Submitted:
10 March 2026
Posted:
11 March 2026
You are already at the latest version
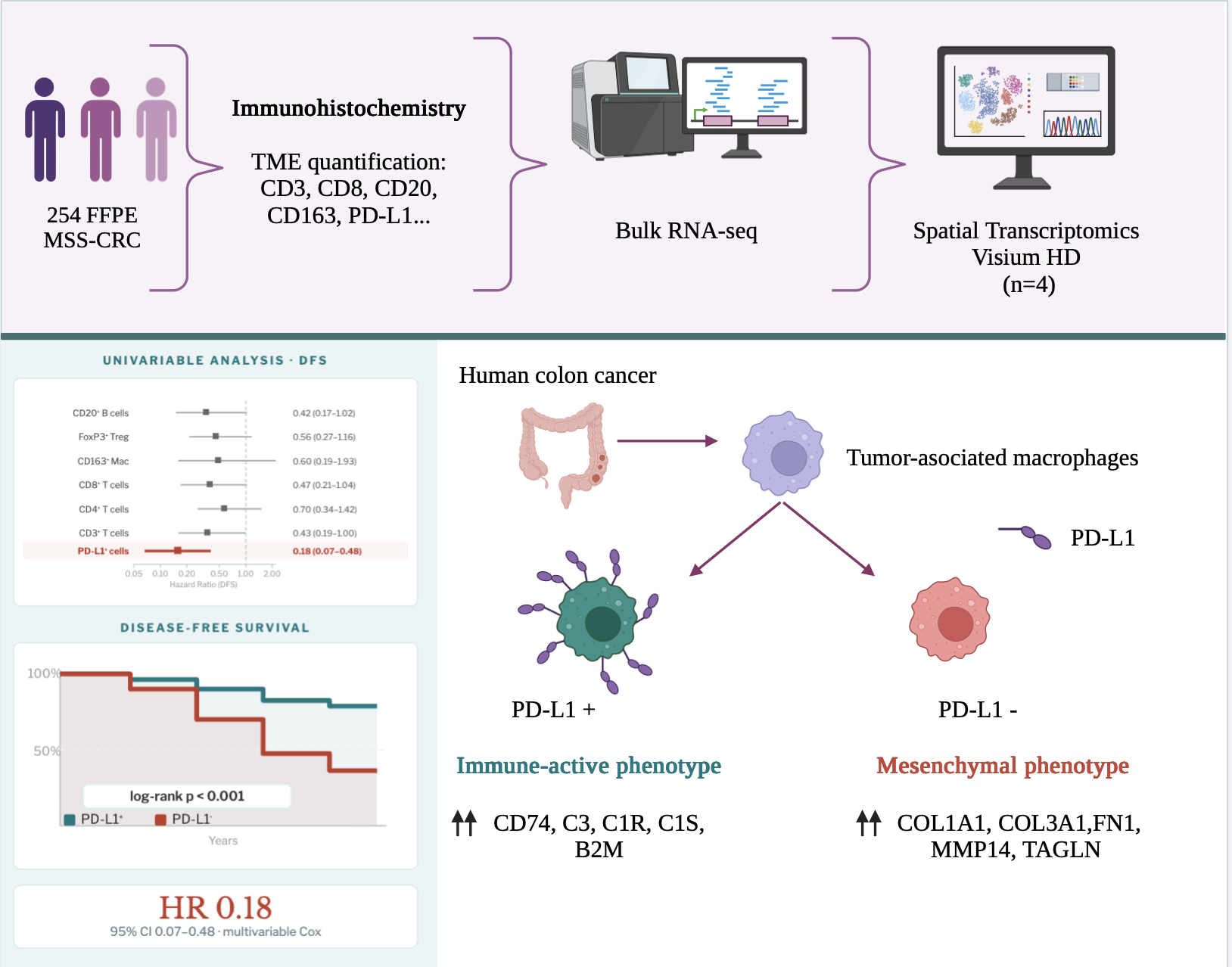
Abstract
Keywords:
1. Introduction
2. Materials and Methods
2.1. Patient Cohorts
2.2. Histological Evaluation
2.2.1. Morphological Assessment
2.2.2. Immune Infiltrate Assessment
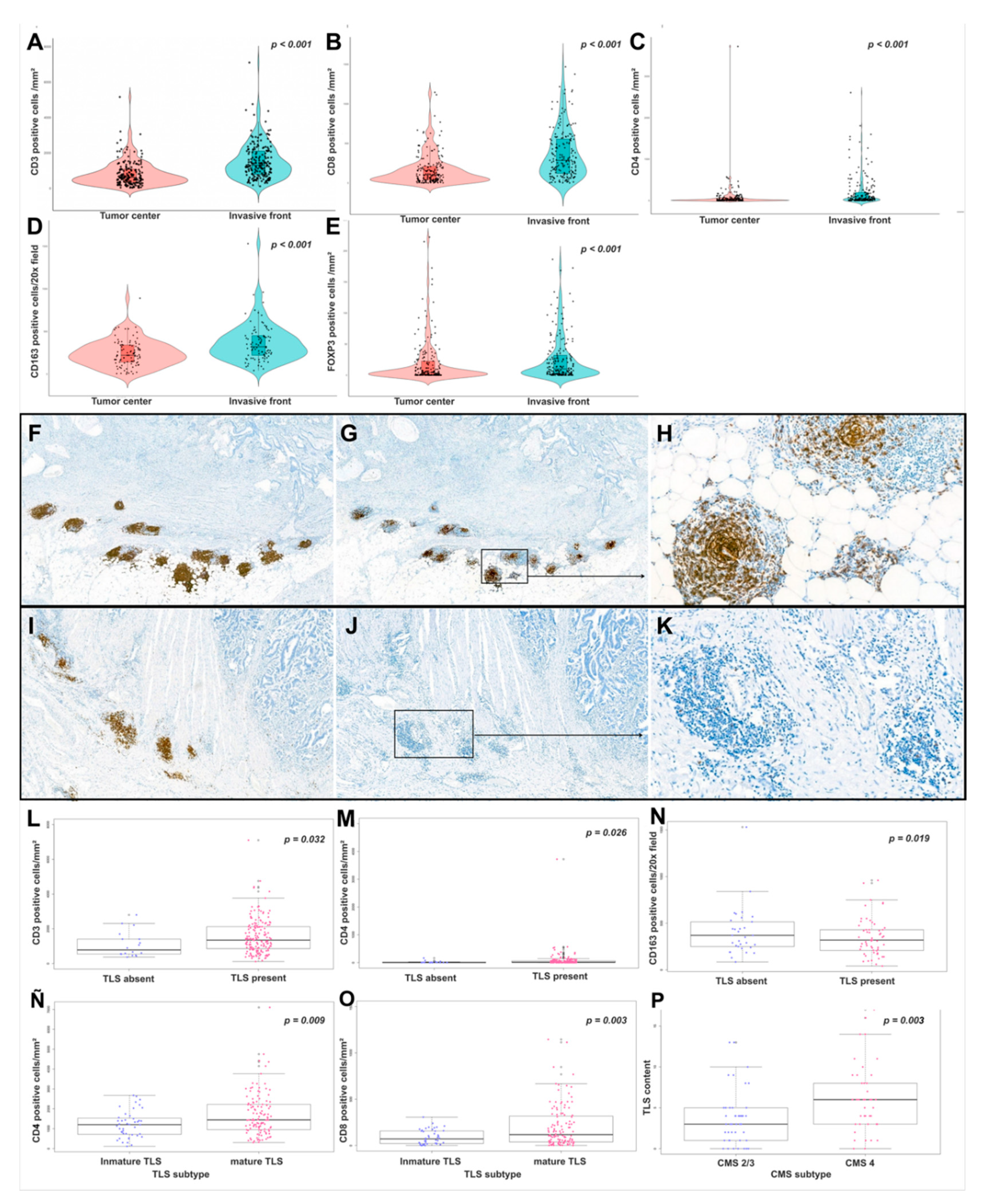
2.2.3. Tertiary Lymphoid Structures
2.2.4. PD-L1 Assessment
2.3. Transcriptomic Analysis
2.3.1. RNA Extraction
2.3.2. CMS Subtype Analysis
2.3.3. mRNA Sequencing
2.3.4. Spatial Transcriptomics
2.3.5. Functional Analysis of Gene Interactions
2.4. Statistical Analysis
2.4.1. Descriptive and Association Analyses
2.4.2. Survival Analysis
3. Results
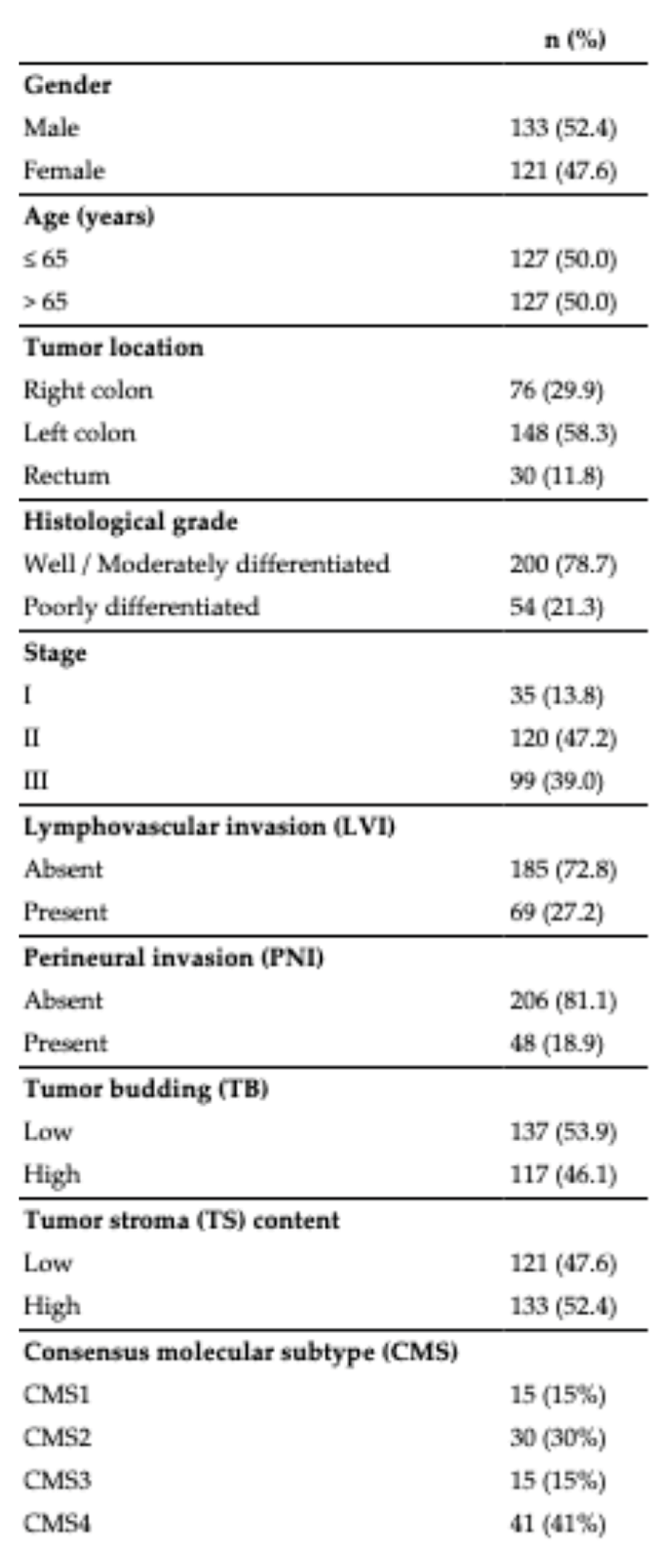
3.1. General Clinicopathological Features
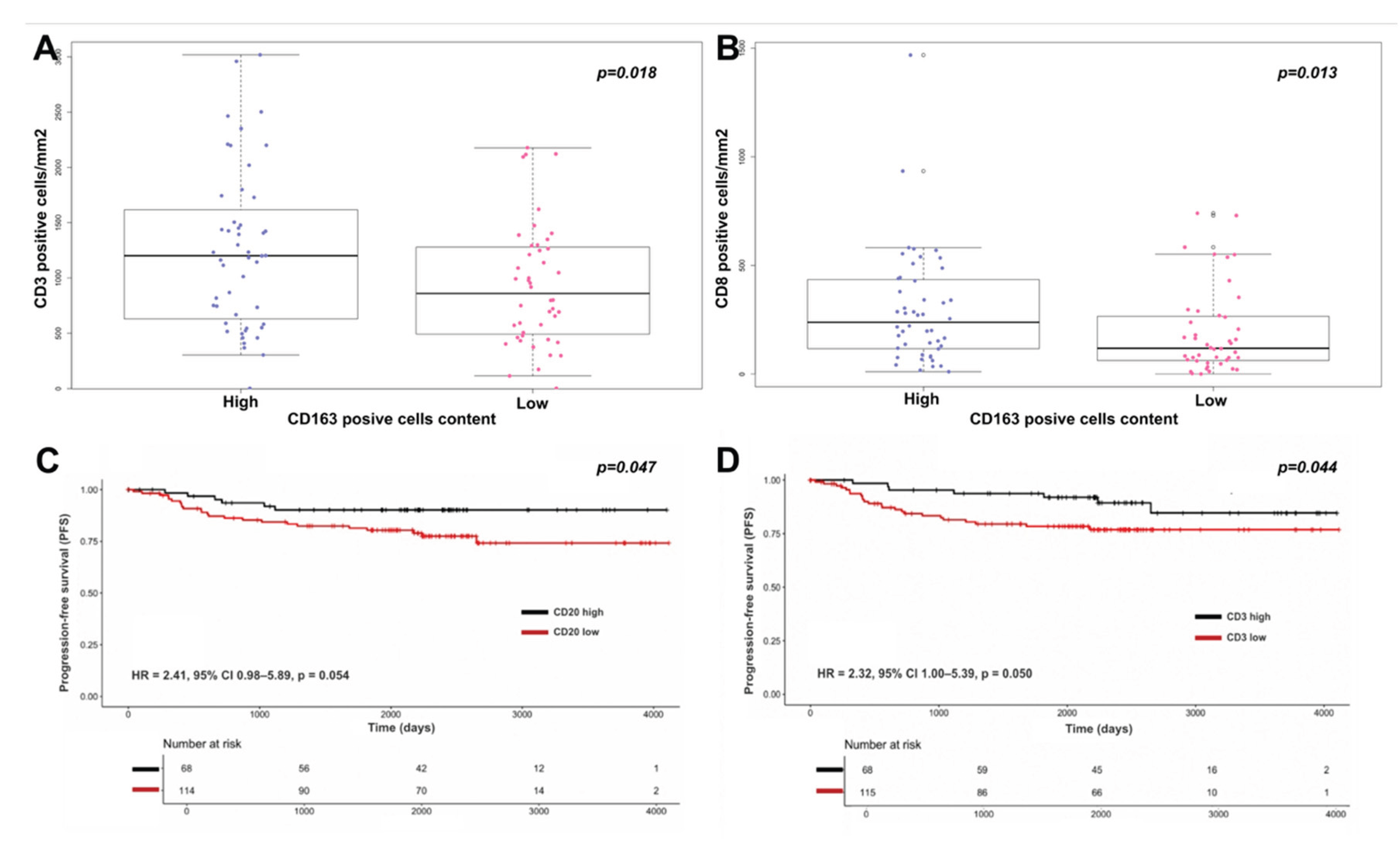
3.2. Tumor Microenvironment Composition
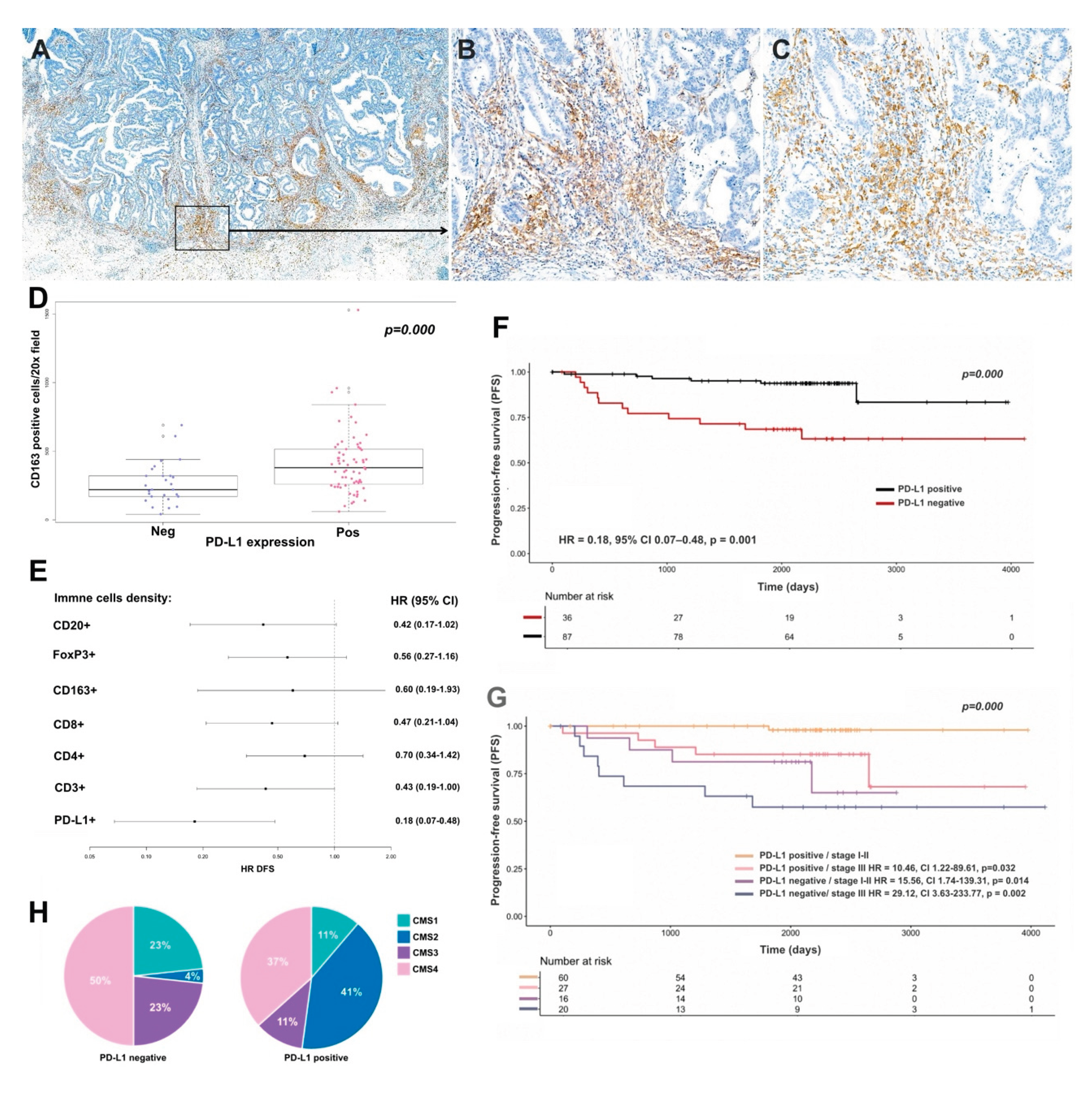
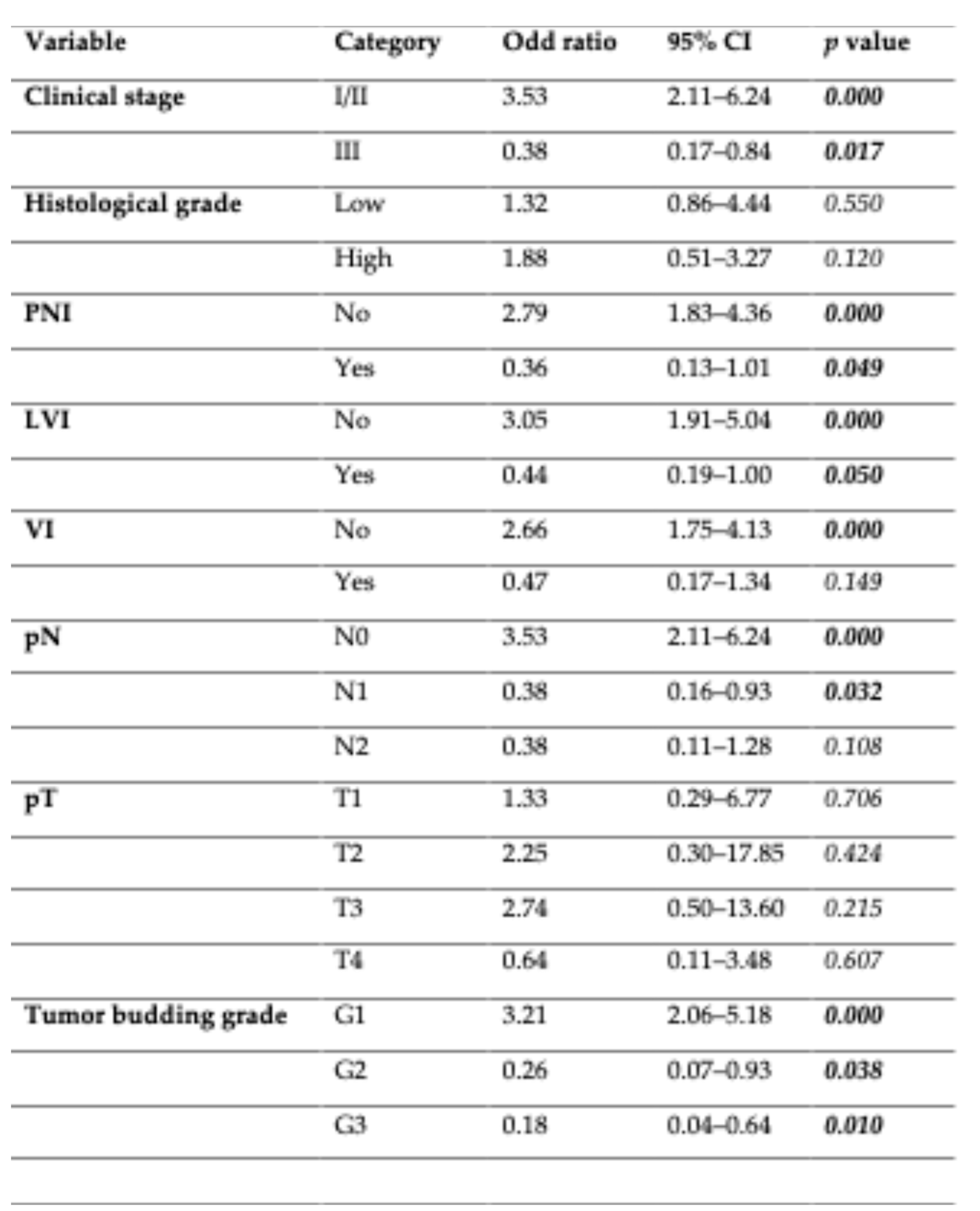
3.3. PD-L1 Expression in Immune Cells: Clinical and Prognostic Implications
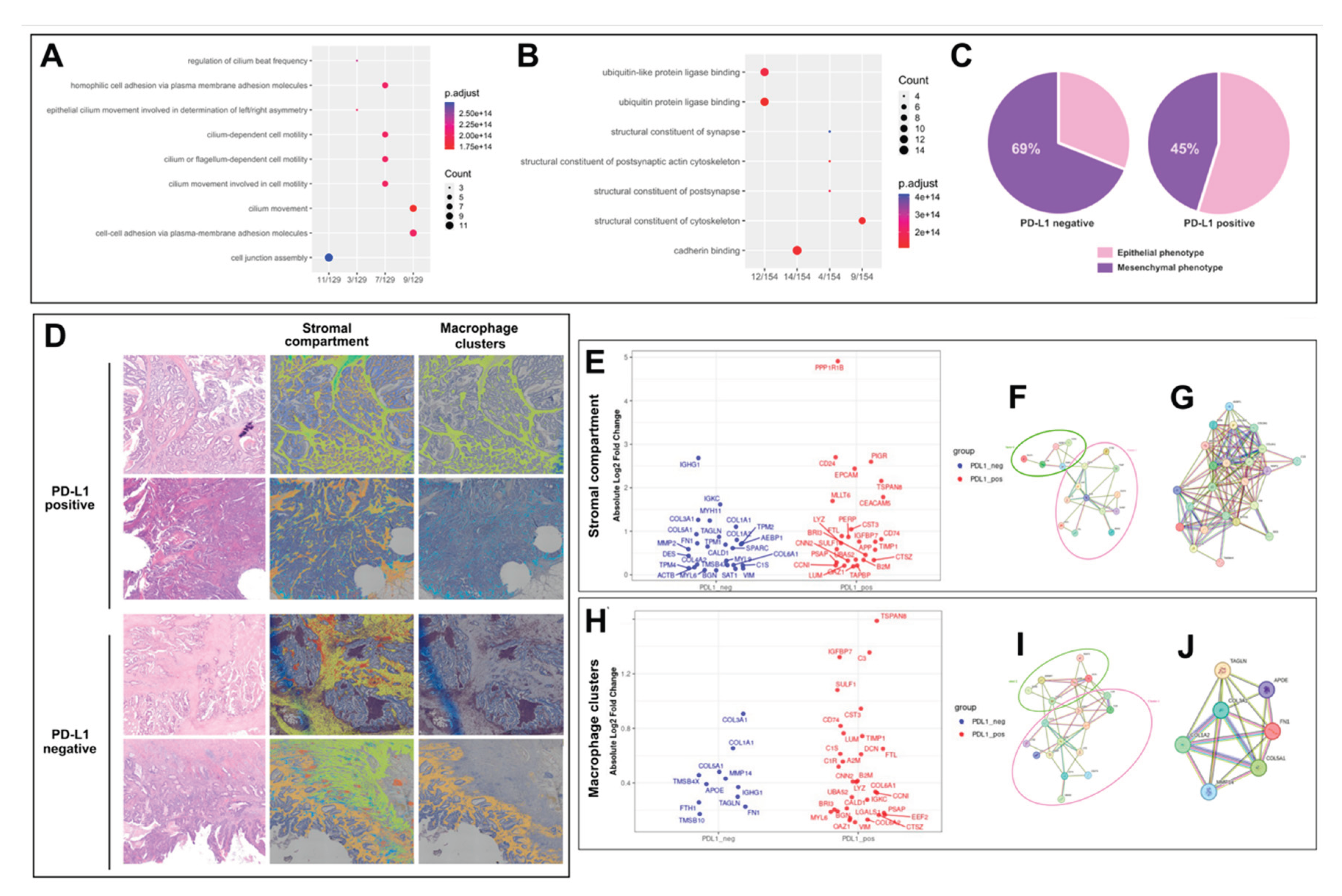
3.4. Bulk Transcriptomic Profiling of PD-L1-Defined Subgroups
3.5. Spatial Transcriptomics Reveals Compartment-Specific EMT–Immune Interactions
3.5.1. Spatial Transcriptomic Profile of the Stromal Compartment
3.5.2. Spatial transcriptomic profile of macrophage-rich stromal clusters
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| CI | Confidence intervals |
| CMS | Consensus molecular subtypes |
| CRC | Colorectal cancer |
| DFS | Disease-free survival |
| EMT | Epithelial to mesenchymal transition |
| FDCs | Follicular dendritic cells |
| FFPE | Formalin-fixed paraffin embedded |
| HE | Hematoxylin and eosin |
| HR | Hazard ratios |
| ICI | Immune checkpoint inhibitors |
| IF | Invasive front |
| IHC | Immunohistochemistry |
| iTLS | Immature tertiary lymphoid structures |
| LVI | Lymphovascular invasion |
| MMRp | Mismatch repair-proficient |
| MSI | Microsatellite instability |
| MSS-CRC | Microsatellite-stable colorectal cancer |
| mTLS | Mature tertiary lymphoid structures |
| OR | Odds ratio |
| ORA | Over-representation analysis |
| pN | Lymph nodes |
| PNI | Perineural invasion |
| pT | Depth of invasion |
| TB | Tumor budding |
| TC | Tumor center |
| TLSs | Tertiary lymphoid structures |
| TME | Tumor microenvironment |
| TS | Tumor stroma |
Appendix A
References
- Bray, F.; Laversanne, M.; Sung, H.; Ferlay, J.; Siegel, R.L.; Soerjomataram, I.; Jemal, A. Global cancer statistics 2022: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J. Clin. 2024, 74, 229–263. [CrossRef]
- Cardoso, R.; Guo, F.; Heisser, T.; Hackl, M.; Ihle, P.; De Schutter, H.; et al. Colorectal cancer incidence, mortality, and stage distribution in European countries in the colorectal cancer screening era: An international population-based study. Lancet Oncol. 2021, 22, 1002–1013. [CrossRef]
- Sung, H.; Siegel, R.L.; Laversanne, M.; Soerjomataram, I.; Jemal, A.; Bray, F. Colorectal cancer incidence trends in younger versus older adults: An analysis of population-based cancer registry data. Lancet Oncol. 2025, 26, 51–63. [CrossRef]
- Siegel, R.L.; Miller, K.D.; Goding Sauer, A.; Fedewa, S.A.; Butterly, L.F.; Anderson, J.C.; et al. Colorectal cancer statistics, 2020. CA Cancer J. Clin. 2020, 70, 145–164. [CrossRef]
- Malvezzi, M.; Carioli, G.; Bertuccio, P.; Boffetta, P.; Levi, F.; La Vecchia, C.; et al. European cancer mortality predictions for the year 2018 with focus on colorectal cancer. Ann. Oncol. 2018, 29, 1016–1022. [CrossRef]
- Gupta, R.; Sinha, S.; Paul, R.N. The impact of microsatellite stability status in colorectal cancer. Curr. Probl. Cancer 2018, 42, 548–559. [CrossRef]
- Taieb, J.; Svrcek, M.; Cohen, R.; Basile, D.; Tougeron, D.; Phelip, J.M. Deficient mismatch repair/microsatellite unstable colorectal cancer: Diagnosis, prognosis and treatment. Eur. J. Cancer 2022, 175, 136–157. [CrossRef]
- Sahin, I.H.; Ciombor, K.K.; Diaz, L.A., Jr.; Yu, J.; Kim, R. Immunotherapy for microsatellite stable colorectal cancers: Challenges and novel therapeutic avenues. Am. Soc. Clin. Oncol. Educ. Book 2022, 42, 242–253. [CrossRef]
- Guven, D.C.; Kavgaci, G.; Erul, E.; Ozkan, M.; Ozturk, M.A.; Yazici, O.; et al. The efficacy of immune checkpoint inhibitors in microsatellite stable colorectal cancer: A systematic review. Oncologist 2024, 29, e580–e600. [CrossRef]
- Galon, J.; Costes, A.; Sanchez-Cabo, F.; Kirilovsky, A.; Mlecnik, B.; Lagorce-Pagès, C.; et al. Type, density, and location of immune cells within human colorectal tumors predict clinical outcome. Science 2006, 313, 1960–1964. [CrossRef]
- Pagès, F.; Mlecnik, B.; Marliot, F.; Bindea, G.; Ou, F.S.; Bifulco, C.; et al. International validation of the consensus Immunoscore for the classification of colon cancer: A prognostic and accuracy study. Lancet 2018, 391, 2128–2139. [CrossRef]
- Wu, X.; Hou, S.; Ye, Y.; Gao, Z. The prognostic value of immunoscore in the early-onset colorectal cancer. BMC Gastroenterol. 2025, 25, 513. [CrossRef]
- Sautès-Fridman, C.; Petitprez, F.; Calderaro, J.; Fridman, W.H. Tertiary lymphoid structures in the era of cancer immunotherapy. Nat. Rev. Cancer 2019, 19, 307–325. [CrossRef]
- Guinney, J.; Dienstmann, R.; Wang, X.; de Reyniès, A.; Schlicker, A.; Soneson, C.; et al. The consensus molecular subtypes of colorectal cancer. Nat. Med. 2015, 21, 1350–1356. [CrossRef]
- Chu, X.; Li, X.; Zhang, Y.; Zhang, Q.; Yu, C.; Hu, Y.; et al. Integrative single-cell analysis of human colorectal cancer reveals patient stratification with distinct immune evasion mechanisms. Nat. Cancer 2024, 5, 1409–1426. [CrossRef]
- Mei, Y.; Xiao, W.; Hu, H.; Zhang, J.; Li, Z.; Zhao, H.; et al. Single-cell analyses reveal suppressive tumor microenvironment of human colorectal cancer. Clin. Transl. Med. 2021, 11, e422. [CrossRef]
- Qian, J.; Olbrecht, S.; Boeckx, B.; Vos, H.; Laoui, D.; Etlioglu, E.; et al. A pan-cancer blueprint of the heterogeneous tumor microenvironment revealed by single-cell profiling. Cell Res. 2020, 30, 745–762. [CrossRef]
- Wang, R.; Li, J.; Zhou, X.; Chen, Q.; Yang, Z.; Lin, X.; et al. Single-cell genomic and transcriptomic landscapes of primary and metastatic colorectal cancer tumors. Genome Med. 2022, 14, 93. [CrossRef]
- Palomar de Lucas, B.; Heras, B.; Tarazona, N.; Martí, R.; Algaba, A.; Gimeno-Valiente, F.; et al. Extended tumor area-based stratification score combining tumor budding and stroma identifies a high-risk, immune-depleted group in localized microsatellite-stable colon cancer patients. Pathol. Res. Pract. 2025, 269, 155871. [CrossRef]
- Bankhead, P.; Loughrey, M.B.; Fernández, J.A.; Dombrowski, Y.; McArt, D.G.; Dunne, P.D.; et al. QuPath: Open source software for digital pathology image analysis. Sci. Rep. 2017, 7, 16878. [CrossRef]
- Vanhersecke, L.; Brunet, M.; Guégan, J.P.; Charon-Barra, C.; Naltet, C.; Leblanc, R.; et al. Mature tertiary lymphoid structures predict immune checkpoint inhibitor efficacy in solid tumors independently of PD-L1 expression. Nat. Cancer 2021, 2, 794–802. [CrossRef]
- Tarazona, N.; Gimeno-Valiente, F.; Gambardella, V.; Font, A.; Heras, B.; Sastre, J.; et al. Detection of postoperative plasma circulating tumour DNA and lack of CDX2 expression as markers of recurrence in patients with localised colon cancer. ESMO Open 2020, 5, e000847. [CrossRef]
- Chakraborty, P.; George, J.T.; Tripathi, S.; Levine, H.; Jolly, M.K. Comparative study of transcriptomics-based scoring metrics for the epithelial-hybrid-mesenchymal spectrum. Front. Bioeng. Biotechnol. 2020, 8, 220. [CrossRef]
- R Core Team. R: A Language and Environment for Statistical Computing; R Foundation for Statistical Computing: Vienna, Austria, 2023. Available online: https://www.R-project.org/.
- Szklarczyk, D.; Kirsch, R.; Koutrouli, M.; Fonseca, N.; Bork, P.; Jensen, L.J.; von Mering, C. The STRING database in 2023: Protein–protein association networks and functional enrichment analyses for any sequenced genome of interest. Nucleic Acids Res. 2023, 51, D638–D646. [CrossRef]
- Stelzer, G.; Rosen, N.; Plaschkes, I.; Zimmerman, S.; Twik, M.; Fishilevich, S.; et al. The GeneCards Suite: From gene data mining to disease genome sequence analyses. Curr. Protoc. Bioinformatics 2016, 54, 1–30. [CrossRef]
- Omatsu, M.; Nakanishi, Y.; Iwane, K.; Tanaka, T.; Matsuda, H.; Akita, H.; et al. THBS1-producing tumor-infiltrating monocyte-like cells contribute to immunosuppression and metastasis in colorectal cancer. Nat. Commun. 2023, 14, 1–15. [CrossRef]
- Cheng, M.; Jiang, Y.; Yang, H.; Zhao, D.; Li, L.; Liu, X. FLNA promotes chemoresistance of colorectal cancer through inducing epithelial-mesenchymal transition and Smad2 signaling pathway. Am. J. Cancer Res. 2020, 10, 1–15. Available online: www.ajcr.us.
- Puig-Blasco, L.; Piotrowski, K.B.; Michaelsen, S.R.; Di Caro, G.; Perren, A.; de Visser, K.E.; et al. Loss of cancer cell-derived ADAM15 alters the tumor microenvironment in colorectal tumors. Int. J. Cancer 2023, 153, 2068–2081. [CrossRef]
- Zheng, L.; Xu, Z.; Zhang, W.; Li, Y.; Wu, J.; Chen, H. Identification and validation of a prognostic signature based on six immune-related genes for colorectal cancer. Discov. Oncol. 2024, 15, 192. [CrossRef]
- Li, J.; Chen, M.; Tong, M.; Cao, Q. TUBA1C orchestrates the immunosuppressive tumor microenvironment and resistance to immune checkpoint blockade in clear cell renal cell carcinoma. Front. Immunol. 2024, 15, 1457691. [CrossRef]
- Hu, X.; Zhu, H.; Chen, B.; Li, Q.; Zhang, L.; Li, W. The oncogenic role of tubulin alpha-1c chain in human tumours. BMC Cancer 2022, 22, 1–13. [CrossRef]
- Palsson-McDermott, E.M.; Dyck, L.; Zaslona, Z.; Menon, D.; McGettrick, A.F.; Mills, K.H.G.; et al. Pyruvate kinase M2 is required for the expression of the immune checkpoint PD-L1 in immune cells and tumors. Front. Immunol. 2017, 8, 1300. [CrossRef]
- Bruneel, K.; Verstappe, J.; Vandamme, N.; Berx, G. Intrinsic balance between ZEB family members is important for melanocyte homeostasis and melanoma progression. Cancers (Basel) 2020, 12, 2248. [CrossRef]
- Berr, A.L.; Wiese, K.; dos Santos, G.; Hendriks, W.; Klemke, M.; et al. Vimentin is required for tumor progression and metastasis in a mouse model of non–small cell lung cancer. Oncogene 2023, 42, 2074–2087. [CrossRef]
- Wolosowicz, M.; Prokopiuk, S.; Kaminski, T.W. The complex role of matrix metalloproteinase-2 (MMP-2) in health and disease. Int. J. Mol. Sci. 2024, 25, 13691. [CrossRef]
- Jiang, X.; Zhang, H.; Zhang, H.; Zhang, Y.; Zhang, Q.; et al. Microcystin-LR-induced interaction between M2 tumor-associated macrophage and colorectal cancer cell promotes colorectal cancer cell migration through regulating the expression of TGF-β1 and CST3. Int. J. Mol. Sci. 2023, 24, 10527. [CrossRef]
- Liu, Z.; Chen, M.; Zheng, W.; Yuan, S.; Zhao, W. Insights into the prognostic value and immunological role of CD74 in pan-cancer. Discover Oncol. 2024, 15, 1–17. [CrossRef]
- Imaoka, M.; Tanese, K.; Masugi, Y.; Hayashi, M.; Sakamoto, M. Macrophage migration inhibitory factor-CD74 interaction regulates the expression of programmed cell death ligand 1 in melanoma cells. Cancer Sci. 2019, 110, 2273–2283. [CrossRef]
- Ruan, W.J.; Lin, J.; Xu, E.P.; Chen, X.; et al. IGFBP7 plays a potential tumor suppressor role against colorectal carcinogenesis with its expression associated with DNA hypomethylation of exon 1. J. Zhejiang Univ. Sci. B 2006, 7, 929–932. [CrossRef]
- Li, Y.; Xi, Y.; Zhu, G.; Zhang, X.; Chen, Y.; et al. Downregulated IGFBP7 facilitates liver metastasis by modulating epithelial-mesenchymal transition in colon cancer. Oncol. Rep. 2019, 42, 1935–1945. [CrossRef]
- Qiu, X.; Quan, G.; Ou, W.; Chen, Y.; et al. Unraveling TIMP1: a multifaceted biomarker in colorectal cancer. Front. Genet. 2023, 14, 1265137. [CrossRef]
- Chen, M.; Liu, H.; Li, Z.; Ming, A.L.; Chen, H. Mechanism of PKM2 affecting cancer immunity and metabolism in tumor microenvironment. J. Cancer 2021, 12, 3566–3574. [CrossRef]
- Rodríguez, D.; Morrison, C.J.; Overall, C.M. Matrix metalloproteinases: What do they not do? New substrates and biological roles identified by murine models and proteomics. Biochim. Biophys. Acta Mol. Cell Res. 2010, 1803, 39–54. [CrossRef]
- Peddle, A.M.; Rasschaert, G.; Tejpar, S. Immunosensitivity cuts across mismatch repair status in colorectal cancer. Cancer Cell 2025, 43, 175–177. [CrossRef]
- Ragland, S.A.; Criss, A.K. From bacterial killing to immune modulation: Recent insights into the functions of lysozyme. PLoS Pathog. 2017, 13, e1006512. [CrossRef]
- Gao, P.; Liu, H.; Huang, H.; Zhang, W.; et al. The Crohn disease-associated ATG16L1T300A polymorphism regulates inflammatory responses by modulating TLR- and NLR-mediated signaling. Autophagy 2022, 18, 2561–2575. [CrossRef]
- Ahn, S.B.; Sharma, S.; Mohamedali, A.; et al. Potential early clinical stage colorectal cancer diagnosis using a proteomics blood test panel. Clin. Proteomics 2019, 16, 1–13. [CrossRef]
- De Martino, D.; Bravo-Cordero, J.J. Collagens in cancer: Structural regulators and guardians of cancer progression. Cancer Res. 2023, 83, 1–13. [CrossRef]
- Elsafadi, M.; Manikandan, M.; Almalki, S.; et al. Transgelin is a poor prognostic factor associated with advanced colorectal cancer (CRC) stage promoting tumor growth and migration in a TGFβ-dependent manner. Cell Death Dis. 2020, 11, 1–16. [CrossRef]
- Lee, H.W.; Park, Y.M.; Lee, S.J.; Kim, S.Y.; et al. Alpha-smooth muscle actin (ACTA2) is required for metastatic potential of human lung adenocarcinoma. Clin. Cancer Res. 2013, 19, 5879–5889. [CrossRef]
- Lee, J.M.; Dedhar, S.; Kalluri, R.; Thompson, E.W. The epithelial-mesenchymal transition: New insights in signaling, development, and disease. J. Cell Biol. 2006, 172, 973–981. [CrossRef]
- Afshar-Kharghan, V. The role of the complement system in cancer. J. Clin. Investig. 2017, 127, 780–789. [CrossRef]
- Vandooren, J.; Itoh, Y. Alpha-2-Macroglobulin in inflammation, immunity and infections. Front. Immunol. 2021, 12, 803244. [CrossRef]
- Almitairi, J.O.M.; Venkatraman Girija, U.; Furze, C.M.; et al. Structure of the C1r–C1s interaction of the C1 complex of complement activation. Proc. Natl. Acad. Sci. USA 2018, 115, 768–773. [CrossRef]
- Zhang, W.; Ge, Y.; Cheng, Q.; Zhang, Q.; Fang, L.; Zheng, J. Decorin is a pivotal effector in the extracellular matrix and tumour microenvironment. Oncotarget 2018, 9, 5480–5491. [CrossRef]
- Chen, X.; Song, E. Turning foes to friends: Targeting cancer-associated fibroblasts. Nat. Rev. Drug Discov. 2019, 18, 99–115. [CrossRef]
- Liu, T.; Zhou, L.; Li, D.; Andl, T.; Zhang, Y. Cancer-associated fibroblasts build and secure the tumor microenvironment. Front. Cell Dev. Biol. 2019, 7, 60. [CrossRef]
- Nobin, H.; Garvin, S.; Hagman, H.; Nodin, B.; Jirström, K.; Brunnström, H. The prognostic value of programmed death-ligand 1 (PD-L1) expression in resected colorectal cancer without neoadjuvant therapy—Differences between antibody clones and cell types. BMC Cancer 2024, 24, 1051. [CrossRef]
- Shan, T.; Chen, S.; Wu, T.; Yang, Y.; Li, S.; Chen, X. PD-L1 expression in colon cancer and its relationship with clinical prognosis. Int. J. Clin. Exp. Pathol. 2019, 12, 1764–1769. PMID: 31933995.
- Zhong, G.; Peng, C.; Chen, Y.; et al. Expression of STING and PD-L1 in colorectal cancer and their correlation with clinical prognosis. Int. J. Clin. Exp. Pathol. 2018, 11, 1256–1264. PMID: 31938220.
- Rosenbaum, M.W.; Bledsoe, J.R.; Morales-Oyarvide, V.; Huynh, T.G.; Mino-Kenudson, M. PD-L1 expression in colorectal cancer is associated with microsatellite instability, BRAF mutation, medullary morphology and cytotoxic tumor-infiltrating lymphocytes. Mod. Pathol. 2016, 29, 1104–1112. [CrossRef]
- Nie, F.; Sun, X.; Sun, J.; Zhang, J.; Wang, Y. Epithelial-mesenchymal transition in colorectal cancer metastasis and progression: Molecular mechanisms and therapeutic strategies. Cell Death Discov. 2025, 11, 336. [CrossRef]
- Gu, Y.; Zhang, Z.; ten Dijke, P. Harnessing epithelial-mesenchymal plasticity to boost cancer immunotherapy. Cell Mol. Immunol. 2023, 20, 318–340. [CrossRef]
- Martinez-Ciarpaglini, C.; Oltra, S.; Roselló, S.; et al. Low miR200c expression in tumor budding of invasive front predicts worse survival in patients with localized colon cancer and is related to PD-L1 overexpression. Mod. Pathol. 2019, 32, 306–313. [CrossRef]
- Fan, S.; Cui, X.; Zheng, L.; et al. Prognostic value of desmoplastic stromal reaction, tumor budding and tumor-stroma ratio in stage II colorectal cancer. J. Gastrointest. Oncol. 2022, 13, 2903–2921. [CrossRef]
- Huijbers, A.; Tollenaar, R.A.E.M.; van Pelt, G.W.; et al. The proportion of tumor-stroma as a strong prognosticator for stage II and III colon cancer patients: Validation in the VICTOR trial. Ann. Oncol. 2013, 24, 179–185. [CrossRef]
- Polack, M.; Smit, M.A.; van Pelt, G.W.; et al. Results from the UNITED study: A multicenter study validating the prognostic effect of the tumor–stroma ratio in colon cancer. ESMO Open 2024, 9, 102988. [CrossRef]
- Marcuello, M.; Mayol, X.; Felipe-Fumero, E.; et al. Modulation of the colon cancer cell phenotype by pro-inflammatory macrophages: A preclinical model of surgery-associated inflammation and tumor recurrence. PLoS ONE 2018, 13, e0192958. [CrossRef]
- Chen, X.; Yang, M.; Yin, J.; et al. Tumor-associated macrophages promote epithelial–mesenchymal transition and the cancer stem cell properties in triple-negative breast cancer through CCL2/AKT/β-catenin signaling. Cell Commun. Signal. 2022, 20, 92. [CrossRef]
- Wei, C.; Yang, C.; Wang, S.; et al. Crosstalk between cancer cells and tumor associated macrophages is required for mesenchymal circulating tumor cell-mediated colorectal cancer metastasis. Mol. Cancer 2019, 18, 64. [CrossRef]
- Yerolatsite, M.; Torounidou, N.; Amylidi, A.L.; Zarkavelis, G.; Hadjigeorgiou, L.; Lampri, E.; Bali, C.; Georvasili, V.; Kampletsas, E.; Mauri, D. The role of tumor-associated macrophages and PD-1/PD-L1 networking in colorectal cancer. Contemp. Oncol. (Pozn) 2025, 29, 123–130. [CrossRef]
- Xiang, J.; Wang, J.; Xiao, H.; Huang, C.; Wu, C.; Zhang, L.; Qian, C.; Xiang, D. Targeting tumor-associated macrophages in colon cancer: Mechanisms and therapeutic strategies. Front. Immunol. 2025, 16, 1573917. [CrossRef]
- Posch, F.; Silina, K.; Leibl, S.; et al. Maturation of tertiary lymphoid structures and recurrence of stage II and III colorectal cancer. Oncoimmunology 2018, 7, e1378844. [CrossRef]
 |
 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.




