4. Discussion
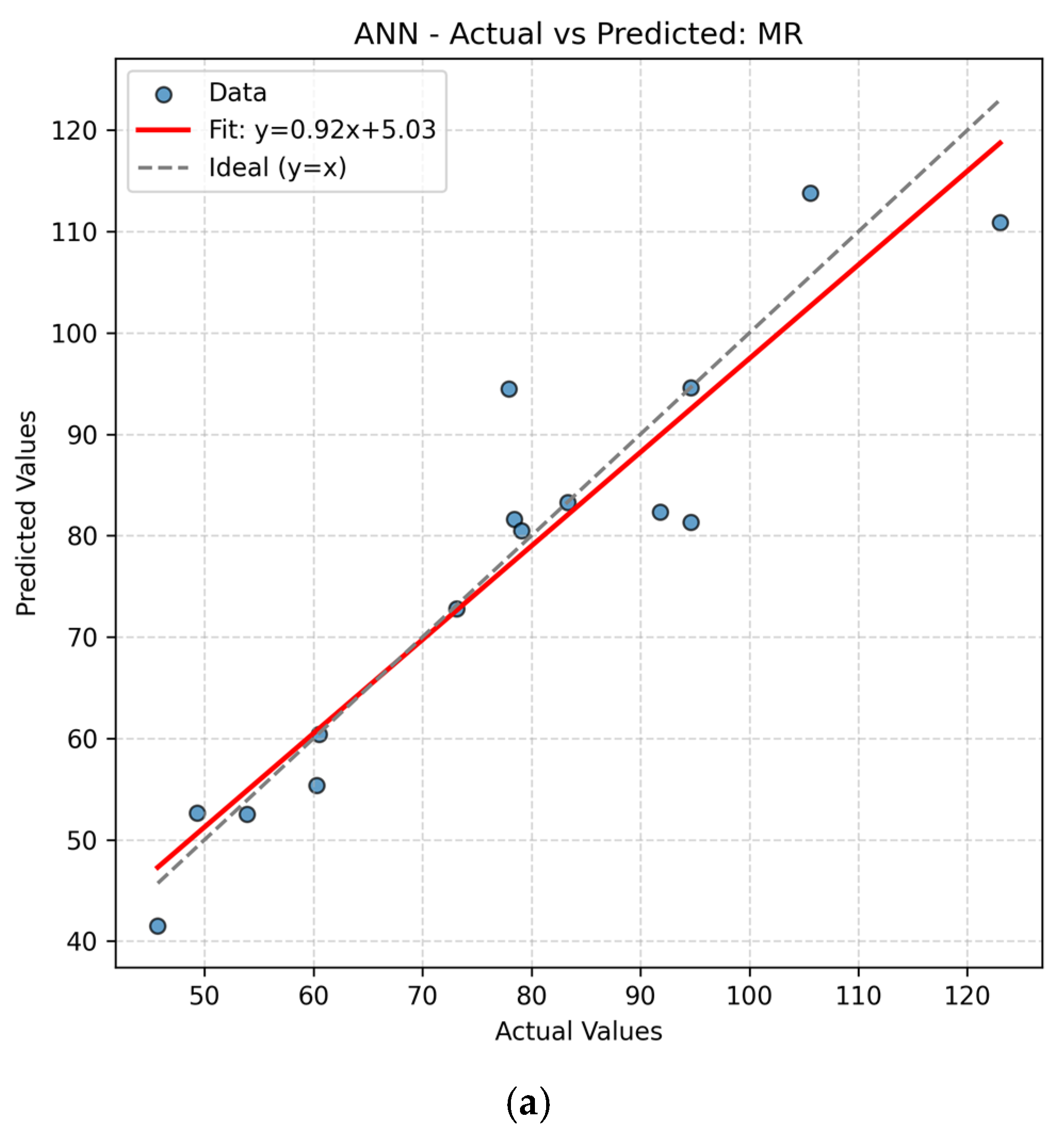
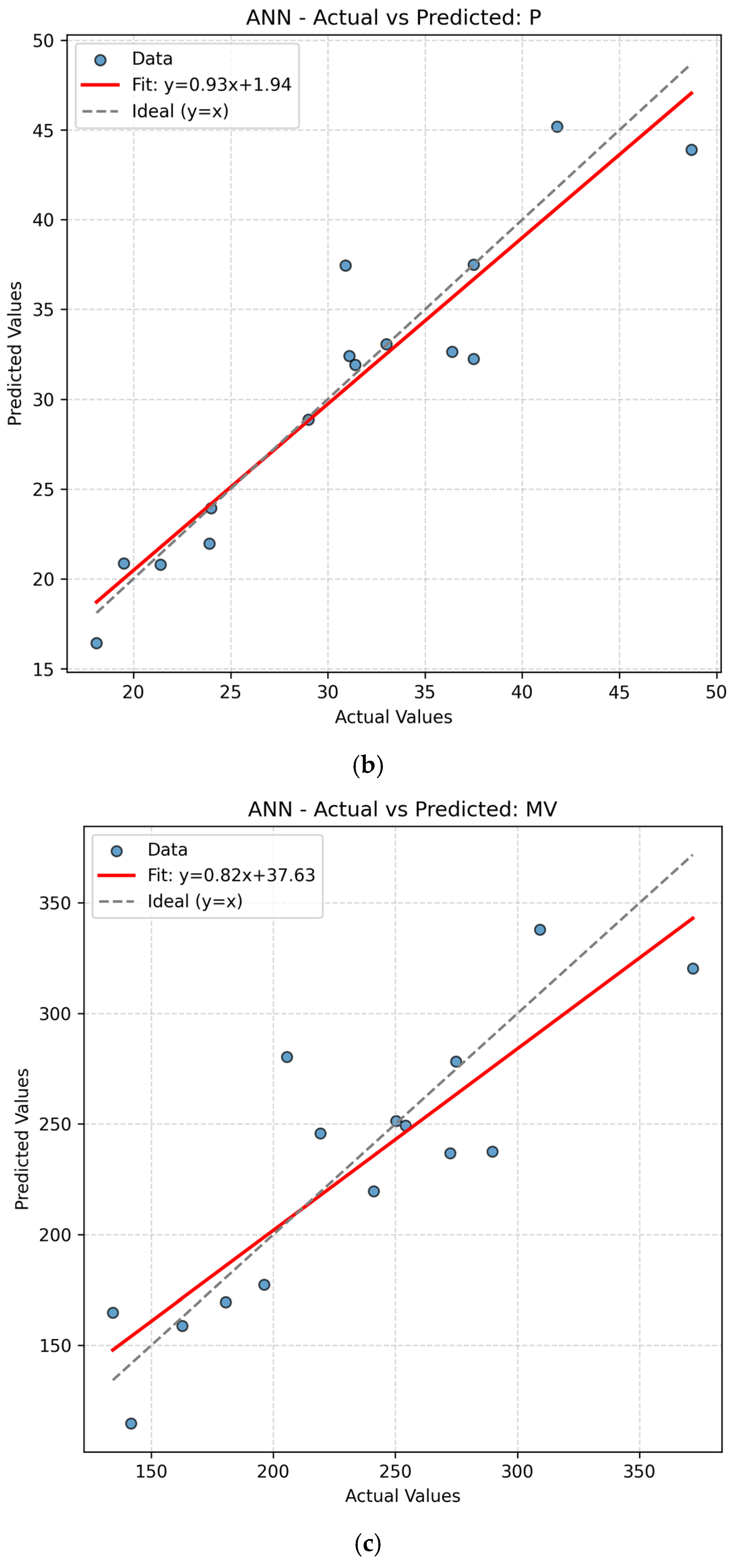
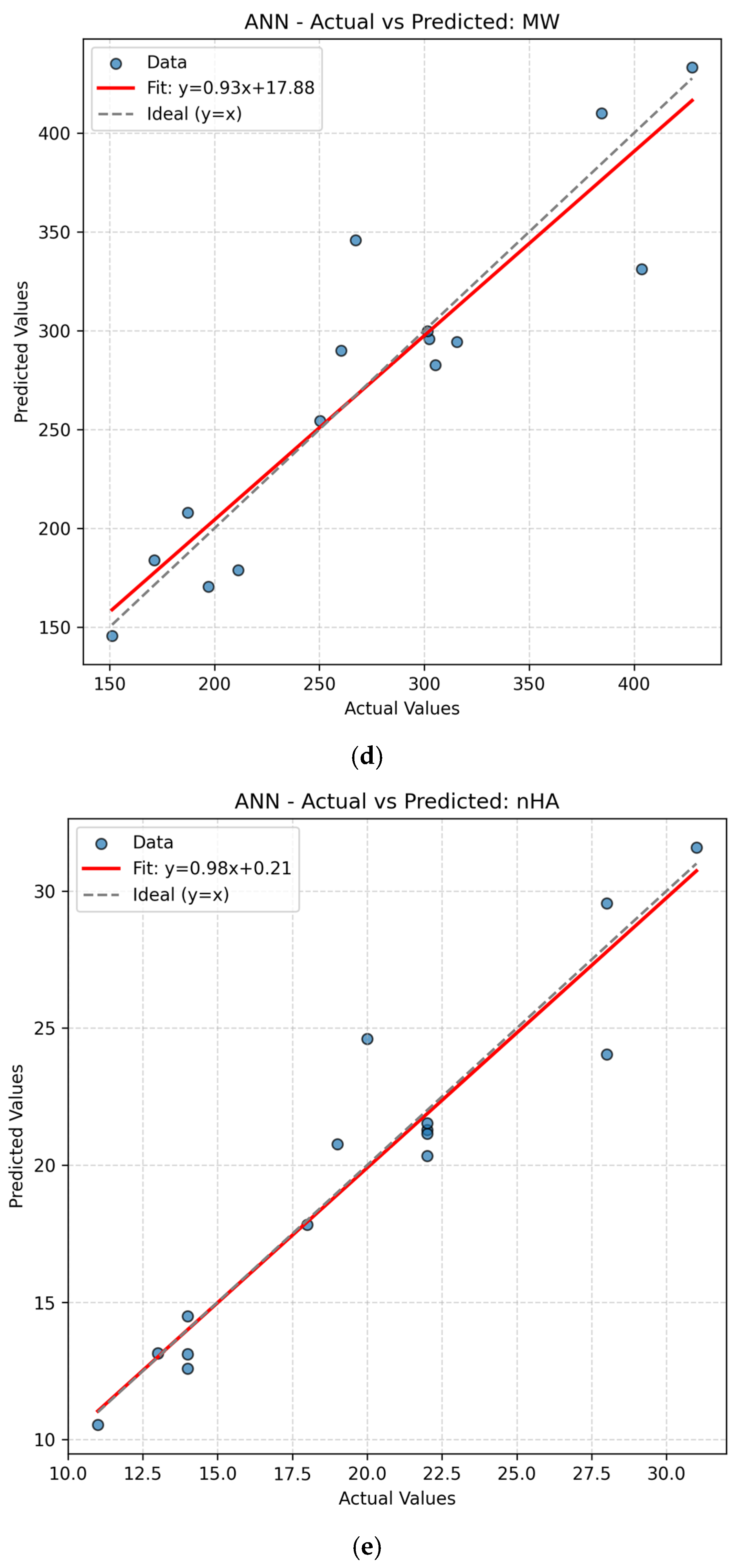
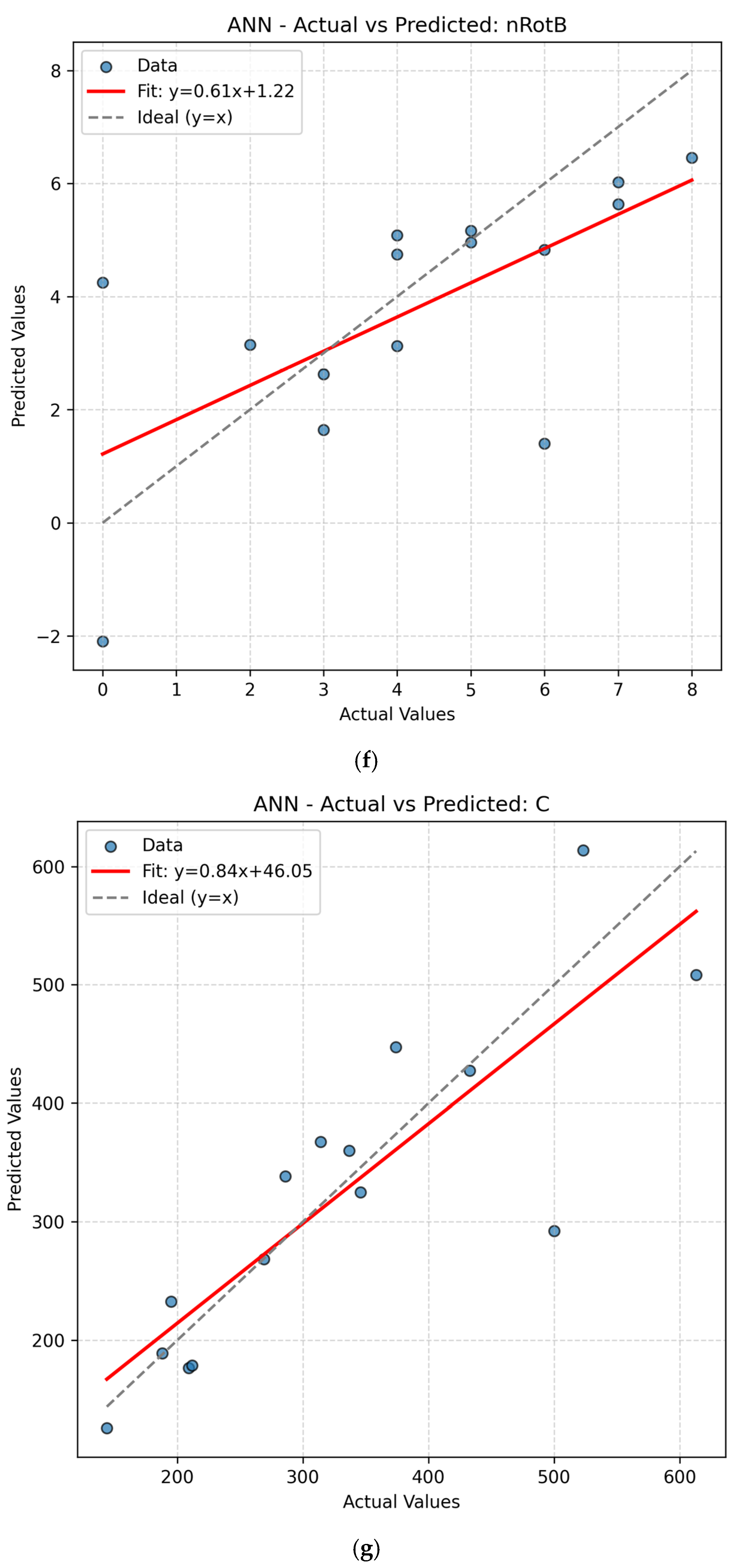
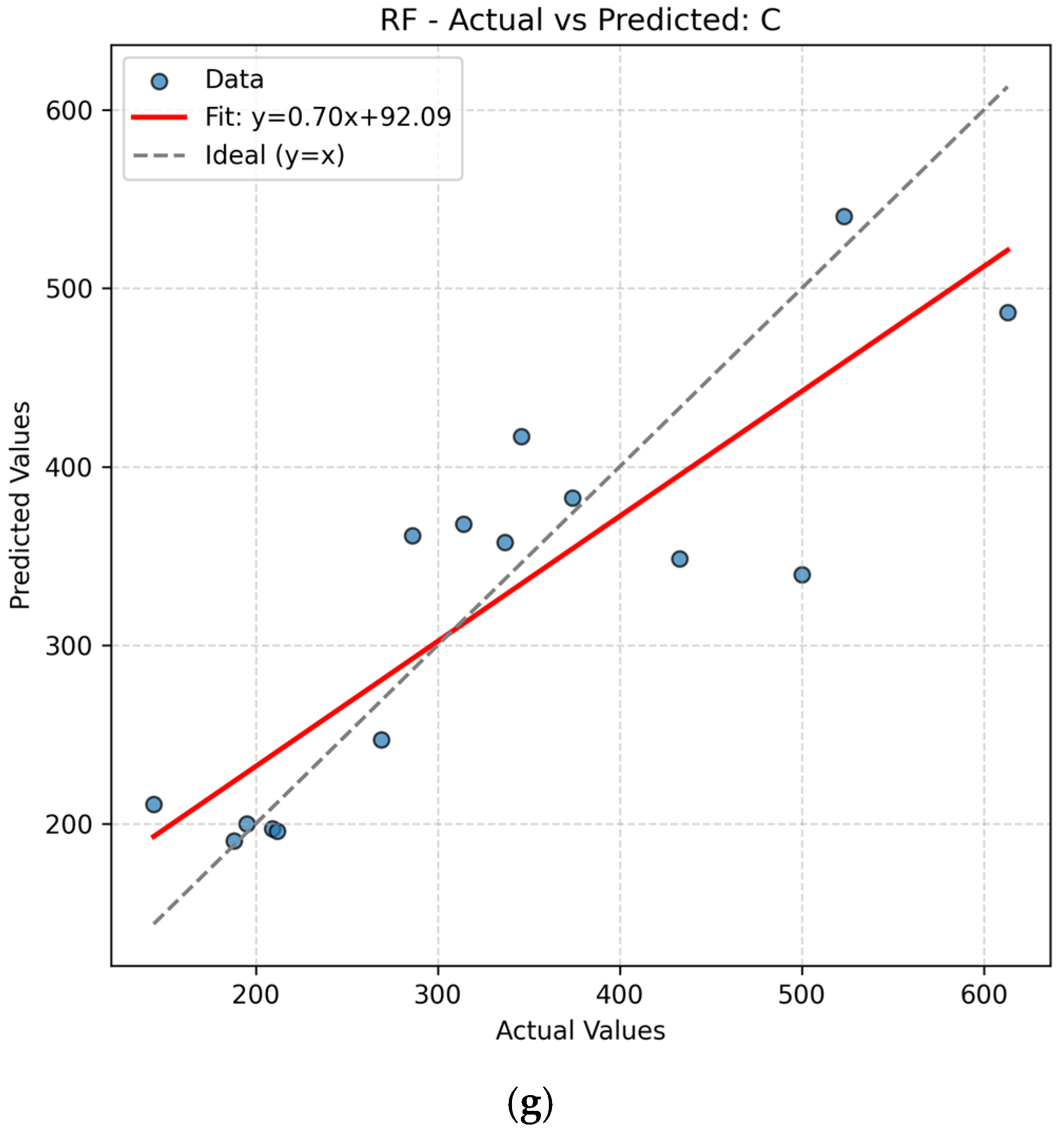
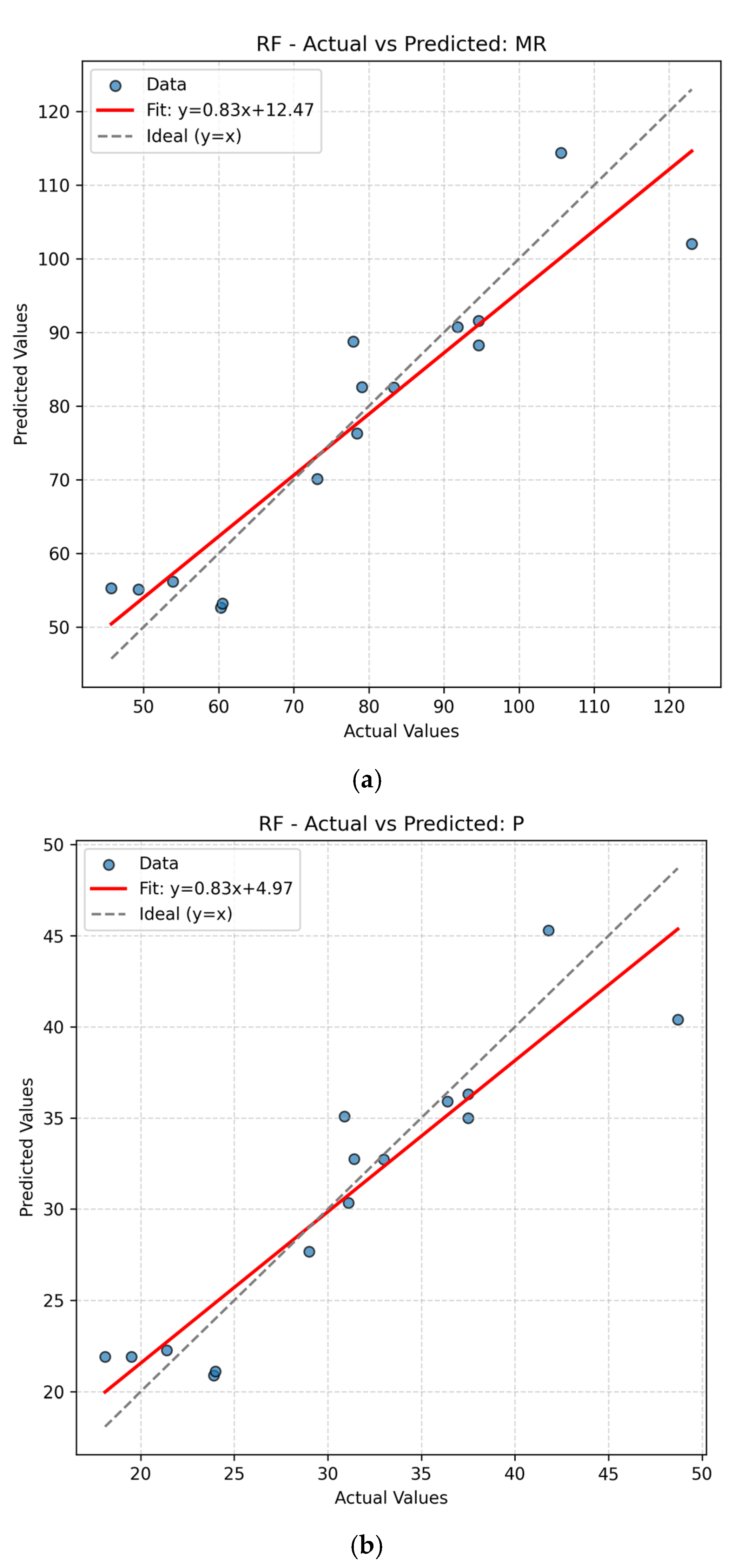
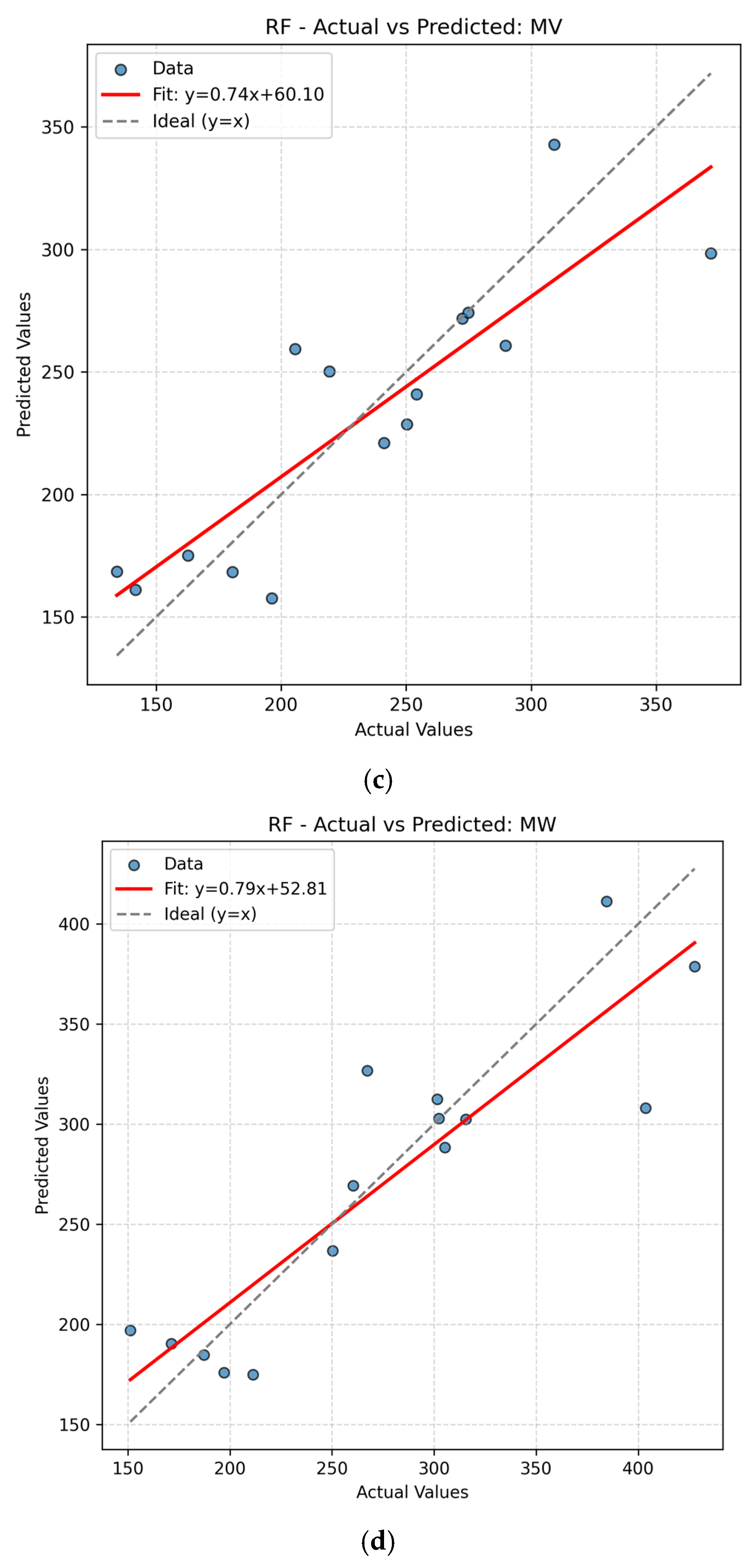
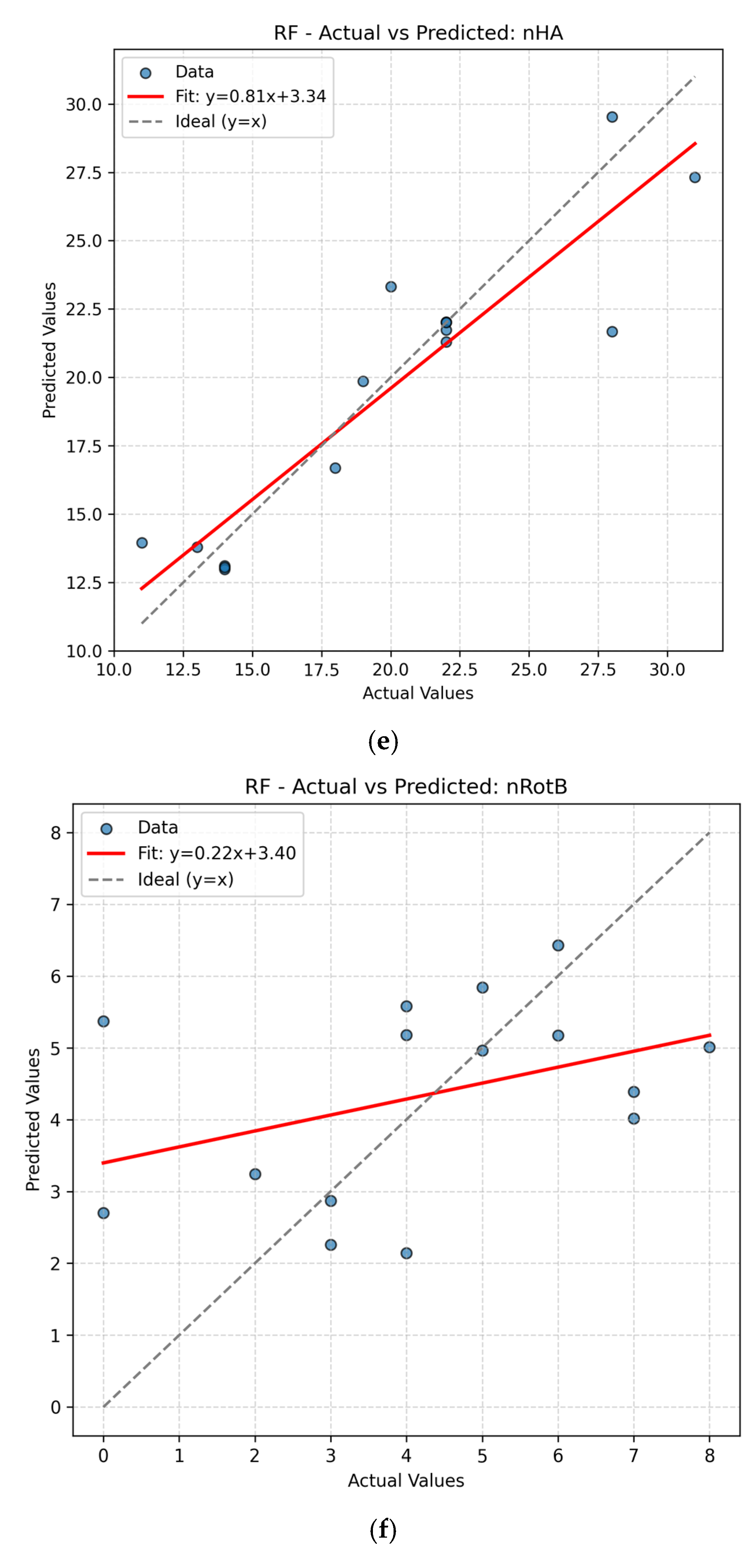
The present work had two linked objectives to assess the supervised machine learning models by predictive accuracy, computational efficiency, and interpretability and show the integration between QSPR descriptors and supervised learning can enhance the drug discovery for Parkinson’s disease (PD) by decreasing the experimental screening. Based on the LOOCV results, ANN and Random Forest (RF) model are able to predict several important properties of PD drugs successfully. MR, Polarizability (P), Molecular Weight (MW) and Heavy Atom Count (nHA) were well predicted with high accuracy, with R2 values always higher than 0.84 for both models, while discrete or structurally complex features [i.e., Rotatable Bond Count (nRotB) and Molecular Complexity (C)] were less well predicted, especially in ANN models (R2 around 0.31 for nRotB). These results represent different abilities of models to represent continuous and discrete or highly variable molecular features. The reported near-perfect R2 values for a large number of endpoints by the ANN reflect outstanding predictive accuracy on the available data set. Both models had a difficult time predicting the correct amount of rotatable bonds. This would imply that the chosen descriptor set does not strongly correlate with the nRotB property and further descriptor engineering or different model architectures might be required for this particular target.
Due to the small dataset size, model performance was assessed using Leave-One-Out Cross-Validation (LOOCV) so as to make a realistic prediction of model performance on unseen compounds. LOOCV enables individual testing of each compound while using the remaining data for training and is ideal for testing small datasets. This strategy guarantees that the reported impact of performance measures such as MSE, RMSE, MAE, and R2, are reported on unseen data, even though the number of data points available constrains the use of more powerful methods such as nested cross-validation, or even external validation.
However, such results need to be put into context of dataset size, dimensionality of the descriptor and the validation strategy. High R2 plus small sample of compounds plus high dimensional topological inputs creates a realistic risk of overfitting, for robust model selection, nested CV plus external holdout sets, y-randomization tests, and reporting out of sample RMSE/MAE rather than in-sample fit alone 8,10 To address this critical concern, Y-randomization test was undertaken with 50 permutations for all models and properties. The resulting average R2rand values for all the randomized models were consistently and strongly negative, representing a value range of -1.708-0.457. This is conclusive evidence that the predictive relationships that were created by the original models are statistically sound and not the result of chance correlation. The test thus establishes the good stability of both these models (ANN and RF) against spurious fitting, which statistically validates the utility of the models for physicochemical properties for which R2 > 0.8.
Comparative evaluation between classes of algorithms reveals some obvious trade-offs. Deep ANNs usually perform well on modeling complex, nonlinear structure-property relationships and, thus, for many tasks in cheminformatics, are often better than linear regressions and simpler tree models, provided the richness of the data.[
11,
14] In this study, ANN slightly outperformed RF for MR, P, and nHA which reflects the ability of ANN to capture nonlinear relationship and RF for MW, MV, and C provided more stable predictions with less extreme errors. Ensemble tree approaches are often on par or exceed ANNs on tabular QSPR problems with significantly reduced tuning cost and improved out-of-the-box stability.[
27,
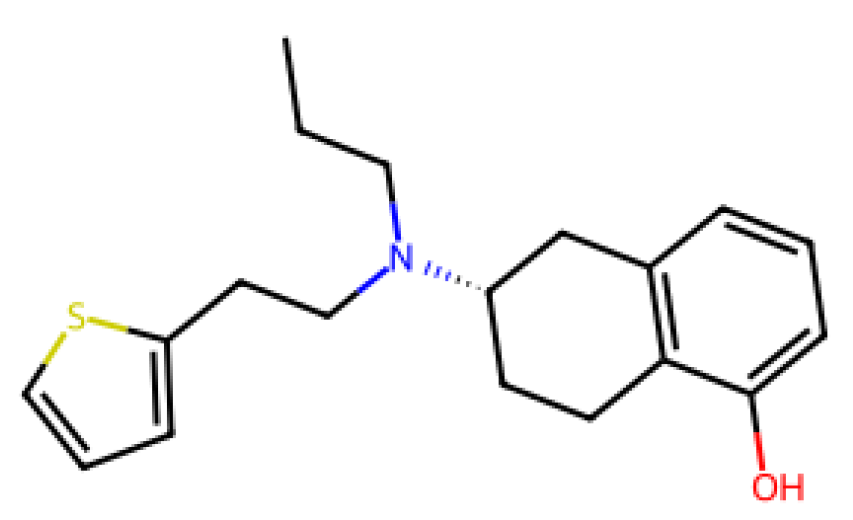
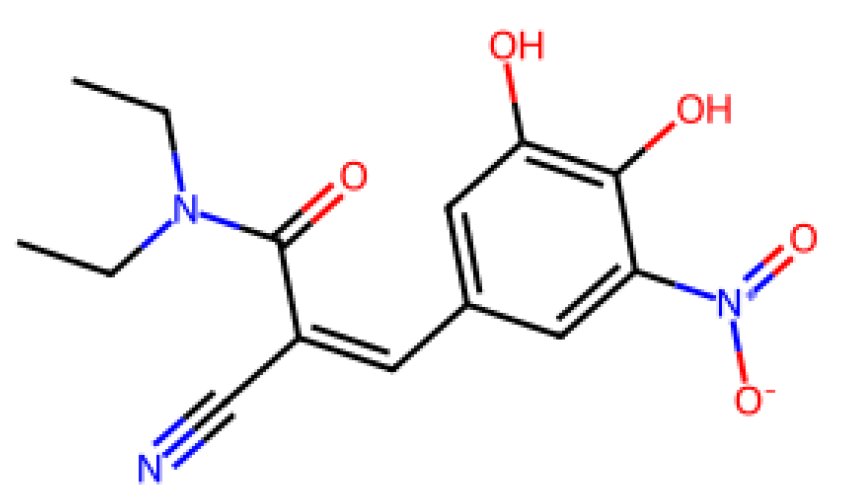
28] The results of the LOOCV analysis showed that there were compounds with larger deviations, such as Pimavanserin and Istradefylline, which could indicate that when small datasets containing compounds with structural diversity are used, the generalization of models can be challenged.
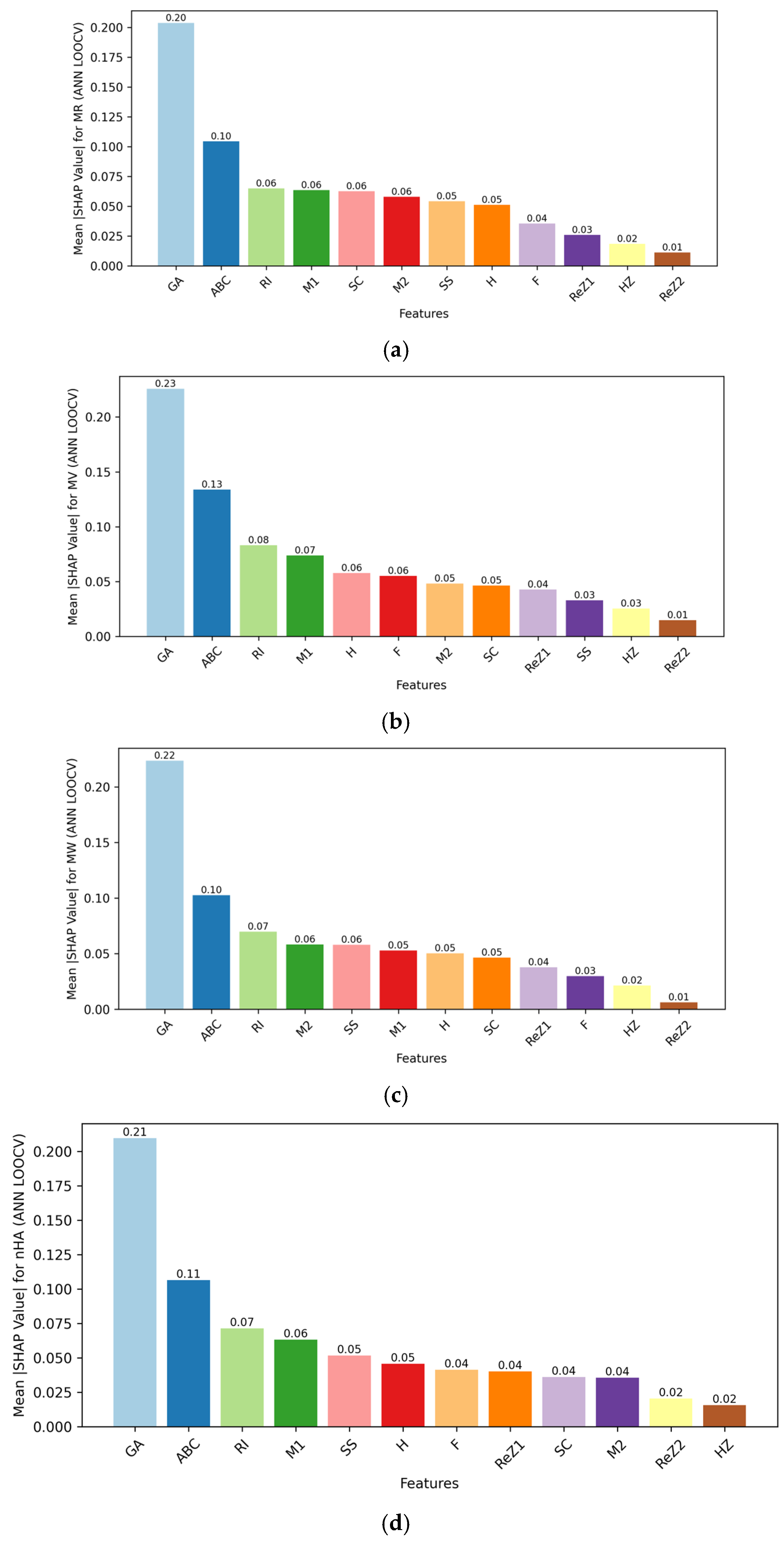
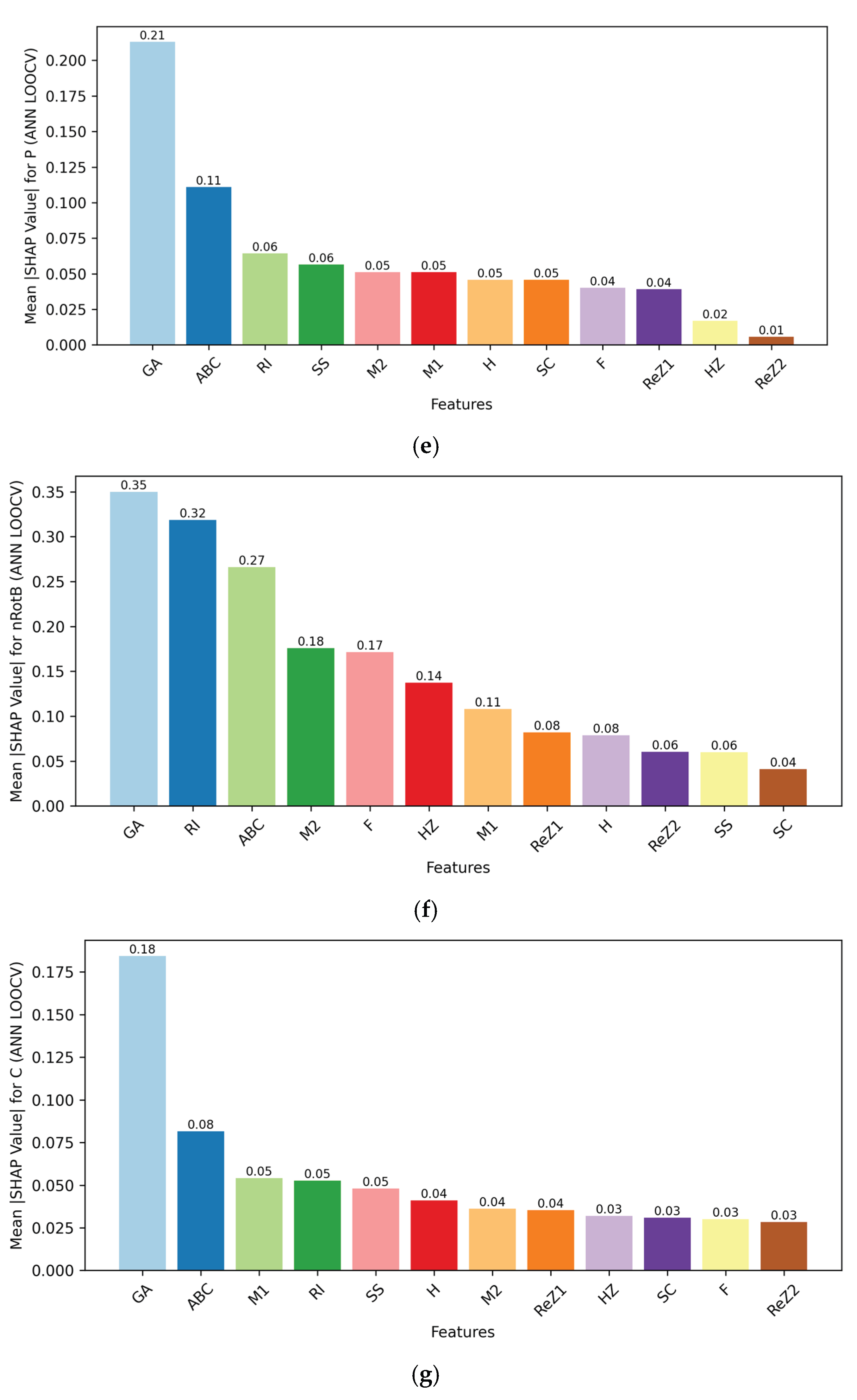
The other decisive axis is interpretability. Interpretability was addressed using Shapley Additive Explanations (SHAP), which identified the most influential topological descriptors for each property, which provides mechanistic insight into model predictions.[
29] This way, it was possible to assess which molecular features were mostly responsible for predicted MR, P, MW, and nHA, increasing transparency even when the ANN is a nonlinear “black box” model. RF models naturally support feature importance measures, which were complementary in a way that gave access to interpretability. The ability to associate certain descriptors with predicted properties is of special interest in medicinal chemistry decision-making and in the early-stage PD drug prioritization.
Integrating QSPR descriptors with supervised learning gives three practical speed advantages to discovery. First, it allows high-throughput in silico prioritization of compounds to balance the BBB permeation, lipophilicity, and pharmacokinetic constraints, thus reducing the pool of candidates subject to experimental assays.[
30,
31] After which, iterative cycles of model-guided selection and targeted synthesis which are augmented by active learning can maximize information per experiment and reduce total wet-lab burden.[
1,
2] This is useful for the combination of QSPR outputs with target-specific predictions (e.g., docking scores or binding affinities for dopamine receptors or alpha-synuclein aggregations) supports multi- optimization and prioritization for PD-relevant mechanisms.[
13] While iterative model-guided selection or active learning was not applied in this study, the current LOOCV results provide a framework to use computational prioritization in PD drug discovery. The findings highlight that descriptor-based supervised learning can be used to speed up the early-stage evaluation, however, caution needs to be taken as a result of small datasets size, high-dimensionality inputs, and endpoints with low-predictability, which might limit the generalization of the model.[
8,
10]
Operational recommendations follow directly from these observations where benchmark models with nested cross-validation, quantification of uncertainty (prediction intervals, ensembles), and definition of an applicability domain are useful before deploying predictions for experimental selection.[
10] To enlarge descriptor space to 3D and quantum chemical features; consider graph neural networks or message passing architectures in case of large datasets, as they learn representations directly from molecular graphs.[
13] This will help in explicitly including PD-specific endpoints and multiobjective criteria so that computational prioritization shares the same therapeutic constraints as central nervous system drug development.