCT05737784 | SCN2A | SCN1A-Associated DEE | Neuromuscular Diseases | RNase H mediated | 2'-MOE gapmer | IT | naked |
| I/II | DYNE-251 | NCT05524883 | Dys Exon 51 | DMD | Neuromuscular Diseases | Steric blocking | PMO | IV | Fab-PMO(AOC) |
| I/II | AOC-1044 (del-zota) |
NCT05670730 | Dys Exon 44 | DMD | Neuromuscular Diseases | Steric blocking | PMO | IV | Fab-PMO(AOC) |
| I/II | ISTH0036 | NCT02406833 | TGF-β2 | POAG | Ophthalmic Diseases | RNase H mediated | LNA Gapmer | IVT | naked |
| I | ASOTARI | NCT06451172 | Essential genes for bacterial | Antibiotic-resistant bacterial keratitis | Ophthalmic & Infectious Diseases | Trojan Horse Strategy | PNA | Eye Drops | GP-SiNPs-asPNA |
| I | STK-002 | ISRCTN41725621 | OPA1 | ADOA | Ophthalmic Diseases | TANGO | 2'-MOE Gapmer | IVT | naked |
| I | NIO752 | NCT05469360 NCT06372821 |
TAU | AD | Neurological Diseases | RNase H mediated | 2'-MOE Gapmer | IT | naked |
| I | ION356 | NCT05786433 | PLP1 | PMD | Neurological Diseases | RNase H mediated | 2'-MOE / cEt Gapmer | IT | naked |
| I | ION716 | NCT06249918 | Prion Protein | CJD | Neurological Diseases | RNase H mediated | 2'-MOE Gapmer | IT | naked |
| I | AMX0114 | NCT06665165 | CAPN2 | ALS (CAPN2) | Neurological Diseases | RNase H mediated | 2'-MOE Gapmer | IT | naked |
| I | Atipeksen | NCT07215416 | ATM Exon 53 | A-T | Neurological Diseases | Steric blocking | 2'-MOE PS | IT | naked |
| I | BP1002(Liposome) | NCT04072458 NCT05190471 |
Bcl-2 | Bcl-2 Positive Malignancies | Oncology & Hematology | RNase H mediated | P-ethoxy-DNA | IV | Liposome |
| I | Danvatirsen (AZD9150) |
NCT03819465 | STAT3 | NSCLC | Oncology & Hematology | RNase H mediated | 2'-cEt Gapmer | IV | naked |
| I | Danvatirsen (AZD9150) |
NCT05986240 | STAT3 | AML / MDS | Oncology & Hematology | RNase H mediated | 2'-cEt Gapmer | IV | naked |
| I | OT-101 | NCT06579196 | TGF-β2 | NSCLC | Oncology & Hematology | RNase H mediated | PS | Intratumoral Perfusion | naked |
| NCT Number | Target | Drug Name / Code | Indication | Sponsor | status |
|---|---|---|---|---|---|
| NCT07197268 | ASXL3 | nL-ASXL3-001 | BRS | n-Lorem Foundation | Active |
| NCT07215416 | ATM | ASO targeting ATM | A-T | academic institution | Active |
| NCT06706388 | ATN1 | nL-ATN1-002 | DRPLA | n-Lorem Foundation | Active |
| NCT07084311 | ATN1 | nL-ATN1-002 | DRPLA | n-Lorem Foundation | Active |
| NCT07221760 | ATN1 | nL-ATN1-001 | DRPLA | n-Lorem Foundation | Not yet recruiting |
| NCT06392126 | CHCHD10 | nL-CHCHD-001 | ALS (CHCHD10 related) | n-Lorem Foundation | Active |
| NCT06977451 | CHCHD10 | nL-CHCHD-001 | ALS (CHCHD10 related) | n-Lorem Foundation | Active |
| NCT07095686 | CHCHD10 | nL-CHCHD-001 | ALS (CHCHD10 related) | n-Lorem Foundation | Enrolling |
| NCT06565572 | FLVCR1 | nL-FLVC-001 | PCARP | academic institution | Enrolling |
| NCT06816498 | LMNB1 | nL-LMNB1-001 | ADLD | n-Lorem Foundation | Active |
| NCT07197294 | MAPK8IP3 | nL-MAPK8-001 | NEDBA | n-Lorem Foundation | Active |
| NCT07177196 | PRPH2 | nL-PRPH2-001 | Retinal Dystrophy | n-Lorem Foundation | Active |
| NCT06314490 | SCN2A | nL-SCN2A-002 | SCN2A-Related Disorders | academic institution | Active |
| NCT07095712 | TARDBP | nL-TARD-001 | ALS (TDP-43 related) | n-Lorem Foundation | Active |
| NCT07222371 | TUBB4A | nL-TUBB4-001 | Leukodystrophy | academic institution | Active |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




