Submitted:
11 March 2026
Posted:
12 March 2026
You are already at the latest version
Abstract
Keywords:
Background
Clinical Relevance and Importance Towards Improving Patient Outcomes
Methods
Aim of the Study
Research Questions
Search Strategy and Reproducibility
- Additional updates: October 2025 – March 10, 2026
- Date of last search: March 10, 2026
- Language: English
Databases Searched
- PubMed
- MEDLINE
- Scopus
- Web of Science Core Collection
Timeframe of Search
Search Concept Framework
-
Disease Context
- o
- Carcinoid tumors
- o
- Neuroendocrine tumors (NETs)
- o
- Small intestinal neuroendocrine tumors (SI-NETs)
-
Metastatic Process
- o
- Liver metastasis
- o
- Hepatic tropism
- o
- Organ-specific metastasis
- o
- Metastatic colonization
-
Developmental Regulators and Signaling Pathways
- o
- Transcription factors: FOXA1/2, GATA4/6, SOX4/9, HES1
- o
- Signaling pathways: Notch, Wnt/β-catenin, Hippo/YAP-TAZ, FGF/HGF, TGF-β
- o
- Ligand signaling: Dll1/4
Search Syntax and Query Development
- Controlled vocabulary terms (e.g., MeSH terms in PubMed/MEDLINE)
- Free-text keywords
- Gene and pathway names
- Synonyms and alternate terminology
- OR to group related synonyms or molecular regulators
- AND to intersect disease context, metastasis, and molecular pathways
- Quotation marks to capture exact phrases
- Wildcards/truncation symbols (e.g., * ) to capture variations of key terms
Search Validation
- Initial searches were refined through pilot testing of queries
- Results were examined to confirm retrieval of key landmark studies in carcinoid liver metastasis
- Additional keyword variations were added iteratively based on retrieved articles
Reproducibility Measures
- Standardized search queries were developed and documented for each database.
- Searches were performed using identical conceptual structures across all databases.
- All search results were exported and archived for verification and deduplication.
- The search strategy was repeated and verified during the revision phase to confirm consistent retrieval patterns.
- Multiple rounds of manual screening ensured that relevant studies were not inadvertently excluded.
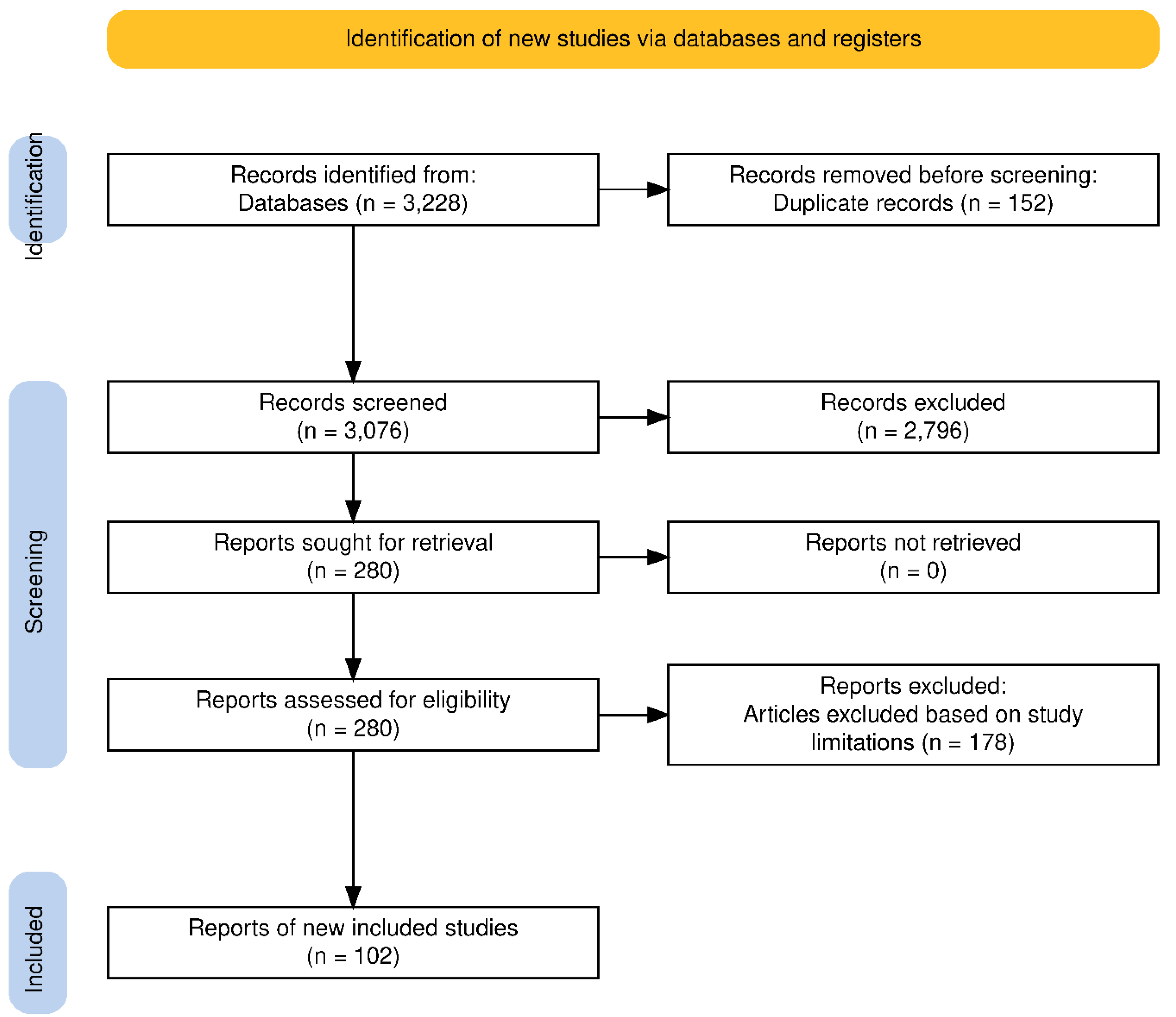
- 3228 records identified
- 3076 after duplicates removed
- 2796 excluded after title/abstract screening
- 178 excluded after full-text review during data extraction
- 102 studies included in the final analysis
Objectives of the Search
- To identify shared transcriptional regulators between SI-NECs and hepatocytes
- To delineate common signaling networks that may favor hepatic colonization
- To understand how developmental mimicry and lineage convergence contribute to organ-specific metastasis
Screening and Eligibility Criteria
Initial Screening
- Titles and abstracts were screened for relevance to SI-NECs, hepatocytes, liver metastasis, and developmental genes/TFs/signaling pathways.
Full-Text Evaluation
- Studies providing insights directly or indirectly into shared pathways or transcriptional programs were selected.
Data Extraction Criteria
- Role of FOXA1/2, GATA4/6, SOX4/9, HES1/NOTCH, Wnt/β-catenin, Hippo/YAP-TAZ, FGF/HGF, TGF-β, and Dll1/4 in hepatic colonization
- Evidence linking these pathways to liver-specific metastasis in neuroendocrine tumors
Inclusion and Exclusion Criteria
- Studies focused on liver metastasis of carcinoid tumor
- Research focusing on shared developmental regulators between SI-NECs and hepatocytes
- Studies focusing on liver tropism of carcinoid tumor mets.
- Studies not involving SI-NECs developmental regulators.
- Studies not involving Hepatic developmental regulators.
- Research unrelated to liver tropism of carcinoid tumor mets.
- Articles lacking insights into the shared developmental regulators between SI-NECs and Hepatocytes.
- Articles that did not conform to the study focus.
- Insufficient methodological rigor.
Assessment of Article Quality and Potential Biases
Quality Assessment
Potential Biases Assessment
- Publication Bias: To address the potential for publication bias, a comprehensive search strategy was adopted to include a balanced representation of both positive and negative results, incorporating a wide range of published articles from databases like Google Scholar.
- Selection Bias: Predefined and transparent inclusion criteria were applied to minimize subjectivity in the selection process. Articles were chosen based on their relevance to the study’s objectives, adhering strictly to these criteria. This approach reduced the risk of subjectivity and ensured that the selection process was objective and consistent.
- Reporting Bias: To mitigate reporting bias, articles were checked for inconsistencies or missing data. Multiple detailed reviews of the methodologies and results were conducted for all selected articles to identify and address any reporting bias.
Results
Investigating Determinants Underlying Liver Tropism in Carcinoid Tumor Metastasis and Its Clinical Implications:
Discussion
Conclusions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
Protocol
Abbreviations
| SI-NECs | Small Intestinal Neuroendocrine Cells |
| TFs | Transcription Factors |
| FOXA1/2 | Forkhead Box A1 / A2 |
| GATA4/6 | GATA Binding Protein 4 / 6 |
| SOX4/9 | SRY-Box Transcription Factor 4 / 9 |
| HES1 | Hairy and Enhancer of Split-1 |
| NOTCH | Neurogenic Locus Notch Homolog |
| Dll1/4 | Delta-like Ligand 1 / 4 |
| Wnt | Wingless-related Integration Site |
| β-catenin | Beta-catenin |
| Hippo | Hippo Signaling Pathway |
| YAP/TAZ | Yes-associated Protein / Transcriptional Coactivator with PDZ-binding Motif |
| FGF | Fibroblast Growth Factor |
| HGF | Hepatocyte Growth Factor |
| TGF-β | Transforming Growth Factor Beta |
| c-MET | Mesenchymal-Epithelial Transition Factor Receptor |
| ECM | Extracellular Matrix |
References
- Veenendaal, L.M.; Borel Rinkes, I.H.; Lips, C.J.; van Hillegersberg, R. Liver metastases of neuroendocrine tumours; early reduction of tumour load to improve life expectancy. World J Surg Oncol. 2006, 4, 35. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Gravante, G.; De Liguori Carino, N.; Overton, J.; Manzia, T.M.; Orlando, G. Primary carcinoids of the liver: a review of symptoms, diagnosis and treatments. Dig Surg. 2008, 25, 364–368. [Google Scholar] [CrossRef] [PubMed]
- Schwartz, G.; Colanta, A.; Gaetz, H.; Olichney, J.; Attiyeh, F. Primary carcinoid tumors of the liver. World J Surg Oncol. 2008, 6, 91. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Vitale, G; Carra, S; Alessi, Y; Campolo, F; Pandozzi, C; Zanata, I; Colao, A; Faggiano, A. On Behalf Of The Nike Group. Carcinoid Syndrome: Preclinical Models and Future Therapeutic Strategies. Int J Mol Sci. 2023, 24, 3610. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Ober, E.A.; Lemaigre, F.P. Development of the liver: Insights into organ and tissue morphogenesis. J Hepatol. 2018, 68, 1049–1062. [Google Scholar] [CrossRef] [PubMed]
- Rubin de Celis Ferrari, A.C.; Glasberg, J.; Riechelmann, R.P. Carcinoid syndrome: update on the pathophysiology and treatment. Clinics 2018, 73, e490s. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Marasco, M.; Romano, E.; Arrivi, G.; Prosperi, D.; Rinzivillo, M.; Caruso, D.; Mercantini, P.; Rossi, M.; Faggiano, A.; Panzuto, F. Exploring Carcinoid Syndrome in Neuroendocrine Tumors: Insights from a Multidisciplinary Narrative Review. Cancers 2024, 16, 3831. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Subash, N.; Papali, M.M.; Bahadur, K.P.; Avanthika, C.; Jhaveri, S.; Thannir, S.; Joshi, M.; Valisekka, S.S. Recent Advances in the Diagnosis and Management of Carcinoid Syndrome. Dis Mon. 2022, 68, 101304. [Google Scholar] [CrossRef] [PubMed]
- Hofland, J.; Herrera-Martínez, A.D.; Zandee, W.T.; de Herder, W.W. Management of carcinoid syndrome: a systematic review and meta-analysis. Endocr Relat Cancer 2019, 26, R145–R156. [Google Scholar] [CrossRef] [PubMed]
- Kimbrough, C.W.; Beal, E.W.; Dillhoff, M.E.; Schmidt, C.R.; Pawlik, T.M.; Lopez-Aguiar, A.G.; Poultsides, G.; Makris, E.; Rocha, F.G.; Crown, A.; Abbott, D.E.; Fisher, A.V.; Fields, R.C.; Krasnick, B.A.; Idrees, K.; Marincola-Smith, P.; Cho, C.S.; Beems, M.; Maithel, S.K.; Cloyd, J.M. Influence of carcinoid syndrome on the clinical characteristics and outcomes of patients with gastroenteropancreatic neuroendocrine tumors undergoing operative resection. Surgery 2019, 165, 657–663. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Ahlman, H.; Westberg, G.; Wängberg, B.; Nilsson, O.; Tylén, U.; Scherstén, T.; Tisell, L.E. Treatment of liver metastases of carcinoid tumors. World J Surg. 1996, 20, 196–202. [Google Scholar] [CrossRef] [PubMed]
- Zuetenhorst, J.M.; Taal, B.G. Metastatic carcinoid tumors: a clinical review. Oncologist 2005, 10, 123–131. [Google Scholar] [CrossRef] [PubMed]
- Lee, C.S.; Friedman, J.R.; Fulmer, J.T.; Kaestner, K.H. The initiation of liver development is dependent on Foxa transcription factors. Nature 2005, 435, 944–947. [Google Scholar] [CrossRef] [PubMed]
- Heslop, J.A.; Duncan, S.A. FoxA factors: the chromatin key and doorstop essential for liver development and function. Genes Dev. 2020, 34(15-16), 1003–1004. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Reizel, Y.; Morgan, A.; Gao, L.; Lan, Y.; Manduchi, E.; Waite, E.L.; Wang, A.W.; Wells, A.; Kaestner, K.H. Collapse of the hepatic gene regulatory network in the absence of FoxA factors. Genes Dev. 2020, 34(15-16), 1039–1050. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Lan, Q.; Cao, M.; Kollipara, R.K.; Rosa, J.B.; Kittler, R.; Jiang, H. FoxA transcription factor Fork head maintains the intestinal stem/progenitor cell identities in Drosophila. Dev Biol. 2018, 433, 324–343. [Google Scholar] [CrossRef] [PubMed]
- Gosalia, N.; Yang, R.; Kerschner, J.L.; Harris, A. FOXA2 regulates a network of genes involved in critical functions of human intestinal epithelial cells. Physiol Genomics 2015, 47, 290–297. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Kerschner, J.L.; Gosalia, N.; Leir, S.H.; Harris, A. Chromatin remodeling mediated by the FOXA1/A2 transcription factors activates CFTR expression in intestinal epithelial cells. Epigenetics 2014, 9, 557–565. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Park, J.W.; Lee, J.K.; Witte, O.N.; Huang, J. FOXA2 is a sensitive and specific marker for small cell neuroendocrine carcinoma of the prostate. Mod Pathol. 2017, 30, 1262–1272. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Han, M.; Li, F.; Zhang, Y.; Dai, P.; He, J.; Li, Y.; Zhu, Y.; Zheng, J.; Huang, H.; Bai, F.; Gao, D. FOXA2 drives lineage plasticity and KIT pathway activation in neuroendocrine prostate cancer. Cancer Cell. 2022, 40, 1306–1323.e8. [Google Scholar] [CrossRef] [PubMed]
- Katoh, M.; Igarashi, M.; Fukuda, H.; Nakagama, H.; Katoh, M. Cancer genetics and genomics of human FOX family genes. Cancer Lett. 2013, 328, 198–206. [Google Scholar] [CrossRef] [PubMed]
- Tosun Yıldırım, H.; Nergiz, D.; Sadullahoğlu, C.; Süren, D.; Yıldırım, Ş.; Üreyen, I.; Sezer, C. FOXA1 is associated with high tumor grade, myometrial invasion and lymph node invasion in endometrial endometrioid carcinoma. Ginekol Pol. 2021, 92, 544–549. [Google Scholar] [CrossRef] [PubMed]
- Zhao, R.; Watt, A.J.; Li, J.; Luebke-Wheeler, J.; Morrisey, E.E.; Duncan, S.A. GATA6 is essential for embryonic development of the liver but dispensable for early heart formation. Mol Cell Biol. 2005, 25, 2622–2631. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Haworth, K.E.; Kotecha, S.; Mohun, T.J.; Latinkic, B.V. GATA4 and GATA5 are essential for heart and liver development in Xenopus embryos. BMC Dev Biol. 2008, 8, 74. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Zheng, R.; Rebolledo-Jaramillo, B.; Zong, Y.; Wang, L.; Russo, P.; Hancock, W.; Stanger, B.Z.; Hardison, R.C.; Blobel, G.A. Function of GATA factors in the adult mouse liver. PLoS One 2013, 8, e83723. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Borok, M.J.; Papaioannou, V.E.; Sussel, L. Unique functions of Gata4 in mouse liver induction and heart development. Dev Biol. 2016, 410, 213–222. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Zhang, L.; He, J.B. Progress of GATA6 in liver development. Yi Chuan 2018, 40, 22–32. [Google Scholar] [CrossRef] [PubMed]
- Aronson, B.E.; Stapleton, K.A.; Krasinski, S.D. Role of GATA factors in development, differentiation, and homeostasis of the small intestinal epithelium. Am J Physiol Gastrointest Liver Physiol. 2014, 306, G474–G490. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Okumura, T.; Takeda, K.; Kuchiki, M.; Akaishi, M.; Taniguchi, K.; Adachi-Yamada, T. GATAe regulates intestinal stem cell maintenance and differentiation in Drosophila adult midgut. Dev Biol. 2016, 410, 24–35. [Google Scholar] [CrossRef] [PubMed]
- Walker, E.M.; Thompson, C.A.; Battle, M.A. GATA4 and GATA6 regulate intestinal epithelial cytodifferentiation during development. Dev Biol. 2014, 392, 283–294. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Aronson, B.E.; Stapleton, K.A.; Krasinski, S.D. Role of GATA factors in development, differentiation, and homeostasis of the small intestinal epithelium. Am J Physiol Gastrointest Liver Physiol. 2014, 306, G474–G490. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Kohlnhofer, B.M.; Thompson, C.A.; Walker, E.M.; Battle, M.A. GATA4 regulates epithelial cell proliferation to control intestinal growth and development in mice. Cell Mol Gastroenterol Hepatol. 2016, 2, 189–209. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Shang, T.; Jiang, T.; Cui, X.; Pan, Y.; Feng, X.; Dong, L.; Wang, H. Diverse functions of SOX9 in liver development and homeostasis and hepatobiliary diseases. Genes Dis. 2023, 11, 100996. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Yin, C. Molecular mechanisms of Sox transcription factors during the development of liver, bile duct, and pancreas. Semin Cell Dev Biol. 2017, 63, 68–78. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Yin, C. Molecular mechanisms of Sox transcription factors during the development of liver, bile duct, and pancreas. Semin Cell Dev Biol. 2017, 63, 68–78. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Fu, L.; Shi, Y.B. The Sox transcriptional factors: Functions during intestinal development in vertebrates. Semin Cell Dev Biol. 2017, 63, 58–67. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Fu, L.; Shi, Y.B. The Sox transcriptional factors: Functions during intestinal development in vertebrates. Semin Cell Dev Biol. 2017, 63, 58–67. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- de Santa Barbara, P.; van den Brink, G.R.; RobertsDJ. Development and differentiation of the intestinal epithelium. Cell Mol Life Sci. 2003, 60, 1322–1332. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Poncy, A.; Antoniou, A.; Cordi, S.; Pierreux, C.E.; Jacquemin, P.; Lemaigre, F.P. Transcription factors SOX4 and SOX9 cooperatively control development of bile ducts. Dev Biol. 2015, 404, 136–148. [Google Scholar] [CrossRef] [PubMed]
- Shang, T.; Jiang, T.; Cui, X.; Pan, Y.; Feng, X.; Dong, L.; Wang, H. Diverse functions of SOX9 in liver development and homeostasis and hepatobiliary diseases. Genes Dis. 2023, 11, 100996. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Kawaguchi, Y. Sox9 and programming of liver and pancreatic progenitors. J Clin Invest. 2013, 123, 1881–1886. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Bukong, T.N.; Lo, T.; Szabo, G.; Dolganiuc, A. Novel developmental biology-based protocol of embryonic stem cell differentiation to morphologically sound and functional yet immature hepatocytes. Liver Int. 2012, 32, 732–741. [Google Scholar] [CrossRef] [PubMed]
- Zhang, K.; Zhang, Y.Q.; Ai, W.B.; Hu, Q.T.; Zhang, Q.J.; Wan, L.Y.; Wang, X.L.; Liu, C.B.; Wu, J.F. Hes1, an important gene for activation of hepatic stellate cells, is regulated by Notch1 and TGF-β/BMP signaling. World J Gastroenterol. 2015, 21, 878–887. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Auguet, T.; Bertran, L.; Binetti, J.; Aguilar, C.; Martínez, S.; Guiu-Jurado, E.; Sabench, F.; Adalid, L.; Porras, J.A.; Riesco, D.; Del Castillo, D.; Richart, C. Hepatocyte Notch Signaling Deregulation Related to Lipid Metabolism in Women with Obesity and Nonalcoholic Fatty Liver. Obesity 2020, 28, 1487–1493. [Google Scholar] [CrossRef] [PubMed]
- Sun, Y.; Tong, H.; Chu, X.; Li, Y.; Zhang, J.; Ding, Y.; Zhang, S.; Gui, X.; Chen, C.; Xu, M.; Li, Z.; Gardiner, E.E.; Andrews, R.K.; Zeng, L.; Xu, K.; Qiao, J. Notch1 regulates hepatic thrombopoietin production. Blood 2024, 143, 2778–2790. [Google Scholar] [CrossRef] [PubMed]
- Köhler, C.; Bell, A.W.; Bowen, W.C.; Monga, S.P.; Fleig, W.; Michalopoulos, G.K. Expression of Notch-1 and its ligand Jagged-1 in rat liver during liver regeneration. Hepatology 2004, 39, 1056–1065. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Chen, G.; Sun, L.; Yu, M.; Meng, D.; Wang, W.; Yang, Y.; Yang, H. The Jagged-1/Notch-1/Hes-1 pathway is involved in intestinal adaptation in a massive small bowel resection rat model. Dig Dis Sci. 2013, 58, 2478–2486. [Google Scholar] [CrossRef] [PubMed]
- Kay, S.K.; Harrington, H.A.; Shepherd, S.; Brennan, K.; Dale, T.; Osborne, J.M.; Gavaghan, D.J.; Byrne, H.M. The role of the Hes1 crosstalk hub in Notch-Wnt interactions of the intestinal crypt. PLoS Comput Biol. 2017, 13, e1005400. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Hu, J.; Li, J.; Dai, C.; Ren, J.; Yang, W.; He, C.; Meng, F.; Dai, C.; Zeng, S. HES1 deficiency impairs development of human intestinal mesenchyme by suppressing WNT5A expression. Biochem Biophys Res Commun. 2023, 655, 50–58. [Google Scholar] [CrossRef] [PubMed]
- Kunnimalaiyaan, M.; Yan, S.; Wong, F.; Zhang, Y.W.; Chen, H. Hairy Enhancer of Split-1 (HES-1), a Notch1 effector, inhibits the growth of carcinoid tumor cells. Surgery 2005, 138, 1137–1142; discussion 1142. [Google Scholar] [CrossRef] [PubMed]
- Harada, K.; Sato, Y.; Ikeda, H.; Hsu, M.; Igarashi, S.; Nakanuma, Y. Notch1-Hes1 signalling axis in the tumourigenesis of biliary neuroendocrine tumours. J Clin Pathol. 2013, 66, 386–391. [Google Scholar] [CrossRef] [PubMed]
- Curry, C.L.; Reed, L.L.; Nickoloff, B.J.; Miele, L.; Foreman, K.E. Notch-independent regulation of Hes-1 expression by c-Jun N-terminal kinase signaling in human endothelial cells. Lab Invest. 2006, 86, 842–852. [Google Scholar] [CrossRef] [PubMed]
- Kim, J.T.; Li, J.; Jang, E.R.; Gulhati, P.; Rychahou, P.G.; Napier, D.L.; Wang, C.; Weiss, H.L.; Lee, E.Y.; Anthony, L.; Townsend, CMJr; Liu, C.; Evers, B.M. Deregulation of Wnt/β-catenin signaling through genetic or epigenetic alterations in human neuroendocrine tumors. Carcinogenesis 2013, 34, 953–961. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Jin, X.F.; Spoettl, G.; Maurer, J.; Nölting, S.; Auernhammer, C.J. Inhibition of Wnt/β-Catenin Signaling in Neuroendocrine Tumors in vitro: Antitumoral Effects. Cancers 2020, 12, 345. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Jin, Y.; Anbarchian, T.; Wu, P.; Sarkar, A.; Fish, M.; Peng, W.C.; Nusse, R. Wnt signaling regulates hepatocyte cell division by a transcriptional repressor cascade. Proc Natl Acad Sci U S A 2022, 119, e2203849119. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Chan, S.C.; Zhang, Y.; Pontoglio, M.; Igarashi, P. Hepatocyte nuclear factor-1β regulates Wnt signaling through genome-wide competition with β-catenin/lymphoid enhancer binding factor. Proc Natl Acad Sci U S A 2019, 116, 24133–24142. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Monga, S.P.; Mars, W.M.; Pediaditakis, P.; Bell, A.; Mulé, K.; Bowen, W.C.; Wang, X.; Zarnegar, R.; Michalopoulos, G.K. Hepatocyte growth factor induces Wnt-independent nuclear translocation of beta-catenin after Met-beta-catenin dissociation in hepatocytes. Cancer Res. 2002, 62, 2064–2071. [Google Scholar] [PubMed]
- Moparthi, L.; Koch, S. Wnt signaling in intestinal inflammation. Differentiation 2019, 108, 24–32. [Google Scholar] [CrossRef] [PubMed]
- Mah, A.T.; Yan, K.S.; Kuo, C.J. Wnt pathway regulation of intestinal stem cells. J Physiol. 2016, 594, 4837–4847. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Gasnier, M.; Lim, H.Y.G.; Barker, N. Role of Wnt signaling in the maintenance and regeneration of the intestinal epithelium. Curr Top Dev Biol. 2023, 153, 281–326. [Google Scholar] [CrossRef] [PubMed]
- Sherwood, R.I.; Maehr, R.; Mazzoni, E.O.; Melton, D.A. Wnt signaling specifies and patterns intestinal endoderm. Mech Dev. 2011, 128(7-10), 387–400. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Pinto, D.; Clevers, H. Wnt control of stem cells and differentiation in the intestinal epithelium. Exp Cell Res. 2005, 306, 357–363. [Google Scholar] [CrossRef] [PubMed]
- Tsai, S.M.; Liu, D.W.; Wang, W.P. Fibroblast growth factor (Fgf) signaling pathway regulates liver homeostasis in zebrafish. Transgenic Res. 2013, 22, 301–314. [Google Scholar] [CrossRef] [PubMed]
- Padrissa-Altés, S.; Bachofner, M.; Bogorad, R.L.; Pohlmeier, L.; Rossolini, T.; Böhm, F.; Liebisch, G.; Hellerbrand, C.; Koteliansky, V.; Speicher, T.; Werner, S. Control of hepatocyte proliferation and survival by Fgf receptors is essential for liver regeneration in mice. Gut 2015, 64, 1444–1453. [Google Scholar] [CrossRef] [PubMed]
- Berg, T.; Rountree, C.B.; Lee, L.; Estrada, J.; Sala, F.G.; Choe, A.; Veltmaat, J.M.; De Langhe, S.; Lee, R.; Tsukamoto, H.; Crooks, G.M.; Bellusci, S.; Wang, K.S. Fibroblast growth factor 10 is critical for liver growth during embryogenesis and controls hepatoblast survival via beta-catenin activation. Hepatology 2007, 46, 1187–1197. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Danopoulos, S.; Schlieve, C.R.; Grikscheit, T.C.; Al Alam, D. Fibroblast Growth Factors in the Gastrointestinal Tract: Twists and Turns. Dev Dyn. 2017, 246, 344–352. [Google Scholar] [CrossRef] [PubMed]
- Katoh, M.; Katoh, M. FGF signaling network in the gastrointestinal tract (review). Int J Oncol. 2006, 29, 163–168. [Google Scholar] [CrossRef] [PubMed]
- Dessimoz, J.; Opoka, R.; Kordich, J.J.; Grapin-Botton, A.; Wells, J.M. FGF signaling is necessary for establishing gut tube domains along the anterior-posterior axis in vivo. Mech Dev. 2006, 123, 42–55. [Google Scholar] [CrossRef] [PubMed]
- Vitale, G.; Cozzolino, A.; Malandrino, P.; Minotta, R.; Puliani, G.; Saronni, D.; Faggiano, A.; Colao, A. Role of FGF System in Neuroendocrine Neoplasms: Potential Therapeutic Applications. Front Endocrinol (Lausanne) 2021, 12, 665631. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Presta, M.; Chiodelli, P.; Giacomini, A.; Rusnati, M.; Ronca, R. Fibroblast growth factors (FGFs) in cancer: FGF traps as a new therapeutic approach. Pharmacol Ther. 2017, 179, 171–187. [Google Scholar] [CrossRef] [PubMed]
- Geske, M.J.; Zhang, X.; Patel, K.K.; Ornitz, D.M.; Stappenbeck, T.S. Fgf9 signaling regulates small intestinal elongation and mesenchymal development. Development 2008, 135, 2959–2968. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Nakamura, T.; Sakai, K.; Nakamura, T.; Matsumoto, K. Hepatocyte growth factor twenty years on: Much more than a growth factor. J Gastroenterol Hepatol. 2011, 26, 188–202. [Google Scholar] [CrossRef] [PubMed]
- Yang, L.; Roh, Y.S.; Song, J.; Zhang, B.; Liu, C.; Loomba, R.; Seki, E. Transforming growth factor beta signaling in hepatocytes participates in steatohepatitis through regulation of cell death and lipid metabolism in mice. Hepatology 2014, 59, 483–495. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Sanchez, A.; Alvarez, A.M.; Benito, M.; Fabregat, I. Transforming growth factor beta modulates growth and differentiation of fetal hepatocytes in primary culture. J Cell Physiol. 1995, 165, 398–405. [Google Scholar] [CrossRef] [PubMed]
- Meindl-Beinker, N.M.; Dooley, S. Transforming growth factor-beta and hepatocyte transdifferentiation in liver fibrogenesis. J Gastroenterol Hepatol. 2008, 23, S122–S127. [Google Scholar] [CrossRef] [PubMed]
- Coulouarn, C.; Factor, V.M.; Thorgeirsson, S.S. Transforming growth factor-beta gene expression signature in mouse hepatocytes predicts clinical outcome in human cancer. Hepatology 2008, 47, 2059–2067. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Beck, P.L.; Rosenberg, I.M.; Xavier, R.J.; Koh, T.; Wong, J.F.; Podolsky, D.K. Transforming growth factor-beta mediates intestinal healing and susceptibility to injury in vitro and in vivo through epithelial cells. Am J Pathol. 2003, 162, 597–608. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Achyut, B.R.; Yang, L. Transforming growth factor-β in the gastrointestinal and hepatic tumor microenvironment. Gastroenterology 2011, 141, 1167–1178. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Gough, N.R.; Xiang, X.; Mishra, L. TGF-β Signaling in Liver, Pancreas, and Gastrointestinal Diseases and Cancer. Gastroenterology 2021, 161, 434–452.e15. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Liao, Y.; Zhang, M.; Lönnerdal, B. Growth factor TGF-β induces intestinal epithelial cell (IEC-6) differentiation: miR-146b as a regulatory component in the negative feedback loop. Genes Nutr. 2013, 8, 69–78. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Kidd, M.; Schimmack, S.; Lawrence, B.; Alaimo, D.; Modlin, I.M. EGFR/TGFα and TGFβ/CTGF Signaling in Neuroendocrine Neoplasia: Theoretical Therapeutic Targets. Neuroendocrinology 2013, 97, 35–44. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Dockray, G.J. Keeping neuroendocrine cells in check: roles for TGFbeta, Smads, and menin? Gut. 2003, 52, 1237–1239. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Nguyen-Lefebvre, A.T.; Selzner, N.; Wrana, J.L.; Bhat, M. The hippo pathway: A master regulator of liver metabolism, regeneration, and disease. FASEB J. 2021, 35, e21570. [Google Scholar] [CrossRef] [PubMed]
- Yimlamai, D.; Christodoulou, C.; Galli, G.G.; Yanger, K.; Pepe-Mooney, B.; Gurung, B.; Shrestha, K.; Cahan, P.; Stanger, B.Z.; Camargo, F.D. Hippo pathway activity influences liver cell fate. Cell. 2014, 157, 1324–1338. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Driskill, J.H.; Pan, D. The Hippo Pathway in Liver Homeostasis and Pathophysiology. Annu Rev Pathol. 2021, 16, 299–322. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Hong, A.W.; Meng, Z.; Guan, K.L. The Hippo pathway in intestinal regeneration and disease. Nat Rev Gastroenterol Hepatol. 2016, 13, 324–337. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Yu, F.X.; Meng, Z.; Plouffe, S.W.; Guan, K.L. Hippo pathway regulation of gastrointestinal tissues. Annu Rev Physiol. 2015, 77, 201–227. [Google Scholar] [CrossRef] [PubMed]
- Xiao, Y.; Tian, Y.; Zhang, J.; Li, Q.; Shi, W.; Huang, X. Small intestinal submucosa promotes angiogenesis via the Hippo pathway to improve vaginal repair. Biomol Biomed. 2023, 23, 838–847. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Harvey, K.F.; Tang, T.T. Targeting the Hippo pathway in cancer. Nat Rev Drug Discov. 2025, 24, 852–869. [Google Scholar] [CrossRef] [PubMed]
- Zygulska, A.L.; Krzemieniecki, K.; Pierzchalski, P. Hippo pathway - brief overview of its relevance in cancer. J Physiol Pharmacol. 2017, 68, 311–335. [Google Scholar] [PubMed]
- Calses, P.C.; Crawford, J.J.; Lill, J.R.; Dey, A. Hippo Pathway in Cancer: Aberrant Regulation and Therapeutic Opportunities. Trends Cancer 2019, 5, 297–307. [Google Scholar] [CrossRef] [PubMed]
- Li, F.L.; Guan, K.L. The two sides of Hippo pathway in cancer. Semin Cancer Biol. 2022, 85, 33–42. [Google Scholar] [CrossRef] [PubMed]
- You, W.K.; Schuetz, T.J.; Lee, S.H. Targeting the DLL/Notch Signaling Pathway in Cancer: Challenges and Advances in Clinical Development. Mol Cancer Ther. 2023, 22, 3–11. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Crabtree, J.S.; Singleton, C.S.; Miele, L. Notch Signaling in Neuroendocrine Tumors. Front Oncol. 2016, 6, 94. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Katoh, M.; Katoh, M. Precision medicine for human cancers with Notch signaling dysregulation (Review). Int J Mol Med. 2020, 45, 279–297. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Noah, T.K.; Shroyer, N.F. Notch in the intestine: regulation of homeostasis and pathogenesis. Annu Rev Physiol. 2013, 75, 263–288. [Google Scholar] [CrossRef] [PubMed]
- Tsai, Y.H.; Wu, A.; Wu, J.H.; Capeling, M.M.; Holloway, E.M.; Huang, S.; Czerwinkski, M.; Glass, I.; Higgins, P.D.R.; Spence, J.R. Acquisition of NOTCH dependence is a hallmark of human intestinal stem cell maturation. Stem Cell Reports 2022, 17, 1138–1153. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Pellegrinet, L.; Rodilla, V.; Liu, Z.; Chen, S.; Koch, U.; Espinosa, L.; Kaestner, K.H.; Kopan, R.; Lewis, J.; Radtke, F. Dll1- and dll4-mediated notch signaling are required for homeostasis of intestinal stem cells. Gastroenterology 2011, 140, 1230–1240.e1–7. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Adams, J.M.; Jafar-Nejad, H. The Roles of Notch Signaling in Liver Development and Disease. Biomolecules 2019, 9, 608. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Zong, Y.; Panikkar, A.; Xu, J.; Antoniou, A.; Raynaud, P.; Lemaigre, F.; Stanger, B.Z. Notch signaling controls liver development by regulating biliary differentiation. Development 2009, 136, 1727–1739. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Wang, T.; You, N.; Tao, K.; Wang, X.; Zhao, G.; Xia, N.; Li, N.; Tang, L.; Liu, W.; Dou, K. Notch is the key factor in the process of fetal liver stem/progenitor cells differentiation into hepatocytes. Dev Growth Differ. 2012, 54, 605–617. [Google Scholar] [CrossRef] [PubMed]
- Morell, C.M.; Strazzabosco, M. Notch signaling and new therapeutic options in liver disease. J Hepatol. 2014, 60, 885–890. [Google Scholar] [CrossRef] [PubMed]
Short Biography of Authors
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).



