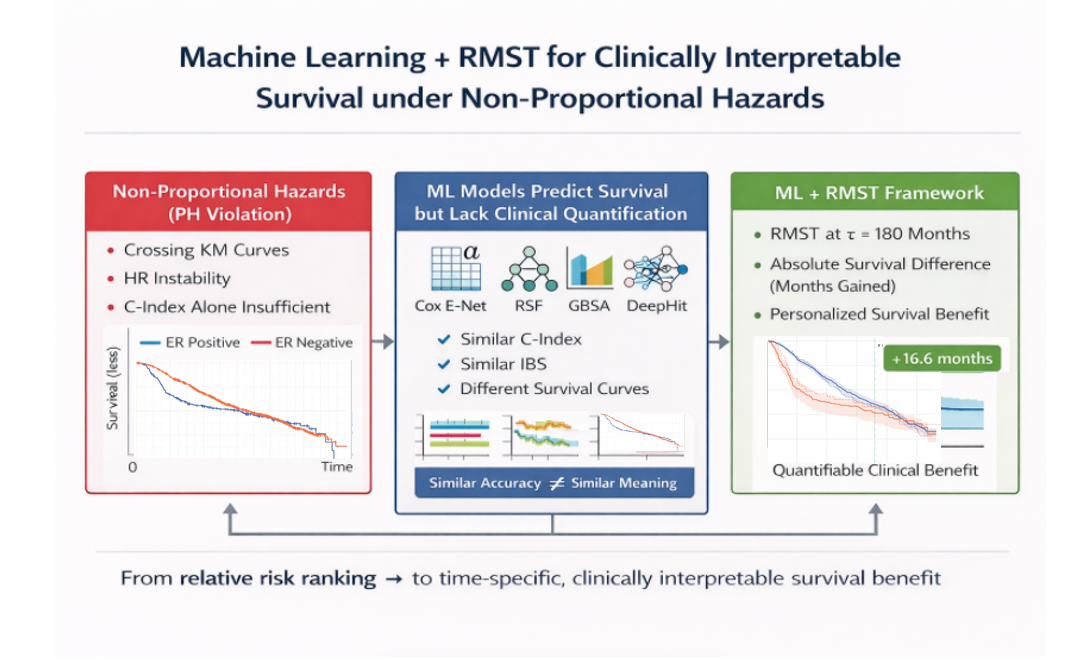
Background: Advances in machine learning (ML) based survival modeling enable the analysis of high-dimensional biomedical data. However, many approaches rely on the proportional hazards (PH) assumption, which is frequently violated in oncology and can limit the interpretability of hazard ratio–based results. Using Estrogen Receptor (ER) status in the METABRIC breast cancer cohort as a case study, we propose a framework that integrates machine learning survival models with Restricted Mean Survival Time (RMST) to provide a more robust and clinically interpretable approach for survival analysis under non-proportional hazards. Methods: Overall survival was analyzed in 1104 patients. PH violations were confirmed using Schoenfeld residuals and Kaplan–Meier inspection. We compared four models: stratified Cox Elastic Net (Cox E-Net), Random Survival Forest (RSF), Gradient Boosting Survival Analysis (GBSA), and DeepHit. Performance was assessed using Harrell’s C-index, time-dependent IPCW C-index, and Integrated Brier Score (IBS). RMST at 180 months was utilized to quantify absolute survival differences between ER subgroups. To improve the stability of the estimates, 200 bootstrap resamples were performed, and 95% confidence intervals were derived from the bootstrap distribution. Results: ER status demonstrated significant PH violation (p < 0.005) with crossing survival curves. Discrimination (C-index 0.664–0.725) and calibration (IBS 0.149–0.169) were comparable across models, with RSF achieving the highest overall performance. Despite similar accuracy, survival curve structures differed substantially. Cox E-Net and RSF reproduced the observed crossing pattern, whereas GBSA generated smoother trajectories and DeepHit showed marked compression of subgroup separation. In the independent test cohort, the empirical RMST difference at 180 months was 16.6 months (ER-positive: 130.4; ER-negative: 113.8). Model-based RMST differences ranged from 1 month (DeepHit) to 27 months (Cox E-Net), with RSF and GBSA (12.8 and 13.8 months) most closely approximating the empirical benchmark. Conclusions: We propose a novel, model-agnostic ML + RMST framework that addresses non-proportional hazards while providing quantifiable, time-specific clinical benefit. Moreover, models with similar discrimination and calibration produced markedly different survival curve behavior and absolute RMST estimates, demonstrating that accuracy metrics alone are insufficient for clinical interpretation. By linking predictive modeling with absolute survival quantification, this framework advances survival evaluation beyond relative risk ranking toward clinically meaningful decision support.