Submitted:
09 March 2026
Posted:
12 March 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
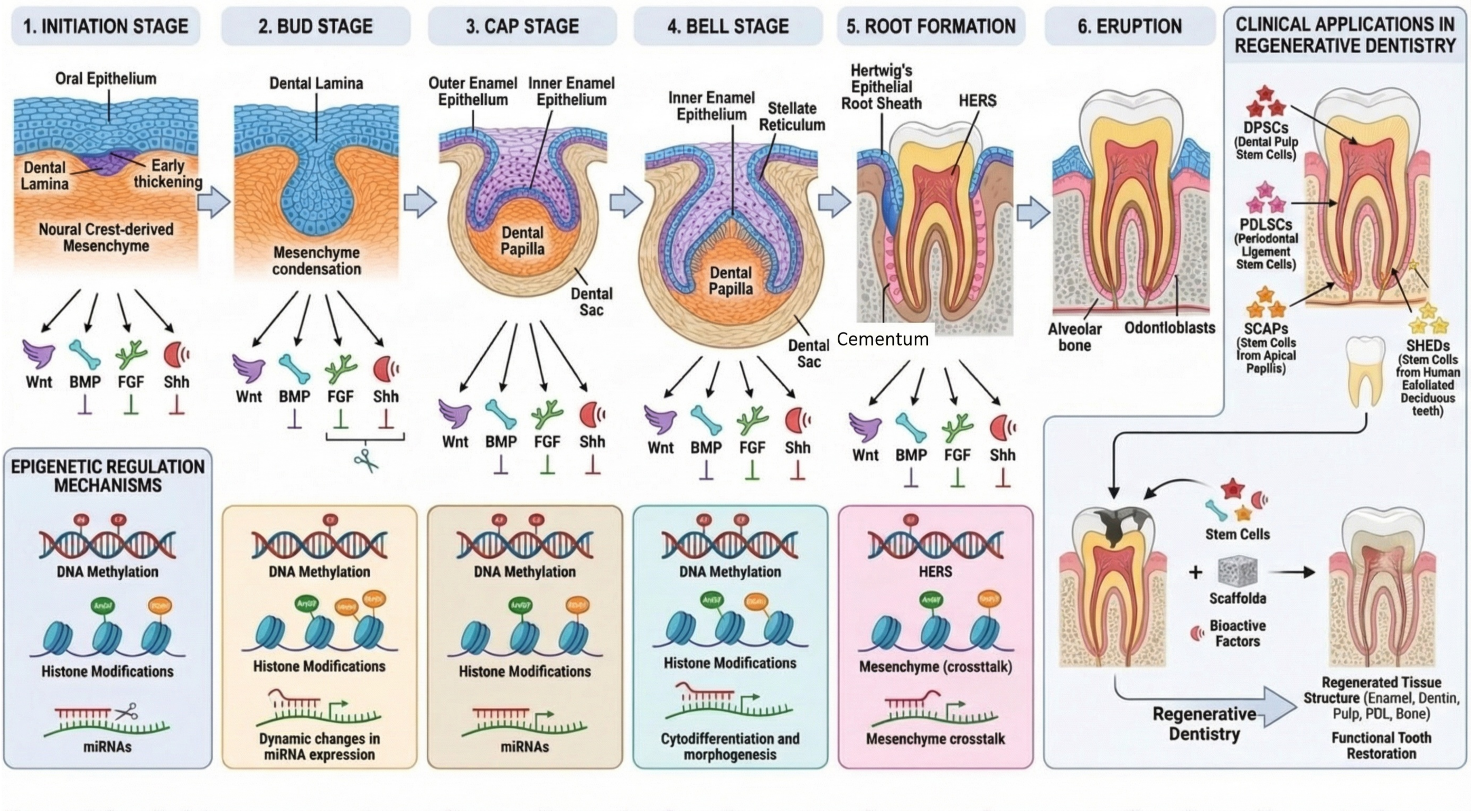
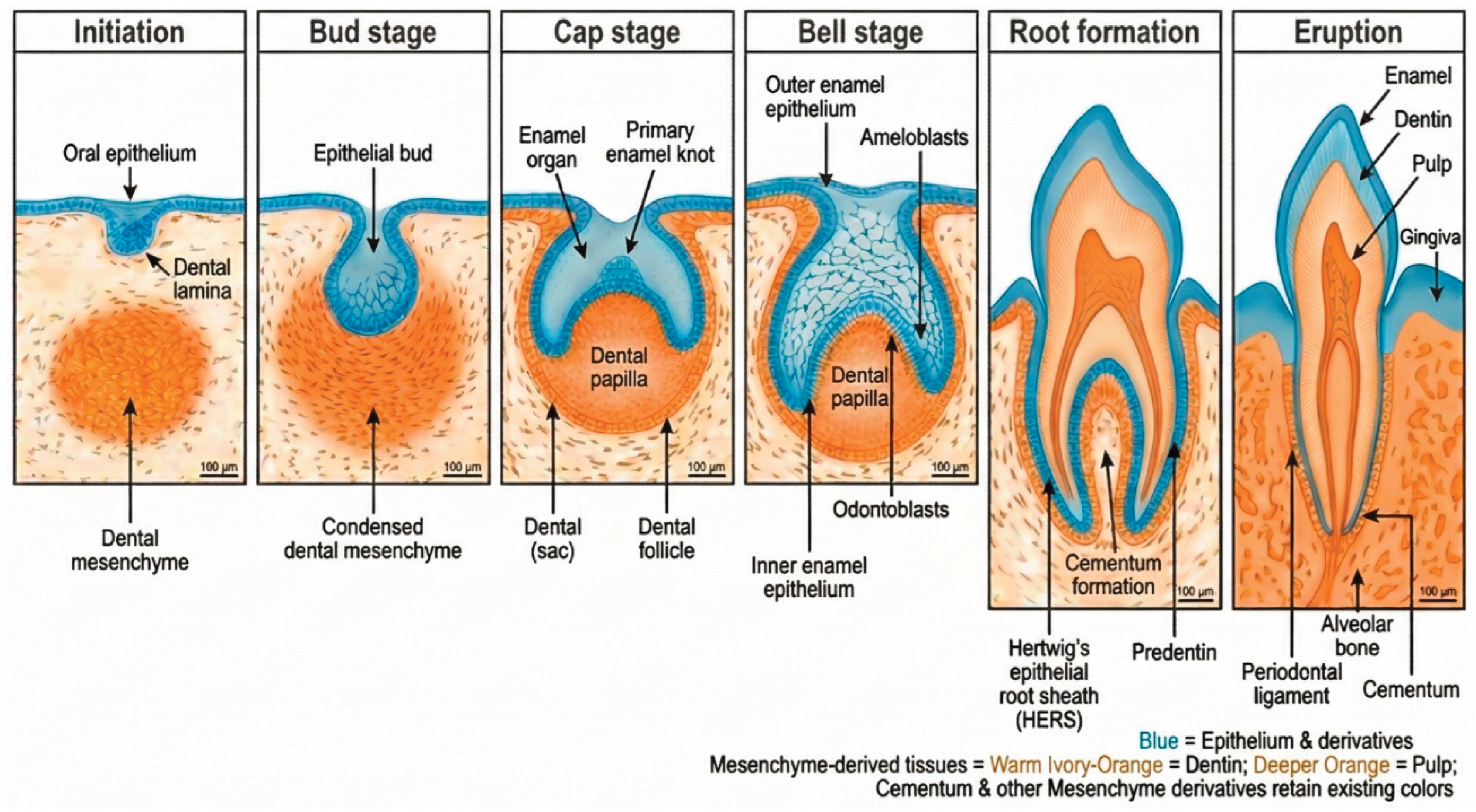
2. Morphological Stages of Tooth Development
2.1. Initiation Stage
2.2. Bud Stage
2.3. Cap Stage
2.4. Bell Stage
2.5. Root Formation and Maturation
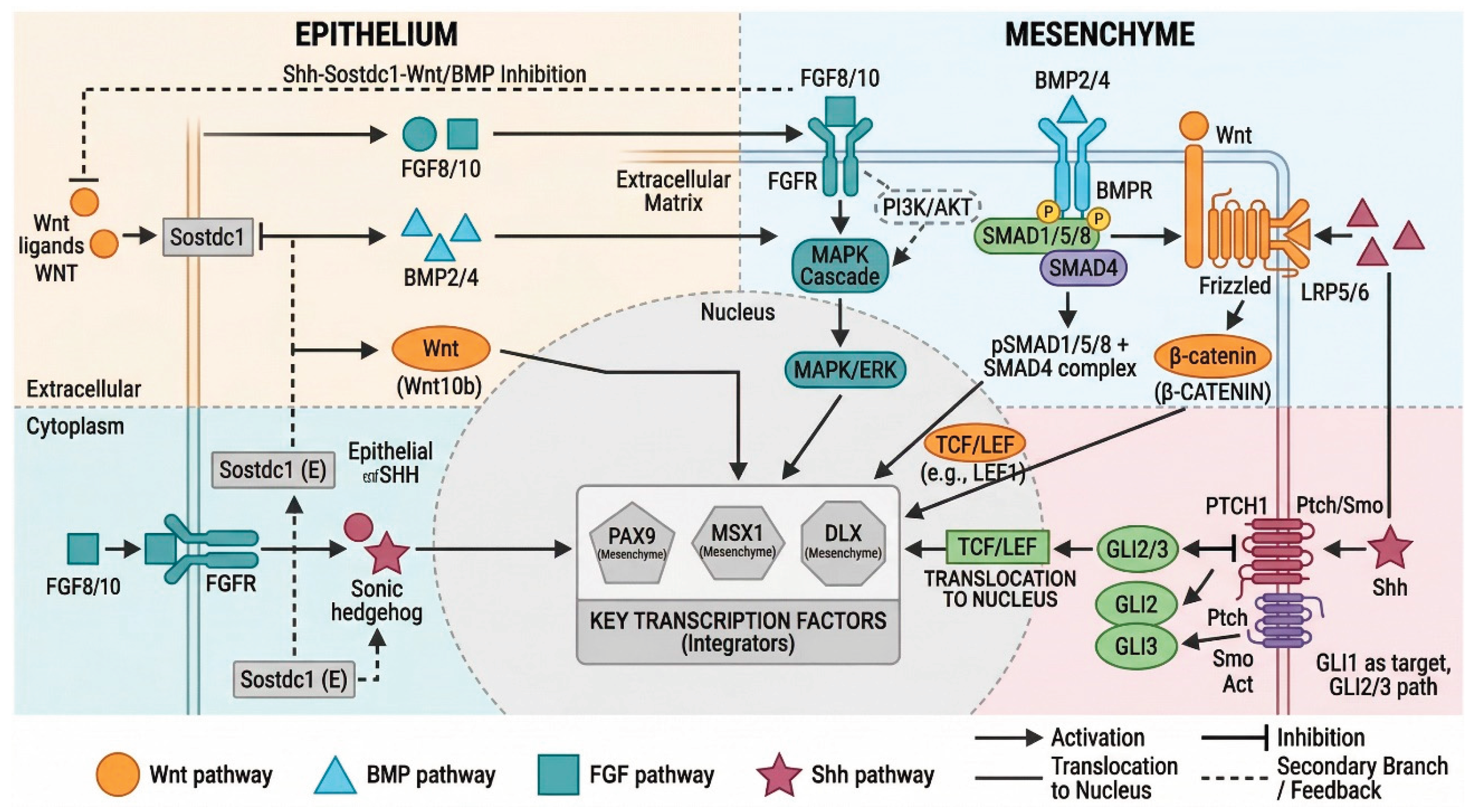
3. Genetic Regulation: Signaling Pathways
3.1. Wnt/β-Catenin Signaling
3.2. Bone Morphogenetic Protein (BMP) Signaling
3.3. Fibroblast Growth Factor (FGF) Signaling
3.4. Sonic Hedgehog (Shh) Signaling
3.5. Pathway Crosstalk and Feedback Regulation
3.6. Additional Regulatory Modules and Mechanical Cues
4. Genetic Regulation: Transcription Factors
4.1. PAX9
4.2. MSX1 and MSX2
4.3. PITX2
4.4. LEF1
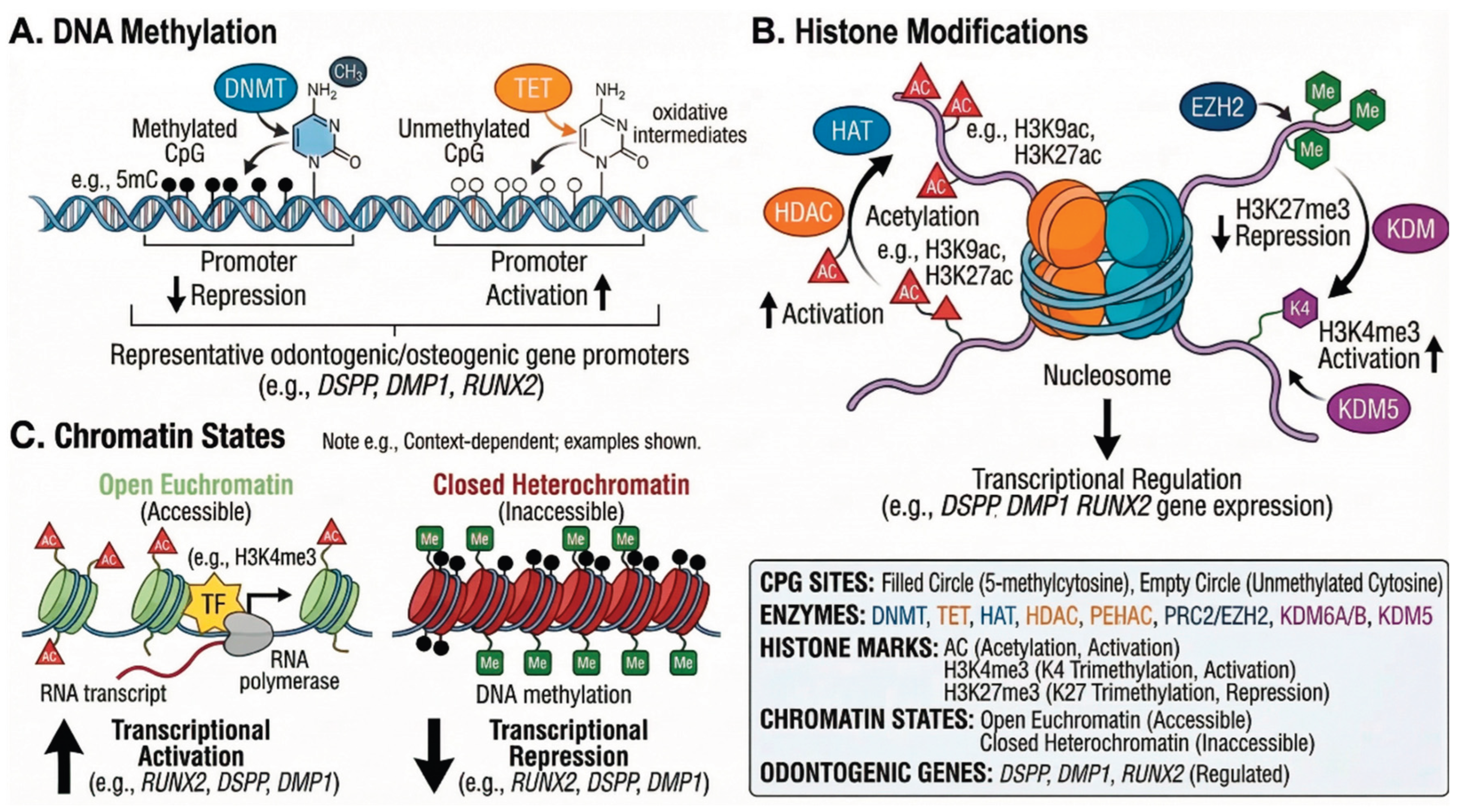
5. Epigenetic Regulation of Tooth Development
5.1. DNA Methylation
5.2. Histone Modifications
- KDM6B: Decreases H3K27me3 levels at the IGFBP5 promoter, promoting odontoblast differentiation and mineralization in PDLSCs (in vitro studies).
- KDM2A: Suppresses differentiation by reducing H3K4 and H3K36 methylation at the epiregulin (EREG) promoter.
- KDM4B: Upregulated by DLX5, promotes osteo-/dentinogenesis in stem cells from apical papilla (SCAPs) in nude mouse xenograft models.
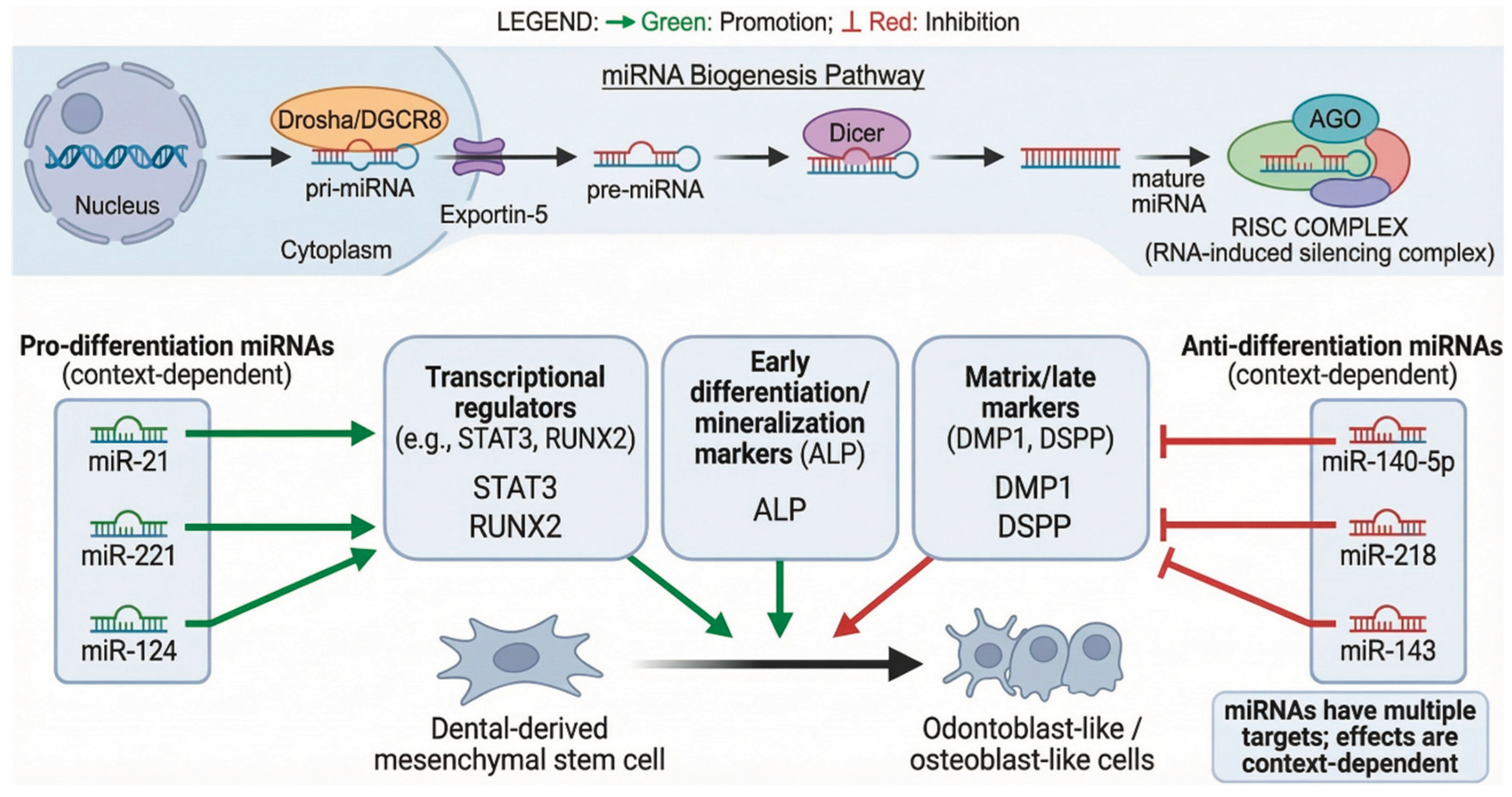
5.3. MicroRNA Regulation
- miR-21: Promotes differentiation by enhancing STAT3 signaling (e.g., a TNF-α–miR-21–STAT3 axis), likely via indirect modulation of STAT3 activity rather than direct STAT3 mRNA targeting; supported by DPSC in vitro studies and transplantation models.
- miR-221 and miR-124: Enhance odontogenic gene expression and mineralization in DPSC cultures.
- miR-140-5p: Inhibits odontoblastic differentiation by targeting the Wnt1/β-catenin signaling pathway, thereby reducing DSPP and DMP-1 expression [40].
- miR-218: Suppresses DMP1 and DSPP expression.
- miR-143: Inhibits multiple osteogenic markers.
| miRNA | Reported target(s)/ pathway (net effect) |
Model system (example) | Effect on odontogenic differentiation | Evidence type (representative) | Key refs | Evidence strength |
|---|---|---|---|---|---|---|
| miR-21 | STAT3 signaling axis (net pro-differentiation; inflammatory priming context) | Human DPSCs | Promotes odontoblast differentiation and mineralization markers | miR-21/STAT3 axis interrogation (gain/loss-of-function; differentiation markers) | [37,38,39] | Primarily in vitro; mechanistic axis defined |
| miR-27a | DKK3 and SOSTDC1 (Wnt/BMP signaling; net pro-differentiation) | Human DPSCs | Enhances odonto/osteoblastic differentiation and mineralization | Mimic/inhibitor + luciferase validation; ARS/ALP assays; in vivo bone formation reported | [37,38,44] | In vitro + in vivo (bone formation); translational relevance |
| miR-34a | NOTCH2 and HES1 (Notch signaling; net pro-differentiation) | Human SCAPs | Promotes odontogenic/osteogenic differentiation (DSPP/RUNX2/OSX/OCN up) | Direct 3′UTR targeting + marker upregulation; Notch–miR feedback | [37,38,43] | Human stem cells in vitro; mechanistic target validation |
| miR-140-5p | Wnt1/β-catenin (net anti-differentiation when overexpressed) | Human DPSCs | Suppresses odontoblastic differentiation by reducing DSPP/DMP-1 via Wnt1 targeting; inhibitor promotes differentiation | Luciferase-validated targeting of Wnt1; marker and mineralization assays | [37,38,40] | In vitro + target validation |
| miR-218 | RUNX2 (net anti-osteogenic/odontogenic when high; decreases upon induction) | Human dental stem cells (PDLSC/DPSC etc.) | Downregulated during mineralized differentiation; targets RUNX2 | MiRNA profiling during induction; functional targeting of RUNX2 | [37,38,41] | Human cells; profiling + functional targeting |
| miR-143-3p | RANK (OPG–RANKL axis; net inhibitory on differentiation when high) | Human DPSCs | Inhibition enhances mineralization and upregulates DSPP/BSP/ALP/OCN/OPN | Dual luciferase validation; differentiation markers + mineralization assays | [37,38,42] | In vitro + mechanistic pathway evidence |
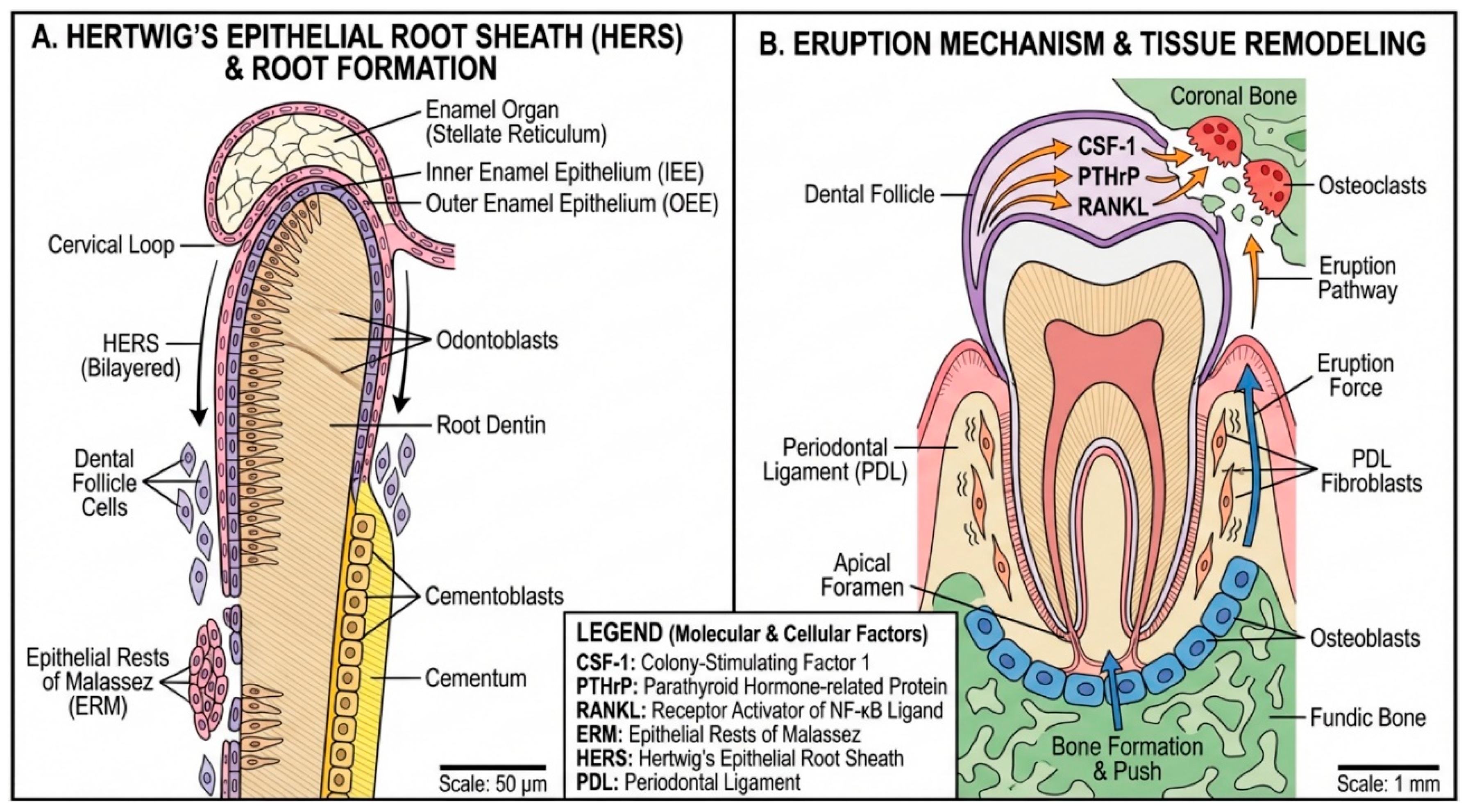
6. Root Formation and Tooth Eruption
6.1. Hertwig’s Epithelial Root Sheath (HERS)
- Root morphogenesis: HERS outlines root shape and number by guiding apical extension.
- Odontoblast induction: HERS induces dental papilla cells to differentiate into root odontoblasts through secreted signaling molecules.
- Cementogenesis: After inducing dentin formation, HERS fragments, allowing dental follicle cells to contact the root surface and differentiate into cementoblasts.
6.2. Tooth Eruption Mechanisms
- CSF-1: Essential for osteoclast differentiation; expressed in the dental follicle overlying the erupting tooth.
- RANKL/OPG axis: RANKL promotes osteoclastogenesis; the dental follicle produces RANKL coronally and OPG basally, creating an asymmetric resorption pattern.
- PTHrP: Regulates RANKL/OPG balance; required for eruption through the PTH1R receptor.
7. Clinical Implications and Regenerative Applications
7.1. Genetic Diagnosis of Dental Anomalies
- AXIN2: Wnt pathway regulator; variants/polymorphisms have been associated with selective tooth agenesis in human genetic studies (note that AXIN2 is also discussed in broader Wnt-related phenotypes in other literature) [35].
- PAX9: Most common cause of nonsyndromic oligodontia, primarily affecting molars.
- MSX1: Causes oligodontia affecting premolars and third molars.
- WNT10A: Major contributor to hypodontia and ectodermal dysplasia.
- PTH1R: Causes primary failure of eruption.
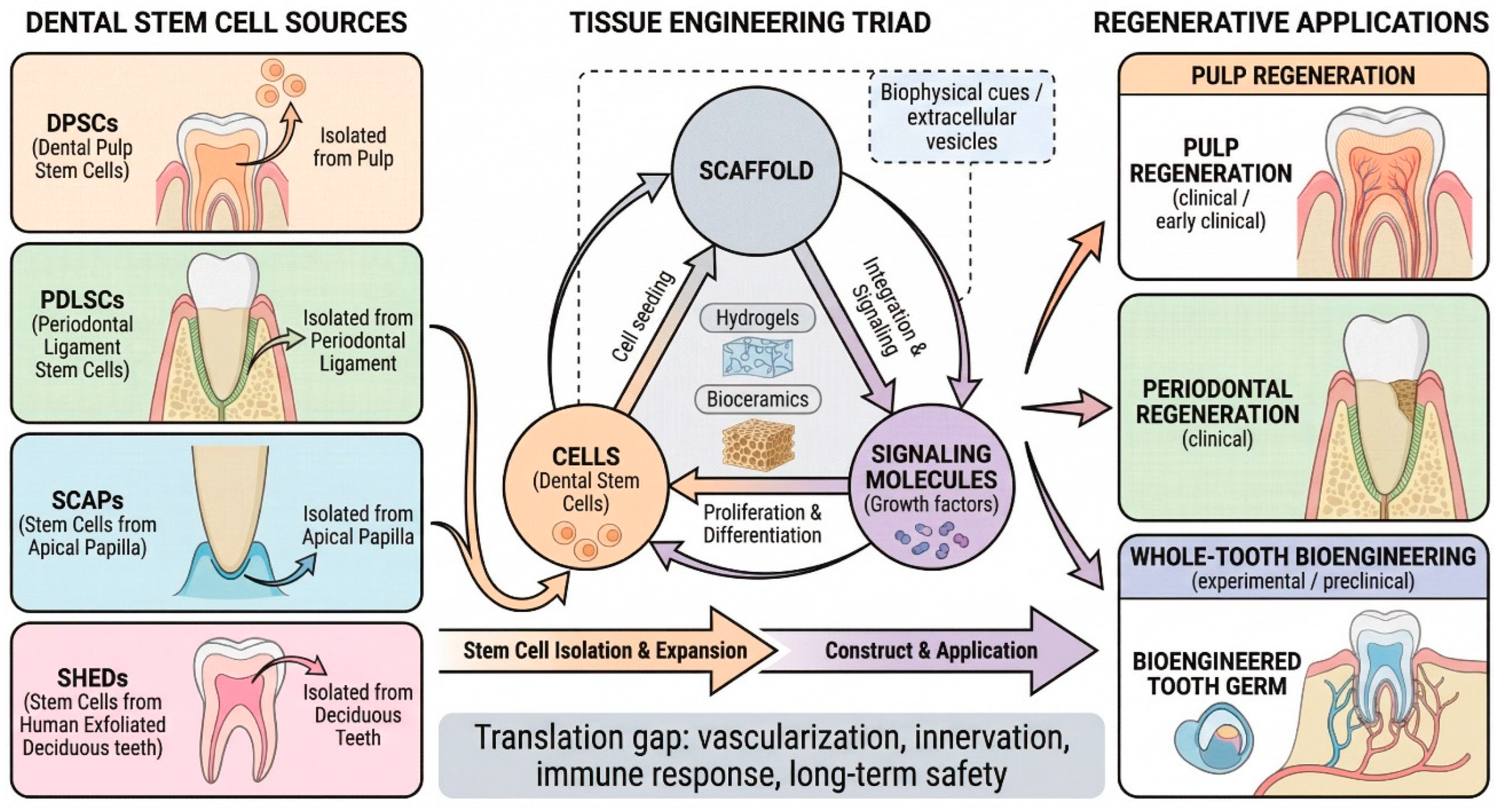
7.2. Dental Stem Cells for Regenerative Medicine
- DPSCs (Dental Pulp Stem Cells): Capable of odontogenic, neurogenic, and angiogenic differentiation in vitro and in animal models.
- SHEDs (Stem cells from Human Exfoliated Deciduous teeth): Higher proliferative capacity than DPSCs in culture.
- PDLSCs (Periodontal Ligament Stem Cells): Can regenerate cementum and periodontal ligament in animal xenograft models.
- SCAPs (Stem Cells from Apical Papilla): Located at developing root apices; important for root maturation.
7.3. Tissue Engineering and Future Directions
- Off-target effects of epigenetic modulators
- Delivery methods for clinical application
- Long-term safety profiles
- Scalability and cost considerations
- Near-term (1–3 years): Single-cell transcriptomic mapping of human dental development to identify cell populations and regulatory networks.
- Medium-term (3–5 years): Development and validation of human iPSC-derived dental cell models and organoid systems for mechanistic studies and drug screening.
- Long-term (5–10 years): Controlled clinical trials of stem cell-based therapies for pulp regeneration, periodontal regeneration, and potentially whole tooth bioengineering. Recent breakthroughs in iPSC-derived dental organoids, such as the generation of human enamel-secreting ameloblast-like cells from iPSCs guided by single-cell transcriptomic atlases [52], underscore the feasibility of this approach.
8. Limitations of Current Evidence
9. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Declaration of Interest Statement
References
- Thesleff, I. Epithelial-mesenchymal signalling regulating tooth morphogenesis. J. Cell Sci. 2003, 116, 1647–1648. [CrossRef]
- Tucker, A.; Sharpe, P. The cutting-edge of mammalian development; how the embryo makes teeth. Nat. Rev. Genet. 2004, 5, 499–508. [CrossRef]
- Jernvall, J.; Thesleff, I. Reiterative signaling and patterning during mammalian tooth morphogenesis. Mech. Dev. 2000, 92, 19–29. [CrossRef]
- Pispa, J.; Thesleff, I. Mechanisms of ectodermal organogenesis. Dev. Biol. 2003, 262, 195–205. [CrossRef]
- Klein, O.D.; Oberoi, S.; Huysseune, A.; Hovorakova, M.; Peterka, M.; Peterkova, R. Developmental disorders of the dentition: an update. Am. J. Med. Genet. C Semin. Med. Genet. 2013, 163, 318–332. [CrossRef]
- Bei, M. Molecular genetics of tooth development. Curr. Opin. Genet. Dev. 2009, 19, 504–510. [CrossRef]
- Balic, A.; Thesleff, I. Tissue interactions regulating tooth development and renewal. Curr. Top. Dev. Biol. 2015, 115, 157–186. [CrossRef]
- Jussila, M.; Thesleff, I. Signaling networks regulating tooth organogenesis and regeneration, and the specification of dental mesenchymal and epithelial cell lineages. Cold Spring Harb. Perspect. Biol. 2012, 4, a008425. [CrossRef]
- Hermans, F.; Hemeryck, L.; Lambrichts, I.; Bronckaers, A.; Vankelecom, H. Intertwined signaling pathways governing tooth development: A Give-and-Take Between Canonical Wnt and Shh. Front. Cell Dev. Biol. 2021, 9, 758203. [CrossRef]
- Tokavanich, N.; Wein, M.N.; English, J.D.; Ono, N.; Ono, W. The role of Wnt signaling in postnatal tooth root development. Front. Dent. Med. 2021, 2, 769134. [CrossRef]
- Chen, Y.; Wang, X.; Wu, Z.; Jia, S.; Wan, M. Epigenetic regulation of dental-derived stem cells and their application in pulp and periodontal regeneration. PeerJ 2023, 11, e14550. [CrossRef]
- Huang, L.; Chen, X.; Yang, X.; Zhang, Y.; Liang, Y.; Qiu, X. Elucidating epigenetic mechanisms governing odontogenic differentiation in dental pulp stem cells: an in-depth exploration. Front. Cell Dev. Biol. 2024, 12, 1394582. [CrossRef]
- Shah, P.; Aghazadeh, M.; Rajasingh, S.; Rajasingh, J.; Jain, V.; Dixon, D. Stem cells in regenerative dentistry: Current understanding and future directions. J. Oral Biosci. 2024, 66, 288–299. [CrossRef]
- Al Madhoun, A.; Sindhu, S.; Haddad, D.; Atari, M.; Ahmad, R.; Al-Mulla, F. Dental pulp stem cells derived from adult human third molar tooth: A brief review. Front. Cell Dev. Biol. 2021, 9, 717624. [CrossRef]
- Gronthos, S.; Mankani, M.; Brahim, J.; Robey, P.G.; Shi, S. Postnatal human dental pulp stem cells (DPSCs) in vitro and in vivo. Proc. Natl. Acad. Sci. USA 2000, 97, 13625–13630. [CrossRef]
- Seo, B.M.; Miura, M.; Gronthos, S.; Bartold, P.M.; Batouli, S.; Brahim, J.; Young, M.; Robey, P.G.; Wang, C.Y.; Shi, S. Investigation of multipotent postnatal stem cells from human periodontal ligament. Lancet 2004, 364, 149–155. [CrossRef]
- Thesleff, I.; Mikkola, M. The role of growth factors in tooth development. Int. Rev. Cytol. 2002, 217, 93–135. [CrossRef]
- Cobourne, M.T.; Sharpe, P.T. Tooth and jaw: molecular mechanisms of patterning in the first branchial arch. Arch. Oral Biol. 2003, 48, 1–14. [CrossRef]
- Andl, T.; Reddy, S.T.; Gaddapara, T.; Millar, S.E. WNT signals are required for the initiation of hair follicle development. Dev. Cell 2002, 2, 643–653. [CrossRef]
- Chen, J.; Lan, Y.; Baek, J.A.; Gao, Y.; Jiang, R. Wnt/beta-catenin signaling plays an essential role in activation of odontogenic mesenchyme during early tooth development. Dev. Biol. 2009, 334, 174–185. [CrossRef]
- Lin, C.R.; Kioussi, C.; O’Connell, S.; Briata, P.; Szeto, D.; Liu, F.; Izpisua-Belmonte, J.C.; Rosenfeld, M.G. Pitx2 regulates lung asymmetry, cardiac positioning and pituitary and tooth morphogenesis. Nature 1999, 401, 279–282. [CrossRef]
- Peters, H.; Neubüser, A.; Kratochwil, K.; Balling, R. Pax9-deficient mice lack pharyngeal pouch derivatives and teeth and exhibit craniofacial and limb abnormalities. Genes Dev. 1998, 12, 2735–2747. [CrossRef]
- Satokata, I.; Maas, R. Msx1 deficient mice exhibit cleft palate and abnormalities of craniofacial and tooth development. Nat. Genet. 1994, 6, 348–356. [CrossRef]
- Cho, S.W.; Kwak, S.; Woolley, T.E.; Lee, M.J.; Kim, E.J.; Baker, R.E.; Kim, H.J.; Shin, J.S.; Tickle, C.; Maini, P.K.; et al. Interactions between Shh, Sostdc1 and Wnt signaling and a new feedback loop for spatial patterning of the teeth. Development 2011, 138, 1807–1816. [CrossRef]
- Diekwisch, T.G. The developmental biology of cementum. Int. J. Dev. Biol. 2001, 45, 695–706. [CrossRef]
- Yamamoto, H.; Cho, S.W.; Kim, E.J.; Kim, J.Y.; Fujiwara, N.; Jung, H.S. Developmental properties of the Hertwig’s epithelial root sheath in mice. J. Dent. Res. 2004, 83, 688–692. [CrossRef]
- Liu, C.; Guo, H.; Shi, C.; Sun, H. BMP signaling in the development and regeneration of tooth roots: from mechanisms to applications. Front. Cell Dev. Biol. 2023, 11, 1272201. [CrossRef]
- Kratochwil, K.; Galceran, J.; Tontsch, S.; Roth, W.; Grosschedl, R. FGF4, a direct target of LEF1 and Wnt signaling, can rescue the arrest of tooth organogenesis in Lef1−/− mice. Genes Dev. 2002, 16, 3173–3185. [CrossRef]
- Wang, X.P.; O’Connell, D.J.; Lund, J.J.; Saadi, I.; Kuraguchi, M.; Turbe-Doan, A.; Cavallesco, R.; Kim, H.; Park, P.J.; Harada, H.; et al. Apc inhibition of Wnt signaling regulates supernumerary tooth formation during embryogenesis and throughout adulthood. Development 2009, 136, 1939–1949. [CrossRef]
- Ahn, Y.; Sanderson, B.W.; Klein, O.D.; Krumlauf, R. Inhibition of Wnt signaling by Wise (Sostdc1) and negative feedback from Shh controls tooth number and patterning. Development 2010, 137, 3221–3231. [CrossRef]
- Dassule, H.R.; Lewis, P.; Bei, M.; Maas, R.; McMahon, A.P. Sonic hedgehog regulates growth and morphogenesis of the tooth. Development 2000, 127, 4775–4785. [CrossRef]
- Lan, Y.; Jia, S.; Jiang, R. Molecular patterning of the mammalian dentition. Semin. Cell Dev. Biol. 2014, 25–26, 61–70. [CrossRef]
- Intarak, N.; Tongchairati, K.; Termteerapornpimol, K.; Chantarangsu, S.; Porntaveetus, T. Tooth agenesis patterns and variants in PAX9: A systematic review. Jpn. Dent. Sci. Rev. 2023, 59, 129–137. [CrossRef]
- Vastardis, H.; Karimbux, N.; Guthua, S.W.; Seidman, J.G.; Seidman, C.E. A human MSX1 homeodomain missense mutation causes selective tooth agenesis. Nat. Genet. 1996, 13, 417–421. [CrossRef]
- Mostowska, A.; Biedziak, B.; Jagodzinski, P.P. Axis inhibition protein 2 (AXIN2) polymorphisms may be a risk factor for selective tooth agenesis. J. Hum. Genet. 2006, 51, 262–266. [CrossRef]
- Zhang, H.; et al. Epigenetic Regulation of Methylation in Determining the Fate of Dental Mesenchymal Stem Cells. Stem Cells Int. 2022, 2022, 5015856. [CrossRef]
- Iranmanesh, P.; Vedaei, A.; Salehi-Mazandarani, S.; Nikpour, P.; Khazaei, S.; Khademi, A.; Galler, K.M.; Nekoofar, M.H.; Dummer, P.M.H. MicroRNAs-mediated regulation of the differentiation of dental pulp-derived mesenchymal stem cells: a systematic review and bioinformatic analysis. Stem Cell Res. Ther. 2023, 14, 76. [CrossRef]
- Giovannetti, A.; Guarnieri, R.; Petrizzelli, F.; Lazzari, S.; Padalino, G.; Traversa, A.; Napoli, A.; Di Giorgio, R.; Pizzuti, A.; Parisi, C.; et al. Small RNAs and tooth development: The role of microRNAs in tooth agenesis and impaction. J. Dent. Sci. 2024, 19, 2150–2156. [CrossRef]
- Xiao, J.; Zheng, K.; Feng, X.; Zhang, J.; Song, D.; Wang, C.; Shen, X.; Zhao, X.; Wei, C.; Huang, D.; Feng, G. MiR-21/STAT3 Signal Is Involved in Odontoblast Differentiation of Human Dental Pulp Stem Cells Mediated by TNF-α. Cell Reprogram. 2018, 20, 107–116. [CrossRef]
- Lu, X.; Chen, X.; Xing, J.; Lian, M.; Huang, D.; Lu, Y.; Feng, G.; Feng, X. miR-140-5p Regulates the Odontoblastic Differentiation of Dental Pulp Stem Cells via the Wnt1/β-Catenin Signaling Pathway. Stem Cell Res. Ther. 2019, 10, 226. [CrossRef]
- Gay, I.; Cavender, A.; Peto, D.; Sun, Z.; Speer, A.; Cao, H.; Amendt, B.A. Differentiation of Human Dental Stem Cells Reveals a Role for microRNA-218. J. Periodontal Res. 2014, 49, 110–120. [CrossRef]
- Yang, C.; Jia, R.; Zuo, Q.; Zheng, Y.; Wu, Q.; Luo, B.; Lin, P.; Yin, L. microRNA-143-3p Regulates Odontogenic Differentiation of Human Dental Pulp Stem Cells through Regulation of the Osteoprotegerin–RANK Ligand Pathway by Targeting RANK. Exp. Physiol. 2020, 105, 876–885. [CrossRef]
- Sun, F.; et al. Crosstalk between miR-34a and Notch Signaling Promotes Differentiation in Apical Papilla Stem Cells (SCAPs). J. Dent. Res. 2014, 93, 589–595. [CrossRef]
- Yu, Z.; Kawashima, N.; Sunada-Nara, K.; Wang, S.; Han, P.; Kieu, T.Q.; Ren, C.; Noda, S.; Tazawa, K.; Okiji, T. MicroRNA-27a Transfected Dental Pulp Stem Cells Undergo Odonto/Osteogenic Differentiation via Targeting DKK3 and SOSTDC1 in Wnt/BMP Signaling In Vitro and Enhance Bone Formation In Vivo. J. Transl. Med. 2025, 23, 189. [CrossRef]
- Wise, G.E.; Frazier-Bowers, S.; D’Souza, R.N. Cellular, molecular, and genetic determinants of tooth eruption. Crit. Rev. Oral Biol. Med. 2002, 13, 323–334. [CrossRef]
- Wise, G.E. Cellular and molecular basis of tooth eruption. Orthod. Craniofac. Res. 2009, 12, 67–73. [CrossRef]
- Lu, K.; Qian, Y.; Gong, J.; Li, Z.; Yu, M.; Wang, H. A novel PTH1R mutation causes primary failure of eruption via the cAMP-PI3K/AKT pathway. Prog. Orthod. 2025, 26, 7. [CrossRef]
- Quigley, R.M.; Kearney, M.; Kennedy, O.D.; Duncan, H.F. Tissue engineering approaches for dental pulp regeneration: The development of novel bioactive materials using pharmacological epigenetic inhibitors. Bioact. Mater. 2024, 40, 182–211. [CrossRef]
- Miura, M.; Gronthos, S.; Zhao, M.; Lu, B.; Fisher, L.W.; Robey, P.G.; Shi, S. SHED: stem cells from human exfoliated deciduous teeth. Proc. Natl. Acad. Sci. USA 2003, 100, 5807–5812. [CrossRef]
- Sonoyama, W.; Liu, Y.; Fang, D.; Yamaza, T.; Seo, B.M.; Zhang, C.; Liu, H.; Gronthos, S.; Wang, C.Y.; Wang, S.; et al. Mesenchymal stem cell-mediated functional tooth regeneration in swine. PLoS ONE 2006, 1, e79. [CrossRef]
- Nakao, K.; Morita, R.; Saji, Y.; Ishida, K.; Tomita, Y.; Ogawa, M.; Saitoh, M.; Tomooka, Y.; Tsuji, T. The development of a bioengineered organ germ method. Nat. Methods 2007, 4, 227–230. [CrossRef]
- Alghadeer, A.; et al. Single-cell census of human tooth development enables generation of human enamel. Dev. Cell 2023, 58, 2163–2180.e9. [CrossRef]
- Shi, Y.; et al. Spatiotemporal cell landscape of human embryonic tooth development. Cell Prolif. 2024, 57, e13653. [CrossRef]
- Laurikkala, J.; Mikkola, M.L.; Mustonen, T.; Åberg, T.; Koppinen, P.; Pispa, J.; Nieminen, P.; Galceran, J.; Grosschedl, R.; Thesleff, I. TNF signaling via the ligand-receptor pair ectodysplasin and Edar controls the function of epithelial signaling centers and is regulated by Wnt and Activin during tooth organogenesis. Dev. Biol. 2001, 229, 443–455. [CrossRef]
- Tucker, A.S.; Headon, D.J.; Courtney, J.M.; Overbeek, P.; Sharpe, P.T. The activation level of the TNF family receptor, Edar, determines cusp number and tooth number during tooth development. Dev. Biol. 2004, 268, 185–194. [CrossRef]
- Häärä, O.; Harjunmaa, E.; Lindfors, P.H.; Huh, S.-H.; Fliniaux, I.; Åberg, T.; Jernvall, J.; Ornitz, D.M.; Mikkola, M.L.; Thesleff, I. Ectodysplasin regulates activator–inhibitor balance in murine tooth development through Fgf20 signaling. Development 2012, 139, 3189–3199. [CrossRef]
- Mitsiadis, T.A.; Graf, D.; Luder, H.; Gridley, T.; Bluteau, G. BMPs and FGFs target Notch signalling via jagged 2 to regulate tooth morphogenesis and cytodifferentiation. Development 2010, 137, 3025–3035. [CrossRef]
- Mitsiadis, T.A.; Regaudiat, L.; Gridley, T. Role of the Notch signalling pathway in tooth morphogenesis. Arch. Oral Biol. 2005, 50, 137–140. [CrossRef]
- Kwon, H.-J.E.; Li, L.; Jung, H.-S. Hippo pathway/Yap regulates primary enamel knot and dental cusp patterning in tooth morphogenesis. Cell Tissue Res. 2015, 362, 447–451. [CrossRef]
- Li, C.-Y.; Hu, J.; Lu, H.; Lan, J.; Du, W.; Galicia, N.; Klein, O.D. αE-catenin inhibits YAP/TAZ activity to regulate signalling centre formation during tooth development. Nat. Commun. 2016, 7, 12133. [CrossRef]
- Ferguson, C.A.; Tucker, A.S.; Christensen, L.; Lau, A.L.; Matzuk, M.M.; Sharpe, P.T. Activin is an essential early mesenchymal signal in tooth development that is required for patterning of the murine dentition. Genes Dev. 1998, 12, 2636–2649. [CrossRef]
- Bloch-Zupan, A.; Mark, M.P.; Weber, B.; Ruch, J.V. In vitro effects of retinoic acid on mouse incisor development. Arch. Oral Biol. 1994, 39, 891–900. [CrossRef]
| Transcription factor | Primary expression (tissue/stage) | Key regulatory role | Mouse genetic phenotype | Human genetic/clinical association | Key refs | Evidence strength |
|---|---|---|---|---|---|---|
| PAX9 | Dental mesenchyme; initiation–bud | Establishes mesenchymal odontogenic competence and supports progression beyond the bud stage | Pax9−/−: tooth development arrested at bud stage; tooth agenesis | PAX9 variants: common cause of nonsyndromic tooth agenesis (often molars); genotype–phenotype correlations reported | [33] | Mouse KO + human LOF variants |
| MSX1 | Dental mesenchyme; bud–cap | Cooperates with PAX9; acts downstream of BMP signaling to maintain odontogenic potential and patterning | Msx1−/−: arrested odontogenesis and cleft palate | MSX1 variants associated with tooth agenesis (sometimes with clefting) | [23,34] | Mouse KO + human LOF variants |
| PITX2 | Dental epithelium; placode–bud | Earliest marker of odontogenic epithelium; regulates epithelial proliferation and epithelial–mesenchymal signaling | Pitx2 loss disrupts tooth morphogenesis (and other developmental processes) | PITX2 mutations: Axenfeld–Rieger syndrome with dental anomalies (agenesis, microdontia) | [21] | Mouse models + syndromic human mutations |
| LEF1 | Dental epithelium/enamel knot; bud–cap | Canonical Wnt effector; activates enamel-knot programs (e.g., FGF4) and links Wnt output to Shh regulation | Lef1−/−: bud-stage arrest | Direct LEF1 dental genotype–phenotype evidence remains limited; pathway-level human genetics supports canonical Wnt requirement | [28] | Mouse KO; limited direct human genetics |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




