4. Discussion
The present study aimed to determine whether modulation of key signaling pathways could improve the developmental competence and lineage specification of bovine haploid parthenogenetic embryos (bhPE) under serum-free conditions. Overall, our findings indicate that although specific small molecules can enhance certain developmental parameters, none of the treatments tested were sufficient to overcome the intrinsic limitations imposed by haploid genome constitution.
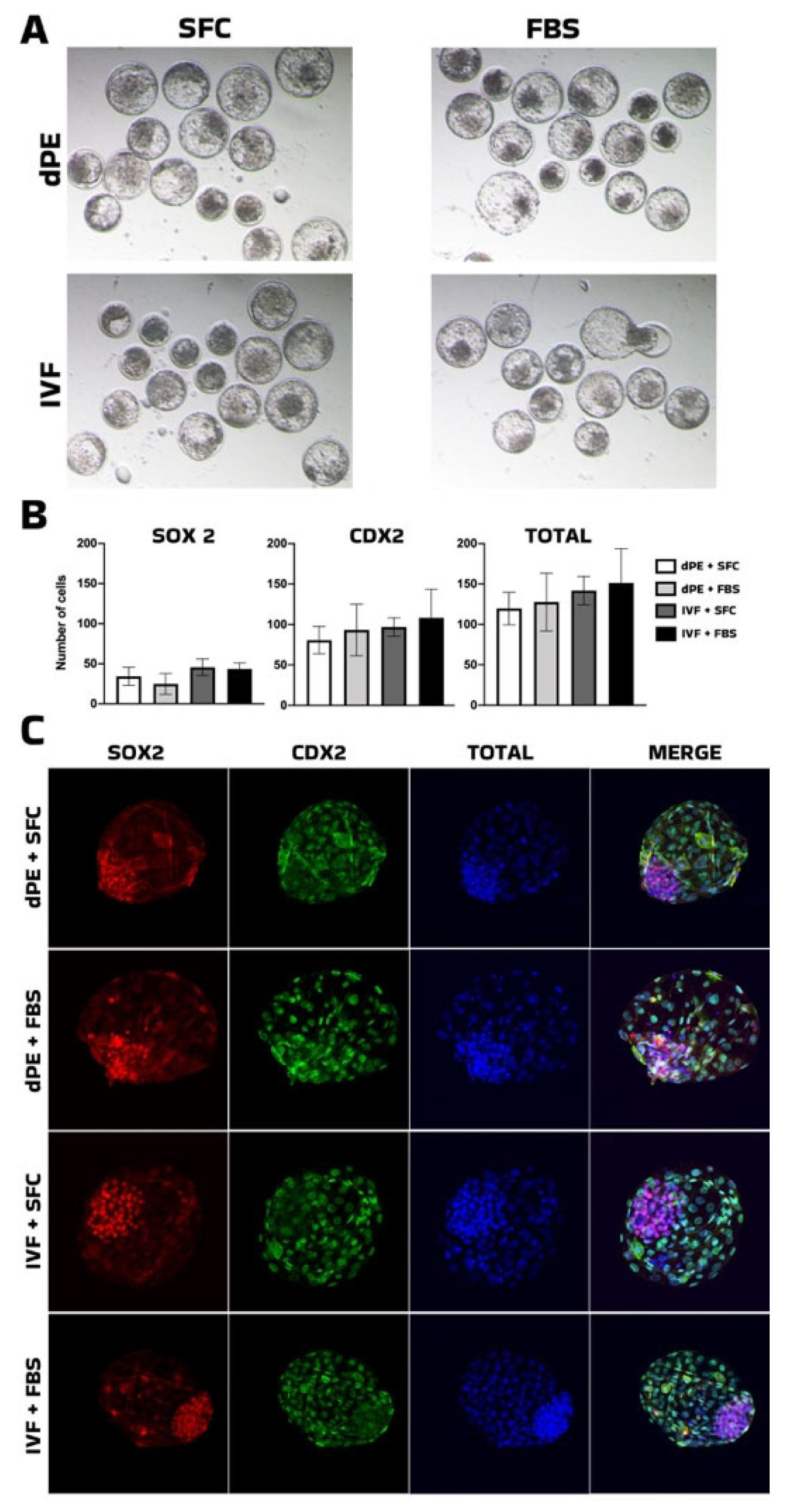
As an initial step, we evaluated the replacement of fetal calf serum (FCS) with insulin–transferrin–selenium (ITS) to establish a chemically defined culture system. The use of fetal bovine serum (FBS) in embryo culture is known to introduce batch-to-batch variability due to its undefined and complex composition. In contrast, ITS is a well-characterized supplement that has been widely used as a serum substitute in cell and embryo culture systems. ITS supports cell proliferation, metabolism, and survival while reducing reliance on undefined serum components [
23,
24].
Pioneering studies have demonstrated that defined culture media supplemented with ITS can support in vitro development of bovine embryos produced by somatic cell nuclear transfer [
25] and IVF [
26]. Moreover, ITS supplementation has been successfully applied in various mammalian embryonic models [
27,
28,
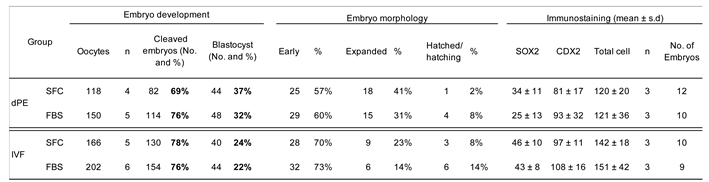
29], supporting its suitability as a serum-free alternative. In the present study, ITS supplementation supported normal developmental progression and pluripotency-associated marker expression in both IVF-derived and diploid parthenogenetic embryos. These findings indicate that ITS provides adequate trophic support in the absence of serum, without introducing detectable bias in developmental outcomes or lineage specification. Importantly, the establishment of this defined culture system enabled subsequent evaluation of signaling pathway modulators under controlled conditions, minimizing potential confounding effects associated with serum-derived factors.
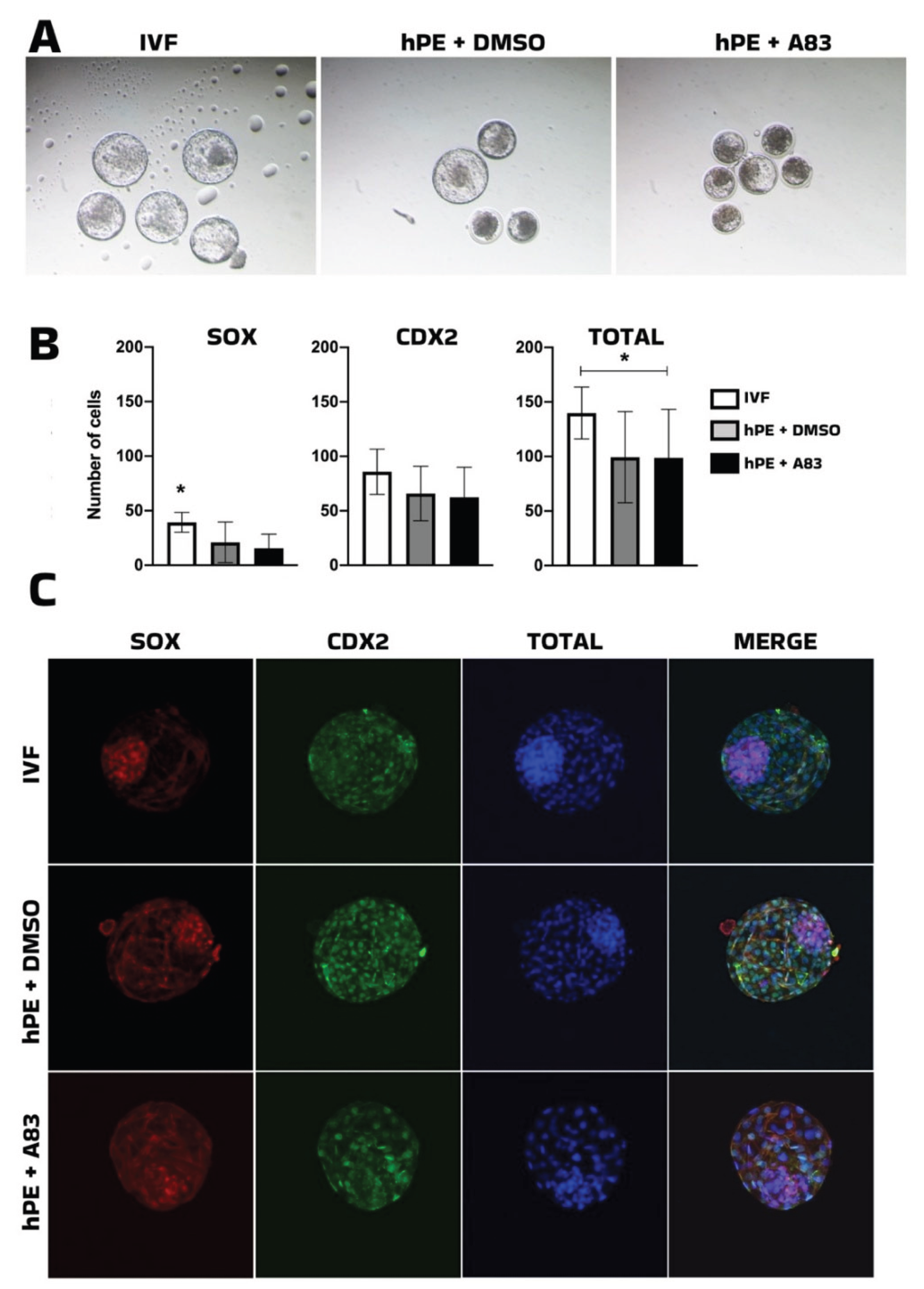
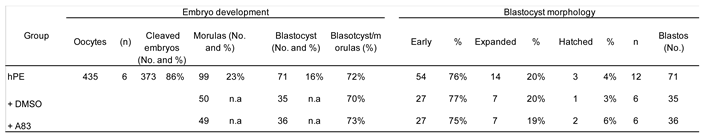
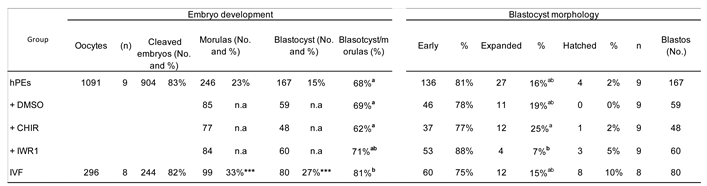
The initial analysis of small molecules to identify factors capable to boost either competence or pluripotency of parthenogenetic haploid embryos showed that inhibition of SMAD-dependent TGF-β signaling using A83-01 from the morula stage onward did not improve blastocyst formation or morphology in bhPE. Although TGF-β pathway inhibition has been associated with enhanced pluripotency maintenance in stem cell systems [
21,
30] , its modulation at post-compaction stages was insufficient to enhance lineage allocation in haploid embryos.
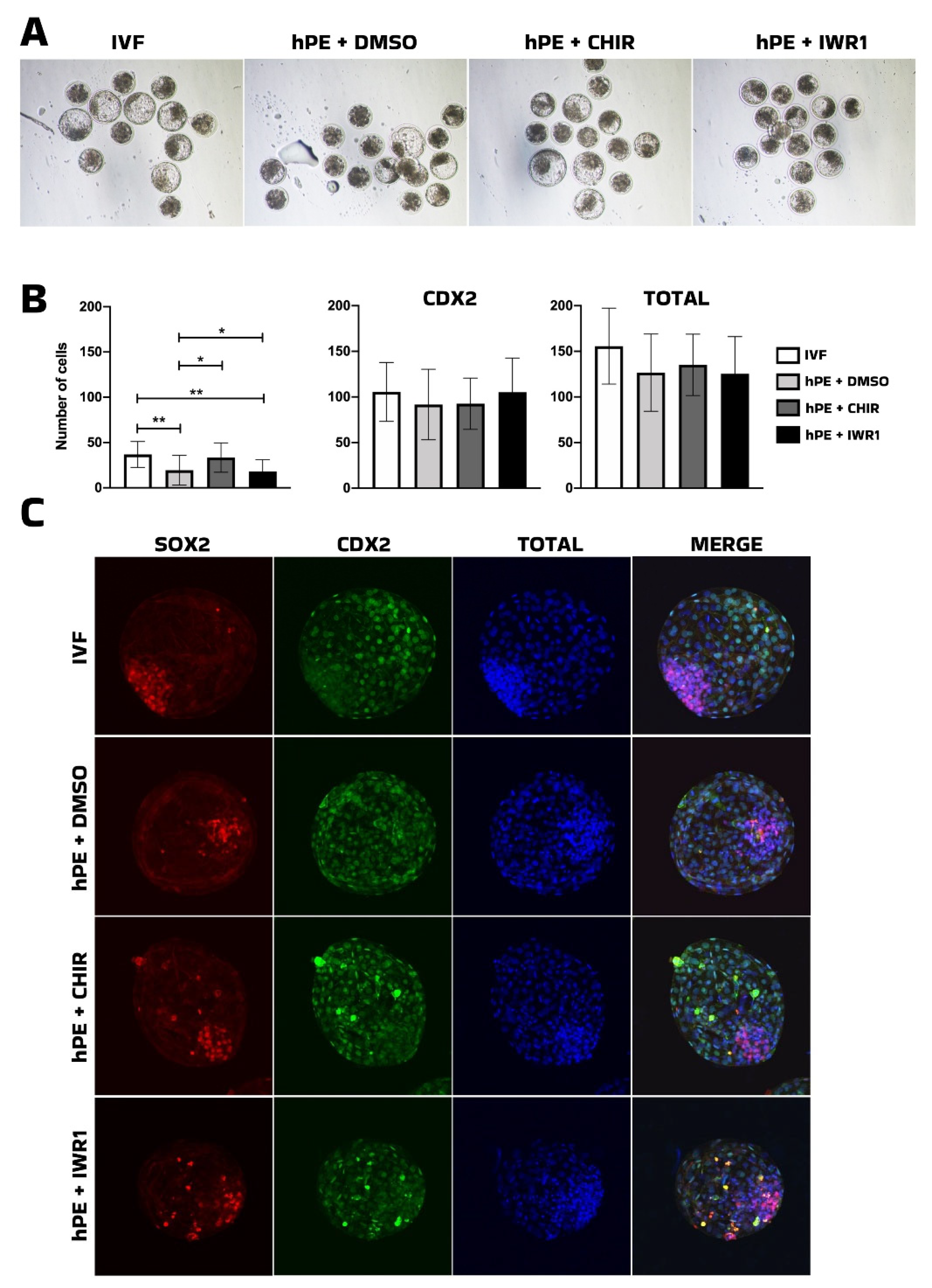
Next, modulation of the WNT pathway revealed a more nuanced response. CHIR99021, a GSK3β inhibitor and WNT agonist, increased the number of SOX2-positive cells in bhPE to levels comparable to IVF embryos, indicating a beneficial effect on pluripotency-associated lineage allocation in this model. Early studies found that the inhibition of GSK3 in conjunction with MEK from zygote stage onward offered enhanced blastocyst development and expression of epiblast NANOG and SOX2 of bovine IVF embryos [
31,
32]. Similarly, inhibition of GSK3 and MAP2K improved blastocyst morphology and expression of pluripotency associated genes (FGF4 and NANOG) [
33]. In contrast, IWR-1, a WNT pathway antagonist, did not produce similar effects. Although the proportion of morulae progressing to the blastocyst stage was comparable to that of the IVF group, no improvements were observed in total cell number or lineage allocation. Tankyrase inhibitors, including IWR-1, have been reported as essential components for the derivation and maintenance of livestock embryonic stem cells (ESCs) [
34]. However, in a subsequent study, the same group demonstrated that bovine ESCs exposed to IWR-1 exhibited reduced protein abundance of key HIPPO pathway components, including TEAD4 and YAP1, along with decreased expression of canonical YAP/TEAD target genes such as CYR61. These findings suggest that while IWR-1 may support ESC stabilization under specific culture conditions, its modulation of WNT and HIPPO signaling does not necessarily enhance lineage allocation in haploid embryos at the preimplantation stage.
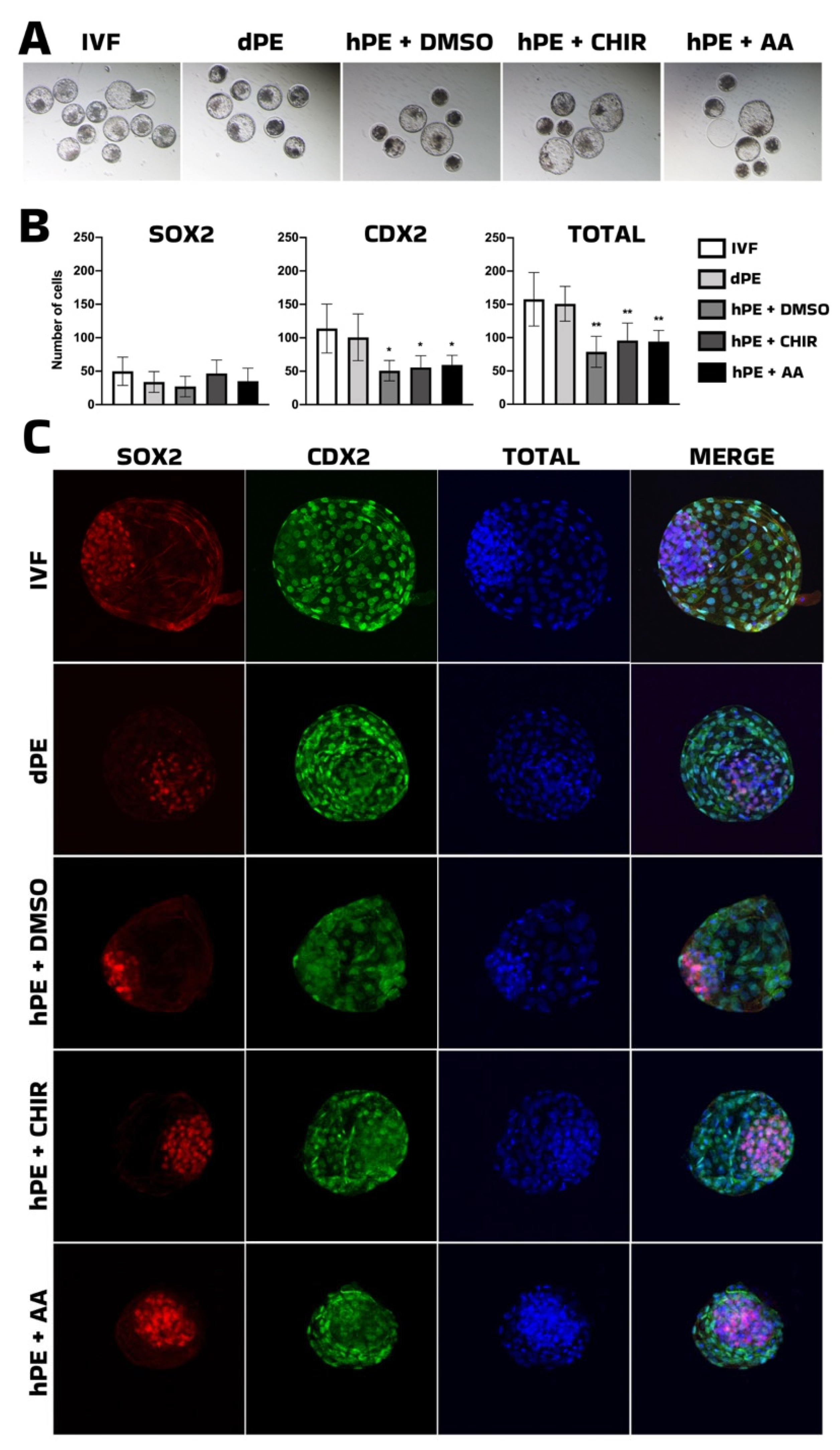
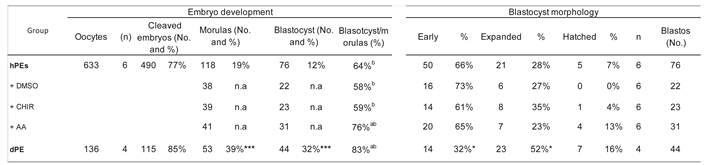
We next evaluated the effects of Activin A supplementation in comparison with CHIR99021. The developmental progression of CHIR99021-treated haploid embryos remained inferior to diploid parthenogenetic control. Thus, while WNT activation may enhance aspects of inner cell mass specification, it does not restore global developmental competence. On the other hand, activin A (AA), a member of the TGF-β superfamily, has previously been described as an embryokine that enhances bovine embryo development to the blastocyst stage [
22]. In the present study, AA supplementation significantly improved blastocyst formation and hatching rates in haploid parthenogenetic embryos, demonstrating that its beneficial effects extend to uniparental embryos. Interestingly, despite the clear improvement in developmental progression, AA supplementation did not alter the number of SOX2-positive cells in haploid embryos compared with control groups. This suggests that AA does not directly enhance SOX2-associated lineage specification within the inner cell mass (ICM), but rather promotes developmental competence through alternative mechanisms, potentially related to improved cell survival or developmental kinetics. In contrast, both the number of CDX2-positive cells and the total cell number remained higher in diploid parthenogenetic and IVF embryos compared with haploid embryos. These findings indicate that although AA improves blastocyst yield in haploid parthenotes, it does not restore proliferative capacity or lineage allocation to diploid levels, which agrees a previous report indicating that AA improved blastocyst development of IVF embryos after supplementation at morula stage [
35]. The reduced total cell number observed in haploid parthenotes likely reflects intrinsic gene dosage limitations and epigenetic imbalances associated with uniparental genome constitution [
10,
11,
12]. Taken together, these results suggest that Activin A enhances developmental efficiency without fundamentally correcting the intrinsic cellular and genomic constraints of haploid parthenogenetic embryos.
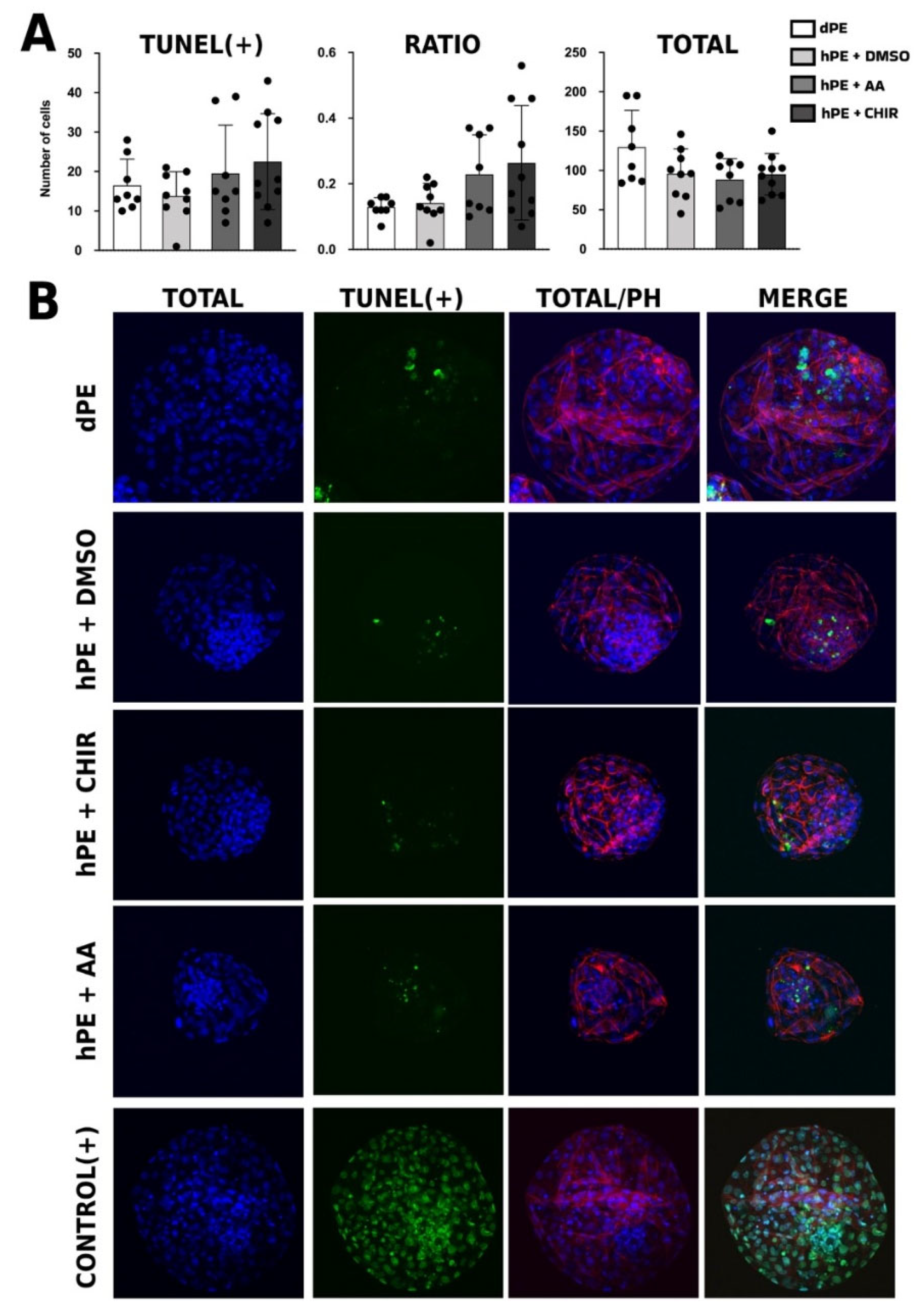
The analysis of DNA fragmentation revealed that the number of TUNEL-positive cells tended to be higher in both AA- and CHIR99021-treated haploid embryos compared with vehicle and diploid controls, suggesting that modulation of these signaling pathways does not mitigate the intrinsic susceptibility of haploid embryos to apoptotic stress. Rather, this trend may reflect persistent genomic instability and altered cell cycle regulation associated with haploidy, which cannot be fully compensated by activation of TGF-β or WNT signaling. Similar findings were reported by Madeja et al., [
36], who observed increased apoptosis in 2i-treated blastocysts without concomitant changes in the transcript abundance of canonical apoptosis-related genes (BAX, BCL2, BAK, and the BAX/BCL2 ratio). Similarly, Trigal et al., [
35] indicated that when added AA at shorter periods activin increased apoptotic rates. These results suggest that elevated DNA fragmentation may occur independently of transcriptional regulation of classical apoptotic markers, potentially involving post-transcriptional mechanisms, mitochondrial dysfunction, replication stress, or alternative cell death pathways. Therefore, the possibility that non-canonical or transcriptionally uncoupled apoptotic pathways contribute to the increased DNA fragmentation observed in treated haploid embryos cannot be excluded.
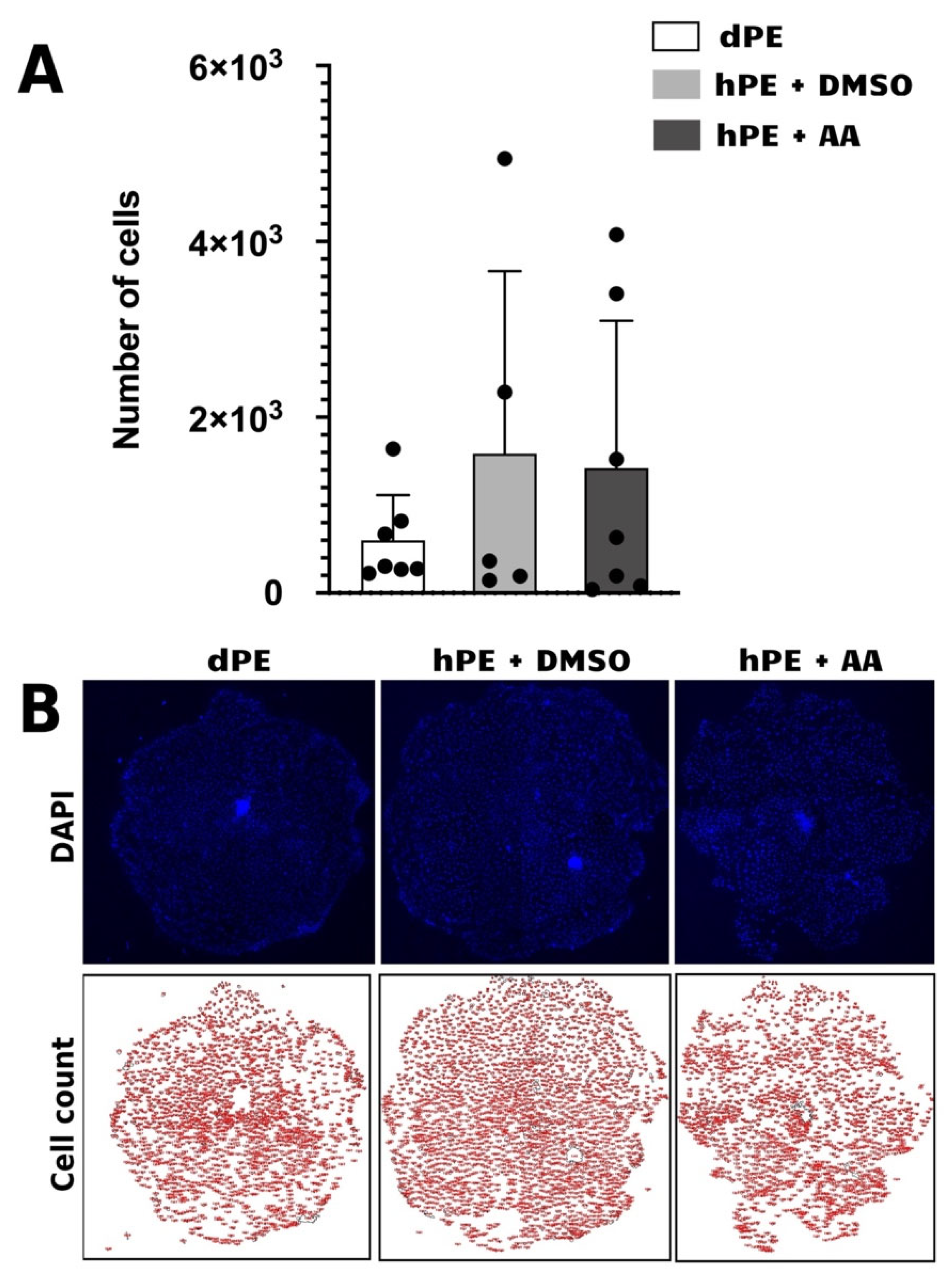
Although Activin A (AA) improved blastocyst formation and hatching rates in haploid parthenogenetic embryos, this benefit did not translate into enhanced outgrowth derivation, indicating that AA treatment during blastocyst formation, does not confer sustained advantages beyond the blastocyst stage. The high variability and absence of improved outgrowth formation further supports the notion that intrinsic limitations associated with haploid genome constitution, including embryonic quality, gene dosage imbalance and epigenetic dysregulation, remain uncorrected despite modulation of TGF-β signaling [
9,
13,
37].