Submitted:
13 March 2026
Posted:
16 March 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
2.1. Recombinant DNA Constructs
2.2. Agrobacterium Infiltration (Agro-Infiltration) of N. benthamiana Leaves
2.3. Infection of Soybean Seedlings with Viruses Pre-Propagated in N. benthamiana
2.4. Evaluation of the Levels of Viral RNA and Plant Gene mRNA with RT-PCR
3. Results and Discussion
3.1. Neutralizing the Toxicity of CPSMV RNA1 cDNA in Agrobacterium Cells with a Fifth Intron
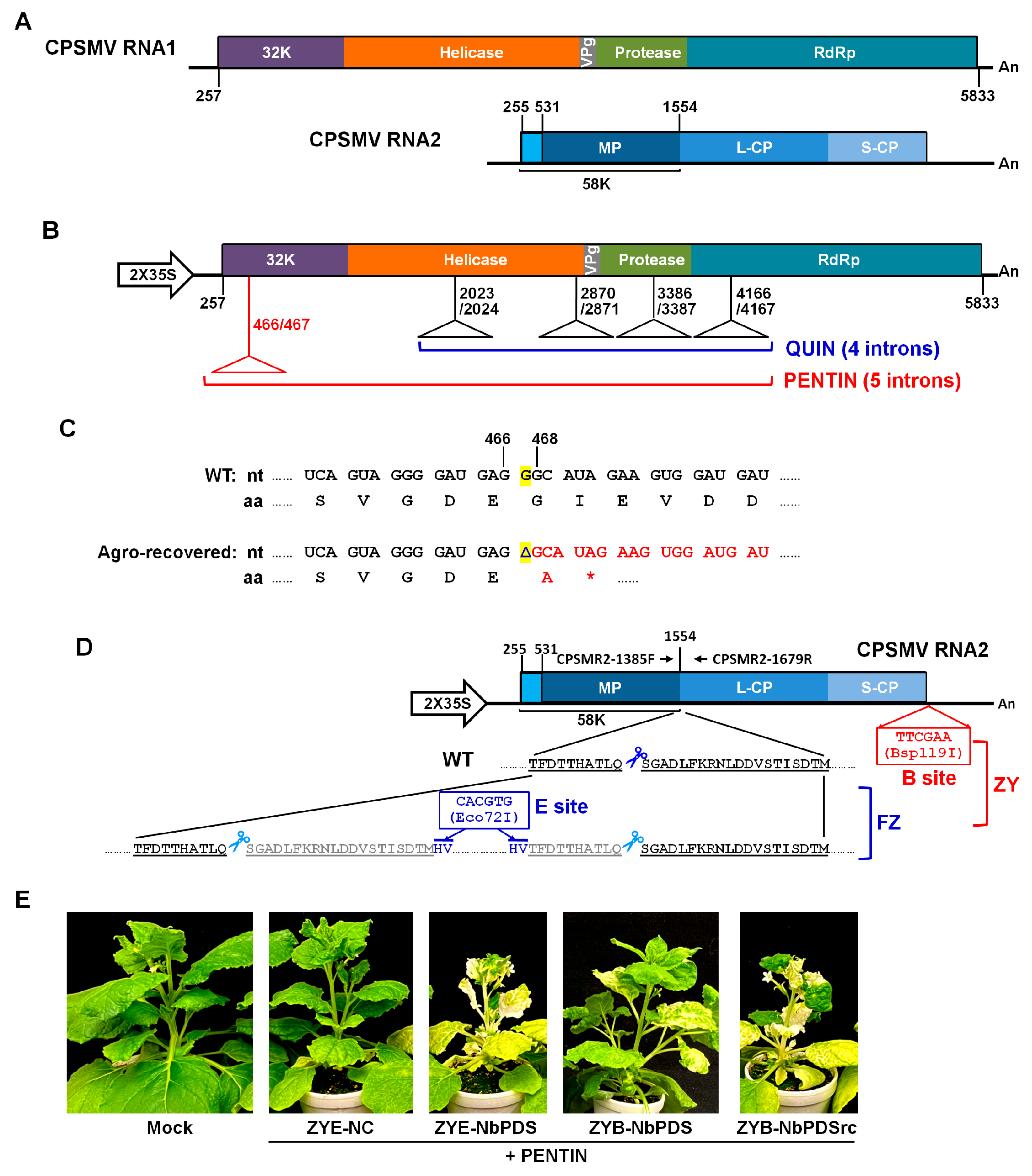
3.2. Upgrading FZ by Adding a Second Cloning Site
3.3. ZYE Constructs Carrying Different Segments of a Putative Protein Kinase cDNA Elicited Unexpected Responses in Soybean
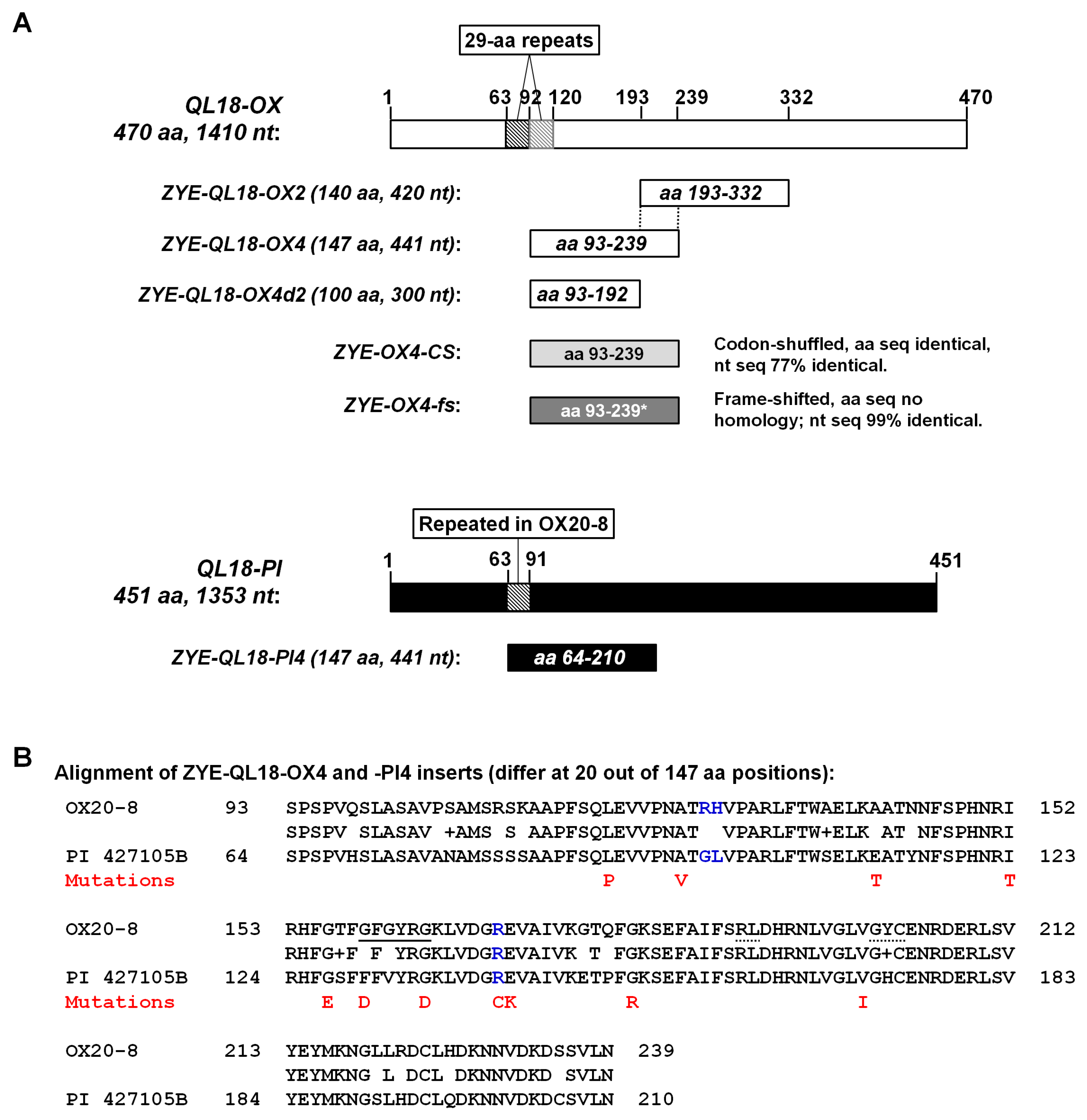

3.4. The Virus-Expressed Protein Fragments, Rather than Their Coding RNAs, Are Responsible for the Differing Symptoms of ZYE-QL18-OX4 and ZYE-QL18-PI4
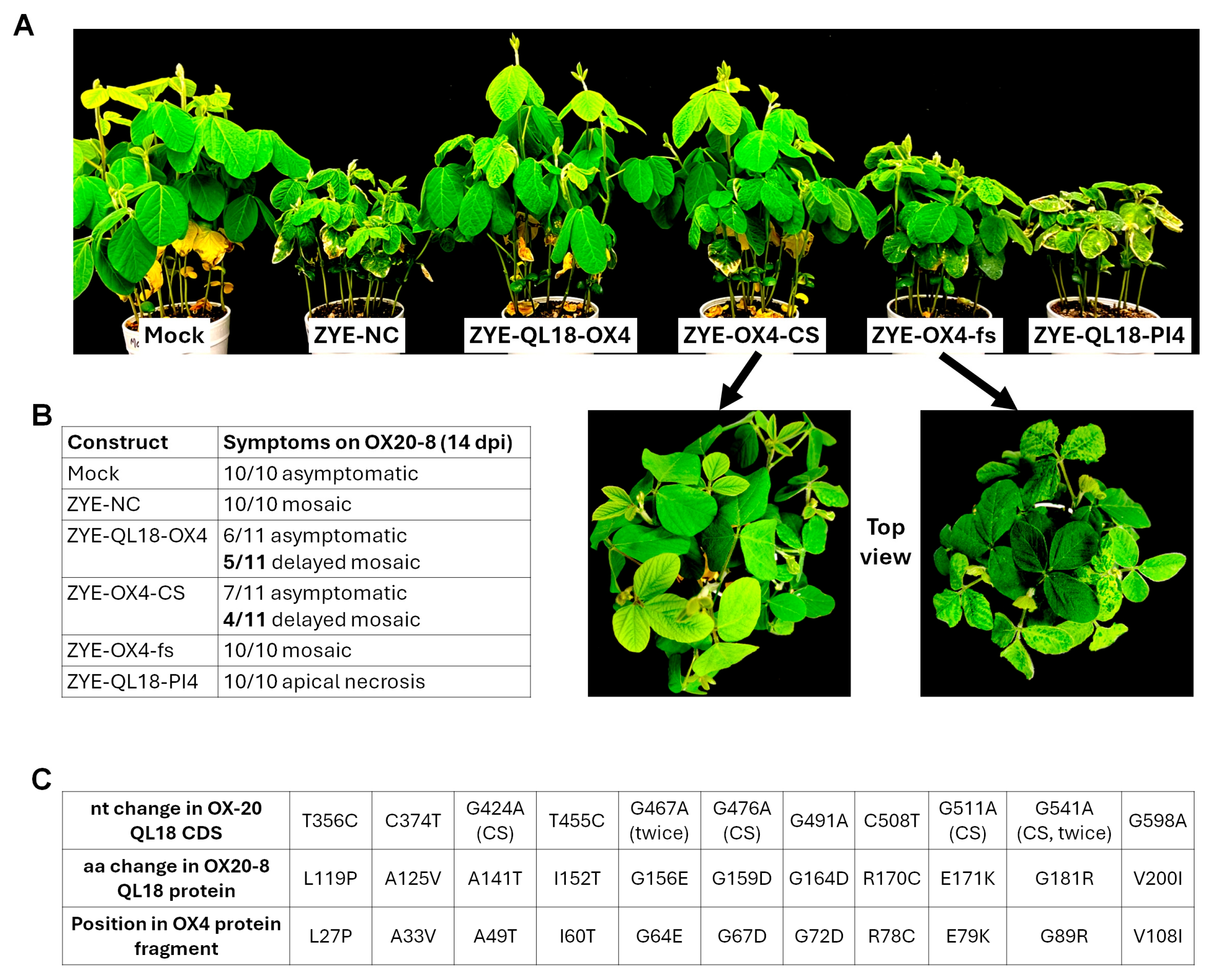
3.5. Viral RNA Recovered from Symptomatic Seedlings Inoculated with ZYE-QL18-OX4 and ZYE-OX4-CS Contain Amino-Acid-Changing Point Mutations Within Inserts
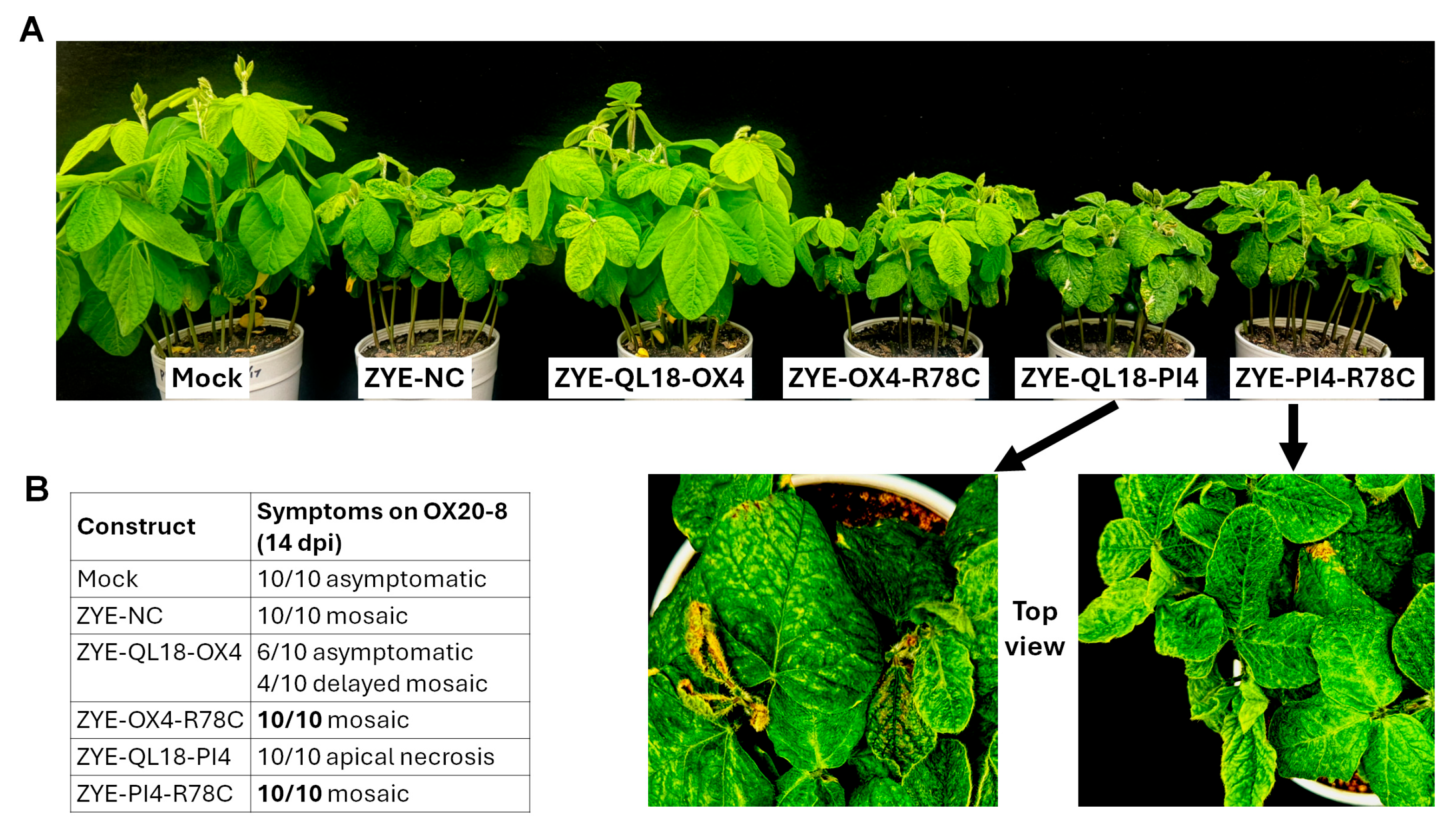
3.6. A Single Mutation Restores Mosaic Symptoms in Soybeans Overexpressing OX4, and Abolishes Apical Necrosis in Those Overexpressing PI4
3.7. A Spectrum of Symptoms Elicited by OX4-PI4 Chimeras in Three Different Soybean Accessions

3.8. OX4-Mediated Partial Failure of Systemic Spread and PI4-Mediated Apical Necrosis May Reflect Varying Strengths of the Same Defense Response
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Chen F, Dong W, Zhang J, Guo X, Chen J, Wang Z, et al. The Sequenced Angiosperm Genomes and Genome Databases. Front Plant Sci. 2018;9: 418. [CrossRef]
- Soltis PS, Soltis DE. Plant genomes: Markers of evolutionary history and drivers of evolutionary change. PLANTS PEOPLE PLANET. 2021;3: 74–82. [CrossRef]
- Kress WJ, Soltis DE, Kersey PJ, Wegrzyn JL, Leebens-Mack JH, Gostel MR, et al. Green plant genomes: What we know in an era of rapidly expanding opportunities. Proc Natl Acad Sci. 2022;119: e2115640118. [CrossRef]
- Gilchrist E, Haughn G. Reverse genetics techniques: engineering loss and gain of gene function in plants. Brief Funct Genomics. 2010;9: 103–110. [CrossRef]
- Paz MM, Martinez JC, Kalvig AB, Fonger TM, Wang K. Improved cotyledonary node method using an alternative explant derived from mature seed for efficient Agrobacterium-mediated soybean transformation. Plant Cell Rep. 2006;25: 206–213. [CrossRef]
- Woodward AW, Bartel B. Biology in Bloom: A Primer on the Arabidopsis thaliana Model System. Genetics. 2018;208: 1337–1349. [CrossRef]
- Zhang M, Liu S, Wang Z, Yuan Y, Zhang Z, Liang Q, et al. Progress in soybean functional genomics over the past decade. Plant Biotechnol J. 2021/08/25 ed. 2022;20: 256–282. [CrossRef]
- Guo Q, Liu Q, Smith NA, Liang G, Wang M-B. RNA Silencing in Plants: Mechanisms, Technologies and Applications in Horticultural Crops. Curr Genomics. 2016;17: 476–489. [CrossRef]
- Zhang C, Ghabrial SA. Development of Bean pod mottle virus-based vectors for stable protein expression and sequence-specific virus-induced gene silencing in soybean. Virology. 2006;344: 401–411. [CrossRef]
- Zhang C, Bradshaw JD, Whitham SA, Hill JH. The Development of an Efficient Multipurpose Bean Pod Mottle Virus Viral Vector Set for Foreign Gene Expression and RNA Silencing. Plant Physiol. 2010;153: 52–65. [CrossRef]
- Liu J-Z, Braun E, Qiu W, Shi Y-F, Marcelino-Guimarães FC, Navarre D, et al. Positive and Negative Roles for Soybean MPK6 in Regulating Defense Responses. Mol Plant-Microbe Interactions®. 2014;27: 824–834. [CrossRef]
- Liu J-Z, Horstman HD, Braun E, Graham MA, Zhang C, Navarre D, et al. Soybean Homologs of MPK4 Negatively Regulate Defense Responses and Positively Regulate Growth and Development. Plant Physiol. 2011;157: 1363–1378. [CrossRef]
- Igarashi A, Yamagata K, Sugai T, Takahashi Y, Sugawara E, Tamura A, et al. Apple latent spherical virus vectors for reliable and effective virus-induced gene silencing among a broad range of plants including tobacco, tomato, Arabidopsis thaliana, cucurbits, and legumes. Virology. 2009;386: 407–416. [CrossRef]
- Gedling CR, Ali EM, Gunadi A, Finer JJ, Xie K, Liu Y, et al. Improved apple latent spherical virus-induced gene silencing in multiple soybean genotypes through direct inoculation of agro-infiltrated Nicotiana benthamiana extract. Plant Methods. 2018;14: 19. [CrossRef]
- Takahashi T, Sugawara T, Yamatsuta T, Isogai M, Natsuaki T, Yoshikawa N. Analysis of the Spatial Distribution of Identical and Two Distinct Virus Populations Differently Labeled with Cyan and Yellow Fluorescent Proteins in Coinfected Plants. Phytopathology. 2007;97: 1200–1206. [CrossRef]
- Sasaki S, Yamagishi N, Yoshikawa N. Efficient virus-induced gene silencing in apple, pear and Japanese pear using Apple latent spherical virus vectors. Plant Methods. 2011;7: 15. [CrossRef]
- Yamagishi N, Yoshikawa N. Virus-induced gene silencing in soybean seeds and the emergence stage of soybean plants with Apple latent spherical virus vectors. Plant Mol Biol. 2009;71: 15–24. [CrossRef]
- Lin J, Guo J, Finer J, Dorrance AE, Redinbaugh MG, Qu F. The Bean Pod Mottle Virus RNA2-Encoded 58-Kilodalton Protein P58 Is Required in cis for RNA2 Accumulation. Simon A, editor. J Virol. 2014;88: 3213–3222. [CrossRef]
- Shin SY, Park M-R, Kim H-S, Moon JS, Lee H-J. Virus-induced gene silencing shows that LATE FLOWERING plays a role in promoting flower development in soybean. Plant Growth Regul. 2023;99: 229–239. [CrossRef]
- Zaulda FA, Yang SH, Han J, Mlotshwa S, Dorrance A, Qu F. A cowpea severe mosaic virus-based vector simplifies virus-induced gene silencing and foreign protein expression in soybean. Plant Methods. 2022;18: 116. [CrossRef]
- Chen X, Bruening G. Nucleotide sequence and genetic map of cowpea severe mosaic virus RNA 2 and comparisons with RNA 2 of other comoviruses. Virology. 1992;187: 682–692. [CrossRef]
- Chen X, Bruening G. Cloned DNA copies of cowpea severe mosaic virus genomic RNAs: Infectious transcripts and complete nucleotide sequence of RNA 1. Virology. 1992;191: 607–618. [CrossRef]
- Karhoff S, Vargas-Garcia C, Lee S, Mian MAR, Graham MA, Dorrance AE, et al. Identification of Candidate Genes for a Major Quantitative Disease Resistance Locus From Soybean PI 427105B for Resistance to Phytophthora sojae. Front Plant Sci. 2022;13: 893652. [CrossRef]
- Karhoff S, Lee S, Mian MAR, Ralston TI, Niblack TL, Dorrance AE, et al. Phenotypic Characterization of a Major Quantitative Disease Resistance Locus for Partial Resistance to Phytophthora sojae. Crop Sci. 2019;59: 968–980. [CrossRef]
- Sun R, Han J, Zheng L, Qu F. The AC2 Protein of a Bipartite Geminivirus Stimulates the Transcription of the BV1 Gene through Abscisic Acid Responsive Promoter Elements. Viruses. 2020;12. [CrossRef]
- Zhang S, Sun R, Perdoncini Carvalho C, Han J, Zheng L, Qu F. Replication-Dependent Biogenesis of Turnip Crinkle Virus Long Noncoding RNAs. Simon AE, editor. J Virol. 2021;95: e00169-21. [CrossRef]
- Cao M, Ye X, Willie K, Lin J, Zhang X, Redinbaugh MG, et al. The Capsid Protein of Turnip Crinkle Virus Overcomes Two Separate Defense Barriers To Facilitate Systemic Movement of the Virus in Arabidopsis. J Virol. 2010;84: 7793–7802. [CrossRef]
- Steinberg SF. Post-translational modifications at the ATP-positioning G-loop that regulate protein kinase activity. Pharmacol Res. 2018;135: 181–187. [CrossRef]
- Reinhardt R, Leonard TA. A critical evaluation of protein kinase regulation by activation loop autophosphorylation. eLife. 2023;12: e88210. [CrossRef]
- Spassov DS, Ruiz-Saenz A, Piple A, Moasser MM. A Dimerization Function in the Intrinsically Disordered N-Terminal Region of Src. Cell Rep. 2018;25: 449-463.e4. [CrossRef]


Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).



