Submitted:
10 March 2026
Posted:
11 March 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
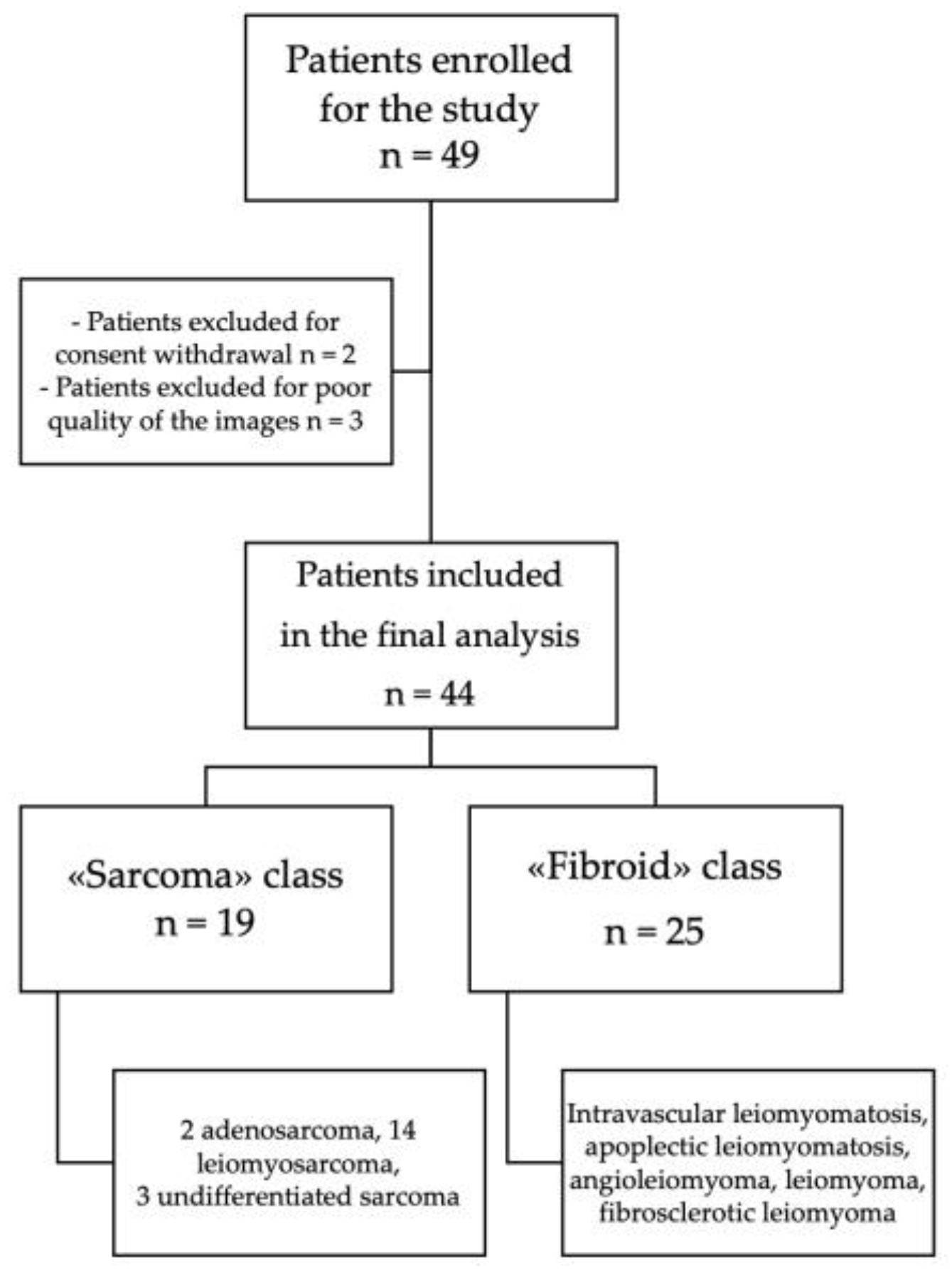
2.1. Dataset
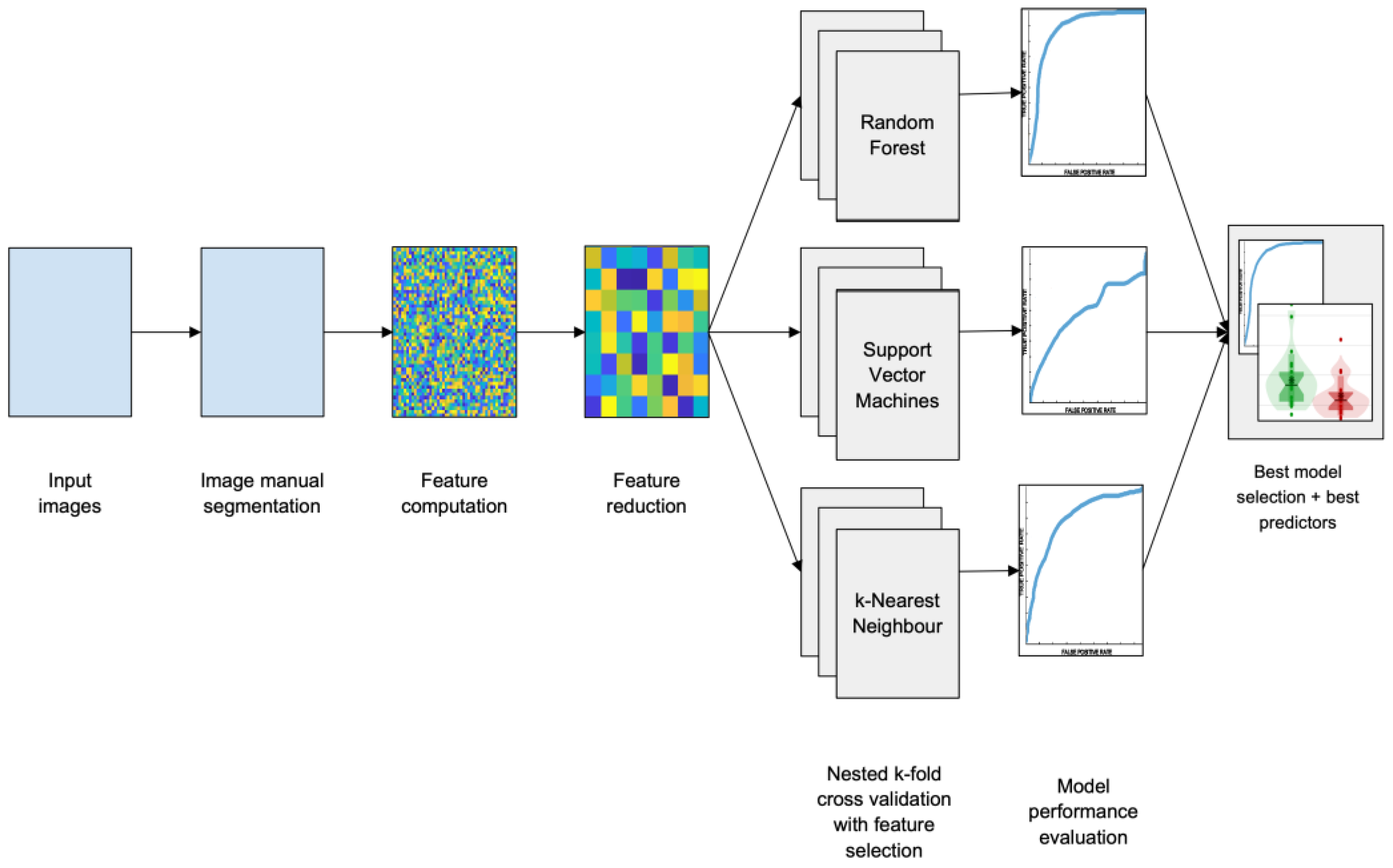
2.2. Machine Learning Model
3. Results
3.1. Dataset
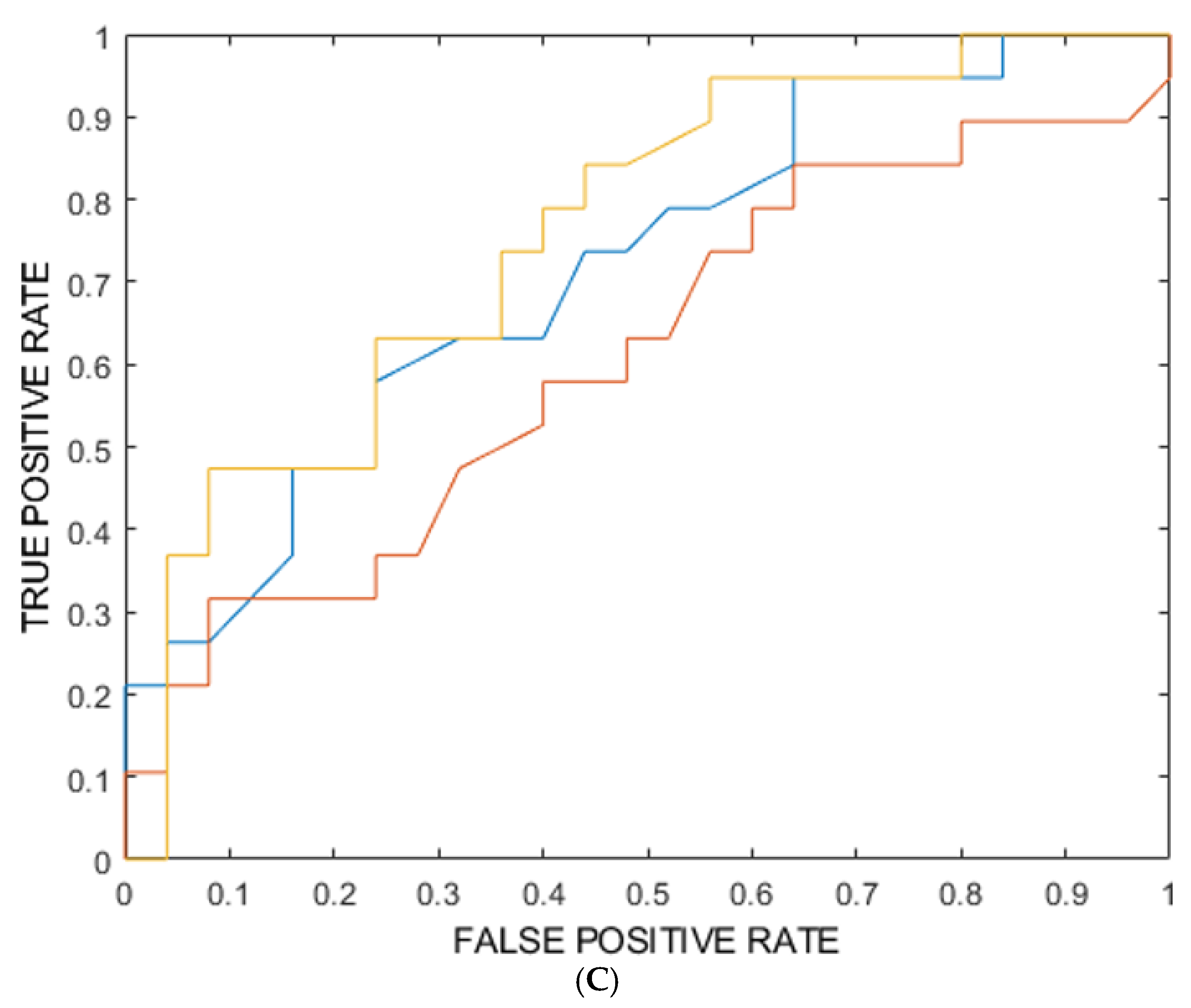
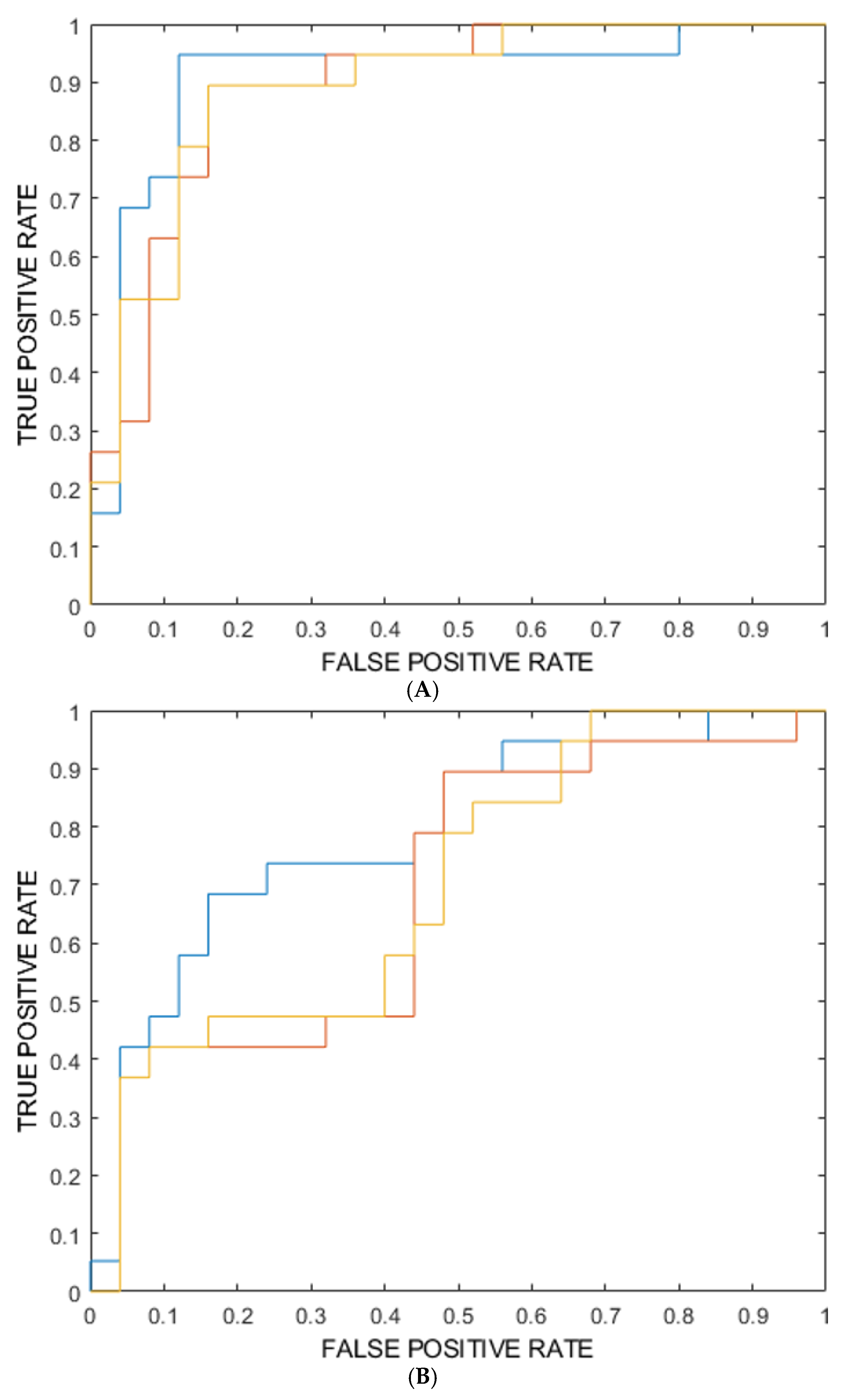
3.2. Machine Learning Model
| Training | Validation | Internal testing (mean) | Internal testing 1 | Internal testing 2 | |
| ROC-AUC (%) [95% CI] | 100* [99-100] | 89** [85-92] | 90** [87-92] | 90 | 90 |
| Accuracy (%) [95% CI] | 100* [99-100] | 82** [81-83] | 83** [80-87] | 82 | 82 |
| Sensitivity (%) [95% CI] | 100* [99-100] | 76** [73-80] | 79** [79-79] | 74 | 95 |
| Specificity (%) [95% CI] | 100* [99-100] | 86** [86-86] | 87** [81-92] | 88 | 72 |
| PPV (%) [95% CI] | 100* [99-100] | 84** [81-87] | 82** [76-88] | 82 | 72 |
| NPV (%) [95% CI] | 100* [99-100] | 87** [85-89] | 84** [84-85] | 81 | 95 |
| Training | Validation | Internal testing (mean) | Internal testing 1 | Internal testing 2 | |
| ROC-AUC (%) [95% CI] | 89** [87-91] | 83** [79-87] | 73** [59-87] | 73 | 73 |
| Accuracy (%) [95% CI] | 80** [77-82] | 70** [64-75] | 61 [32-91] | 66 | 59 |
| Sensitivity (%) [95% CI] | 75** [71-78] | 64** [57-71] | 56 [18-94] | 53 | 95 |
| Specificity (%) [95% CI] | 83** [81-85] | 73** [66-80] | 65* [42-88] | 76 | 32 |
| PPV (%) [95% CI] | 79** [77-82] | 69** [61-78] | 55 [22-87] | 63 | 51 |
| NPV (%) [95% CI] | 80** [78-82] | 77** [70-83] | 66 [39-94] | 68 | 89 |
| Training | Validation | Internal testing (mean) | Internal testing 1 | Internal testing 2 | |
| ROC-AUC (%) [95% CI] | 88** [86-89] | 62** [57-68] | 69* [50-88] | 73 | 73 |
| Accuracy (%) [95% CI] | 80** [78-82] | 59** [52-65] | 62** [50-74] | 68 | 66 |
| Sensitivity (%) [95% CI] | 75** [73-77] | 51** [47-55] | 42** [19-65] | 47 | 95 |
| Specificity (%) [95% CI] | 84** [81-88] | 64** [53-74] | 77** [72-83] | 84 | 44 |
| PPV (%) [95% CI] | 82** [78-86] | 53** [44-62] | 58** [40-76] | 69 | 56 |
| NPV (%) [95% CI] | 81** [79-83] | 66** [58-74] | 64** [54-74] | 68 | 92 |
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
Abbreviations
| CI | Confidence interval |
| FDR | Fisher Discriminant Ratio |
| GLCM | Gray-Level Co-occurrence Matrix |
| GLRLM | Gray-Level Run Length Matrix |
| GLSZM | Gray-Level Size Zone Matrix |
| IBSI | Image Biomarker Standardization Initiative |
| IRB | Institutional Review Board |
| kNN | K-nearest neighbor |
| MR | Magnetic resonance |
| NGLDM | Neighboring Gray Level Dependence Matrix |
| NGTDM | Neighborhood Gray Tone Difference Matrix |
| NPV | Negative predictive value |
| PCA | Principal components analysis |
| PPV | Positive predictive value |
| RF | Random forest |
| ROC-AUC | Area Under the Receiver Operating Characteristic Curve |
| STUMP | Smooth muscle tumors of uncertain malignant potential |
| SVM | Support vector machines |
| T2WI | T2 weighted image |
| US | Ultrasound |
| VOI | Volume of interest |
| WHO | 2020 World Health Organization |
References
- Bell SW, Kempson RL, Hendrickson MR. Problematic uterine smooth muscle neoplasms. A clinicopathologic study of 213 cases. Am J Surg Pathol. 1994;18(6):535-58.
- Taghzouti H, Saoud MK. 2022-RA-1158-ESGO Uterine sarcoma. International Journal of Gynecological Cancer2022.
- Ferrandina G, Aristei C, Biondetti PR, Cananzi FCM, Casali P, Ciccarone F, et al. Italian consensus conference on management of uterine sarcomas on behalf of S.I.G.O. (Societa' italiana di Ginecologia E Ostetricia). Eur J Cancer. 2020;139:149-68. [CrossRef]
- D'Angelo E, Prat J. Uterine sarcomas: a review. Gynecol Oncol. 2010;116(1):131-9. [CrossRef]
- Ludovisi M, Moro F, Pasciuto T, Di Noi S, Giunchi S, Savelli L, et al. Imaging in gynecological disease (15): clinical and ultrasound characteristics of uterine sarcoma. Ultrasound Obstet Gynecol. 2019;54(5):676-87. [CrossRef]
- Exacoustos C, Romanini ME, Amadio A, Amoroso C, Szabolcs B, Zupi E, et al. Can gray-scale and color Doppler sonography differentiate between uterine leiomyosarcoma and leiomyoma? J Clin Ultrasound. 2007;35(8):449-57. [CrossRef]
- Aviram R, Ochshorn Y, Markovitch O, Fishman A, Cohen I, Altaras MM, et al. Uterine sarcomas versus leiomyomas: gray-scale and Doppler sonographic findings. J Clin Ultrasound. 2005;33(1):10-3. [CrossRef]
- Namimoto T, Yamashita Y, Awai K, Nakaura T, Yanaga Y, Hirai T, et al. Combined use of T2-weighted and diffusion-weighted 3-T MR imaging for differentiating uterine sarcomas from benign leiomyomas. Eur Radiol. 2009;19(11):2756-64. [CrossRef]
- Valletta R, Corato V, Lombardo F, Avesani G, Negri G, Steinkasserer M, et al. Leiomyoma or sarcoma? MRI performance in the differential diagnosis of sonographically suspicious uterine masses. Eur J Radiol. 2024;170:111217. [CrossRef]
- Raffone A, Raimondo D, Neola D, Travaglino A, Giorgi M, Lazzeri L, et al. Diagnostic accuracy of MRI in the differential diagnosis between uterine leiomyomas and sarcomas: A systematic review and meta-analysis. Int J Gynaecol Obstet. 2024;165(1):22-33. [CrossRef]
- DeMulder D, Ascher SM. Uterine Leiomyosarcoma: Can MRI Differentiate Leiomyosarcoma From Benign Leiomyoma Before Treatment? AJR Am J Roentgenol. 2018;211(6):1405-15. [CrossRef]
- Tamai K, Koyama T, Saga T, Morisawa N, Fujimoto K, Mikami Y, et al. The utility of diffusion-weighted MR imaging for differentiating uterine sarcomas from benign leiomyomas. Eur Radiol. 2008;18(4):723-30. [CrossRef]
- Rosa F, Martinetti C, Magnaldi S, Rizzo S, Manganaro L, Migone S, et al. Uterine mesenchymal tumors: development and preliminary results of a magnetic resonance imaging (MRI) diagnostic algorithm. Radiol Med. 2023;128(7):853-68. [CrossRef]
- Gillies RJ, Kinahan PE, Hricak H. Radiomics: Images Are More than Pictures, They Are Data. Radiology. 2016;278(2):563-77. [CrossRef]
- Dai M, Liu Y, Hu Y, Li G, Zhang J, Xiao Z, et al. Combining multiparametric MRI features-based transfer learning and clinical parameters: application of machine learning for the differentiation of uterine sarcomas from atypical leiomyomas. Eur Radiol. 2022;32(11):7988-97. [CrossRef]
- Technologies D: https://www.deeptracetech.com/files/TechnicalSheet__TRACE4.pdf : 9-11. Accessed.
- Zwanenburg A, Vallieres M, Abdalah MA, Aerts H, Andrearczyk V, Apte A, et al. The Image Biomarker Standardization Initiative: Standardized Quantitative Radiomics for High-Throughput Image-based Phenotyping. Radiology. 2020;295(2):328-38. [CrossRef]
- Sbaraglia M, Bellan E, Dei Tos AP. The 2020 WHO Classification of Soft Tissue Tumours: news and perspectives. Pathologica. 2021;113(2):70-84. [CrossRef]
- Raspagliesi F, Maltese G, Bogani G, Fuca G, Lepori S, De Iaco P, et al. Morcellation worsens survival outcomes in patients with undiagnosed uterine leiomyosarcomas: A retrospective MITO group study. Gynecol Oncol. 2017;144(1):90-5. [CrossRef]
- Bogani G, Chiappa V, Ditto A, Martinelli F, Donfrancesco C, Indini A, et al. Morcellation of undiagnosed uterine sarcoma: A critical review. Crit Rev Oncol Hematol. 2016;98:302-8. [CrossRef]
- Bogani G, Cliby WA, Aletti GD. Impact of morcellation on survival outcomes of patients with unexpected uterine leiomyosarcoma: a systematic review and meta-analysis. Gynecol Oncol. 2015;137(1):167-72. [CrossRef]
- Nagai T, Takai Y, Akahori T, Ishida H, Hanaoka T, Uotani T, et al. Novel uterine sarcoma preoperative diagnosis score predicts the need for surgery in patients presenting with a uterine mass. Springerplus. 2014;3:678. [CrossRef]
- Nagai T, Takai Y, Akahori T, Ishida H, Hanaoka T, Uotani T, et al. Highly improved accuracy of the revised PREoperative sarcoma score (rPRESS) in the decision of performing surgery for patients presenting with a uterine mass. Springerplus. 2015;4:520. [CrossRef]
- Chiappa V, Interlenghi M, Salvatore C, Bertolina F, Bogani G, Ditto A, et al. Using rADioMIcs and machine learning with ultrasonography for the differential diagnosis of myometRiAL tumors (the ADMIRAL pilot study). Radiomics and differential diagnosis of myometrial tumors. Gynecol Oncol. 2021;161(3):838-44. [CrossRef]
- Suzuki A, Aoki M, Miyagawa C, Murakami K, Takaya H, Kotani Y, et al. Differential Diagnosis of Uterine Leiomyoma and Uterine Sarcoma using Magnetic Resonance Images: A Literature Review. Healthcare (Basel). 2019;7(4). [CrossRef]
- Nakagawa M, Nakaura T, Namimoto T, Iyama Y, Kidoh M, Hirata K, et al. Machine Learning to Differentiate T2-Weighted Hyperintense Uterine Leiomyomas from Uterine Sarcomas by Utilizing Multiparametric Magnetic Resonance Quantitative Imaging Features. Acad Radiol. 2019;26(10):1390-9. [CrossRef]
- Xie H, Hu J, Zhang X, Ma S, Liu Y, Wang X. Preliminary utilization of radiomics in differentiating uterine sarcoma from atypical leiomyoma: Comparison on diagnostic efficacy of MRI features and radiomic features. Eur J Radiol. 2019;115:39-45. [CrossRef]
- Roller LA, Wan Q, Liu X, Qin L, Chapel D, Burk KS, et al. MRI, clinical, and radiomic models for differentiation of uterine leiomyosarcoma and leiomyoma. Abdom Radiol (NY). 2024;49(5):1522-33. [CrossRef]
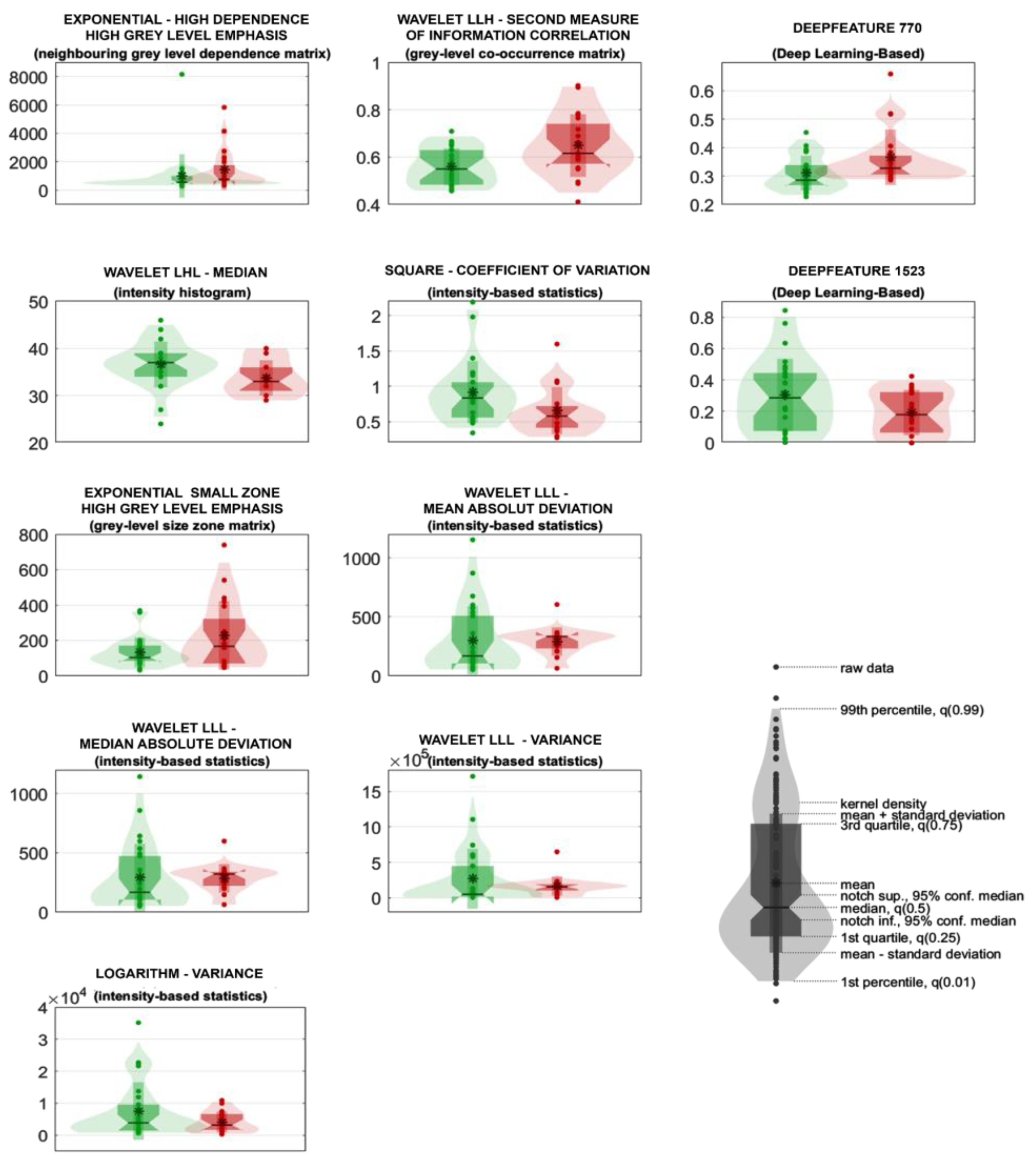
| # | Feature family | Feature nomenclature | Median in the positive class (95% CI) | Median in the negative class (95% CI) | Uncorrected p-value | Corrected p-value |
| 1 | Texture - Neighbouring Grey Level Dependence Matrix | High Dependence High Grey Level Emphasis - Exponential filter |
761.86 [316.76 - 1206.96] | 562.32 [400.75 - 723.89] | < 0.05 | 0.12 |
| 2 | Texture - Grey-Level Co-Occurrence Matrix | Second Measure Of Information Correlation - Wavelet LLH filter | 0.62 [0.55 - 0.68] | 0.55 [0.5 - 0.6] | < 0.05 | 0.17 |
| 3 | Deep Learning-Based | DeepFeature 770 | 0.33 [0.3 - 0.35] | 0.29 [0.26 - 0.31] | < 0.05 | 0.22 |
| 4 | Intensity Histogram | Median - Wavelet LHL filter | 33 [31.02 - 34.98] | 37 [35.43 - 38.57] | < 0.05 | 0.29 |
| 5 | Intensity-Based Statistics | Coefficient Of Variation - Square filter | 0.58 [0.46 - 0.69] | 0.83 [0.67 - 1] | < 0.05 | 0.3 |
| 6 | Deep Learning-Based | DeepFeature 1523 | 0.18 [8.06e-02 - 0.28] | 0.29 [0.17 - 0.4] | 0.11 | 1 |
| 7 | Texture - Grey-Level Size Zone Matrix | Small Zone High Grey Level Emphasis - Exponential filter |
166.66 [62.09 - 271.23] | 102.89 [74.87 - 130.9] | 0.19 | 1 |
| 8 | Intensity-Based Statistics | Mean Absolut Deviation - Wavelet LLL filter | 333.36 [289.84 - 376.87] | 168.41 [36.94 - 299.88] | 0.25 | 1 |
| 9 | Intensity-Based Statistics | Median Absolute Deviation - Wavelet LLL filter | 321.55 [276.28 - 366.82] | 167.4 [47.2 - 287.59] | 0.27 | 1 |
| 10 | Intensity-Based Statistics | Variance - Wavelet LLL filter | 1.55e+05 [1.17e+05 - 1.93e+05] | 5.00e+04 [-9.63e+04 - 1.96e+05] | 0.29 | 1 |
| 11 | Intensity-Based Statistics | Variance - Logarithm filter | 3159.3 [1362.72 - 4955.88] | 3843.17 [1091.45 - 6594.89] | 0.46 | 1 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




