Submitted:
05 March 2026
Posted:
10 March 2026
You are already at the latest version
Abstract
Keywords:
Introduction
Methods
Review Design and Question
Eligibility Criteria
| ID | Database | Date | Query | Filters | Results | Exported | Selected |
| I1 | PubMed (humans) | 21/11/2025 | ( “epigenetic clock”[tiab] OR “epigenetic clocks”[tiab] OR “DNA methylation age”[tiab] OR “DNAm age”[tiab] OR “epigenetic age”[tiab] OR “methylation clock”[tiab] OR “methylation clocks”[tiab] ) AND ( trial[tiab] OR randomized[tiab] OR randomised[tiab] OR intervention[tiab] OR interventional[tiab] OR “pre-post”[tiab] OR “before and after”[tiab] ) AND (“2010/01/01”[PDAT] : “2025/12/31”[PDAT]) AND (english[la] OR spanish[la]) AND humans[mh] | 2010–2025; EN/ES; Humans; Article | 145 | 145 | See PRISMA |
| I2 | PubMed (animals) | 22/11/2025 | (“epigenetic clock”[tiab] OR “epigenetic clocks”[tiab] OR”DNA methylation age”[tiab] OR “DNAm age”[tiab] OR”epigenetic age”[tiab] OR “methylation clock”[tiab] OR”methylation clocks”[tiab])AND(trial[tiab] OR randomized[tiab] OR randomised[tiab] ORintervention[tiab] OR treatment[tiab] OR”pre-post”[tiab] OR “before and after”[tiab])AND(Mice[MeSH Terms] OR mouse[tiab] OR murine[tiab] ORrat[tiab] OR rats[tiab] OR mammal*[tiab])AND(“2010/01/01”[PDAT] : “2025/12/31”[PDAT])AND(english[la] OR spanish[la])NOT humans[mh] | 2010–2025; EN/ES; Article; NOT humans | 5 | 5 | See PRISMA |
| I3 | Scopus | 21/11/2025 | TITLE-ABS-KEY( (“epigenetic clock” OR “epigenetic clocks” OR “DNA methylation age” OR “DNAm age” OR “epigenetic age” OR “methylation clock” OR “methylation clocks”) AND (trial OR randomized OR randomised OR intervention OR interventional OR “pre-post” OR “before and after”) ) | 2010–2025; EN/ES; Article | 507 | 507 | See PRISMA |
| I4 | Cochrane CENTRAL | 21/11/2025 | ( “epigenetic clock”:ti,ab,kw OR “epigenetic clocks”:ti,ab,kw OR “DNA methylation age”:ti,ab,kw OR “DNAm age”:ti,ab,kw OR “epigenetic age”:ti,ab,kw OR “methylation clock”:ti,ab,kw OR “methylation clocks”:ti,ab,kw ) AND ( trial:ti,ab,kw OR randomized:ti,ab,kw OR randomised:ti,ab,kw OR intervention:ti,ab,kw OR “pre-post”:ti,ab,kw OR “before and after”:ti,ab,kw ) | 2010–2025; EN/ES | 5 | 5 | See PRISMA |
Study Selection
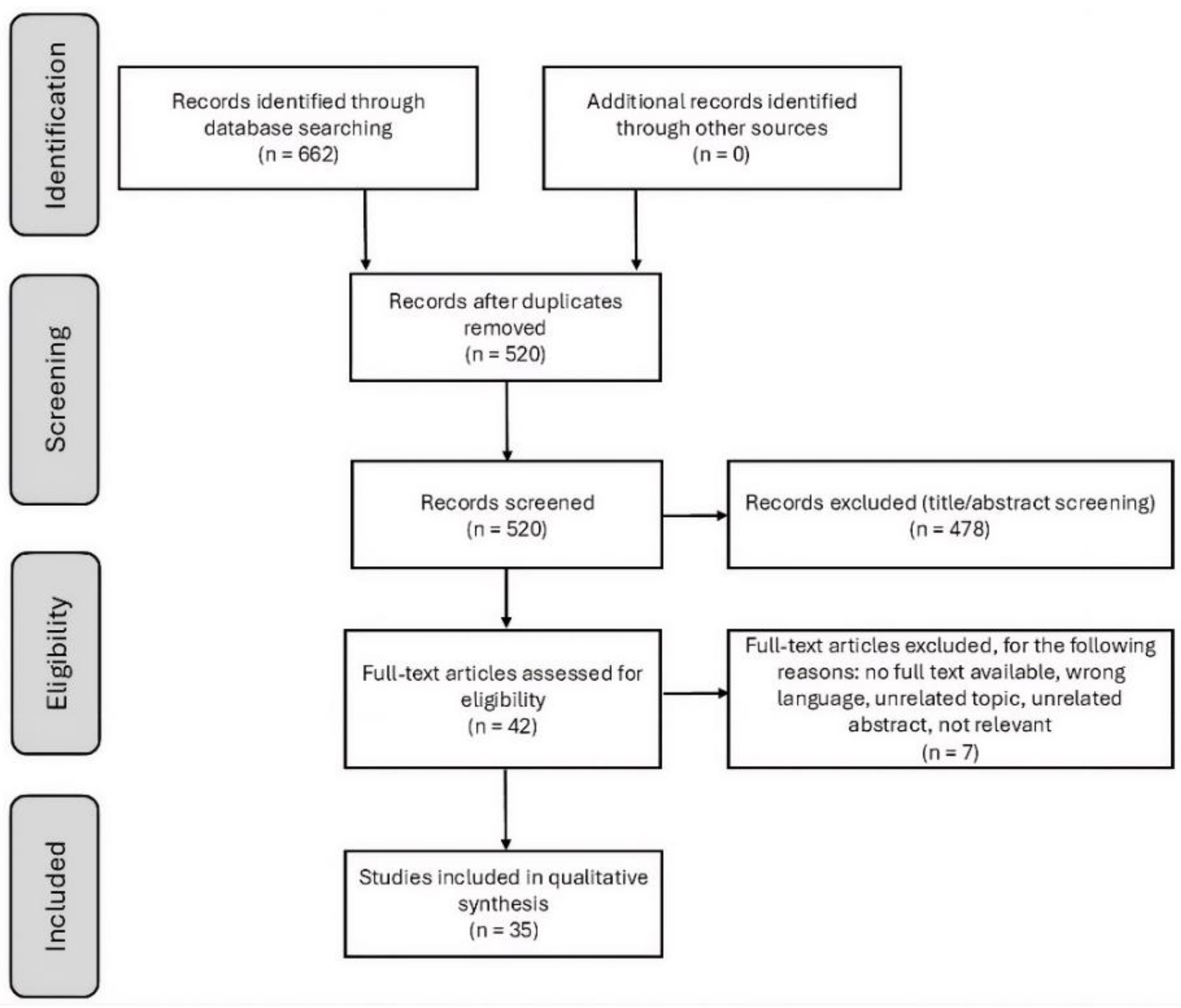
Data Extraction
Risk of bias (RoB) Assessment
Synthesis
Results
Overview of Included Studies
Lifestyle Interventions
Dietary Interventions
Physical Activity Interventions
Multicomponent Programs
Exogenous Compounds (Humans and Preclinical)
Pharmacological Interventions
Vitamins and Omega-3
Multinutrient Formulations
Polyphenols/Extracts and Nucleotide Supplementation
NAD⁺ Modulators
Blood/Circulation-Derived Interventions (Humans and Rodents)
Human Studies
Preclinical Studies
Cellular Reprogramming (Preclinical)
Discussion
Lifestyle Interventions: Modest Effects, More Coherent Under Sustained Exposure
Exogenous Compounds: High Heterogeneity and Vulnerability to Non-Reproducible Signals
Blood/Circulation-Derived Interventions: Striking Effects, Yet Transient and Interpretively Ambiguous
Partial Reprogramming: Early, Conceptual Evidence with Small Effects
Limitations
Conclusions
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
Use of generative AI tools
Appendix A. Full Search Strategy (Search Terms and Strings by Database)
Appendix A1. Mechanistic Evidence Search Strategy (PubMed)Appendix A2. Intervention Search Strategy (PubMed, Scopus, and CENTRAL)
| ID | Database | Date | Query | Filters | Results | Exported | Selected |
| M0 | PubMed | 17/11/2025 | (“hallmarks of aging”[tiab] OR “hallmarks of ageing”[tiab]) AND (update[tiab] OR review[pt] OR framework[tiab]) AND (2010:2025[dp]) | 2010–2025[dp]; review[pt] | 475 | 2 | 36599349; 23746838 |
| M1 | PubMed | 17/11/2025 | (epigenetic*[tiab] AND (aging[tiab] OR ageing[tiab])) AND (review[pt] OR mechanisms[tiab] OR framework[tiab]) AND (2010:2025[dp]) | 2010–2025[dp]; review[pt] | 3831 | 3 | 36522308; 29643443; 36336680 |
| M2 | PubMed | 17/11/2025 | (“DNA methylation”[tiab] OR “DNA Methylation”[Mesh]) AND (dynamics[tiab] OR remodeling[tiab] OR turnover[tiab] OR demethylation[tiab] OR “epigenetic drift”[tiab]) AND (aging[tiab] OR ageing[tiab] OR “Aging”[Mesh]) AND (review[pt] OR “Review”[Publication Type] OR mechanisms[tiab]) AND (2010:2025[dp]) NOT (atherosclero*[tiab] OR cardiomyocyte*[tiab] OR cancer*[tiab] OR tumor*[tiab] OR Huntington*[tiab] OR plaque[tiab]) | 2010–2025[dp]; review/Review; NOT (cardio/cáncer/etc.) | 180 | 3 | 25913071; 32356238; 29268958 |
| M3 | PubMed | 17/11/2025 | (chromatin[tiab] OR heterochromatin[tiab] OR “H3K9me3”[tiab] OR “H3K27me3”[tiab] OR sirtuin*[tiab] OR polycomb[tiab] OR histone*[tiab]) AND (aging[tiab] OR ageing[tiab]) AND (review[pt] OR mechanisms[tiab]) AND (2010:2025[dp]) | 2010–2025[dp]; review[pt] | 2976 | 2 | 27518561; 27482540 |
| M4 | PubMed | 17/11/2025 | (“DNA methylation age”[tiab] OR “epigenetic clock”[tiab] OR “epigenetic age”[tiab] OR PhenoAge[tiab] OR GrimAge[tiab] OR DunedinPACE[tiab]) AND (human*[tiab] OR tissue*[tiab]) AND (2010:2025[dp]) | 2010–2025[dp]; human*/tissue* | 718 | 4 | 24138928; 29676998; 36516495; 35029144 |
| ID | Database | Date | Query | Filters | Results | Exported | Selected |
| I1 | PubMed (humans) | 21/11/2025 | ( “epigenetic clock”[tiab] OR “epigenetic clocks”[tiab] OR “DNA methylation age”[tiab] OR “DNAm age”[tiab] OR “epigenetic age”[tiab] OR “methylation clock”[tiab] OR “methylation clocks”[tiab] ) AND ( trial[tiab] OR randomized[tiab] OR randomised[tiab] OR intervention[tiab] OR interventional[tiab] OR “pre-post”[tiab] OR “before and after”[tiab] ) AND (“2010/01/01”[PDAT] : “2025/12/31”[PDAT]) AND (english[la] OR spanish[la]) AND humans[mh] | 2010–2025; EN/ES; Humans; Article | 145 | 145 | See PRISMA |
| I2 | PubMed (animals) | 22/11/2025 | (“epigenetic clock”[tiab] OR “epigenetic clocks”[tiab] OR”DNA methylation age”[tiab] OR “DNAm age”[tiab] OR”epigenetic age”[tiab] OR “methylation clock”[tiab] OR”methylation clocks”[tiab])AND(trial[tiab] OR randomized[tiab] OR randomised[tiab] ORintervention[tiab] OR treatment[tiab] OR”pre-post”[tiab] OR “before and after”[tiab])AND(Mice[MeSH Terms] OR mouse[tiab] OR murine[tiab] ORrat[tiab] OR rats[tiab] OR mammal*[tiab])AND(“2010/01/01”[PDAT] : “2025/12/31”[PDAT])AND(english[la] OR spanish[la])NOT humans[mh] | 2010–2025; EN/ES; Article; NOT humans | 5 | 5 | See PRISMA |
| I3 | Scopus | 21/11/2025 | TITLE-ABS-KEY( (“epigenetic clock” OR “epigenetic clocks” OR “DNA methylation age” OR “DNAm age” OR “epigenetic age” OR “methylation clock” OR “methylation clocks”) AND (trial OR randomized OR randomised OR intervention OR interventional OR “pre-post” OR “before and after”) ) | 2010–2025; EN/ES; Article | 507 | 507 | See PRISMA |
| I4 | Cochrane CENTRAL | 21/11/2025 | ( “epigenetic clock”:ti,ab,kw OR “epigenetic clocks”:ti,ab,kw OR “DNA methylation age”:ti,ab,kw OR “DNAm age”:ti,ab,kw OR “epigenetic age”:ti,ab,kw OR “methylation clock”:ti,ab,kw OR “methylation clocks”:ti,ab,kw ) AND ( trial:ti,ab,kw OR randomized:ti,ab,kw OR randomised:ti,ab,kw OR intervention:ti,ab,kw OR “pre-post”:ti,ab,kw OR “before and after”:ti,ab,kw ) | 2010–2025; EN/ES | 5 | 5 | See PRISMA |
Appendix B. Extraction Table
| Paper | Intervention | Comparator | Epigenetic_outcome | Secondary_outcome | Risk_of_bias | ||||||||||||||||
| ID | First_author | Year | Title | Country | Specie | Design | N_total | N_per_group | Population_description | Intervention_family | Intervention_description | Dose_schedule | Comparator_type | Comparator_description | Clock_name | Tissue | Timepoints | Effect_on_DNAmAge | Effect_on_pace_of_aging | Safety_AEs | RoB_overall |
| 7 | Fitzgerald, Kara N | 2021 | Potential reversal of epigenetic age using a diet and lifestyle intervention: a pilot randomized clinical trial. | USA | Human | Randomized controlled parallel-group pilot trial; 8-week multimodal lifestyle intervention vs no-intervention control | 43 randomized (38 analysed for primary outcome) | Intervention 21 randomized (18 analysed); Control 22 randomized (20 analysed) | Community-dwelling men aged 50–72 years without recent or chronic disease (cardiovascular, diabetes, autoimmune, cancer, neurodegenerative, etc.), non-smokers, limited alcohol, recruited around Portland, Oregon. | Lifestyle | 8-week plant-centered, methylation-supportive diet plus PhytoGanix® and UltraFlora® with structured exercise. | 8-week intervention; diet emphasizing high intake of dark leafy greens, overnight 12-hour fast (~7pm–7am), reduced refined carbohydrates, avoidance of added sugar and most processed foods; supplements: PhytoGanix® 2 servings/day and UltraFlora® Intensive Care 2 capsules/day (L. plantarum 299v); lifestyle: ≥30 minutes moderate-intensity exercise 5 days/week target ≥7 hours sleep per night. | No-intervention / usual lifestyle control | Control group received no specific dietary, supplemental, exercise, sleep or stress-management intervention; continued usual habits and any pre-existing low-dose supplements allowed by protoco | Horvath DNAmAge | Saliva | Baseline and end of 8-week intervention (study visit at ~week 9) | Compared with controls, intervention participants were on average 3.23 years younger at end of study (between-group difference, p=0.018). Within the intervention arm, DNAmAge decreased by mean 1.96 years vs baseline (p=0.066), while controls increased by 1.27 years (p=0.153). No net change in overall methylation level of the 353 Horvath clock CpGs; change reflected repositioning of methylation patterns toward a younger profile | NR | NR (no explicit adverse event or safety data reported) | (RoB 2) Overall some concerns: randomization and concealment were adequate with similar baselines, and DNAmAge (Illumina EPIC) was analysed blinded despite unblinded participants/coaches, but attrition was moderate and slightly unbalanced (18/21 vs 20/22 analysed) with limited reasons reported, and the small pilot sample plus no detailed pre-published analysis plan keep the rating at some concerns. |
| 9 | Bischoff-Ferrari, Heike A | 2025 | Individual and additive effects of vitamin D, omega-3 and exercise on DNA methylation clocks of biological aging in older adults from the DO-HEALTH trial. | Switzerland | Human | Post hoc analysis of randomized, double-blind, placebo-controlled 2×2×2 factorial RCT (DO-HEALTH) with 3-year vitamin D, omega-3 and home exercise interventions | 777 | Placebo 95; Vitamin D only 101; Omega-3 only 98; SHEP only 92; Vitamin D + Omega-3 95; Vitamin D + SHEP 104; Omega-3 + SHEP 95; Vitamin D + Omega-3 + SHEP 97 | Community-dwelling adults ≥70 years (mean age ~75.5, ~60% women), generally healthy and active (no major cardiovascular events or cancer in prior 5 years, MMSE ≥24, independent mobility) | Exogenous compound–based | Daily vitamin D3 and omega-3 supplementation with or without a simple home strength exercise program (SHEP) | Vitamin D3 2,000 IU/day; omega-3 1 g/day (330 mg EPA + 660 mg DHA from marine algae) given as two identical gel capsules per day; SHEP strength-training 30 min three times per week for 3 years versus attention-control joint-flexibility exercises 30 min three times per week | Placebo | Vitamin D and omega-3 placebos identical in appearance, taste and weight; attention-control flexibility exercise program 30 min three times weekly; factorial design comparing vitamin D vs placebo, omega-3 vs placebo and SHEP vs control exercise plus their combinations over 3 years | PC-PhenoAge | Whole blood | Baseline and 3-year follow-up | Omega-3 vs no omega-3 reduced PhenoAge age-acceleration change (standardized d ≈ −0.16; 95% CI −0.30 to −0.02), corresponding to ~3 months slower biological aging over 3 years; vitamin D alone and SHEP alone showed no significant effects; combinations including omega-3 plus vitamin D and/or SHEP showed additive benefits on PhenoAge (d ~ −0.24 to −0.32) | NR | NR (no explicit adverse event or safety data reported) | RoB2 – overall low to some concerns. Parent DO-HEALTH trial was a large, multicenter, randomized, double-blind, placebo-controlled 2×2×2 factorial RCT with centralized randomization and matching placebos; exercise arms used an attention-control flexibility program, with blinding less feasible. DNAm assays were performed on stored blood with laboratory staff blinded to treatment allocation. |
| 11 | Fiorito, Giovanni | 2021 | DNA methylation-based biomarkers of aging were slowed down in a two-year diet and physical activity intervention trial: the DAMA study. | Italy | Human | 24-month randomized 2×2 factorial lifestyle RCT (diet, PA, diet+PA, control); DNAm outcomes = secondary analysis | 219 paired samples (post-QC) | Diet 57 → 56; PA 56 → 56; Diet+PA 53 → 53; Control 58 → 54 (final pairs aggregated to N=219) | Healthy postmenopausal women (50–69 y) with high mammographic density (>50%), nonsmokers, attending Florence breast cancer screening program | Lifestyle | Plant-based, low-glycemic-load diet; structured physical activity program (~1 h/day moderate + 6–10 MET-h/week vigorous); combined arm = both; control = minimal lifestyle advice | 24 ± 3 months; diet: counseling + 6 group sessions + 8 cooking classes; PA: supervised weekly 1-h session + home exercises + walks | Active minimal-intervention lifestyle advice | – Diet effect = diet-containing arms (diet + diet+PA) vs no-diet arms (PA + control) – PA effect = PA-containing arms (PA + diet+PA) vs no-PA arms (diet + control) |
DNAmGrimAge, DNAmGrimAge Acceleration (DNAmGrimAA), Epigenetic Mutation Load (EML; stochastic epigenetic mutations), GrimAge components (PAI-1, Leptin, GDF-15, etc.) | Whole blood (buffy coat) | Baseline and 24 months | Diet slowed aging: – Within-group: diet −0.41 y (95% CI −0.79, −0.03) vs controls +0.25 y – DiD (diet vs control): β = −0.66 y (95% CI −1.15, −0.17; p=0.01) – WBC-adjusted: β = −0.42 y (p=0.05) PA: no DNAmGrimAA effect (β = +0.09 y; p=0.73) |
No formal pace clock; EML used PA slowed epigenetic mutation accumulation: – DiD (PA vs control): β = −2.06 “years” (p < 0.001), robust to WBC adjustment Diet: no EML effect (β = −0.37; p=0.39) |
NR (no explicit adverse event or safety data reported) | (RoB 2) Low risk for randomization and minimal missing DNAm data, but some concerns due to the open-label lifestyle intervention, unclear blinding, and secondary outcomes with modelling flexibility.– Low risk: randomization, minimal missing DNAm data – Some concerns: open-label lifestyle, unclear blinding, secondary outcomes with modelling flexibility |
| 12 | Orr, Miranda E | 2024 | A randomized placebo-controlled trial of nicotinamide riboside in older adults with mild cognitive impairment. | USA | Human | 10-week double-blind RCT, NR vs placebo, n=20 analyzed | 20 | NR 10, placebo 10 | Adults ≥65 y with mild cognitive impairment (MoCA<26); mostly Hispanic; mean age ~75–77 | Exogenous compound–based | Nicotinamide riboside (NIAGEN®) 1 g/day | Escalation 250→1000 mg/day; total 10 weeks | Placebo | matched capsules, identical escalation | IEAA, EEAA, PhenoAge, GrimAge | PBMCs | Baseline, 10 weeks | No significant within-group or between-group changes in any epigenetic-age metric Exploratory bootstrap: AgeAccelPheno & Grim: subtle decrease with NR; placebo ≈0 (Pheno) or slight ↑ (Grim) EEAA (Hannum): slight increase (accelerated aging) with NR IEAA: no change Overall: very small, inconsistent, non-significant effects |
NR | Mild–moderate AEs similar in NR vs placebo (7/10 each); one placebo stroke; one NR severe nausea resolved with dose reduction | (RoB 2) Some concerns due to the small sample size (n=10 per group), short study duration, and exploratory testing across multiple epigenetic clocks. |
| 25 | Fuentealba, Matias | 2025 | Multi-Omics Analysis Reveals Biomarkers That Contribute to Biological Age Rejuvenation in Response to Single-Blinded Randomized Placebo-Controlled Therapeutic Plasma Exchange. | USA | Human | Single-site single-blinded randomized placebo-controlled 4-arm trial; exploratory biological aging endpoints; primary aim = safety & feasibility; 3-month intervention, 3 timepoints | 42 completed (DNA methylation analyzed for all) | TPE+IVIG 10; Bi-weekly TPE 11; Monthly TPE 11; Sham 10 | Healthy adults ≥50 y (one in 40s allowed), no major clinical disease; exclusions for cardiovascular/pulmonary disease, active cancer/infection, GH/stem cells, psychiatric illness, anti-aging supplements (1 exception taking rapamycin) | Blood-derived | Therapeutic plasma exchange (1× plasma volume, 5% albumin replacement) ± 2 g IVIG; devices: Spectra Optia; sham procedure with water-filled lines + 250 mL saline | – Bi-weekly TPE+IVIG: 2 sessions first week of each month × 3 months (6 TPE sessions total) – Bi-weekly TPE (no IVIG): same schedule – Monthly TPE: 1 session/month × 6 – Sham: identical appearance, no plasma removal |
Placebo / sham-controlled | Sham apheresis with realistic noise/flow simulation; participants, caregivers, and raters blinded | ~35 TruAge clocks: Horvath, Hannum, PhenoAge, GrimAge, SystemsAge family, fitness clocks (FitAge, Gait, Grip, VO2max), PC clocks (PCGrimAge, PCHorvath/Hannum/PhenoAge), organ/system clocks (immune, inflammatory, metabolic, kidney, liver, heart, musculoskeletal), stochastic/drift clocks | Peripheral whole blood (EPIC 850k) | Baseline (tp1), before 4th session (tp2), before 6th session (tp3) | At timepoint 2 (peak effect): – TPE+IVIG: −2.61 y (FDR 6.2e-5) vs sham – Monthly TPE: −1.32 y (FDR 2.4e-2) vs sham – Bi-weekly TPE: negative direction; significant vs sham (mean not specified) Timepoint 3: No significant differences vs sham (attenuation/compensation) Clock families: SystemsAge immune/inflammatory clocks show largest effects (≈7–10 y decreases in TPE+IVIG; ≈2.5–5 y in monthly TPE) Overall: 15 clocks show significant rejuvenation at tp2 across active arms |
NR | 1 mild allergic reaction to albumin (0.42%); two total AEs requiring discontinuation (one IVIG-related). No major events; well tolerated | (RoB 2) Some concerns: allocation was first-come-first-served, raising high concern for sequence generation and possible baseline imbalance; however, blinding of patients, caregivers, and raters was reported, missing data were low (42/44 completed), and outcomes were measured with objective arrays likely analysed blinded, while testing multiple clocks (35+) raises concerns about multiplicity and selective reporting. |
| 27 | Hernández-Arciga, Ulalume | 2025 | Dietary methionine restriction started late in life promotes healthy aging in a sex-specific manner. | Norway | Human and mouse (Only human part was taken) | Double-blind randomized 8-week dietary RCT (SAAR vs high-SAA control), but this paper analyzes only a subset of the SAAR arm (n=20) with pre/post DNAm; no control-arm DNAm presented → effectively a single-arm pre–post epigenetic study. | 20 SAAR participants (paired baseline + 8-week samples) | Single analyzed group: SAAR/MetR n=20 (17F / 3M); no control methylation data | Overweight/obese but otherwise healthy adults; mean age 32.9 ± 6.1 y, BMI 31.5 ± 2.3 kg/m²; non-smoking, diet-controlled living conditions per parent trial | Lifestyle | Dietary sulfur amino acid restriction (human analogue of methionine restriction) | 8 weeks; SAAR vs high-SAA control in parent RCT, but only SAAR analyzed here | None for epigenetic outcomes (control arm not analyzed) | N/A — only SAAR arm methylation measured; therefore effects are within-group pre–post | Universal mammalian clocks (Horvath multi-species panel); mammalian blood clock; other mammalian composite clocks (exact names not fully itemized but derived from Epigenetic Clock Development Foundation)* | Whole blood (EDTA) | Baseline and 8 weeks | No effect – Authors explicitly state: “8 weeks of the sulfur amino acids diet did not affect the epigenetic age.” – No mean Δ reported; visual inspection shows no significant shift across mammalian clocks. – Direction = null, magnitude = not detectably different from 0. |
NR | NR (no explicit adverse event or safety data reported) | (RoB 2) Some concerns because only the SAAR arm was analysed, undermining the randomized comparison, and the reasons for missing methylation data (available for 20/31 SAAR participants) were not described. Measurement risk is low given objective lab-based clocks, and transparent reporting of a null result reduces concerns about selective reporting. |
| 43 | McGee, Kirsty C | 2024 | A combination nutritional supplement reduces DNA methylation age only in older adults with a raised epigenetic age. | UK | Human | Uncontrolled open-label pre–post 12-week intervention; no comparator arm; DNAm age (blood + saliva) + inflammation + function | Blood: 79; Saliva: 75 (paired baseline + 12-week) | Single arm only (supplement group; 80 completers) | Healthy older adults ≥60 y; mean age 71.9 ± 6.2 y; BMI 25.8 ± 3.9; 49F / 31M; mostly White | Exogenous compound–based | Daily combination supplement (vitamins + polyphenols + omega-3) | 12 weeks daily; ingredients/day: Vit D3 20 µg; Niacinamide 50 mg; Vit C 85 mg; Omega-3 (EPA+DHA) 250 mg; Olive extract (10 mg hydroxytyrosol); Resveratrol 30 mg; Astaxanthin 3.2 mg | None (no placebo or control) | N/A – single-arm pre/post only | Blood: Horvath, Hannum, PhenoAge, GrimAge, Mean EpiAge (composite) Saliva: InflammAge (age & acceleration) |
Whole blood (EPIC 850K) | Baseline and 12 weeks | Effect_on_DNAmAge — whole cohort: Blood clocks: No significant changes in Horvath, Hannum, PhenoAge, GrimAge, or Mean EpiAge; no significant change in epigenetic age acceleration. Effect_on_DNAmAge — subgroups: Raised Horvath acceleration (≥2 y, n=23): Horvath EAA ↓ ~1.98 y (−21.5%), p=0.069 (trend) |
NR | NR (no explicit adverse event or safety data reported) | (RoB 2) High risk because there was no randomization or control group, creating high susceptibility to regression to the mean and time effects; the study used an open-label design, included subgroup analyses with multiplicity concerns, and although missingness was small, the reasons were not detailed. |
| 54 | Sae-Lee, Chanachai | 2018 | Dietary Intervention Modifies DNA Methylation Age Assessed by the Epigenetic Clock. | Netherlands | Human | Double-blind, randomized, placebo-controlled 2-year RCT in older adults, but this epigenetic analysis includes only the supplemented arm (n=44) → effectively single-arm pre–post DNAm with sex × MTHFR genotype stratification; secondary analysis | 44 (all in folic acid + B12 group; paired baseline + 2-year samples) | By sex: 19 males, 25 females | Community-dwelling older adults aged 65–75 y, generally healthy, non-smokers, not heavy drinkers | Exogenous compound–based | Folic acid + vitamin B12 supplementation | Folic acid 400 µg/day + vitamin B12 500 µg/day, orally, daily for 2 years | None for DNAm analysis (placebo arm not used) | Within-group comparison: baseline vs 2-year follow-up in supplemented individuals; analyses stratified by MTHFR C677T genotype (CC vs TT) and sex | Horvath 2013 pan-tissue DNAmAge (age acceleration residual) | Whole blood | Baseline and 2 years | Whole supplemented group (n=44): – Age acceleration residual: no significant change – Mean Δ ≈ −0.77 ± 1.44 years; p = 0.60 |
NR | NR (no explicit adverse event or safety data reported) | (RoB 2) Some concerns: although the parent trial was a well-conducted double-blind RCT, the epigenetic analysis included only the active/supplemented arm, losing the randomized comparison. Multiple small-n subgroup analyses (sex × genotype) with one positive finding (female 677CC) raise multiplicity/chance-finding concerns, while objective DNAm measures and minimal missing data suggest low risk for measurement and attrition. |
| 65 | Salas-Huetos, Albert | 2021 | Sperm DNA methylation changes after short-term nut supplementation in healthy men consuming a Western-style diet. | Spain | Human | 14-week randomized controlled parallel-group diet trial; this paper is a pre-planned methylation substudy of ejaculated sperm; ~72 men with paired DNAm data; germline epigenetic aging assessed | 72 (paired baseline + 14-week sperm methylation) | ≈35–37 per arm (nuts vs control; exact split not essential for clock outcomes) | Healthy men ~18–35 y, consuming Western-style diets; BMI normal/overweight; no fertility problems; no nut allergies | Lifestyle | Mixed tree-nut supplementation | 60 g/day nuts (30 g walnuts + 15 g almonds + 15 g hazelnuts) for 14 weeks, consumed atop habitual diet | Active dietary control (Western diet without nuts) | Control group maintained habitual Western diet and were instructed not to consume nuts | Sperm-specific germ line epigenetic age predictor (Jenkins sperm clock); includes epigenetic age and age acceleration | Ejaculated sperm (purified sperm fraction) | Baseline and 14 weeks | No effect – No significant within-group change in sperm germ line age for either nuts or control – No significant between-group difference in Δ germ line age – Authors explicitly state: no detectable effect on sperm epigenetic aging, despite many nut-related DMRs |
NR | NR (no explicit adverse event or safety data reported) | (RoB 2) Some concerns: the parent randomization was sound, but the methylation subset (~72/119) may not be fully representative, and participants were unblinded in the diet trial. Measurement bias is likely low due to objective sperm DNAm arrays, and the methylation outcomes were prespecified with a transparently reported null result. |
| 67 | Kou, Minghao | 2025 | Epigenetic Age Acceleration and Cardiometabolic Biomarkers in Response to Weight-Loss Dietary Interventions Among Obese Individuals: The MACRO Trial. | USA | Human | 12-month parallel-arm RCT (low-carb vs low-fat). This Aging Cell paper is a secondary analysis of DNAm-based biological aging + cardiometabolic biomarkers at 0, 3, 12 months. |
Baseline 144; 3 months 129; 12 months 112 | Baseline: 71 LF / 73 LC 3 mo: 62 LF / 67 LC 12 mo: 54 LF / 58 LC |
Adults 22–75 y with obesity (BMI 30–45); mean age ~47 y; ~89% women; predominantly White and African-American; no diabetes or CVD | Lifestyle | Low-carbohydrate diet vs low-fat diet | • Low-carb: <40 g/day digestible carbohydrate • Low-fat: <30% energy from fat; <7% saturated fat Duration 12 months |
Active comparator (two diet arms) | Low-carb vs low-fat; no calorie targets for either arm | PCPhenoAge (AA), PCGrimAge (AA), DunedinPACE | Whole blood (EPIC 850K) | Baseline, 3 months, 12 months | No substantial diet-specific effect on epigenetic aging • DunedinPACE: ~1.00 → 0.99 (very small decrease; similar in both diets) • PCPhenoAge AA: little change; moves with chronological aging • PCGrimAge AA: similar minimal change; no diet group separation • No significant group×time interactions for any clock |
DunedinPACE decreased slightly (~0.01), but not different by diet; considered negligible effect size | NR (no explicit adverse event or safety data reported) | (RoB 2) Some concerns: the randomized RCT design supports low risk for randomization, but the open-label diet raises concerns about deviations, and 12-month attrition (144→112) was not fully explored for prognostic imbalance. Objective DNAm arrays reduce measurement bias, yet as a secondary analysis using multiple clocks, multiplicity remains a concern. |
| 72 | Carreras-Gallo, Natalia | 2025 | Effects of a natural ingredients-based intervention targeting the hallmarks of aging on epigenetic clocks, physical function, and body composition: a single-arm clinical trial. | USA | Human | 12-month single-arm, pre–post clinical trial (no randomization, no placebo); 1–4 DNAm measurements per participant | 51 participants (baseline + 3m + 6m + 12m variable availability) | Single cohort (no comparator arms) | Adults ≥55 y; 49% female; age 54–84; relatively healthy with low baseline EAA compared to external aging-biobank reference; self-selected supplement users | Exogenous compound–based | “Cel System” (Cel1, Cel2, Cel3) + 10-min walking + 5-min mindfulness daily | Daily supplementation × 12 months – Cel1: 2-HOBA (hobamine), astragalus extract, rutin, vitamin C, levomefolate, B12, zinc, selenium – Cel2: NMN, pterostilbene, astaxanthin, L-carnosine, vitamin D, riboflavin – Cel3: Apigenin, fisetin, oleuropein, EGCG, berberine, ALA, withaferin A Exact pill counts/frequency not reported |
None (pre–post only) | Each participant compared to own baseline | – First-generation / PC clocks: PCHorvath pan-tissue, PCHorvath skin&blood, PCHannum, IntrinClock, stochastic clocks – Second-generation: PCPhenoAge, PCGrimAge, OMICmAge, Marioni cAge, DNAmTL – Causal-framework clocks: DamAge, CausAge, AdaptAge – Third-generation pace: DunedinPACE – SystemsAge (overall + organ-specific: lung, immune, metabolic, etc.) – Fitness clocks: DNAmFitAge, DNAmGrip, DNAmGait, DNAmVO2max, DNAmFEV1 |
Whole blood (EPIC 850K) | Baseline, 3 months, 6 months, 12 months | PC Horvath pan-tissue EAA 0.60 → −0.15 at 12m (Δ ≈ −0.75 y; p=0.048) Strongest reduction at 6m: −0.36 (p=6.1×10⁻⁴) PC Horvath skin & blood EAA −1.23 → −0.31 (Δ ≈ +0.92 y “older”; p=0.045) PCHannum EAA Worsening at 6m (−0.45 → +0.29; p=0.027), resolves by 12m (−0.15; p=0.80) DamAge (damage-related “causal” clock) 2.46 → −0.66 (3m) and −0.74 (6m) (p≈0.003–0.0014) Partial rebound by 12m: −0.12 (p=0.12) SystemsAge overall −0.72 → −0.49 (ns; p=0.73) SystemsAge-lung 0.56 → −0.45 (Δ ≈ −1.01 y; p=0.0061) Fitness clocks Example: DNAmGrip EAA −0.24 → −0.87 (Δ ≈ −0.63 y; p=0.0042) |
DunedinPACE 0.94 → 0.99 at 12m (Δ ≈ +0.05, ~5% faster pace; p = 7.4×10⁻⁵) Short-term slowing at 3m (0.96) but rebound and overshoot by 12m |
NR (no explicit adverse event or safety data reported) | (ROBINS-I) High risk because the study had no randomization, control group, or blinding, and the multi-component supplement plus lifestyle intervention makes attribution to any single component impossible. Additional concerns include possible selection bias from self-selected supplement users, missing timepoints (1–4 per participant), industry involvement (authors employed by TruDiagnostic/SRW), and the inability to rule out regression to the mean and time effects. |
| 73 | Borsky, Pavel | 2025 | Human clinical trial of plasmapheresis effects on biomarkers of aging (efficacy and safety trial). | Czech Republic | Human | Prospective stratified randomized cross-over (G1 = 8 sessions; G2 = delayed-start 4 sessions) Analyzed primarily as dose–response per plasmapheresis session Duration 18 weeks |
41 enrolled → 38 at 9w: 34 completers | G1: 28 allocated G2: 13 allocated |
Healthy adult first-time plasma donors 40–60 y; screened per standard blood-donor rules; no major chronic disease; median age ~49.6 | Blood-derived | Standard donor plasmapheresis (Haemonetics PCS2) | Plasma volume 570–830 mL/session; anticoagulant citrate (CITRASOL 4%); q≥14 days • G1: 8 sessions (0–18 wks) • G2: 0–9 wks control, then 4 sessions (9–18 wks) |
Dose–response (per-session) within a partially cross-over design; no untreated control | Effect modeled as change in clock value per additional session adjusting for age, sex, monocyte %, naive CD4 T-cells | Horvath1, Horvath2, PhenoAge, GrimAge, GrimAge2, GrimAge2_tuned, GrimAge2_calibrated, Hannum, RobustHannum, PC clocks (PC Horvath1/2/Hannum/PhenoAge/GrimAge/DNAmTL), DunedinPACE, GrimAge component surrogates (ADM, B2M, Cystatin C, GDF-15, Leptin, PACKYRS, PAI-1, TIMP-1, COX) | Whole blood (buffy coat DNA) | Baseline, 9 weeks, 18 weeks | Significant positive per-session increases: (Estimate = increase in epigenetic age per plasmapheresis session) DNAmGrimAgeBasedOnRealAge: +0.26 ± 0.05 y/session, p=5×10⁻⁷ DNAmGrimAge2BasedOnRealAge: +0.22 ± 0.05, p=2×10⁻⁴ DNAmGrimAge2_Tuned: +0.16 ± 0.03, p=1.26×10⁻⁵ DNAmGrimAge2_Calibrated: +0.22 ± 0.05, p=2×10⁻⁴ Hannum DNAmAge: +0.17 ± 0.04, p=2×10⁻⁴ RobustHannum: +0.13 ± 0.03, p=2.42×10⁻⁵ DunedinPACE: +0.003 ± 0.001, p=0.0058 (≈ 0.3% faster pace/session) |
DunedinPACE increases with each session (+0.003/session). Cumulative effect over 8 sessions ≈ +0.024 (≈2.4% faster pace), directionally adverse. |
One participant discontinued early due to hypotension | (RoB 2) High risk: although stratified randomization was used, there was no non-treatment control and the cross-over design only partly mitigates confounding. The study was not blinded, had a small sample with attrition, tested many clocks with multiple comparisons, and although the mixed-model dose–response analysis was appropriate, time effects, procedure stress, and lifestyle confounding cannot be ruled out. |
| 75 | Yaskolka Meir, Anat | 2021 | Lifestyle weight-loss intervention may attenuate methylation aging: the CENTRAL MRI randomized controlled trial. | Israel | Human | 18-month randomized controlled lifestyle trial (LF vs MED/LC diets; at 6 months split to ±PA). This Aging Cell paper is a secondary DNAm substudy with n=120 (baseline + 18 months). |
120 (complete paired methylation) | Originally 4 groups (LF, LF+PA, MED/LC, MED/LC+PA ≈30 each), but DNAm analyses pool main diet contrasts (LF vs MED/LC). Subgroup effects (weight-loss success, liver fat status) emphasized. | Sedentary adults with abdominal obesity or dyslipidemia; mean age 48.6±9.3 y, BMI 30.2±3.3; workplace cohort; 58.8% with fatty liver at baseline. | Lifestyle | Low-fat (LF) diet vs Mediterranean/low-carb (MED/LC) diet; after 6 months each split into diet alone vs diet+PA. | Daily hypocaloric diet for 18 months; PA program moderate-intensity aerobic (after 6 months in the PA arms). | Active comparator (LF vs MED/LC; ±PA). Epigenetic results mainly within-subject and by subgroup, not between diets. | LF vs MED/LC; PA vs no PA; but no significant diet-group effects on DNAm aging. | Primary: Li 2018 240-CpG blood DNAmAge Secondary: Horvath 353-CpG (baseline only) |
Whole blood (EPIC 850K) | Baseline and 18 months | Cohort (240-CpG clock) mAge increased +1.1 ± 1.9 years over 18 months (≈ chronological aging). No difference between LF (+1.3 y) and MED/LC (+0.9 y), p=0.2. No effect of ±PA. |
NR | NR (no explicit adverse event or safety data reported) | (RoB 2) Some concerns because the DNAm substudy included only 120/278 participants, making selection bias possible, and the open-label lifestyle intervention with variable PA adherence raises concerns about deviations; subgroup findings also increase multiplicity concerns. Objective methylation assays reduce measurement bias, and there was no selective reporting of diet-group null results. |
| 78 | Vetter, Valentin Max | 2022 | Vitamin D supplementation is associated with slower epigenetic aging. | Germany | Human | Prospective two-wave longitudinal cohort (BASE-II → GendAge). Quasi-interventional, non-randomized, with optimal pair matching of treated vs untreated vitamin-D–deficient participants. |
~1,070 with epigenetic clocks at follow-up (exact n varies slightly by clock). | • Treated deficient → sufficient: 63 • Untreated deficient: 63 (matched) • Healthy controls (sufficient→sufficient): 63 (matched) |
Community-dwelling older adults (60–85 y baseline; 68.3±3.5 → 75.6±3.8 at follow-up), ~52% female. Vitamin D deficiency common at baseline (46%). | Exogenous compound–based | Vitamin D supplementation | Dose, preparation, and adherence not recorded; defined only via self-report + medication lists. | Matched quasi-control groups | Untreated deficient: remained deficient, no supplementation Healthy controls: always sufficient, no supplementation. Follow-up ~7.4 years. |
7-CpG (Vetter) Horvath 2013 Hannum 2013 PhenoAge GrimAge |
Whole blood | DNAm clocks evaluated at follow-up (T1). 7-CpG also available longitudinally but intervention effect analyzed cross-sectionally at T1. |
Among initially vitamin-D–deficient older adults, those who started supplementation and became sufficient had about 2.6 years lower 7-CpG DNAm age acceleration and 1.3 years lower Horvath DNAm age acceleration at follow-up than matched deficient non-supplementers, while Hannum, PhenoAge, and GrimAge showed no significant differences. | NR | NR (no explicit adverse event or safety data reported) | (ROBINS-I) High risk of bias because the study was non-randomized with self-selected supplementation, and dose and adherence were not known. Matching only partly reduces confounding, possible lifestyle co-interventions were not measured, and the use of multiple clocks with subgroup analyses raises concerns about multiplicity. |
| 81 | Nwanaji-Enwerem, Jamaji C | 2021 | An epigenetic aging analysis of randomized metformin and weight loss interventions in overweight postmenopausal breast cancer survivors. | USA | Human | 6-month, 2×2 factorial, randomized, double-blind, placebo-controlled RCT Arms (each n=48 in EA subset): Metformin only Weight-loss only Weight-loss + metformin Placebo only Epigenetic clocks are a post-hoc secondary analysis. |
192 | 48 per arm | Overweight/obese postmenopausal breast cancer survivors, mean age ~63 y, clinically stable, no active chemo/radiotherapy. | Exogenous compound–based | Metformin (parent trial: titrated up to ~850 mg BID) Phone-based weight-loss program (calorie restriction + PA target ~300 min/week) |
Metformin daily for 6 months Weight-loss coaching throughout 6 months |
Placebo (for metformin) and active lifestyle comparators (factorial design) | Placebo tablets + no weight-loss program (true control arm) | Hannum EAA, Horvath EAA, SkinBlood EAA, IEAA, EEAA, PhenoAge EAA, GrimAge EAA, DNAmTL, EpiTOC, EpiTOC2, MiAge | Peripheral blood (buffy coat) | Baseline and 6 months | Weight loss only vs placebo: • Hannum EAA: +0.98 y (p=0.19) • Horvath EAA: +0.74 y (p=0.42) • SkinBlood EAA: +0.86 y (p=0.21) • PhenoAge EAA: +2.02 y (p=0.05) — nominal, opposite direction, not interpreted as real • GrimAge EAA: +0.76 y (p=0.27) Metformin only vs placebo: All EAA differences −0.91 to +0.82 y (all p ≥ 0.18). Weight loss + metformin vs placebo: All EAA differences −0.63 to +0.45 y (all p ≥ 0.50). |
NR | NR (no explicit adverse event or safety data reported) | (RoB 2) Some concerns because the epigenetic analysis included only 192/333 randomized participants, so the subset may be subject to selection bias despite the rigorous factorial RCT design. Testing many clocks across many contrasts raises multiplicity concerns, though objective EPIC measures reduce measurement bias and consistently reported null results lower selective reporting concerns. |
| 82 | Clement, James | 2022 | Umbilical cord plasma concentrate has beneficial effects on DNA methylation GrimAge and human clinical biomarkers. | USA | Human | Phase I single-arm, open-label pre–post trial 10 weekly intramuscular injections of umbilical cord plasma concentrate; outcome measures before vs after |
18 (paired baseline & post-treatment) | Adults 60–95 y (mean ~74), generally in age-typical health; some with hypertension, hyperlipidemia, diabetes; 1 with early dementia & RA. No active cancer. 10 women, 8 men. | Blood-derived | Human umbilical cord blood plasma concentrate (hUCBP secretome) | • 1 mL IM injection weekly × 10 weeks • Each vial derived from 100 mL cord plasma (≈5 mg pellet of concentrated extracellular vesicles/proteins resuspended in 1 mL saline) Total exposure ≈ 1 L cord plasma equivalent |
None | Pre–post within-subject (baseline vs 10 wks) | Horvath 2013, Hannum 2013, Skin&Blood 2018, PhenoAge, DNAmTL, DNAmGrimAge | Whole blood | Baseline and post-treatment (10 weeks) | Primary signal GrimAge acceleration: 0.04 → −0.78 years → Δ = −0.82 years, p = 0.0093 (paired t-test) GrimAge protein surrogates: • DNAm Cystatin C ↓ (p = 0.024) • DNAm GDF-15 ↓ (p = 0.0024) Other clocks Horvath, Hannum, Skin&Blood, PhenoAge, DNAmTL: No significant changes |
NR | 2 participants had mild injection-site redness/heat, resolved with diphenhydramine; tolerated further injections | (ROBINS-I) High risk because there was no control group, leaving high confounding and regression-to-the-mean concerns, and the sample was very small (N=18) and convenience-based rather than population-based. The study was open-label with industry involvement, tested multiple clocks and biomarkers, and the GrimAge effect was only nominal (uncorrected). | |
| 88 | Campisi, Manuela | 2025 | Unveiling the geroprotective potential of Monarda didyma L.: insights from in vitro studies and a randomized clinical trial on slowing biological aging and improving quality of life. | Italy | Human | Double-blind placebo-controlled randomized parallel-group trial | 81 | Monarda 40; Placebo 41 | Adults aged 45–65, university employees undergoing occupational health checks, generally healthy, non-smokers, free of major chronic disease. | Exogenous compound–based | Monarda didyma L. extract | 100 mg/day oral capsule for 12 weeks | Placebo | Identical 100 mg maltodextrin capsule daily for 12 weeks. | 5-CpG DNAmAge (ELOVL2, C1orf132, KLF14, TRIM59, FHL2) | Whole blood | Baseline and week 12 | DNAmAge remained stable in intervention (p=0.45) and increased in placebo (p<0.0001); DNAmAge significantly lower in intervention vs placebo at week 12 (p=0.016). | NR | NR (no explicit adverse event or safety data reported) | (RoB 2) Some concerns: although the study used a randomized, double-blind, placebo-controlled RCT design with objective biomarkers, concerns remain due to industry funding, multiple endpoints, modest sample size, and short duration. |
| 92 | Loh, Kah Poh | 2023 | Exercise and epigenetic ages in older adults with myeloid malignancies. | USA | Human | Single-arm, pre–post pilot trial; ~8–12 weeks of exercise during chemotherapy. DNAm measured at baseline and post-intervention (after 2 chemo cycles). Exploratory; no control group. |
20 (complete paired DNAm data) | Adults ≥60 with myeloid malignancies receiving outpatient chemotherapy (AML 55%, MDS 40%, MDS/MPN 5%); mean age 71.2 y, 65% male, 90% White; mixed chemo stages (HMA±venetoclax, others). |
Lifestyle | GO-EXCAP (mobile-health–supported home exercise) | Low–moderate intensity progressive walking + resistance bands; daily step tracking via Garmin; ~26 min/day of resistance work; ~3 days/week; duration 8–12 weeks (two chemo cycles). | None (pre–post only) | Baseline vs post-intervention within-subject comparison | Horvath, Hannum, IEAA, EEAA, GrimAge, PhenoAge, DunedinPACE | Whole blood | Baseline and 8–12 weeks | • GrimAge: mean Δ −0.7 y (median −1.4), p = 0.17 • PhenoAge: mean Δ −0.8 y (median −1.4), p = 0.35 • DunedinPACE: mean Δ +0.02 (median −0.1), p = 0.47 • Horvath/Hannum/IEAA/EEAA: all non-significant (p ≥ 0.43) |
DunedinPACE: Mean Δ +0.02 (ns); median −0.1 (ns). No statistically significant effect. |
NR (no explicit adverse event or safety data reported) | (ROBINS-I) High risk because the study was single-arm and uncontrolled, with strong confounding from chemotherapy, disease course, and regression to the mean, and it had a very small sample (n=20 paired). Multiple exploratory correlations were tested without multiple-testing correction, although objective DNAm measurement suggests low measurement bias. | |
| 96 | Petersen, Curtis L | 2021 | Weight management intervention identifies association of decreased DNA methylation age with improved functional age measures in older adults with obesity. | USA | Human | Single-arm, pre–post 12-week multi-component weight-loss program (diet + aerobic/resistance exercise); pilot substudy with paired DNAm data. | 16 (paired baseline + 12-week samples) | Community-dwelling adults ≥65 y with obesity (BMI >30); mean age 73.5 y, 87.5% women, all White, non-Hispanic. | Lifestyle | 12-week structured weight-management program | 12 weeks; group sessions; exercise dose not fully quantified (reported adherence data only) | None (pre–post only) | Baseline vs post-intervention within-subject comparison | Horvath, Hannum, PhenoAge (via ENmix) | Whole blood | Baseline and 12 weeks | Mean within-group change (post – pre) • Hannum: −0.8 ± 4.8 y (p = 0.51) • Horvath: −1.1 ± 2.8 y (p = 0.14) • PhenoAge: −0.5 ± 4.1 y (p = 0.64) |
NR | NR (no explicit adverse event or safety data reported) | (ROBINS-I) High risk because there was no control group, leaving confounding and regression-to-the-mean concerns, and only 16/28 had paired DNAm data, suggesting possible selection bias (those with blood samples had lower baseline grip). The sample was very small with no correction for multiple testing, although objective DNAm measurement reduces measurement bias. | |
| 103 | Pavanello, Sofia | 2019 | Exploring Epigenetic Age in Response to Intensive Relaxing Training: A Pilot Study to Slow Down Biological Age. | Italy | Human | Prospective before–after pilot (no control). Two groups (both receive intervention): post-MI patients and healthy subjects. Duration: 60 days of twice-daily relaxation practices (after 4 supervised training days). |
20 | 14 MI; 6 healthy | Recent MI patients with carotid atherosclerosis (on standard cardiac rehab + meds) and age/sex-matched healthy adults; Caucasian; smokers more common among MI at baseline but stopped after MI. | Lifestyle | Relaxation Response (RR) training | • 4 days supervised RR sessions • Then 20 min twice daily (morning + evening) at home for 60 days • Both MI and healthy subjects received identical protocol |
None (pre–post only) | Baseline vs post-intervention within-subject comparison | 5-CpG DNAmAge (Zbiec-Piekarska model; ELOVL2, C1orf132, KLF14, TRIM59, FHL2) | Whole blood | Baseline (T0) and 60 days (T1) | Mean change (T1–T0) • All subjects: −1.50 ± 4.36 y (p = 0.143) • MI patients: −0.14 ± 2.88 y (p = 0.428) • Healthy subjects: −4.67 ± 5.78 y (p = 0.053) |
NR | NR (no explicit adverse event or safety data reported) | (ROBINS-I) High risk because there was no control group and MI patients were also undergoing cardiac rehabilitation and medication changes, so time effects and confounding cannot be separated. The sample was extremely small (especially the healthy group, n=6), results relied on borderline p-values without multiplicity correction, and although objective DNAm and qPCR assays reduce measurement bias, these design limitations remain substantial. |
| 107 | Wang, Shuyue | 2025 | Nucleotides as an Anti-Aging Supplementation in Older Adults: A Randomized Controlled Trial (TALENTs study). | China | Human | 19-week, double-blind, randomized, placebo-controlled parallel RCT | 121 | 59 NTs, 62 placebo | Community adults 60–70 y, generally healthy; ~66% female. Balanced comorbidities (HTN 30%, diabetes 14%, CVD 54%, renal disease 40%). Extensive COVID-19 infections during trial (~75–80% both arms by midpoint). | Exogenous compound–based | Mixed 5′-NTs supplementation | • 1.2 g/day total NTs (4 × capsules/day) • Each capsule: 0.3 g NTs + 0.1 g starch • Composition (modeled on breast milk): AMP:CMP:GMP:UMP = 16:41:19:24 • Duration 19 weeks |
Placebo control | Placebo capsules (0.4 g starch) | PC-corrected Horvath, Hannum, GrimAge, PhenoAge, plus Median DNAmAge = clock-median of the four PC ages (primary) |
Whole blood | Baseline (T0), week 11 (T1), week 19 (T2) | NTs vs placebo Δ at 19 weeks: β = −3.08 years (95% CI −5.07, −1.10), p = 0.0023 → ~3-year epigenetic age reduction vs placebo over ~4.5 months. Midpoint (week 11): β = −1.94 y (p = 0.11) — trend only. |
NR | Transient rise in uric acid at week 11 (+31.6 µmol/L, p<0.001), attenuated at week 19 | (RoB 2) Overall low risk: randomization, allocation concealment, and blinding were appropriate, attrition was very low with ITT GEE analyses, and objective molecular outcomes were assessed by blinded lab staff, limiting measurement bias. Some concerns remain due to sponsor involvement (supplement provided), while COVID infections may add noise but were balanced across arms. |
| 118 | Gensous, Noemie | 2020 | One-year Mediterranean diet promotes epigenetic rejuvenation with country- and sex-specific effects: a pilot study from the NU-AGE project. | Italy and Poland | Human | 1-year randomized controlled dietary intervention (NU-AGE RCT), but epigenetic substudy includes only intervention arm → uncontrolled pre–post design for DNAm outcomes | 120 | Single group | Older adults 65–79 y, community-dwelling, generally healthy (no cancer, severe organ disease, dementia, frailty); Italy 27M/33F; Poland 24M/36F | Lifestyle | NU-AGE individualized Mediterranean-style diet: high vegetables, fruits, legumes, whole grains, olive oil; low red meat; age-tailored micronutrients | 12-month continuous dietary counseling + adherence monitoring (NU-AGE score ↑ from ~52 to 65–67) | None for epigenetic outcomes (control arm not included in methylation analysis) | N/A — pre–post only | Horvath DNAmAge (pan-tissue) and its acceleration measures: AgeAccel, IEAA (intrinsic), EEAA (extrinsic) | Whole blood | Baseline (T0) and 12 months (T1) | AgeAccel Italy (all): NS (p=0.182) Italy males: NS trend (p=0.063) Italy females: NS Poland (all): −AgeAccel (p=0.031; adj p=0.094) Poland females: significant rejuvenation → p=0.0013; BH adj p=0.008 Poland males: NS IEAA Italy (all): small decrease (p=0.035; adj p=0.104) Poland females: significant decrease (adj p=0.042) Others: NS EEAA No significant effects in any subgroup |
NR | NR (no explicit adverse event or safety data reported) | (RoB 2) High risk for causal interpretation because the methylation analysis lacked a control arm, appears limited to the intervention arm (possible selection bias), used small subgroups, and involved multiple tests with modest effect sizes. Measurement bias is likely low given objective array-based assays. |
| 123 | Obeid, R | 2018 | Effect of adding B-vitamins to vitamin D and calcium supplementation on CpG methylation of epigenetic aging markers. | Germany | Human | 12-month double-blind randomized controlled trial; two arms (D+Ca vs D+Ca+B); this is a secondary epigenetic analysis (n=63) using targeted CpG methylation and the Weidner 3-CpG clock | 63 | D+Ca: 31 D+Ca+B: 32 |
Older adults, mean age 68.4 ± 10.1 y; majority female (imbalance toward B-vitamin group); community-dwelling; screened to exclude major illness or medication changes | Exogenous compound–based | Vitamin D3 + Calcium + B-vitamins (folate, B6, B12) | Daily × 12 months – Both arms: Vitamin D3 1200 IU/day, Calcium carbonate 800 mg/day – B-vitamin arm only: Folic acid 0.5 mg, Pyridoxine 50 mg, Cyanocobalamin 0.5 mg (3 capsules/day) |
Active control | Vitamin D3 + Calcium only (B-vitamin capsules replaced with inert filler) | Weidner 3-CpG clock (ASPA, ITGA2B, PDE4C) via pyrosequencing | Whole blood | Baseline and 12 months | CpG-specific methylation changes ASPA (normally decreases with age): – D+Ca: −0.96% – D+Ca+B: +1.40% → significant between-group effect (p=0.046) → younger PDE4C (normally increases with age): – D+Ca: +0.22% – D+Ca+B: +1.95%, trend (p=0.062) → older |
NR | NR (no explicit adverse event or safety data reported) | (RoB 2) High risk for epigenetic aging inference due to the small sample, sex/age imbalance between arms, and reliance on a semi-validated 3-CpG clock with internal calibration (risk of circularity). Additional concerns include lower pyrosequencing precision for PDE4C, small effect sizes, and borderline p-values. |
| 148 | Horvath, Steve | 2025 | Cognitive rejuvenation in old rats by hippocampal OSKM gene therapy. | USA and Argentina | Rat (female Sprague–Dawley) | In vivo gene-therapy intervention; 3 groups: young intact, old control (GFP vector), old OSKM vector; ~39-day expression; behavioral + DNAm outcomes; non-randomized, parallel-group | 20 hippocampal samples (6 young, 6 old control, 8 old OSKM) | Young 6; Old control 6; Old OSKM 8 | Female SD rats; young 3.5 mo; old 25.3 mo; standard housing; ad libitum food/water | Celular reprogrammation | Bilateral stereotaxic delivery of OSKM-GFP high-capacity adenovector (Tet-Off) into dorsal dentate gyrus | Single bilateral 3 µL injection per hemisphere; OSKM expression for 39 days; GFP-only vector for controls | Active (GFP-only) + age comparison to young intact | Old controls received identical surgery and GFP-only vector; young rats left intact | Rat brain DNAmAge clock; human–rat relative age clock; mouse brain clock (MammalMethylChip40) | Hippocampus | Single terminal measure after 39 days | All comparisons: old OSKM vs old controls Rat brain clock – DNAmAge slightly younger in OSKM vs control – p = 0.064 (two-sided), p < 0.05 one-sided Human–rat relative age clock – OSKM slightly younger – p = 0.076 (two-sided) Mouse brain clock – OSKM slightly younger – p = 0.088 (two-sided) Estimated magnitude: <1 “rat-year”. |
NR | – No seizures, teratoma-like lesions, or behavioral toxicity observed – Long-term safety unknown |
(RoB 2) High risk (exploratory) because allocation was not clearly randomized, blinding of behavioral scoring was not reported, sample sizes were small, and the study involved multiple hypothesis testing (clocks, chromatin states, EWAS) over a short duration. |
| 149 | Horvath, Steve | 2023 | Reversal of Biological Age in Multiple Rat Organs by Young Porcine Plasma Fraction. | India (treatment), with collaborators in USA, Argentina, Croatia | Rat (male Sprague–Dawley; replication includes both sexes) | Main experiment: non-randomized 3-arm parallel-group (young vs old control vs old + E5); 155-day intervention with terminal multi-organ DNAm profiling Replication experiment: randomized E5 vs saline in 26-month-old rats (blood DNAm only) |
Main experiment: 6 young, 6 old control, 6 old + E5 (multi-organ) Replication: 9 E5 vs 8 saline (blood) |
Young 6; Old control 6; Old + E5 6 (main) E5 9; Saline 8 (replication) |
Old male SD rats (109 weeks ≈ 2+ years old); young reference 30 weeks; replication cohort 26-month mixed-sex SD rats | Blood-derived | E5 — exosome-rich plasma fraction from 6-month-old pigs (Yorkshire breed) | Two IV series, each consisting of 4 injections on alternate days (8 total), separated by 95 days; tail-vein administration. | Active (old untreated control) + age reference (young) | Old control rats received saline; young rats untreated | Rat pan-tissue clock; rat blood clock; rat liver clock; rat brain clock; human–rat pan-tissue clocks (absolute & relative age) | Whole blood | Single terminal timepoint at day 155 (main); baseline + day 15 (replication) | Liver: −77.6% epigenetic age Blood: −68.2% Heart: −56.5% Hypothalamus: −29.6% Mean across 4 organs: −67.4% (≈ halving of biological age) Clock ranges (original models): liver 68.6–78.6%; blood 52.5–74.5%; heart 46.5%; hypothalamus 24.4%. Replication experiment (baseline → day 15) 26-month-old rats: Rat blood clock: significant rejuvenation (p = 0.0094) Rat pan-tissue clock: trend (p = 0.054), significant after removing 1 outlier (p = 0.00086); significant in females (p = 0.014) |
NR | No overt toxicity observed over 155 days | (RoB 2) High risk (exploratory) because the main study was not explicitly randomized or blinded, group sizes were very small (n=6), there were many endpoints with multiple comparisons, and generalizability is limited by species specificity. Despite this, the reported effects were large, consistent, and replicated across tissues, clocks, and a second cohort. |
| 150 | Chiavellini, Priscila | 2024 | Young Plasma Rejuvenates Blood DNA Methylation Profile, Extends Mean Lifespan, and Improves Physical Appearance in Old Rats. | Argentina | Rat (female Sprague–Dawley) | Non-randomized 2-arm longitudinal intervention in very old rats, with q2-week blood DNAm until natural death; additional younger reference cohorts for clock trajectory | 17 old rats: Control 8, Treated 9 (187 total repeated blood samples) |
25.6-month-old female SD rats (late-life), housed standard conditions; age-reference cohorts at 3.7, 8, 15.7 months used for DNAm-age curve only | Blood-derived | Biweekly young-plasma injections (from 2-month-old donors) | 1 mL plasma intraperitoneal every 2 weeks, from 25.6 months until death | Untreated old control group | Control rats handled but did not receive plasma | Horvath rat blood DNAm-age clock | Whole blood | Repeated q2-week blood draws from 25.6 months → death; aggregated into age bands 27–31.5 mo and 32.5–35.5 mo for primary comparison | Treated vs control: Immediate post-intervention: DNAm age lower in treated rats at nearly all timepoints (NS per-timepoint). Primary positive finding (age-banded): 27–31.5 mo: NS 32.5–35.5 mo: Significantly lower DNAm age in treated rats (~3–4 months younger, p<0.05) |
NR | NR (no explicit adverse event or safety data reported) | (RoB 2) High risk because the study was non-randomized with a small sample, unblinded design, and subjective survival and appearance outcomes, with exploratory DMC analysis; allocation and detection biases are therefore high, although DNAm measurement itself presents low risk. | |
| 219 | Izquierdo, A G | 2025 | Epigenetic Aging Acceleration in Obesity Is Slowed Down by Nutritional Ketosis Following Very Low-Calorie Ketogenic Diet (VLCKD): A New Perspective to Reverse Biological Age | Spain | Human | Mixed design: Cross-sectional (obesity vs normal-weight) Single-arm longitudinal VLCKD (n=10 obese adults), no control, pre–post at 0 / 30 / 180 days |
Cross-sectional: 48 (28 obese, 20 normal-weight) VLCKD: 10 obese adults |
Adults with obesity (BMI>30), mean age ~49 y; European Caucasian; excluded recent major weight change, meds except antidiabetics; both sexes | Lifestyle | PNK® Very Low-Calorie Ketogenic Diet (VLCKD), structured 5-stage commercial program | 6-month diet: strict ketogenic phase → stepwise food reintroduction Assessments: baseline (BL), nutritional ketosis (NK ~day 30), endpoint (EP ~day 180) |
Pre–post within-subject (no external control) | Baseline vs NK vs EP within same individuals | Horvath, Hannum, Levine PhenoAge Outcome = AgeAccel (DNAmAge − chronological age) |
Whole blood | Day 0 (BL), Day 30 (NK), Day 180 (EP) | Cross-sectional (obesity vs normal weight) Obesity: AgeAccel ≈ +4.4 y Normal weight: AgeAccel ≈ −3.1 y All clocks significantly higher AgeAccel in obesity (p<0.0001) Strong BMI correlation (r=0.76–0.81) |
NR | NR (no explicit adverse event or safety data reported) | (ROBINS-I) High risk because there was no control group for epigenetic outcomes and the sample was very small (n=10), so regression to the mean and time effects cannot be ruled out. Multiple exploratory metabolic and ketone correlations with limited multiplicity correction and industry involvement (PNK® advisory relationships) add further bias concerns, although standardized DNAm methods suggest low measurement bias. | |
| 244 | Reading, C L | 2025 | An exploratory analysis of bezisterim treatment associated with decreased biological age acceleration, and improved clinical measure and biomarker changes in mild-to-moderate probable Alzheimer’s disease. | USA and Argentina | Human | Phase 3 randomized, double-blind, placebo-controlled parallel-group trial (30 weeks); epigenetic analysis = exploratory subset with post-treatment single timepoint only (between-group comparison, no baseline DNAm) | 33 (17 bezisterim, 16 placebo) | Bezisterim 17; Placebo 16 (per-protocol completers with QC-passing DNAm) | Age 60–85 y; probable AD by NIA-AA 2011; CDR global 1–2; MMSE 14–24; MRI/CT excluding other pathology; balanced baseline demographics and metabolic/cognitive profiles between arms | Exogenous compound–based | Bezisterim (NE3107/HE3286) – 17α-ethynyl-androst-5-ene-3β,7β,17β-triol | 20 mg oral BID for 30 weeks (≈210 days) | Placebo | Matching placebo, BID, same schedule and procedures | SBCAge (Horvath Skin&Blood), PhenoAge, GrimAge, Hannum, InflammAge | Whole blood | Single post-treatment sample at completion (day 150–210); no baseline DNAm | Outcome metric: dAge = Biological Age − Chronological Age (years); negative = biologically younger. Values below are bezisterim − placebo: SBCAge: −3.68 y, p = 0.017 PhenoAge: −3.71 y, p = 0.081 (trend) GrimAge: −1.92 y, p = 0.068 (trend) Hannum: −5.00 y, p = 0.006 InflammAge: −4.77 y, p = 0.022 |
NR | Any TEAE: Placebo 72.7% (24/33) vs Bezisterim 62.5% (15/24) Headache: more common with bezisterim (12.5% vs 0%) Treatment-related TEAEs: Bezisterim 12.5% (3/24) vs Placebo 18.2% (6/33) SAEs: 1 in bezisterim (pneumonia, not related); 7 events in 3 placebo participants (none related) Discontinuations due to AE: 3 in placebo, 0 in bezisterim |
(RoB 2) Some concerns because the epigenetic subset was small and restricted to selected completers (33/439), potentially healthier or late-enrolled participants, raising possible selection bias. Measurement bias is likely low given standard EPIC v2.0 assays with explicit batch processing, while BioVie’s design and sponsorship may introduce interpretive bias despite transparent, explicitly exploratory analyses. |
| 314 | Perlmutter, A | 2024 | The impact of a polyphenol-rich supplement on epigenetic and cellular markers of immune age: a pilot clinical study | USA | Human | Single-arm, open-label pre–post pilot; no control group; 90-day supplementation; paired DNAm baseline → day 90 | 47 with paired DNAm; per-protocol n=40 for primary analysis | Single-arm (n=40 PP) | Generally healthy adults, age 18–85 (mean 54 ± 11 y), BMI 24.2 ± 3.3, 40% male; excluded BMI ≥40, major recent lifestyle changes; recruited remotely across US | Exogenous compound–based | HTB Rejuvenate (Himalayan Tartary buckwheat–based polyphenol formulation) | 4 capsules/day (2 AM, 2 PM with food) × 90 days; | None (pre–post only) | Baseline DNAm → day-90 DNAm within same participants | OMICmAge, PCPhenoAge, PCGrimAge v1, DunedinPACE | Whole blood | Baseline (day 0) and day 90 | OMICmAge EAA: NS (p=0.740) PCPhenoAge EAA: NS (p=0.690) PCGrimAge EAA: NS (p=0.320) |
DunedinPACE: No significant change reported | NR (no explicit adverse event or safety data reported) | (RoB 2) High risk of bias due to the open-label design with no comparator, small sample size, subgroup analyses without multiplicity correction, sponsor involvement, and uncontrolled lifestyle confounding. |
| 350 | Zhang, B | 2023 | Multi-omic rejuvenation and life span extension on exposure to youthful circulation | USA | Mouse (female C57BL/6J) | Long-term heterochronic parabiosis (old 20 mo + young 3 mo) vs old isochronic controls; 3 months parabiosis, with attached and 2-month detached cohorts; also 5-week short-term HPB; includes multi-omic profiling + longevity | Typically 5–7 per group per tissue per platform (RRBS & array) | Old female C57BL/6J (20 mo) paired with young 3 mo | Blood-derived | 3-month heterochronic parabiosis (vs old–old isochronic) | Old isochronic (O:O) | – RRBS clocks: 2 multi-tissue (Meer, Thompson), 1 blood clock (Petkovich), 1 scAge – Array clocks: Universal relative-age, Universal log-linear, Mouse liver, Mouse liver developmental | Blood and Liver | End of 3 mo attachment; 2 mo post-detachment; short-term 5-week HPB | Blood (post-detachment): ~16–32% younger epigenetic age Liver: • Attached: ~17–27% younger (arrays), ~5–26% (RRBS) • Detached: ~11–26% younger (sustained rejuvenation) Short-term 5-week HPB: 0–11% reduction, often non-significant Interpretation: Robust, multi-platform evidence of systemic epigenetic rejuvenation, durable after detachment. |
Surgical risk; no long-term toxicity reported | NR (no explicit adverse event or safety data reported) | (RoB 2) High risk (exploratory) due to the preclinical surgical model with small sample size, but the reported rejuvenation effects were highly consistent across eight clocks, multiple tissues, and omics layers. | |||
| 364 | Waziry, R | 2023 | Effect of long-term caloric restriction on DNA methylation measures of biological aging in healthy adults from the CALERIE trial | USA | Human | 24-month randomized controlled trial (2:1 CR vs ad-libitum), post-hoc DNAm aging analysis; n=197 with DNAm | 128 CR, 69 AL | Healthy non-obese adults, age 21–50 (men) / 21–47 (women), BMI 22–27.9 | Lifestyle | 25% caloric restriction (achieved ≈12%) for 2 years | Ad-libitum diet (no CR) | PC PhenoAge, PC GrimAge, DunedinPACE (primary) | Whole blood | Baseline, 12 mo, 24 mo | PC PhenoAge: No CR vs AL difference (d = −0.03 @12 mo; +0.05 @24 mo; NS) PC GrimAge: No difference (d = −0.04 @12 mo; +0.05 @24 mo; NS) DunedinPACE: Significant slowing 12 mo: d = −0.29, P < 0.003 24 mo: d = −0.25, P < 0.003 ≈ 2–3% reduction in pace of aging TOT (20% CR): DunedinPACE d ≈ −0.40 to −0.43 (stronger effect) |
Not reported in this paper; parent CALERIE RCT showed acceptable 2-year safety | NR (no explicit adverse event or safety data reported) | (RoB 2) Low to moderate risk: the study used a proper RCT design with a high-quality DNAm assay, but the epigenetic analysis was post hoc, adherence was low (<25%), and the population was a healthy, selective sample. | |||
| 436 | Horvath, S | 2021 | DNA methylation age analysis of rapamycin in common marmosets | USA | Non-human primate (common marmoset) | Chronic rapamycin vs vehicle cross-sectional comparison after ~2–3.5 y exposure; not randomized | 37 | 20 control, 17 rapamycin | Middle-aged captive marmosets (~9–10 y), relatively healthy | Exogenous compound–based | Rapamycin ~1 mg/kg/day, oral in yogurt, 5 days/week | ~2–3.5 years exposure | Vehicle (Eudragit) | Marmoset blood epigenetic clock (trained in separate 58-sample set) | Whole blood | Single post-treatment sample (no baseline DNAm) | Multivariate regression (DNAmAge ~ age + sex + treatment): Rapamycin coefficient: −0.18 y, p = 0.686 → NS; no effect EWAS: No genome-wide significant CpGs; only 48 CpGs at p<0.005 (exploratory) Rapamycin does not alter blood DNAmAge in these middle-aged marmosets |
NR | NR (no explicit adverse event or safety data reported) | (ROBINS-I) High risk because the study was non-randomized and cross-sectional with no baseline DNAm measurement, limiting causal interpretation, although measurement quality was high. | |
| 457 | Demidenko, O | 2021 | Rejuvant®, a potential life-extending compound formulation with alpha-ketoglutarate and vitamins, conferred an average 8 year reduction in biological aging, after an average of 7 months of use, in the TruAge DNA methylation test | USA | Human | Uncontrolled before–after in supplement customers; no placebo/no randomization | 42 | Healthy adults, mean age ~63 y; low BMI; heavy supplement users | Exogenous compound–based | (Ca-AKG 1 g/day + Vitamin A (men) or D (women)) | 2 tablets/day for 4–10 months (mean ~7 months) | None | 9-CpG targeted Sanger (TruMe “TruAge” proprietary clock) | Saliva | Baseline → ~7 months | Full cohort: Mean TruAge −7.96 y (biological age decrease ≈8 y) Lifestyle-stable subset (n=13): Mean −7.69 y Stats: One-sided paired t-tests (p≈10⁻⁵–10⁻¹²); no control group Very large apparent rejuvenation by proprietary clock; cannot infer causality |
NR | NR (no explicit adverse event or safety data reported) | (ROBINS-I) Very high risk because there was no control group, the study relied on a proprietary 9-CpG saliva clock, and there were concerns about selection bias and sponsor involvement. | ||
| 509 | Wang, T | 2017 | Epigenetic aging signatures in mice livers are slowed by dwarfism, calorie restriction and rapamycin treatment | USA and UK | Mouse | Preclinical comparison of long-lived models vs age-matched WT; n=4 per group | 32 | ~4/group × 8 groups across ages & interventions | Ames dwarfs (male), UM-HET3 females; 2 mo & 22 mo | Exogenous compound–based | Prop1df/df dwarfism, 40% calorie restriction, rapamycin 42 mg/kg diet (4–22 mo) | Genetic; CR at 60% of ad-lib; rapamycin lifelong (4→22 mo) | Age-matched WT (same strain) | Mouse liver DNAm age (ElasticNet, 148 CpGs) | Liver | 2 mo and 22 mo | Ames dwarf: −10.1 months epigenetic age Calorie restriction: −9.4 months Rapamycin: −6.0 months Young dwarfs (2 mo): −1.5 mo (developmentally younger) All significant (p < 0.05–10⁻⁴) |
NR | NR (no explicit adverse event or safety data reported) | (ROBINS-I) Moderate to high risk due to the very small sample size (n=4) and lack of randomization for DNAm outcomes, although the direction of effects was consistent across all interventions. | |
| 515 | Reynolds, Lindsay | 2025 | A tree nut and extra virgin olive oil intervention to improve cardiometabolic health – a feasibility study incorporating epigenetic aging | USA | Human | Single-arm 4-week dietary feasibility study with randomized education vs no-education sub-comparison; no diet control group | 29 | Adults 48–81 y (mean 68) with metabolic syndrome (≥3 MetS criteria) | Lifestyle | Daily 1 oz tree nuts + 2 Tbsp EVOO | Nuts 1 oz/day; EVOO 2 Tbsp/day; 4 weeks | Within-subject pre–post (no dietary control) | DunedinPACE, AgeAccelGrim | Whole blood | Baseline, 4 weeks | DunedinPACE: Δ = −0.002 ± 0.070, p = 0.86 (NS) AgeAccelGrim: Δ = −0.04 ± 1.34, p = 0.89 (NS) Between-arm (education vs no education): not analyzed for DNAm outcomes Baseline: DunedinPACE mean 1.179 (all >1.0 → accelerated aging in MetS) |
NR | NR (no explicit adverse event or safety data reported) | (ROBINS-I) High risk because there was no control group, the sample was very small, the duration was only 4 weeks, and the feasibility design relied on pre–post comparisons only. | ||
References
- López-Otín C, Blasco MA, Partridge L, Serrano M, Kroemer G. Hallmarks of aging: An expanding universe. Cell. 2023;186(2):243-278. [CrossRef]
- López-Otín C, Blasco MA, Partridge L, Serrano M, Kroemer G. The Hallmarks of Aging. Cell. 2013 Jun;153(6):1194–217.
- Sen P, Shah PP, Nativio R, Berger SL. Epigenetic Mechanisms of Longevity and Aging. Cell. 2016 Aug;166(4):822–39.
- Braga DL, Mousovich-Neto F, Tonon-da-Silva G, Salgueiro WG, Mori MA. Epigenetic changes during ageing and their underlying mechanisms. Biogerontology. 2020;21:423-443. [CrossRef]
- Pal S, Tyler JK. Epigenetics and aging. Sci Adv. 2016;2(7):e1600584. [CrossRef]
- Wang K, Liu H, Hu Q, et al. Epigenetic regulation of aging: implications for interventions of aging and diseases. Signal Transduct Target Ther. 2022;7(1):374. [CrossRef]
- Ciccarone F, Tagliatesta S, Caiafa P, Zampieri M. DNA methylation dynamics in aging: how far are we from understanding the mechanisms? Mech Ageing Dev. 2018;174:3-17. [CrossRef]
- Horvath S, Raj K. DNA methylation-based biomarkers and the epigenetic clock theory of ageing. Nat Rev Genet. 2018;19(6):371-384. [CrossRef]
- Belsky DW, Caspi A, Corcoran DL, et al. DunedinPACE, a DNA methylation biomarker of the pace of aging. eLife. 2022;11:e73420. [CrossRef]
- Jones MJ, Goodman SJ, Kobor MS. DNA methylation and healthy human aging. Aging Cell. 2015;14:924-932. [CrossRef]
- Horvath S. DNA methylation age of human tissues and cell types. Genome Biol. 2013;14(10):R115. [CrossRef]
- Levine ME, Lu AT, Quach A, et al. An epigenetic biomarker of aging for lifespan and healthspan. Aging (Albany NY). 2018;10(4):573-591. [CrossRef]
- Lu AT, Binder AM, Zhang J, et al. DNA methylation GrimAge version 2. Aging (Albany NY). 2022;14(23):9484-9549.
- Ulalume Hernández-Arciga, Ceda Stamenkovic, Yadav S, Nicoletti C, Albalawy WN, Farazdaq Al Hammood, et al. Dietary methionine restriction started late in life promotes healthy aging in a sex-specific manner. Science advances. 2025 Apr;11(16).
- Salas-Huetos A, James ER, Jordi Salas-Salvadó, Mònica Bulló, Aston KI, Carrell DT, et al. Sperm DNA methylation changes after short-term nut supplementation in healthy men consuming a Western-style diet. Andrology. 2021 Jan;9(1):260–8.
- Kou M, Li X, Yoriko Heianza, Dorans K, Bazzano L, Qi L. Epigenetic age acceleration and cardiometabolic biomarkers in response to weight-loss dietary interventions among obese individuals: The MACRO trial. Aging cell. 2025 Nov;24(11).
- Izquierdo AG, Lorenzo PM, N Costa-Fraga, Primo D, G Rodríguez-Carnero, Nicoletti CF, et al. Epigenetic aging acceleration in obesity is slowed down by nutritional ketosis following very low-calorie ketogenic diet (VLCKD): A new perspective to reverse biological age. Nutrients [Internet]. 2025;17(6). Available from: [https://www.scopus.com/inward/record.uri?eid=2-s2.0-105001235593&doi=10.3390%2Fnu17061060&partnerID=40&md5=092fee851bef241ad29428ecbd8ef480].
- Noémie Gensous, Paolo Garagnani, Santoro A, Giuliani C, Ostan R, Fabbri C, et al. One-year Mediterranean diet promotes epigenetic rejuvenation with country- and sex-specific effects: a pilot study from the NU-AGE project. GeroScience. 2020 Apr;42(2):687–701.
- Reynolds LM, Howard TD, Langefeld CD, Vitolins MZ. A tree nut and extra virgin olive oil intervention to improve cardiometabolic health – a feasibility study incorporating epigenetic aging. Preprints.org [Preprint]. 2025 Feb 7. [CrossRef]
- R Waziry, Ryan CP, Corcoran DL, Huffman KM, Kobor MS, Kothari M, et al. Effect of long-term caloric restriction on DNA methylation measures of biological aging in healthy adults from the CALERIE trial. Nature Aging [Internet]. 2023;3(3):248–57. Available from: [https://www.scopus.com/inward/record.uri?eid=2-s2.0-85147758471&doi=10.1038%2Fs43587-022-00357-y&partnerID=40&md5=32f05ca439f47daaa689609f53602555].
- Loh KP, Chandrika Sanapala, Jensen-Battaglia M, Rana A, Sohn MB, Watson E, et al. Exercise and epigenetic ages in older adults with myeloid malignancies. European journal of medical research. 2023 May;28(1):180.
- Fiorito G, Saverio Caini, Palli D, Bendinelli B, Saieva C, Ilaria Ermini, et al. DNA methylation-based biomarkers of aging were slowed down in a two-year diet and physical activity intervention trial: the DAMA study. Aging cell. 2021 Oct;20(10).
- Fitzgerald KN, Hodges R, Hanes D, Stack E, Cheishvili D, Szyf M, et al. Potential reversal of epigenetic age using a diet and lifestyle intervention: a pilot randomized clinical trial. Aging. 2021 Apr;13(7):9419–32.
- Meir AY, Keller M, Bernhart SH, Ehud Rinott, Gal Tsaban, Hila Zelicha, et al. Lifestyle weight-loss intervention may attenuate methylation aging: the CENTRAL MRI randomized controlled trial. Clinical epigenetics. 2021;13(1):48.
- Petersen CL, Christensen BC, Batsis JA. Weight management intervention identifies association of decreased DNA methylation age with improved functional age measures in older adults with obesity. Clinical epigenetics. 2021 Mar;13(1):46.
- Pavanello S, Campisi M, Tona F, Lin CD, Sabino Iliceto. Exploring epigenetic age in response to intensive relaxing training: A pilot study to slow down biological age. International journal of environmental research and public health. 2019 Aug;16(17).
- Nwanaji-Enwerem JC, Chung FFL, Van der Laan L, Novoloaca A, Cuenin C, Johansson H, et al. An epigenetic aging analysis of randomized metformin and weight loss interventions: two clinical trials. Clinical Epigenetics. 2021;13(1):180. [CrossRef]
- Reading CL, Yan J, Testa MA, Simonson DC, Javaid H, Schmunk LJ, et al. An exploratory analysis of bezisterim treatment effects on epigenetic age biomarkers in older adults with mild-to-moderate probable Alzheimer’s disease. Aging (Albany NY). 2025;17(4):825-848. [CrossRef]
- Horvath S, Zoller JA, Haghani A, Lu AT, Raj K, Jasińska AJ, et al. DNA methylation age analysis of rapamycin in common marmosets. GeroScience. 2021;43(5):2411-2423. [CrossRef]
- Wang T, Tsui B, Kreisberg JF, Robertson NA, Gross AM, Yu MK, et al. Epigenetic aging signatures in mice livers are slowed by dwarfism, calorie restriction and rapamycin treatment. Genome Biol. 2017;18(1):57. [CrossRef]
- Bischoff-Ferrari HA, Gängler S, Wieczorek M, Belsky DW, Ryan J, Kressig RW, et al. Individual and additive effects of vitamin D, omega-3, and exercise on biological aging in DO-HEALTH. Nature Aging. 2025;5(3):313-326. [CrossRef]
- Vetter VM, Sommerer Y, Kalies CH, Spira D, Bertram L, Demuth I. Vitamin D supplementation is associated with slower epigenetic aging. GeroScience. 2022;44(4):1843-1859. [CrossRef]
- Chanachai Sae-Lee, Corsi S, Barrow TM, Kuhnle GGC, Bollati V, Mathers JC, et al. Dietary intervention modifies DNA methylation age assessed by the epigenetic clock. Molecular nutrition & food research. 2018 Dec;62(23).
- Obeid R, U Hübner, Bodis M, Graeber S, Geisel J. Effect of adding B-vitamins to vitamin D and calcium supplementation on CpG methylation of epigenetic aging markers. Nutrition, metabolism, and cardiovascular diseases : NMCD. 2018 Apr;28(4):411–7.
- McGee KC, Sullivan J, Hazeldine J, Schmunk LJ, Martin-Herranz DE, Jackson T, et al. A combination nutritional supplement reduces DNA methylation age only in older adults with a raised epigenetic age. GeroScience. 2024 Oct;46(5):4333–47.
- Carreras-Gallo N, Dargham R, Thorpe SP, Warren S, Mendez TL, Smith R, et al. Effects of a natural ingredients-based intervention targeting the hallmarks of aging on epigenetic clocks, physical function, and body composition: a single-arm clinical trial. Aging. 2025 Mar;17(3):699–725.
- Demidenko O, Barardo D, Budovskii V, Finnemore R, Palmer FR, Kennedy BK, et al. Rejuvant®, a potential life-extending compound, reduces the epigenetic age in several cell types. Aging. 2021;13(20):24008-24030. [CrossRef]
- Perlmutter A, Bland JS, Chandra A, Malani SS, Smith R, Mendez TL, et al. The impact of a polyphenol-rich supplement on epigenetic and cellular markers of immune age: a pilot clinical study. Frontiers in Nutrition [Internet]. 2024;11. Available from: [https://www.scopus.com/inward/record.uri?eid=2-s2.0-85211133065&doi=10.3389%2Ffnut.2024.1474597&partnerID=40&md5=2faf468d3005a2487e6e5acd5bcd4335].
- Campisi M, Cannella L, Paccagnella O, Brazzale AR, Agnolin A, Grothe T, et al. Unveiling the geroprotective potential of a novel combination of senolytics with natural compounds: A clinical and in vitro study. Biogerontology. 2025;26(2):129-147. [CrossRef]
- Wang S, Song L, Fan R, Chen Q, Fu R, You M, et al. Nucleotides as an anti-aging supplementation in older adults: A randomized controlled trial (TALENTs study). Advanced science (Weinheim, Baden-Wurttemberg, Germany). 2025 Sep;12(33).
- Orr ME, Kotkowski E, Ramirez P, Bair-Kelps D, Liu Q, Brenner C, et al. A randomized placebo-controlled trial of nicotinamide riboside in older adults with mild cognitive impairment. GeroScience. 2024 Feb;46(1):665–82.
- Fuentealba M, Kiprov D, Schneider K, Mu WC, Kumaar PA, Kasler H, et al. Multi-Omics Analysis Reveals Biomarkers That Contribute to Biological Age Rejuvenation in Response to Single-Blinded Randomized Placebo-Controlled Therapeutic Plasma Exchange. Aging Cell. 2025 Aug;24(8):e70103. [CrossRef]
- Borsky P, Holmannova D, Parova H, Horvath S, Sramek P, Brooke RT, et al. Human clinical trial of plasmapheresis effects on biomarkers of aging (efficacy and safety trial). Sci Rep. 2025 Jul 1;15(1):21059. [CrossRef]
- Clement J, Yan Q, Agrawal M, Coronado RE, Sturges JA, Horvath M, et al. Umbilical cord plasma concentrate has beneficial effects on DNA methylation GrimAge and human clinical biomarkers. Aging Cell. 2022 Oct;21(10):e13696. [CrossRef]
- Horvath S, Singh K, Raj K, Khairnar S, Sanghavi A, Shrivastava A, et al. Reversal of Biological Age in Multiple Rat Organs by Young Porcine Plasma Fraction. bioRxiv [Preprint]. 2023 Aug 7:2023.08.06.552148. [CrossRef]
- Chiavellini P, Lehmann M, Gallardo MD, Canatelli Mallat M, Pasquini DC, Zoller JA, et al. Young plasma rejuvenates blood DNA methylation profile, extends mean lifespan, and improves physical appearance in old rats. J Gerontol A Biol Sci Med Sci. 2024 May 1;79(5):glae071. [CrossRef]
- Zhang B, Lee DE, Trapp A, Tyshkovskiy A, Lu AT, Bareja A, et al. Multi-omic rejuvenation and life span extension on exposure to youthful circulation. Nat Aging. 2023 Aug;3(8):948-964. [CrossRef]
- Horvath S, Lacunza E, Canatelli Mallat M, Portiansky EL, Gallardo MD, Brooke RT, et al. Cognitive rejuvenation in old rats by hippocampal OSKM gene therapy. GeroScience. 2025;47(1):809-823. [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the author. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.




