Submitted:
10 March 2026
Posted:
11 March 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
2.1. DNA Extraction and Genome Sequencing
2.2. Data Processing and Genome Assembly
2.2.1. Quality Control, Adapter Removal, Draft Genome Assembly, and Refinement
2.2.2. Gene Prediction and Functional Annotation, Resistance and Virulence Genes, Biosynthetic Clusters, and Plasmid Detection
3. Results and Discussion
3.1. Quality Control and Draft Genome Assembly
3.2. Prediction and Functional Annotation, Resistance and Virulence Genes, Biosynthetic Clusters, and Plasmid Detection
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| MDPI | Multidisciplinary Digital Publishing Institute |
| DOAJ | Directory of open access journals |
| TLA | Three letter acronym |
| LD | Linear dichroism |
References
- Tingle, C.C.D.; Rother, J.A.; Dewhurst, C.F.; Lauer, S.; King, W.J. Fipronil: Environmental fate, ecotoxicology, and human health concerns. Rev. Environ. Contam. Toxicol. 2007, 176, 1–66. [Google Scholar]
- Silva, M.R.; Sartori, S.; Castro, A.P.; Krüger, R.H.; et al. Soil bacterial communities in the Brazilian Cerrado: Response to vegetation type and management. Acta Oecol 2019, 100, 103463. [Google Scholar] [CrossRef]
- Provase, M.; Boeing, G.A.N.S.; Tsukada, E.; Salla, R.F.; Abdalla, F.C. Impact of environmental concentrations of fipronil on DNA integrity and brain structure of Bombus atratus bumblebees. Environ. Toxicol. Pharmacol. 2024, 110, 104536. [Google Scholar] [CrossRef] [PubMed]
- Goulart, L.M.; et al. Fate and toxicity of 2,4-D and fipronil in mesocosm systems. Environ. Pollut. 2024, 332, 121899. [Google Scholar] [CrossRef]
- ANVISA. F43 Fipronil. Monografias Autorizadas; Agência Nacional de Vigilância Sanitária: Brasília, Brazil, 2022; Available online: https://encurtador.com.br/dlyF6 (accessed on 15 January 2026).
- Lozano, V.L.; Defarge, N.; Mesnage, R.; Hennequin, D.; Cassier, R.; de Vendômois, J.S.; Panoff, J.M.; Séralini, G.E.; Amiel, C. Sex-dependent impacts of fipronil on non-target organisms: Oxidative stress and genotoxicity. Ecotoxicol. Environ. Saf. 2021, 208, 111491. [Google Scholar] [CrossRef]
- Essack, S.Y. Environment: The neglected component of the One Health triad. Lancet Planet. Health 2018, 2, e238–e239. [Google Scholar] [CrossRef]
- Zhou, Z.; Wu, X.; Lin, Z.; Pang, S.; Mishra, S.; Chen, S. Biodegradation of fipronil: Current state of mechanisms of biodegradation and future perspectives. Appl. Microbiol. Biotechnol. 2021, 105, 7695–7708. [Google Scholar] [CrossRef]
- Bhatt, P.; Gangola, S.; Ramola, S.; Bilal, M.; Bhatt, K.; Huang, Y.; Zhou, Z.; Chen, S. Insights into the toxicity and biodegradation of fipronil in contaminated environment. Microbiol. Res. 2023, 266, 127247. [Google Scholar] [CrossRef]
- Gupta, A.; Kumar, V.; Chauhan, A.; Singh, N.; Chandra, R. Progress in bioremediation of pesticide residues in soil and water: Microbial consortia and enzyme-assisted degradation. Environ. Eng. Res. 2020, 25, 446–461. [Google Scholar] [CrossRef]
- Bonfá, M.R.L.; Durrant, L.R.; Piubeli, F.A.; Prado, C.C.A.; Guima, S.E.S.; Pereira, R.M. Bioprospecting of microbial enzymes with application in environmental biotechnology: An omics approach. In Bioremediation and Bioeconomy; Wiley: Hoboken, NJ, USA, 2024; p. Chapter 17. [Google Scholar] [CrossRef]
- Bonfá, M.R.L.; Prado, C.C.A.; Piubeli, F.A.; Durrant, L.R. Fipronil microbial degradation: An overview from bioremediation to metabolic pathways. In Pesticides Bioremediation; Siddiqui, S., Meghvansi, M.K., Chaudhary, K.K., Eds.; Springer: Cham, Switzerland, 2022. [Google Scholar] [CrossRef]
- Haque, M.A.; et al. Unveiling chlorpyrifos mineralizing and tomato plant-growth activities of Enterobacter sp. Front. Microbiol. 2022, 13, 1060554. [Google Scholar] [CrossRef]
- Aswathi, A.; et al. Plant growth–promoting traits of rhizosphere bacteria under pesticide stress. Appl. Soil Ecol. 2024, 193, 104941. [Google Scholar] [CrossRef]
- Ali, S.; Khan, I.; Ali, M.; Shah, N.S.; Shin, J.H. Biodegradation and detoxification of pesticides: Insight into microorganisms and enzymes. Biodegradation 2021, 32, 271–296. [Google Scholar] [CrossRef]
- Wu, W.; Feng, Y.; Zong, Z. Characterization of a strain representing a new Enterobacter species, Enterobacter chengduensis sp. nov. Antonie van Leeuwenhoek 2019, 112, 491–500. [Google Scholar] [CrossRef] [PubMed]
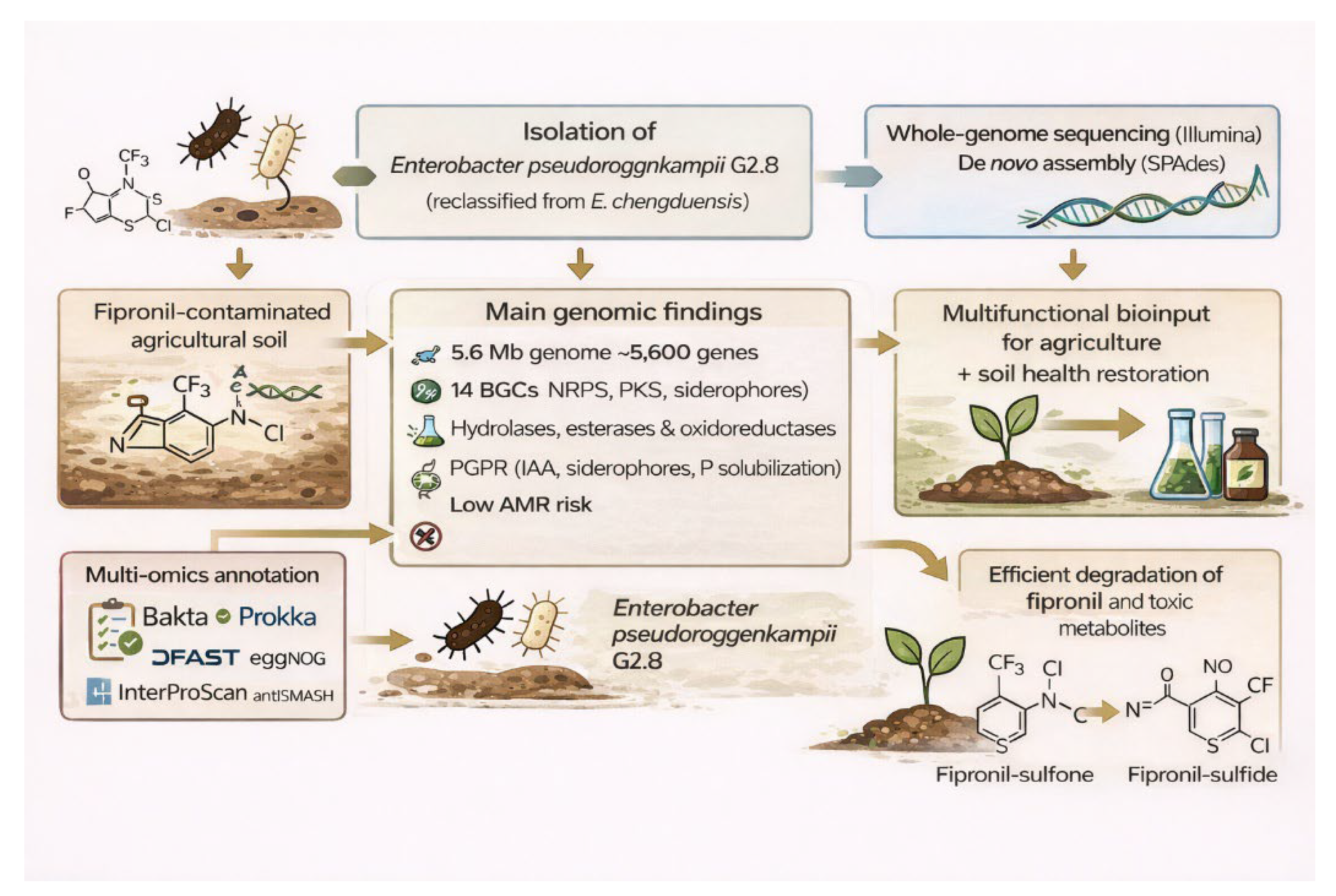
- Prado, C.; et al. Fipronil degradation in soil by Enterobacter chengduensis strain G2.8. Life 2023, 13, 1935. [Google Scholar] [CrossRef]
- Zagui, G.S.; Moreira, N.C.; Santos, D.V.; Paschoalato, C.F.P.R.; Sierra, J.; Nadal, M.; Domingo, J.L.; Darini, A.L.C.; Andrade, L.N.; Segura-Muñoz, S.I. Multidrug-resistant Enterobacter spp. in wastewater and surface water: Molecular characterization of β-lactam resistance and metal tolerance genes. Environ. Res. 2023, 233, 116443. [Google Scholar] [CrossRef]
- Gonçalves, D.L.D.R.; Chang, M.R.; Nobrega, G.D.; Venancio, F.A.; Higa Júnior, M.G.; Fava, W.S. Hospital sewage in Brazil: A reservoir of multidrug-resistant carbapenemase-producing Enterobacteriaceae. Braz. J. Biol. 2024, 84, e277750. [Google Scholar] [CrossRef]
- Wu, W.; et al. Taxonogenomic insights into pathogenic and environmental Enterobacter lineages. Microb. Genom. 2023, 9, mgen000932. [Google Scholar] [CrossRef]
- Taboadela-Hernanz, J.; et al. Genomic differentiation of plant-associated and clinical Enterobacter strains. Microorganisms 2025, 13, 114. [Google Scholar] [CrossRef]
- Chun, J.; et al. Proposed minimal standards for the use of genome data for the taxonomy of prokaryotes. Int. J. Syst. Evol. Microbiol. 2018, 68, 461–466. [Google Scholar] [CrossRef]
- Didelot, X.; et al. Transforming clinical microbiology with bacterial genome sequencing. Nat. Rev. Genet. 2016, 17, 601–612. [Google Scholar] [CrossRef]
- Medema, M.H.; et al. Minimum information about a biosynthetic gene cluster (MIBiG). Nat. Chem. Biol. 2015, 11, 625–631. [Google Scholar] [CrossRef] [PubMed]
- Płaza, G.A.; Banat, I.M.; Mrozik, W.; Briguglio, N.; Kaczorek, E. Biodegradation of xenobiotics by bacterial and fungal isolates: A review of mechanisms and environmental applications. Int. J. Environ. Res. Public Health 2020, 17, 196. [Google Scholar] [CrossRef]
- Blin, K.; et al. antiSMASH 7.0: Improved detection and comparative analysis of biosynthetic gene clusters. Nucleic Acids Res. 2023, 51, W46–W50. [Google Scholar] [CrossRef] [PubMed]
- Galperin, M.Y.; et al. The dark matter of bacterial genomes. Nat. Rev. Microbiol. 2019, 17, 491–504. [Google Scholar] [CrossRef]
- Andrews, S. FastQC: A quality control tool for high throughput sequence data; Babraham Bioinformatics: Cambridge, UK, 2010; Available online: http://www.bioinformatics.babraham.ac.uk/projects/fastqc/ (accessed on 15 January 2026).
- Krueger, F. Trim Galore! 2021. Available online: https://www.bioinformatics.babraham.ac.uk/projects/trim_galore/ (accessed on 15 January 2026).
- Bankevich, A.; Nurk, S.; Antipov, D.; Gurevich, A.A.; Dvorkin, M.; Kulikov, A.S.; Lesin, V.M.; Nikolenko, S.I.; Pham, S.; Prjibelski, A.D.; Pyshkin, A.V.; Sirotkin, A.V.; Vyahhi, N.; Tesler, G.; Alekseyev, M.A.; Pevzner, P.A. SPAdes: A new genome assembly algorithm and its applications to single-cell sequencing. J. Comput. Biol. 2012, 19, 455–477. [Google Scholar] [CrossRef]
- Zerbino, D.R.; Birney, E. Velvet: Algorithms for de novo short read assembly using de Bruijn graphs. Genome Res. 2008, 18, 821–829. [Google Scholar] [CrossRef]
- Seemann, T. Shovill: Faster SPAdes assembly of Illumina reads. 2017. Available online: https://github.com/tseemann/shovill (accessed on 15 January 2026).
- Li, D.; Liu, C.-M.; Luo, R.; Sadakane, K.; Lam, T.-W. MEGAHIT: An ultra-fast single-node solution for large and complex metagenomics assembly via succinct de Bruijn graph. Bioinformatics 2015, 31, 1674–1676. [Google Scholar] [CrossRef]
- Gurevich, A.; Saveliev, V.; Vyahhi, N.; Tesler, G. QUAST: Quality assessment tool for genome assemblies. Bioinformatics 2013, 29, 1072–1075. [Google Scholar] [CrossRef]
- Walker, B.J.; Abeel, T.; Shea, T.; Priest, M.; Abouelliel, A.; Sakthikumar, S.; Cuomo, C.A.; Zeng, Q.; Wortman, J.; Young, S.K.; Earl, A.M. Pilon: An integrated tool for comprehensive microbial variant detection and genome assembly improvement. PLoS ONE 2014, 9, e112963. [Google Scholar] [CrossRef]
- Simão, F.A.; Waterhouse, R.M.; Ioannidis, P.; Kriventseva, E.V.; Zdobnov, E.M. BUSCO: Assessing genome assembly and annotation completeness with single-copy orthologs. Bioinformatics 2015, 31, 3210–3212. [Google Scholar] [CrossRef]
- Wood, D.E.; Salzberg, S.L. Kraken: Ultrafast metagenomic sequence classification using exact alignments. Genome Biol. 2014, 15, R46. [Google Scholar] [CrossRef]
- Seemann, T. Prokka: Rapid prokaryotic genome annotation. Bioinformatics 2014, 30, 2068–2069. [Google Scholar] [CrossRef] [PubMed]
- Schwengers, O.; et al. Bakta: Rapid and standardized annotation of bacterial genomes. Microb. Genom. 2021, 7, 000685. [Google Scholar] [CrossRef] [PubMed]
- Tanizawa, Y.; Fujisawa, T.; Nakamura, Y. DFAST: A flexible prokaryotic genome annotation pipeline for faster genome publication. Bioinformatics 2018, 34, 1037–1039. [Google Scholar] [CrossRef] [PubMed]
- Jain, C.; Rodriguez-R, L.M.; Phillippy, A.M.; Konstantinidis, K.T.; Aluru, S. High throughput ANI analysis of 90K prokaryotic genomes reveals clear species boundaries. Nat. Commun. 2018, 9, 5114. [Google Scholar] [CrossRef]
- Meier-Kolthoff, J.P.; Göker, M. TYGS is an automated high-throughput platform for state-of-the-art genome-based taxonomy. Nat. Commun. 2019, 10, 2182. [Google Scholar] [CrossRef]
- Meier-Kolthoff, J.P.; Sardà Carbasse, J.; Peinado-Olarte, R.L.; Göker, M. TYGS and LPSN: A database tandem for fast and reliable genome-based classification and nomenclature of prokaryotes. Nucleic Acids Res. 2022, 50, D801–D807. [Google Scholar] [CrossRef]
- Huerta-Cepas, J.; Szklarczyk, D.; Forslund, K.; Cook, H.; Heller, D.; Walter, M.C.; Rattei, T.; Mende, D.R.; Sunagawa, S.; Kuhn, M.; Jensen, L.J.; von Mering, C.; Bork, P. eggNOG 4.5: A hierarchical orthology framework with improved functional annotations. Nucleic Acids Res. 2016, 44, D286–D293. [Google Scholar] [CrossRef]
- Huerta-Cepas, J.; Forslund, K.; Coelho, L.P.; Szklarczyk, D.; Jensen, L.J.; von Mering, C.; Bork, P. Fast genome-wide functional annotation through orthology assignment by eggNOG-Mapper. Mol. Biol. Evol. 2017, 34, 2115–2122. [Google Scholar] [CrossRef]
- Jones, P.; Binns, D.; Chang, H.-Y.; Fraser, M.; Li, W.; McAnulla, C.; McWilliam, H.; Maslen, J.; Mitchell, A.; Nuka, G.; Pesseat, S.; Quinn, A.F.; Sangrador-Vegas, A.; Scheremetjew, M.; Yong, S.-Y.; Lopez, R.; Hunter, S. InterProScan 5: Genome-scale protein function classification. Bioinformatics 2014, 30, 1236–1240. [Google Scholar] [CrossRef]
- Grant, J.R.; Enns, E.; Marinier, E.; Mandal, A.; Herman, E.K.; Chen, C.; Graham, M.; Van Domselaar, G.; Stothard, P. Proksee: In-depth characterization and visualization of bacterial genomes. Nucleic Acids Res. 2023, 51, W484–W490. [Google Scholar] [CrossRef] [PubMed]
- Couvin, D.; Bernheim, A.; Toffano-Nioche, C.; Touchon, M.; Michalik, J.; Néron, B.; Rocha, E.P.C.; Vergnaud, G.; Gautheret, D.; Pourcel, C. CRISPRCasFinder, an update of CRISRFinder, includes a portable version, enhanced performance and integrates search for Cas proteins. Nucleic Acids Res. 2018, 46, W246–W251. [Google Scholar] [CrossRef] [PubMed]
- Seemann, T. ABRicate: Mass screening of contigs for antibiotic resistance genes. 2016. Available online: https://github.com/tseemann/abricate (accessed on 15 January 2026).
- Alcock, B.P.; Huynh, W.; Chalil, R.; Van, V.; Hong, Y.G.; McArthur, A.G.; et al. CARD 2023: Expanded curation, support for machine learning, and resistome prediction at the Comprehensive Antibiotic Resistance Database. Nucleic Acids Res. 2023, 51, D690–D699. [Google Scholar] [CrossRef] [PubMed]
- Robertson, J.; Quick, J.; Taylor, D.; Woodall, C.; Akpan, I.; Kontoravdi, C.; Loman, N.J. MOB-suite: Software tools for clustering, reconstruction and typing of plasmids from draft assemblies. Microb. Genom. 2018, 4, e000206. [Google Scholar] [CrossRef]
- Carattoli, A.; Hasman, H. PlasmidFinder and in silico pMLST: Identification and typing of plasmid replicons in whole-genome sequencing (WGS). In Horizontal Gene Transfer; Springer: New York, NY, USA, 2019; pp. 285–294. [Google Scholar] [CrossRef]
- Wu, W.; Wei, L.; Feng, Y.; Kang, M.; Zong, Z. Enterobacter huaxiensis sp. nov. and Enterobacter chuandaensis sp. nov., recovered from human blood. Int. J. Syst. Evol. Microbiol. 2019, 69, 708–714. [Google Scholar] [CrossRef]
- Guima, S.E.S.; Piubeli, F.; Bonfá, M.R.L.; Pereira, R.M. New insights into the effect of fipronil on the soil bacterial community. Microorganisms 2023, 11, 52. [Google Scholar] [CrossRef]
- Prado, C.C.A.; et al. Fipronil biodegradation and metabolization by Bacillus megaterium strain E1. J. Chem. Technol. Biotechnol. 2021, 97, 474–481. [Google Scholar] [CrossRef]
- Destoumieux-Garzón, D.; et al. The One Health concept: 10 years old and a long road ahead. Front. Vet. Sci. 2018, 5, 14. [Google Scholar] [CrossRef]
- Hernández, S.; et al. One Health approach to antimicrobial resistance: Environmental perspective. Sci. Total Environ. 2019, 681, 476–488. [Google Scholar] [CrossRef]
- FAO/WHO. One Health approach to food safety. Food and Agriculture Organization of the United Nations/World Health Organization, 2017.
- Makarova, K.S.; et al. Evolutionary classification of CRISPR–Cas systems: A burst of class 2 and derived variants. Nat. Rev. Microbiol. 2020, 18, 67–83. [Google Scholar] [CrossRef]
- Antipov, D.; Hartwick, N.; Shen, M.; Raiko, M.; Lapidus, A.; Pevzner, P.A. plasmidSPAdes: Assembling plasmids from whole genome sequencing data. Bioinformatics 2016, 32, 3380–3387. [Google Scholar] [CrossRef]
- Unknown genes in bacterial genomes: distribution, diversity and association with environmental niches. Microbial Genomics 2023, 9, 000996. [CrossRef]

| Method | Comparison | Results | Reference Thresholds | Interpretation |
|---|---|---|---|---|
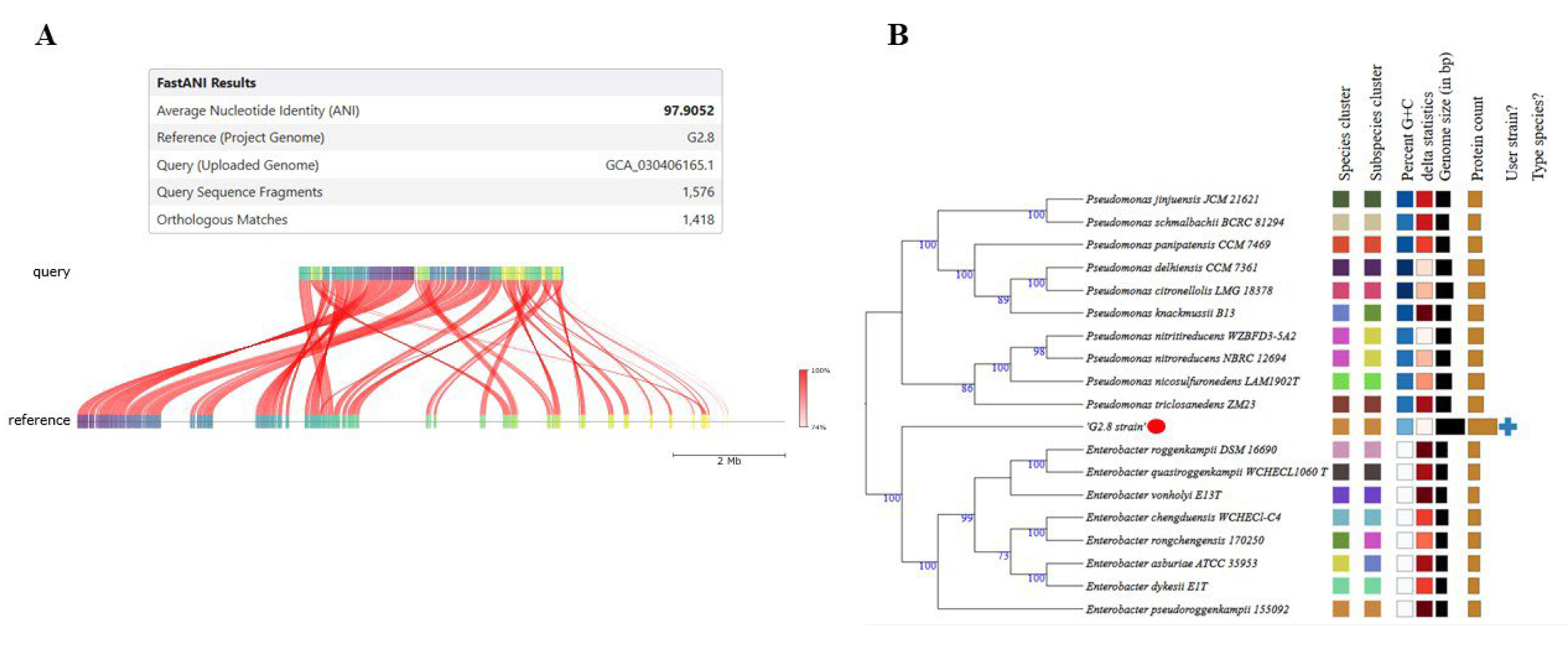
| FastANI | G2.8 × E. pseudoroggenkampii (ASM3040616v1) | 97,86% | ≥95% | Same species |
| dDDH (d4, TYGS) | G2.8 × type strain | 80,50% | ≥70% | Same species |
| Syntenia | G2.8 × E. pseudoroggenkampii | High collinearity | — | Genomic conservation |
| Comparison with E. chengduensis | ANI / dDDH | < threshold | ≥95 / ≥70 | Distinct species |
| 1. Main statistics | 2. Protein and enzyme prediction by tool | 3. Secondary metabolites (antiSMASH) | 4. Resistance genes (AMR) | 5. Metabolic functionality | |||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| Categorias | N° of genes | Program |
Nº of genes/ proteins |
BGC type | Nº of genes | CARD-RGI (36 genes) | Nº of loci (occurrences) | ABRicate (9 genes) | Nº of loci (occurrences) | Functional category | N° of genes |
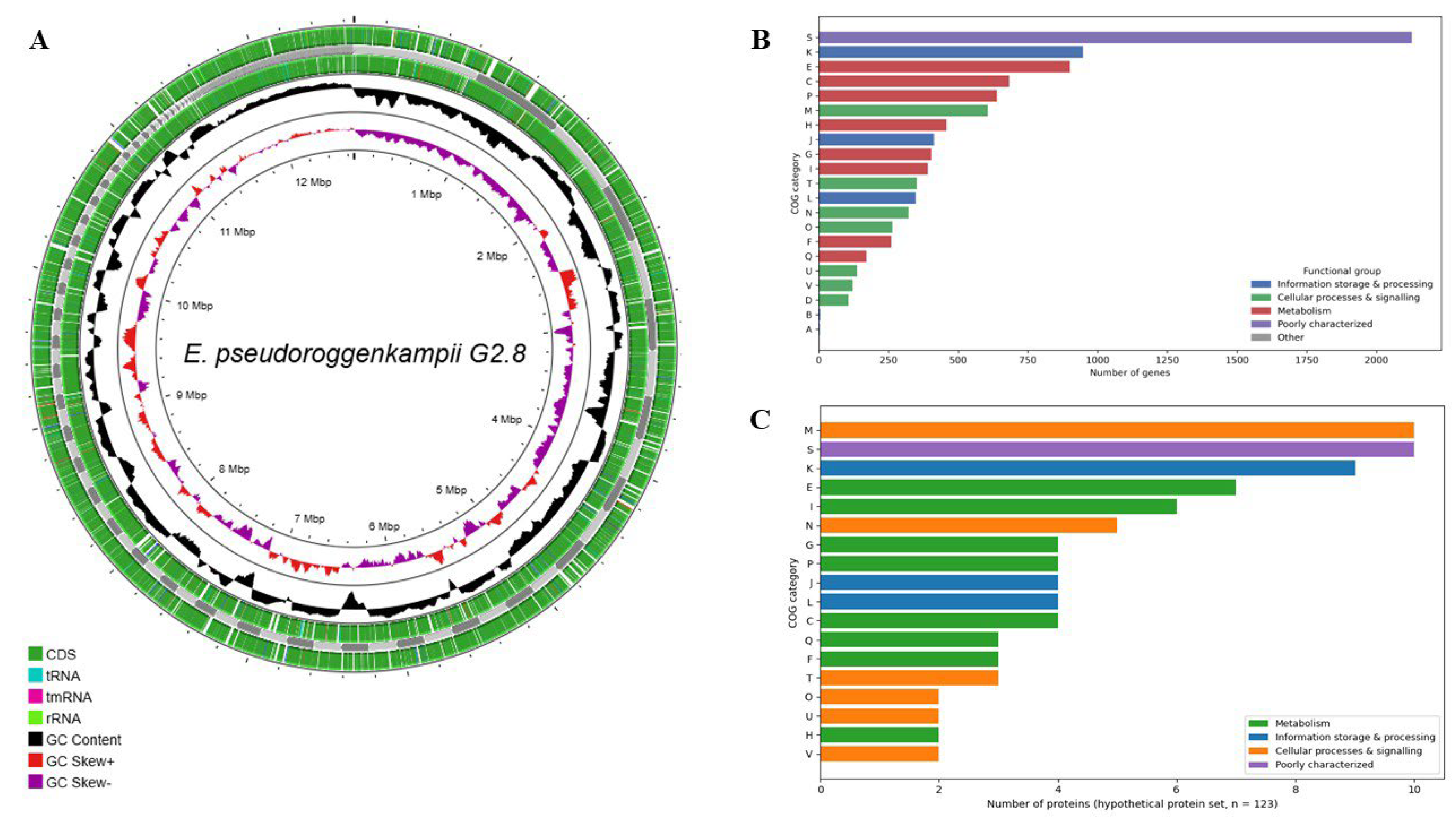
| Total de features (CDS + RNAs) | 5.666 | PROKKA – total CDS | 10.794 | CDS in BGC | 195 | acrD | 4 | oqxB_1 | 3 | COG | 5.542 genes with functional categories |
| Proteins annotated (Prokka) | 5.666 | Hypothetical proteins (Prokka) | 3.491 | Total clusters | 14 | msbA | 3 | qnrE1_1 | 2 | KEGG KO | 5.542 genes with annotated metabolic pathways |
| Proteins annotated (Bakta) | 5.662 | BAKTA – total CDS | 11.018 | NRPS | 60 | leuO | 3 | fosA_1 | 2 | InterProScan | 10.983 functional domains identified |
| Genes with eggNOG | 5.541 | Hypothetical proteins (Bakta) | 482 | NRPS + T1PKS | 45 | adeF | 3 | oqxB_1; oqxA_1 | 1 | - | - |
| Genes with COG | 5.542 | DFAST – total CDS | 1.470 | β-lactone | 26 | PBP3 (H. influenzae) | 3 | blaACT-4_2 | 1 | - | - |
| Genes with KEGG KO | 5.542 | Hypothetical proteins (DFAST) | 157 | RiPP | 21 | oqxB | 2 | - | - | - | - |
| - | - | - | - | hserlactone | 22 | soxR | 2 | - | - | - | - |
| - | - | - | - | siderophore | 13 | vanX | 2 | - | - | - | - |
| - | - | - | - | butyrolactone | 8 | vanH | 2 | - | - | - | - |
| emrA / emrB | 2 | - | - | - | - | ||||||
| CDS, coding DNA sequence; BGC, biosynthetic gene cluster; COG, Clusters of Orthologous Groups; KEGG KO, Kyoto Encyclopedia of Genes and Genomes Orthology; CARD-RGI, Comprehensive Antibiotic Resistance Database – Resistance Gene Identifier; NRPS, non-ribosomal peptide synthetase; T1PKS, type I polyketide synthase; RiPP, ribosomally synthesized and post-translationally modified peptide. | |||||||||||
| Functional category | Genes | Function | Evidence tool |
|---|---|---|---|
| Iron acquisition (siderophores) | entA, entB, entC, entD, entE, entF | Enterobactin biosynthesis | antiSMASH / eggNOG / KEGG |
| Iron acquisition (siderophores) | fepA, fepB, fepC, fepD, fepG | Siderophore transport | eggNOG / KEGG |
| Phytohormone production (IAA) | ipdC | Indole-3-pyruvate pathway (IAA) | eggNOG / KEGG |
| Phosphate solubilization | pqqB, pqqC, pqqD, pqqE | PQQ biosynthesis | eggNOG |
| Phosphate solubilization | gcd | Glucose dehydrogenase | KEGG |
| Rhizosphere colonization | motA, motB | Flagellar motilit | eggNOG |
| Rhizosphere colonization | cheA, cheY | Bacterial chemotaxis | eggNOG |
| Environmental stress tolerance | katG, sodA, sodB | Detoxification of reactive oxygen species | InterPro / KEGG |
| Microbial competition | NRPS-like BGC | Antimicrobial metabolite production | antiSMASH |
| Aromatic compound metabolism | pcaH, pcaG | Phenolic compound catabolism | KEGG |
| PGPR, plant growth-promoting rhizobacteria; IAA, indole-3-acetic acid; PQQ, pyrroloquinoline quinone; BGC, biosynthetic gene cluster; NRPS, non-ribosomal peptide synthetase; KEGG, Kyoto Encyclopedia of Genes and Genomes; eggNOG, evolutionary genealogy of genes: Non-supervised Orthologous Groups; InterPro, integrated protein signature database; antiSMASH, Antibiotics & Secondary Metabolite Analysis Shell (tool for prediction of secondary metabolite biosynthetic gene clusters). | |||
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




