1. Introduction: From Population Screening to Personalized Baseline Medicine
Cancer screening has historically been structured around population-level risk stratification, most commonly defined by age and sex. Mammography, fecal immunochemical testing (FIT), prostate-specific antigen (PSA) measurement, Pap smears, and low-dose computed tomography (LDCT) for lung cancer exemplify this paradigm. These approaches have undoubtedly reduced mortality in selected malignancies, yet they remain fundamentally episodic and threshold-based: disease is typically recognized only after anatomical or biochemical alterations exceed predefined cut-offs. Such a structure implicitly assumes that pathology becomes detectable only once it has crossed a visible boundary.
However, large-scale implementation studies of multi-cancer early detection (MCED) assays have exposed both the promise and the structural limits of this logic. The PATHFINDER study demonstrated the feasibility of blood-based multi-cancer detection in asymptomatic populations, but also underscored the complexity of resolving positive findings in the absence of clear anatomical correlates [
1]. Similarly, circulating cell-free DNA (cfDNA)-based assays have shown high specificity and encouraging sensitivity across tumor types in both screening and diagnostic contexts [
2,
3,
4]. Yet their performance remains strongly influenced by tumor fraction, stage, and biological heterogeneity [
5]. Even highly sophisticated assays remain constrained by the biological fact that early tumors shed minimal material into circulation. A single time-point measurement, no matter how refined, captures only a transient biological state within a continuous evolutionary process.
Cancer itself unfolds longitudinally. Genomic analyses reveal that driver mutations may accumulate years before clinical manifestation [
6]. During this preclinical phase, somatic alterations, epigenetic drift, clonal expansions, and microenvironmental remodeling progressively reshape tissue biology. Against this backdrop, a screening strategy based exclusively on isolated measurements appears inherently limited. It asks whether a deviation is already large enough to be detected, rather than whether change is underway.
A conceptual shift is therefore emerging in oncology: instead of comparing individuals to population reference ranges, it may be more informative to compare each individual to their own biological baseline. If biomarker trajectories are established early in adulthood—when disease prevalence is lowest—subtle but persistent deviations might become detectable before overt threshold crossing occurs [
7]. This perspective mirrors preventive cardiology, where longitudinal patterns of lipid levels, blood pressure, and inflammatory markers provide more meaningful risk estimation than single-point values [
8]. The central idea is not simply earlier testing, but more context-aware interpretation.
Traditional tumor markers illustrate the implications of this reframing. Analytes such as CEA, CA-125, CA19-9, PSA, and AFP have long been criticized for insufficient sensitivity and specificity when used in isolation for screening. Yet their limited performance may reflect interpretive strategy rather than intrinsic inadequacy. Each individual maintains a relatively stable physiological set point shaped by genetics, metabolism, immune tone, and environmental exposure. A modest but sustained upward drift from this personal baseline—even if still within laboratory reference intervals—may carry more biological meaning than a single value that transiently exceeds a population threshold.
Evidence supporting dynamic interpretation already exists. In ovarian cancer, longitudinal CA-125 monitoring interpreted through risk algorithms improved detection performance compared with fixed cut-offs [
9]. In prostate cancer, PSA velocity and doubling time often provide greater prognostic insight than absolute PSA concentration alone [
10]. These examples suggest that trajectory conveys information that magnitude alone cannot capture.
As understanding of tumor biology has deepened, it has become clear that malignancy perturbs multiple biological systems simultaneously, including inflammatory signaling, metabolic regulation, angiogenesis, and immune modulation. This multidimensional disruption provides the rationale for multi-analyte platforms that integrate circulating proteins with ctDNA mutations or methylation patterns. Studies such as CancerSEEK have demonstrated that combining analytes improves specificity relative to single markers [
11], and cfDNA methylation classifiers have shown the capacity to detect multiple tumor types with high specificity [
2]. Nevertheless, most of these approaches remain anchored to cross-sectional sampling. The next conceptual advance lies in training algorithms not only on absolute values but on within-person temporal change, allowing artificial intelligence systems to detect coordinated drift across biomarker arrays that would otherwise remain clinically silent [
12].
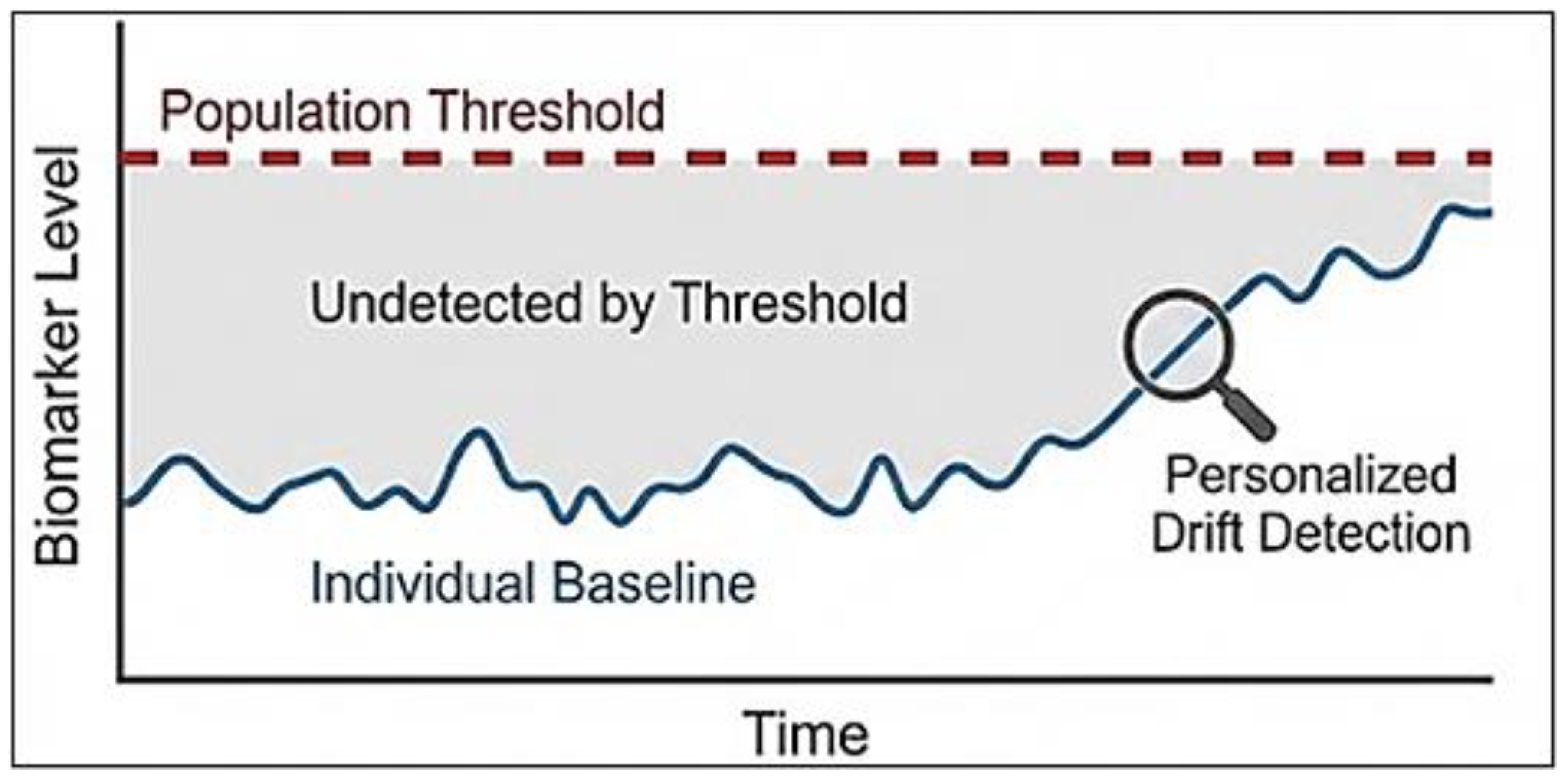
Initiating biomarker mapping early in adulthood offers several theoretical advantages. At a time when cancer prevalence is low, longitudinal data can establish individualized variance patterns and define the boundaries of normal biological fluctuation. In such a framework, early detection ceases to be a binary event triggered by threshold crossing and instead becomes an inference drawn from statistically meaningful deviation. The question shifts from “Is this value abnormal?” to “Has this individual’s biological trajectory diverged from its stable norm?” (
Figure 1).
Transitioning from population-based screening to personalized baseline medicine will require methodological rigor, standardized sampling protocols, and careful management of false positives and overdiagnosis. Yet recent advances in multi-analyte blood testing and risk-adapted surveillance strategies [
2,
11,
13] suggest that oncology is approaching a structural transformation. Early detection may increasingly be understood not as a moment of discovery, but as a process of calibrated longitudinal interpretation—an approach that aligns biological evolution with dynamic monitoring rather than static thresholds (
Table 1).
2. Germline Susceptibility: Predicting First and Subsequent Events
If longitudinal biomarker modeling reframes early detection as a process of identifying biological drift, germline susceptibility reshapes it as a problem of prior probability. The integration of inherited genetic information into oncology has transformed cancer from a purely somatic event into a systemic risk condition that unfolds across the life course. In this perspective, the first tumor is not simply an isolated pathological episode but may represent the earliest clinically visible manifestation of a deeper constitutional vulnerability.
Large population-based sequencing efforts have demonstrated that pathogenic germline variants are present in approximately 8–12% of patients with solid tumors, even in the absence of a striking family history [
14]. This observation has led to the development of “tumor-to-germline” strategies, in which somatic findings identified during therapeutic profiling prompt confirmatory germline testing and structured long-term surveillance. The diagnostic pathway thus acquires a bidirectional quality: treatment-oriented molecular testing may uncover inherited predisposition, while germline results recalibrate screening intensity, modality, and follow-up frequency for the individual and potentially for relatives.
Hereditary Breast and Ovarian Cancer (HBOC) syndrome illustrates the life-course implications of this shift. Pathogenic variants in
BRCA1 and
BRCA2 confer cumulative lifetime breast cancer risks approaching 72% and 69%, respectively, by age 80, with substantial risks of ovarian and contralateral breast cancer [
15]. Importantly, risk persists even after successful treatment of an initial tumor, underscoring that the biological context remains altered. The clinical implications extend beyond breast and ovarian malignancies, as carriers also demonstrate elevated risks of pancreatic and prostate cancers [
16]. These data have prompted the development of integrated prevention frameworks that combine earlier MRI-based breast screening, risk-reducing salpingo-oophorectomy, and, in selected contexts, pancreatic surveillance [
17].
The therapeutic landscape has further reinforced this connection between treatment and prevention. PARP inhibitors exploit synthetic lethality in homologous recombination–deficient tumors, offering effective targeted therapy in BRCA-associated malignancies [
18]. Yet identification of a BRCA alteration during tumor profiling often leads to germline confirmation testing, thereby transforming a therapeutic decision into an opportunity for anticipatory surveillance. The biological information extracted from the tumor thus becomes a gateway to long-term risk prediction.
A similar logic applies to Lynch syndrome, caused by germline pathogenic variants in mismatch repair (MMR) genes including
MLH1,
MSH2,
MSH6, and
PMS2 [
19]. Universal screening of colorectal cancers for MMR deficiency, typically through immunohistochemistry or microsatellite instability testing, has enabled systematic identification of patients requiring germline confirmation. Individuals with Lynch syndrome face markedly elevated lifetime risks of colorectal, endometrial, gastric, small bowel, pancreatic, and urothelial cancers. Even after segmental colectomy, the risk of metachronous colorectal cancer remains significant, justifying intensified colonoscopic surveillance [
20]. Extra-colonic risks further necessitate structured monitoring strategies [
21]. In this context, molecular characterization of a tumor does not merely inform immunotherapy eligibility; it reveals a constitutional repair defect that reshapes the patient’s long-term screening trajectory.
Beyond these high-penetrance syndromes, contemporary genomic practice increasingly identifies actionable variants in additional susceptibility genes. Pathogenic
CDH1 variants define hereditary diffuse gastric cancer (HDGC), associated with high lifetime risks of diffuse gastric carcinoma and lobular breast cancer [
22]. Because early diffuse lesions are difficult to detect endoscopically, prophylactic total gastrectomy remains the most reliable preventive strategy in high-risk carriers [
23]. Here, germline information directly dictates preemptive intervention rather than intensified surveillance alone.
Similarly, pathogenic variants in
STK11 underlie Peutz–Jeghers syndrome (PJS), characterized by gastrointestinal hamartomatous polyps and elevated risks of pancreatic, gastrointestinal, breast, and gynecologic cancers. Multimodal surveillance strategies, including early pancreatic imaging, are recommended to mitigate these risks [
24]. These examples underscore how germline findings recalibrate both the timing and the intensity of early detection.
The growing use of multigene panel testing has also expanded the recognition of moderate-risk genes such as
PALB2,
CHEK2, and
ATM. Interpreting these variants often requires integration of polygenic risk scores (PRS), which aggregate the effects of numerous common alleles to refine risk estimation. Large association analyses have demonstrated that PRS can stratify individuals within the general population to risk levels comparable to those conferred by certain high-penetrance variants [
25,
26]. As summarized in
Table 2, this broader genetic architecture transforms early detection into a graded, probability-based framework rather than a binary classification of “hereditary” versus “sporadic.”
Within the integrated architecture proposed in this manuscript, germline genetics functions as a dynamic prior that modulates surveillance thresholds and testing frequency. Individuals with high-penetrance variants may require earlier and more frequent imaging or molecular testing, whereas those with lower inherited risk may be appropriately monitored with less intensive strategies. The first tumor, when it occurs, becomes a biological inflection point that refines this prior and informs subsequent surveillance logic.
Thus, germline susceptibility extends the concept of early detection beyond the identification of nascent tumors. It situates cancer risk within a life-course continuum, in which inherited vulnerability shapes both the probability of first occurrence and the likelihood of subsequent events. When integrated with longitudinal biomarker modeling and somatic evolutionary analysis, germline data help transform early detection from an age-triggered intervention into a biologically calibrated strategy.
3. Somatic Evolution and Field Cancerization
If germline susceptibility defines inherited vulnerability, somatic evolution describes the biological terrain upon which malignant transformation unfolds. Cancer rarely emerges as an isolated cellular accident. Rather, it represents the culmination of a prolonged evolutionary process occurring within a tissue environment that has gradually accumulated genetic and epigenetic alterations. The concept of field cancerization, first articulated by Slaughter in 1953 in the context of oral squamous epithelium [
27], proposed that large areas of histologically normal tissue might already harbor pre-neoplastic changes. What began as a morphologic observation has since acquired molecular confirmation.
The advent of next-generation sequencing has revealed that clonal expansions carrying canonical driver mutations are detectable in ostensibly normal tissues years before clinical malignancy becomes apparent [
28]. Analyses leveraging data from the Genotype-Tissue Expression (GTEx) project demonstrated that somatic mutations accumulate across multiple normal tissues and increase with age and environmental exposure [
29,
30]. These findings fundamentally alter our understanding of carcinogenesis: the tumor mass is often only the most advanced clone within a broader field of genetically conditioned cells.
Such observations suggest that early detection cannot be limited to identifying a discrete anatomical lesion. Instead, it must consider the evolutionary dynamics of tissue ecosystems that progressively drift toward malignant potential.
Mutational signature analysis has provided a mechanistic framework for understanding these dynamics. Comprehensive whole-genome analyses by the Pan-Cancer Analysis of Whole Genomes (PCAWG) consortium cataloged more than forty reproducible mutational signatures reflecting distinct endogenous and exogenous processes [
31]. Some signatures correspond to well-defined carcinogenic exposures, such as ultraviolet radiation or tobacco smoke, whereas others reflect defective DNA repair pathways, including homologous recombination deficiency or mismatch repair loss. Still others, notably SBS1 and SBS5, represent age-associated processes that accumulate steadily over time.
Crucially, these signatures are not confined to tumors. Ultra-deep sequencing studies have demonstrated that driver mutations, including
TP53,
NOTCH1, and
KRAS, can be detected in histologically normal skin, esophagus, colon, and endometrium, with mutation burden increasing as a function of age [
32,
33,
34]. In sun-exposed epidermis, positively selected clones carrying cancer-associated mutations may occupy substantial fractions of normal tissue [
32]. Such data provide molecular validation of the field cancerization concept: tissues chronically exposed to mutagenic stress accumulate genomic scars long before tumor formation.
Mutational signatures also bridge etiologic insight with therapeutic consequence. Homologous recombination deficiency (HRD) signatures—most classically associated with
BRCA1/2 alterations but also observed in other DNA repair defects—identify tumors sensitive to Poly (ADP-ribose) Polymerase (PARP) inhibition, a therapeutic strategy that exploits defective DNA repair through synthetic lethality [
35]. In parallel, mismatch repair deficiency signatures predict responsiveness to immune checkpoint blockade [
36]. In this way, the same mutational processes that condition tissue fields before tumor emergence later guide treatment selection once malignancy becomes clinically manifest.
Among driver genes,
TP53 occupies a central position in the biology of field evolution. As a key regulator of genomic integrity, p53 coordinates cellular responses to DNA damage, including cell-cycle arrest, DNA repair, and apoptosis [
37,
38]. Loss of p53 function permits survival and expansion of genetically unstable clones, making
TP53 mutation one of the earliest and most frequent events in epithelial carcinogenesis. The detection of identical
TP53 mutations in histologically normal surgical margins of head and neck squamous cell carcinoma predicted local recurrence despite negative pathology, providing one of the earliest molecular demonstrations of clonal persistence beyond visible tumor boundaries [
39]. Comparable findings in Barrett’s esophagus show that
TP53-mutant clones may precede dysplasia and adenocarcinoma by several years [
40].
These observations support the notion of a molecular field memory, whereby repeated cycles of DNA damage and clonal selection gradually reshape epithelial landscapes. Chronic exposure to ultraviolet radiation, tobacco smoke, alcohol, or inflammatory injury does not produce isolated mutations; it creates a mosaic of expanding subclones, each carrying partial elements of malignant potential [
32,
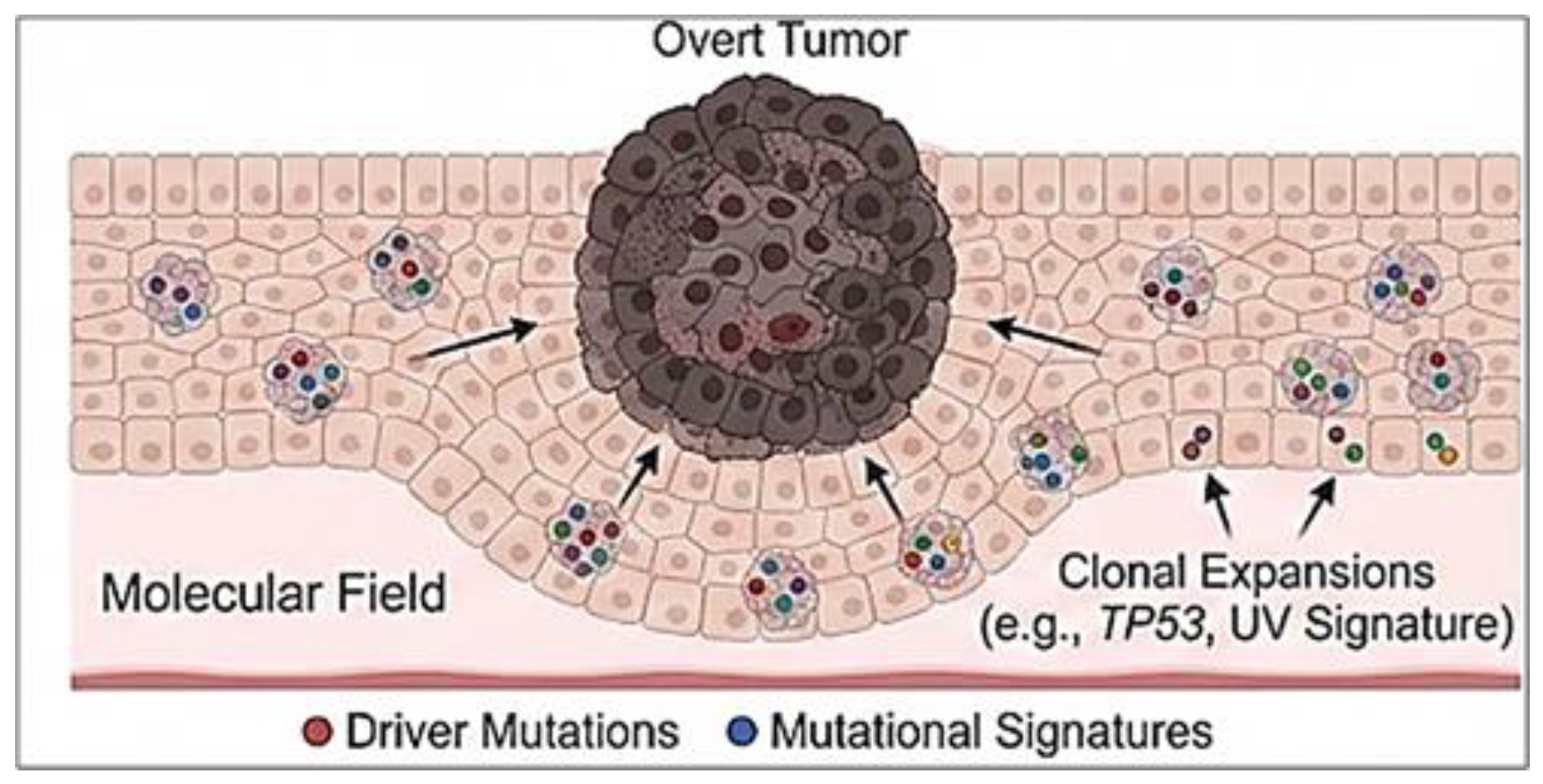
34]. The overt tumor is therefore better understood as the dominant clone emerging from a preconditioned field rather than as a solitary stochastic event. This broader evolutionary architecture is illustrated schematically in
Figure 2.
Recognition of field cancerization carries direct implications for early detection. First, it provides a biological rationale for intensified surveillance in tissues known to harbor widespread clonal alterations. Second, it explains the occurrence of metachronous tumors within the same anatomical region even after apparently complete surgical excision. Third, it suggests that molecular assessment of adjacent or at-risk tissue compartments may eventually refine recurrence risk stratification beyond conventional histopathology.
The phenomenon of clonal hematopoiesis further reinforces this principle. Age-associated expansion of mutant hematopoietic clones can precede overt hematologic malignancy by years and is associated with increased cancer risk [
41]. Although occurring in the blood compartment, clonal hematopoiesis exemplifies the broader concept that somatic evolution within normal tissues may serve as an early harbinger of malignant transformation.
Taken together, somatic evolution and field cancerization extend the early detection paradigm from lesion identification to ecosystem surveillance. Germline genetics defines inherited susceptibility, whereas mutational signatures and driver mutations trace the evolutionary conditioning of tissues exposed to cumulative stress. In this integrated perspective, early detection becomes the recognition of instability within a spatially distributed and temporally evolving biological field.
4. Liquid Biopsy and Molecular Residual Disease
If field cancerization describes the spatial and evolutionary conditioning of tissues before tumor emergence, liquid biopsy introduces a temporal dimension to this framework. The analysis of circulating tumor DNA (ctDNA) has transformed post-treatment surveillance by allowing real-time monitoring of tumor-derived genomic material in the bloodstream [
42]. While cfDNA encompasses all extracellular DNA fragments released from both normal and pathological cellular turnover, ctDNA represents the tumor-derived fraction of this broader pool. Because ctDNA fragments harbor the same somatic mutations as the primary tumor, they function as molecular tracers of clonal persistence [
43].
Unlike imaging modalities, which detect structural lesions only after they reach macroscopic size, ctDNA can reveal residual disease months before radiologic recurrence becomes apparent. In colorectal cancer, the randomized DYNAMIC trial demonstrated that ctDNA-guided postoperative management safely reduced adjuvant chemotherapy use without compromising recurrence-free survival, thereby establishing ctDNA as a clinically actionable marker of molecular residual disease (MRD) [
44]. Similar findings across lung, breast, and bladder cancers have confirmed that postoperative ctDNA detection strongly predicts relapse and often precedes clinical or radiologic progression [
45,
46].
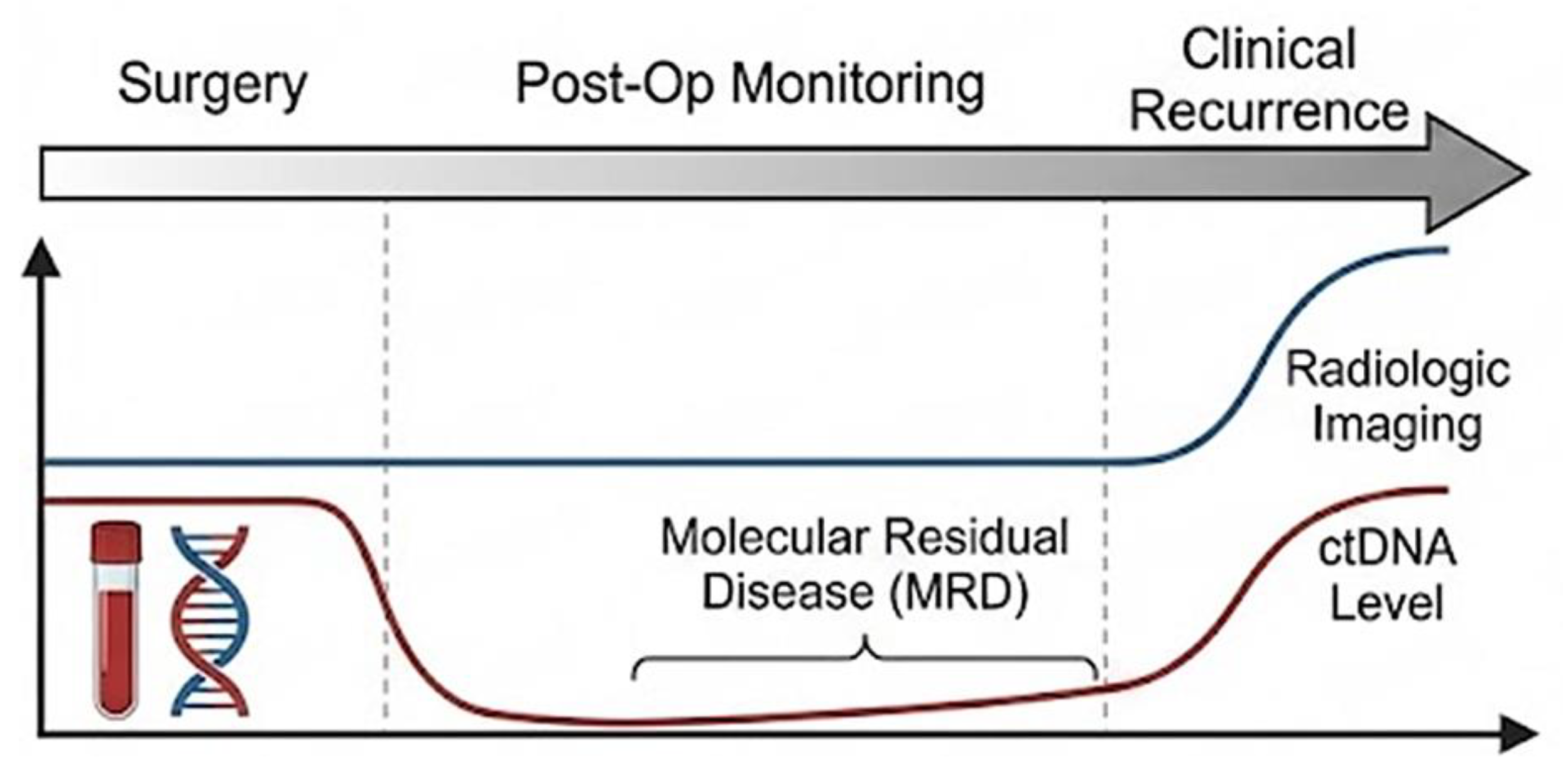
In this context, liquid biopsy does more than signal disease presence; it captures the evolutionary activity of residual clones. The bloodstream becomes a dynamic sampling window into ongoing tumor biology. This temporal surveillance role is schematically illustrated in
Figure 3, where ctDNA functions as a “molecular echo” of clonal persistence between apparent remission and overt relapse.
The interpretive power of ctDNA extends beyond relapse detection. High-depth sequencing permits comparison between plasma-detected variants and the genomic landscape of the original tumor, enabling distinction between true recurrence and independent second primary malignancies. Shared truncal mutations indicate clonal continuity and metastatic resurgence, whereas a divergent mutational profile supports independent tumorigenesis within a preconditioned field [
47,
48,
49]. Such discrimination carries significant clinical consequences, influencing therapeutic choice and surgical strategy. Liquid biopsy thus bridges evolutionary biology with real-world decision-making.
Yet the sensitivity of ctDNA assays introduces new interpretive challenges. Plasma sequencing interrogates total cfDNA rather than ctDNA in isolation, and this broader pool includes DNA fragments released from normal hematopoietic cells. Clonal hematopoiesis of indeterminate potential (CHIP), characterized by age-associated expansion of hematopoietic stem cell clones carrying mutations in genes such as
DNMT3A,
TET2, and
ASXL1, represents a major biological confounder [
41,
50]. Because some of these genes overlap with those mutated in solid tumors, CHIP-associated variants may be mistakenly attributed to tumor-derived ctDNA, leading to false-positive MRD interpretations.
The clinical implications of CHIP extend beyond technical artifact. Clonal hematopoiesis has been associated with increased risks of hematologic malignancy and cardiovascular disease, reflecting systemic aging-related genomic remodeling [
51]. Within the context of liquid biopsy, however, its principal significance lies in the necessity of contextual interpretation. Parallel sequencing of matched white blood cells markedly improves specificity by distinguishing hematopoietic-origin variants from genuine tumor-derived alterations [
52]. Without such controls, ultra-sensitive assays risk detecting age-related clonal evolution rather than true residual cancer.
These considerations underscore an essential principle of the integrated detection framework proposed here: no single modality operates in isolation. ctDNA provides a high-specificity longitudinal signal of clonal persistence, but its sensitivity depends on tumor fraction and biological shedding dynamics. It excels in the postoperative and minimal residual disease setting, where the genomic template of the tumor is already known. Its role in primary screening, by contrast, remains biologically constrained by low tumor burden and variable shedding.
Thus, liquid biopsy occupies a defined but powerful position within the broader architecture of early detection. Germline genetics defines inherited risk; somatic evolutionary analysis characterizes spatial conditioning; ctDNA supplies temporal resolution by tracking clonal activity over time. Together, these elements form a triad that integrates probability, biology, and surveillance. In this synthesis, liquid biopsy does not replace conventional imaging or pathology, but complements them by revealing molecular events that precede structural manifestation.
5. Colorectal Cancer and Volatile Organic Compound Analysis: Toward Continuous “Volatomics”
Among solid tumors, colorectal cancer (CRC) occupies a distinctive position in early detection research. Unlike malignancies developing within relatively sterile or anatomically inaccessible compartments, CRC arises within a metabolically dense luminal ecosystem in constant interaction with the gut microbiota. Tumorigenesis in this context does not unfold in isolation but is embedded within a biochemical dialogue between epithelial cells, microbial communities, dietary substrates, and inflammatory mediators. If mutational signatures capture the genomic imprint of carcinogenesis and ctDNA reflects clonal persistence, volatile organic compounds (VOCs) may represent the functional metabolic expression of tumor–microbiome co-evolution.
The biological rationale for VOC-based detection rests on converging processes. Neoplastic transformation in the colon is accompanied by metabolic reprogramming, oxidative stress, and lipid peroxidation, while dysbiosis reshapes microbial fermentation pathways and sulfur metabolism. Large-scale microbiome analyses have demonstrated reproducible enrichment of
Fusobacterium nucleatum and other taxa in CRC tissue [
53,
54,
55]. Emerging biomechanical data further suggest that early colorectal tumorigenesis is not solely a genomic or microbial phenomenon but also involves measurable stromal and tissue-level remodeling. Biomechanical phenotyping studies have identified distinctive mechanobiological signatures in early-onset colorectal cancer, including increased tissue stiffness and altered mechanotransduction pathways, supporting the concept that field conditioning encompasses structural as well as molecular alterations [
56]. These findings reinforce the view that preclinical carcinogenesis reflects a multidimensional tissue ecosystem shift rather than an isolated mutational event. Metagenomic and metabolomic studies further reveal progressive shifts along the adenoma–carcinoma sequence, suggesting that ecosystem remodeling precedes invasive disease [
57]. Alterations in short-chain fatty acid production, particularly reduced butyrate, together with increased protein-derived metabolites and inflammatory signaling, reinforce the concept of a metabolically conditioned field [
58,
59,
60,
61]. These biochemical perturbations generate volatile metabolites detectable in feces, urine, or breath [
62]. Early proof-of-concept studies demonstrated that fecal VOC profiling could discriminate CRC from controls using gas chromatography–mass spectrometry (GC–MS) platforms [
63], while electronic nose systems achieved sensitivities and specificities in the 85–87% range [
64]. More recent meta-analytic evidence reports pooled sensitivity near 0.86 and specificity near 0.90 for fecal VOC-based CRC detection [
65], and exhaled VOC analysis has shown discriminatory performance in independent cohorts [
66]. These studies collectively demonstrate biological plausibility and diagnostic potential.
Mechanistically, CRC-associated volatile alterations reflect coordinated host–microbiome interactions. Sulfur-containing compounds such as methyl mercaptan and dimethyl trisulfide arise from altered sulfur amino acid metabolism and microbial hydrogen sulfide production, a molecule with documented genotoxic and redox-modulating effects [
67,
68,
69]. Increased lipid peroxidation under oxidative stress generates aldehydes and ketones detectable in fecal headspace [
65,
70]. Microbial tryptophan metabolism yields indolic compounds capable of modulating epithelial and immune signaling through the aryl hydrocarbon receptor axis [
71,
72]. Collectively, these metabolic shifts suggest that VOC signatures may represent the functional extension of field cancerization into the luminal biochemical environment.
Current VOC-based approaches, however, rely almost exclusively on single time-point sampling. Fecal headspace analysis or breath testing captures a transient metabolic state influenced by diet, circadian rhythm, transit time, and short-term microbiome variability. Such cross-sectional strategies are biologically informative but remain episodic, mirroring the threshold-based logic discussed earlier in this manuscript.
The conceptual advance proposed here is longitudinal. Rather than seeking a unique “cancer gas,” the hypothesis is that colorectal tumorigenesis induces sustained alterations in composite volatile patterns relative to an individual’s physiological baseline. In this framework, the relevant signal is not the presence of a specific compound but persistent deviation in a multidimensional volatile profile.
Technological developments provide early proof of feasibility for continuous intestinal gas monitoring. Wearable hydrogen-sensing systems integrated into garments have demonstrated stable real-time detection of flatus-associated gases over several days in ambulatory settings [
73]. Ingestible and miniaturized sensing technologies capable of characterizing luminal chemical environments further illustrate that real-time gastrointestinal monitoring is technically attainable [
74,
75]. However, it must be emphasized that a medical device capable of comprehensive, multi-analyte detection and analysis of intestinal gas composition for oncologic purposes does not yet exist. The proposal advanced here therefore represents a translational hypothesis grounded in current biological and technological evidence, rather than an immediately deployable clinical tool.
In principle, a future multi-gas detection platform could integrate electrochemical sensors for hydrogen and hydrogen sulfide [
76], metal oxide semiconductor arrays for broader VOC responsiveness [
77], and surface acoustic wave technologies for trace compound detection [
78], coupled with machine-learning algorithms capable of recognizing stable composite deviations. Continuous monitoring—particularly during metabolically stable intervals such as nocturnal fasting—could enable construction of individualized volatile baselines. Persistent deviation from that baseline, rather than isolated spikes, would constitute the relevant alert (
Table 3).
Such an approach would align conceptually with the broader trajectory-based detection paradigm articulated in earlier chapters. If mutational signatures reveal genomic conditioning and ctDNA captures clonal persistence, continuous volatile profiling may detect functional metabolic activation within a conditioned epithelial field. In this sense, volatomics would serve as an orthogonal layer within an integrated surveillance architecture, complementing rather than replacing colonoscopy, FIT, or molecular assays.
Importantly, this framework remains hypothesis-driven. Rigorous prospective validation would be required to establish sensitivity, specificity, predictive value, and robustness against confounders such as diet, antibiotics, inflammatory bowel disease, and microbiome variability. Standardization of sensor calibration and analytic pipelines would be essential before clinical translation. Yet the convergence of microbiome science, metabolomics, and wearable sensing technology suggests that continuous metabolic field surveillance is biologically plausible and technically conceivable.
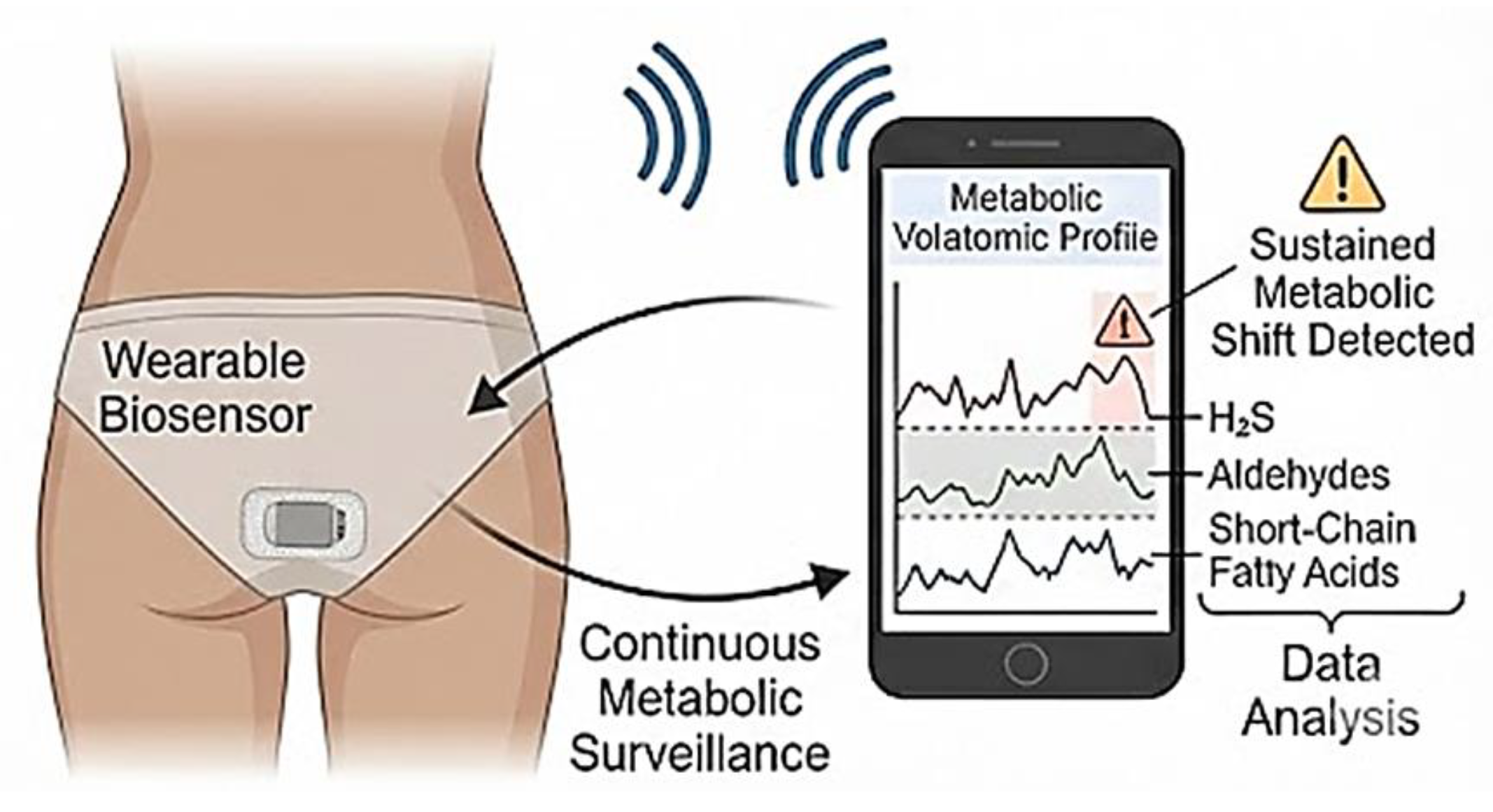
Colorectal cancer thus provides a uniquely accessible model in which tumor evolution, microbial ecology, and metabolomic output intersect. While the proposed multi-gas volatomic platform is not yet realized as a clinical device, its conceptual foundation reflects a logical extension of existing evidence. If validated, such an approach could introduce a dynamic, non-invasive triage layer capable of identifying sustained metabolic deviations that warrant prioritized diagnostic evaluation (
Figure 4).
6. Toward Integrated Early Detection Platforms
If each of the preceding chapters describes a distinct dimension of early detection—baseline biomarker trajectories, inherited susceptibility, somatic field evolution, molecular residual disease, and metabolic ecosystem shifts—the central proposition of this manuscript is that meaningful progress will emerge not from further refinement of any single modality, but from their integration. Early detection, in this view, becomes a longitudinal decision system rather than a stand-alone test.
A recurring limitation of population screening lies in its reliance on isolated measurements interpreted against population-derived thresholds. As discussed earlier, laboratory medicine has long recognized the concept of the reference change value (RCV), which formalizes when serial measurements in the same individual exceed expected analytical and biological variation [
79,
80]. Personalized reference intervals extend this logic by defining individual-specific normality envelopes derived from longitudinal data [
81,
82]. Multi-omics profiling studies further demonstrate that many molecular traits remain remarkably stable within individuals over time while varying substantially between individuals, a statistical structure that strongly favors baseline-based interpretation [
83,
84].
Within an integrated platform, early-life biomarker mapping therefore serves not as a “cancer test,” but as a calibration layer. It defines the background noise against which future deviations are evaluated. This calibration becomes particularly powerful when combined with germline genomic information. High-penetrance variants in genes such as
BRCA1/2 or Lynch-associated genes substantially elevate prior probability, justifying earlier and more frequent surveillance. At the population level, polygenic risk scores (PRS) refine stratification and identify subsets in whom intensified screening may yield higher diagnostic return [
25,
26] (
Figure 5). Emerging implementation studies exploring PRS-guided screening illustrate both feasibility and translational complexity in real-world settings [
85,
86].
In this framework, germline risk does not simply classify individuals into hereditary and non-hereditary categories; it functions as a dynamic prior probability that modulates the weight assigned to emerging biological signals. A subtle biomarker drift or low-level ctDNA detection carries different implications in a high-risk carrier compared with an individual at average genetic risk. Integration therefore enhances interpretability rather than merely increasing data volume.
The architecture becomes even more refined once a tumor has occurred. Molecular profiling of a primary malignancy generates a patient-specific template that captures driver mutations, mutational signatures, and repair deficiencies [
31]. This template informs therapeutic selection in the short term, but it also provides a blueprint for long-term surveillance. Tumor-informed ctDNA assays can track recurrence with high specificity [
44], and comparative genomic analysis can distinguish clonal resurgence from independent second primaries [
47] (
Figure 6). In this way, a clinical event becomes a predictive asset that enhances subsequent monitoring precision.
Liquid biopsy, particularly in the context of molecular residual disease, occupies a central position within this platform because it operationalizes temporal surveillance. However, its limitations—including tumor fraction constraints and confounding by clonal hematopoiesis—underscore the importance of contextual interpretation [
5,
87]. No single assay should be expected to function as a universal early detection solution. Instead, each modality contributes complementary information: baseline biomarkers reduce noise, germline data calibrate prior probability, tumor templates refine lineage interpretation, and ctDNA supplies high-specificity longitudinal alerts.
The volatomics concept introduced in the preceding chapter adds an orthogonal dimension. Continuous metabolic sensing, if technically realized and prospectively validated, would capture functional ecosystem perturbations that precede structural or genomic manifestation. Importantly, this layer should be conceptualized as a triage and prioritization mechanism rather than a diagnostic replacement for colonoscopy, imaging, or tissue pathology. Ingestible and wearable sensing technologies demonstrate that in vivo gastrointestinal chemical monitoring is technically feasible [
74,
75], but translation into oncologic screening requires rigorous validation and careful calibration.
The integration of these heterogeneous streams of data inevitably demands advanced computational frameworks. Machine-learning approaches capable of multi-omics data fusion have shown promise in oncology, yet their clinical credibility depends less on algorithmic novelty than on calibration, harmonization, and transparent validation across populations [
88,
89]. For such a platform to be clinically acceptable, it must generate calibrated risk probabilities linked to predefined escalation pathways, rather than opaque binary outputs. Interpretability and clinical trust are essential components of translational success.
Taken together, these components define early detection as a layered architecture. Baseline biomarker mapping establishes individual reference stability; germline genomics sets probabilistic context; tumor molecular fingerprinting provides lineage specificity; ctDNA offers temporal resolution; and metabolic sensing introduces ecosystem-level surveillance. None of these elements alone is sufficient. Their convergence, however, creates the possibility of calibrated, longitudinal monitoring that adapts to biological risk rather than relying solely on chronological age.
The translational challenge now lies in defining where and how each layer should be deployed. Prospective validation must determine which combinations meaningfully improve outcomes, under what conditions, and at what cost [
44,
86]. Integration should be incremental and evidence-driven, with clear definitions of escalation thresholds and explicit acknowledgment of uncertainty. If achieved, such a platform could shift early detection from sporadic screening toward dynamic risk calibration, aligning surveillance intensity with evolving biological context.
7. Challenges and Ethical Considerations
The promise of integrated early-detection platforms is inseparable from a parallel expansion of ethical, clinical, and societal complexity. As screening becomes more sensitive, more frequent, and increasingly multidimensional, the fundamental question shifts from whether earlier detection is technically possible to whether it delivers net benefit. Sensitivity alone is not a virtue if it leads to overdiagnosis, psychological harm, inequitable access, or unsustainable healthcare burdens. In this context, technological sophistication must be matched by conceptual restraint.
Overdiagnosis remains one of the most persistent hazards of intensified screening. It reflects not merely statistical artifact but biological heterogeneity: some detected lesions would never have progressed to clinical significance during an individual’s lifetime [
90,
91]. As detection moves earlier—potentially into pre-neoplastic or molecularly indeterminate states—the boundary between beneficial interception and unnecessary medicalization becomes increasingly delicate. Multi-marker assays, ctDNA-based monitoring, and future metabolic sensing platforms may reveal borderline or low-penetrance signals whose natural history is uncertain. Without clearly defined diagnostic-resolution pathways, such signals risk converting healthy individuals into patients-in-waiting, exposed to cascades of surveillance, invasive procedures, and overtreatment.
The psychological impact of intensified monitoring must also be acknowledged. Although systematic reviews suggest that most screening-related anxiety is modest and transient at the population level, subsets of individuals—particularly those receiving false-positive results or facing prolonged diagnostic ambiguity—may experience sustained distress [
92]. In a longitudinal, baseline-driven detection paradigm, risk communication itself becomes a clinical intervention. Transparent explanation of uncertainty, probabilistic framing of results, and structured follow-up are essential safeguards rather than optional additions [
93].
Ultra-sensitive assays introduce an additional layer of interpretive complexity. False-positive signals may arise from analytical variation, benign inflammatory states, or biological phenomena such as clonal hematopoiesis. The latter illustrates how age-related somatic evolution can confound plasma-based genomic testing, potentially triggering unnecessary downstream investigations if not properly contextualized [
87,
94]. Even assays with high analytical specificity may generate substantial diagnostic workload when deployed in low-prevalence populations. Ethical analyses of multi-cancer early detection (MCED) testing emphasize that downstream diagnostic cascades, resource allocation, and health-system strain are integral components of clinical utility evaluation [
95,
96]. In an integrated platform, a positive signal must be explicitly defined as a triage event rather than a diagnosis, with predefined, evidence-based escalation pathways.
Economic considerations further complicate the landscape. Lifelong longitudinal monitoring—particularly if it incorporates genomic profiling, repeated molecular assays, and wearable sensing—raises questions of cost-effectiveness and equitable access. Modeling studies suggest that multi-cancer detection strategies may achieve favorable value under certain performance and cost assumptions, yet outcomes remain highly sensitive to prevalence, test characteristics, and downstream management intensity [
97,
98]. If advanced detection technologies become available primarily through subscription-like models accessible to affluent populations, disparities in cancer outcomes may widen. Incremental implementation grounded in robust evidence is therefore essential.
Continuous sensing technologies introduce additional ethical dimensions related to privacy and data governance. Wearable or ingestible devices capable of generating granular physiological data create new vulnerabilities, including re-identification risks, cybersecurity concerns, and secondary data use beyond clinical intent [
99,
100]. The volatomics concept, in particular, touches upon intimate biological signals that may carry social stigma. Responsible implementation requires explicit data-minimization strategies, secure storage architectures, transparent consent processes, and clear separation between clinical use and research applications [
99,
101]. Trust becomes a foundational requirement for adoption.
At its core, the ethical challenge of integrated early detection lies in balancing benefit against medicalization. A longitudinal, multi-layered surveillance architecture risks normalizing constant monitoring and expanding the domain of pre-disease. Ethical frameworks for early detection consistently identify uncertainty, autonomy, privacy, justice, and proportionality as central themes [
93]. The goal is not maximal detection, but maximal net benefit—identifying biological change early enough to improve outcomes while minimizing unnecessary intervention and harm.
For integrated platforms to mature into routine clinical practice, they must evolve under the same evidentiary standards applied to therapeutics. Prospective trials, transparent reporting of false-positive pathways, clearly defined endpoints, and equitable implementation strategies are prerequisites rather than afterthoughts [
95,
101]. Early detection must protect patients not only from advanced cancer but also from the unintended consequences of knowing too early without actionable clarity.
In this light, ethical rigor does not constrain innovation; it disciplines it. The sophistication of emerging detection technologies makes careful governance indispensable. Without it, progress risks outpacing prudence.
8. Conclusion
Early cancer detection is entering a conceptual transition. For decades, screening has been structured around population-based thresholds, episodic testing, and structural lesion identification. While this paradigm has reduced mortality in selected malignancies, it remains fundamentally reactive, detecting disease only after biological deviation has reached macroscopic or biochemical visibility. The central argument of this manuscript is that the next evolution in early detection will not arise from a single transformative assay, but from the calibrated integration of complementary biological layers interpreted longitudinally within individuals.
Cancer does not begin at diagnosis. It unfolds over years through the interplay of inherited susceptibility, somatic mutation accumulation, clonal expansion, microenvironmental remodeling, and metabolic adaptation. Germline variants establish probabilistic vulnerability across the life course. Somatic evolution and field cancerization condition tissues long before overt malignancy appears. Once a tumor emerges, its molecular fingerprint becomes a template for monitoring recurrence through circulating tumor DNA. Emerging metabolomic approaches suggest that functional ecosystem perturbations may precede structural transformation. Each modality captures a different dimension of tumorigenesis—probability, spatial conditioning, temporal persistence, or metabolic activation.
The novelty of the framework proposed here lies not in any individual component, but in their convergence. Longitudinal biomarker mapping establishes individualized baselines against which deviation acquires meaning. Germline information modulates prior probability and calibrates surveillance intensity. Tumor-informed molecular profiling refines lineage interpretation and recurrence tracking. ctDNA supplies high-specificity temporal resolution. The proposed volatomics layer, though not yet realized as a clinical device, extends this architecture by hypothesizing continuous metabolic field surveillance grounded in existing microbiome and metabolomic evidence. Together, these elements define early detection as a dynamic system rather than a static test.
Importantly, this integrated approach does not seek to replace established screening modalities such as colonoscopy, mammography, or low-dose CT. Instead, it introduces a stratified surveillance architecture in which emerging biological signals serve as triage events that prioritize diagnostic evaluation. In this model, detection is no longer triggered solely by age or population averages, but by calibrated deviation from an individual’s biological trajectory.
The implementation of such platforms must proceed incrementally and under rigorous evidentiary standards. Prospective validation, transparent reporting of false-positive pathways, equitable access strategies, and ethical governance are indispensable. Sensitivity must be balanced against overdiagnosis; innovation must remain proportional to demonstrable benefit. The objective is not maximal detection, but maximal meaningful interception—identifying disease at a stage where intervention alters outcome while minimizing harm.
If realized responsibly, integrated longitudinal detection could reframe oncology from episodic screening toward adaptive biological monitoring. Cancer interception would then become less a matter of discovering lesions and more a matter of recognizing sustained biological drift within conditioned tissue ecosystems. The transformation would be structural: from age-triggered screening toward biologically calibrated surveillance, from isolated tests toward integrated decision systems, and from reactive diagnosis toward anticipatory precision prevention.
In this evolving landscape, early detection ceases to be a moment and becomes a process.
Author Contributions
DR: Conceptualization, Methodology, Software, Validation, Formal analysis, Investigation, Resources, Data Curation, Writing - Original Draft, Writing - Review & Editing, Visualization, Project administration.
Funding
This research received no external funding.
Acknowledgments
The author declares that Generative AI (ChatGPT-5.0) was used in the creation of this manuscript. Generative AI was used to revise the manuscript for language, clarity, and correct idiomatic expression, and to help generating figures and tables. The author has reviewed and edited the output and take full responsibility for the content of this publication.
Conflicts of Interest
The author declares no conflicts of interest.
References
- Marinac, C. R., E. K. O’Donnell, C. A. Clarke, M. Lopatin, E. T. Fung, and E. A. Klein. 2025. “Multicancer early detection blood test performance in cancer survivors from the PATHFINDER study.” JCO Precision Oncology 9: e2500498. [CrossRef]
- Liu, M. C., G. R. Oxnard, E. A. Klein, C. Swanton, M. V. Seiden, and CCGA Consortium. 2020. “Sensitive and specific multi-cancer detection and localization using methylation signatures in cell-free DNA.” Annals of Oncology 31: 745–759. [CrossRef]
- Nguyen, L. H. D., T. H. H. Nguyen, V. H. Le, V. Q. Bui, L. H. Nguyen, N. H. Pham, et al. 2025. “Prospective validation study: A non-invasive circulating tumor DNA-based assay for simultaneous early detection of multiple cancers in asymptomatic adults.” BMC Medicine 23: 90. [CrossRef]
- Bryce, A. H., D. D. Thiel, M. V. Seiden, D. Richards, Y. Luan, M. Coignet, et al. 2023. “Performance of a Cell-Free DNA-Based Multi-cancer Detection Test in Individuals Presenting With Symptoms Suspicious for Cancers.” JCO Precision Oncology 7: e2200679. [CrossRef]
- Jamshidi, A., M. C. Liu, E. A. Klein, et al. 2022. “Evaluation of cell-free DNA approaches for multi-cancer early detection.” Cancer Cell 40: 1537–1549.e12. [CrossRef]
- Martincorena, I. 2019. “Somatic mutation and clonal expansions in human tissues.” Genome Medicine 11: 35. [CrossRef]
- Irajizad, E., J. F. Fahrmann, I. Toumazis, J. Vykoukal, J. B. Dennison, Y. Shen, et al. 2024. “Biomarker trajectory for earlier detection of lung cancer.” EBioMedicine 108: 105377. [CrossRef]
- Lloyd-Jones, D. M., M. D. Huffman, K. N. Karmali, D. M. Sanghavi, J. S. Wright, C. Pelser, et al. 2017. “Estimating longitudinal risks and benefits from cardiovascular preventive therapies among Medicare patients: The Million Hearts longitudinal ASCVD risk assessment tool.” Journal of the American College of Cardiology 69: 1617–1636. [CrossRef]
- Menon, U., A. Ryan, J. Kalsi, A. Gentry-Maharaj, A. Dawnay, M. Habib, et al. 2015. “Risk algorithm using serial biomarker measurements doubles the number of screen-detected cancers compared with a single-threshold rule in the UKCTOCS.” Journal of Clinical Oncology 33: 2062–2071. [CrossRef]
- Vickers, A. J., and S. F. Brewster. 2012. “PSA velocity and doubling time in diagnosis and prognosis of prostate cancer.” British Journal of Medical and Surgical Urology 5: 162–168. [CrossRef]
- Cohen, J. D., L. Li, Y. Wang, C. Thoburn, B. Afsari, L. Danilova, et al. 2018. “Detection and localization of surgically resectable cancers with a multi-analyte blood test.” Science 359: 926–930. [CrossRef]
- Zhuang, L., S. H. Park, S. J. Skates, A. E. Prosper, D. R. Aberle, and W. Hsu. 2026. “Advancing precision oncology through modeling of longitudinal and multimodal data.” IEEE Reviews in Biomedical Engineering 19: 182–200. [CrossRef]
- Lennon, A. M., A. H. Buchanan, I. Kinde, A. Warren, A. Honushefsky, A. T. Cohain, et al. 2020. “Feasibility of blood testing combined with PET-CT to screen for cancer and guide intervention.” Science 369: eabb9601. [CrossRef]
- Huang, K. L., R. J. Mashl, Y. Wu, D. I. Ritter, J. Wang, C. Oh, et al. 2018. “Pathogenic Germline Variants in 10,389 Adult Cancers.” Cell 173: 355–370.e14. [CrossRef]
- Kuchenbaecker, K. B., J. L. Hopper, D. R. Barnes, et al. 2017. “Risks of Breast, Ovarian, and Contralateral Breast Cancer for BRCA1 and BRCA2 Mutation Carriers.” JAMA 317: 2402–2416. [CrossRef]
- Golan, T., P. Hammel, M. Reni, E. Van Cutsem, T. Macarulla, M. J. Hall, et al. 2019. “Maintenance Olaparib for Germline BRCA-Mutated Metastatic Pancreatic Cancer.” New England Journal of Medicine 381: 317–327. [CrossRef]
- Ueki, A., R. Yoshida, T. Kosaka, and H. Matsubayashi. 2023. “Clinical risk management of breast, ovarian, pancreatic, and prostatic cancers for BRCA1/2 variant carriers in Japan.” Journal of Human Genetics 68: 517–526. [CrossRef]
- Lord, C. J., and A. Ashworth. 2017. “PARP inhibitors: Synthetic lethality in the clinic.” Science 355: 1152–1158. [CrossRef]
- Curtius, K., S. Gupta, and C. R. Boland. 2022. “Lynch Syndrome—a mechanistic and clinical management update.” Alimentary Pharmacology & Therapeutics 55: 960–977. [CrossRef]
- Seppälä, T. T., G. Möslein, M. Dominguez-Valentin, et al. 2025. “Metachronous colorectal cancer risks after extended or segmental resection in MLH1, MSH2, and MSH6 Lynch syndrome: Multicentre study from the Prospective Lynch Syndrome Database.” British Journal of Surgery 112: znaf061. [CrossRef]
- Møller, P., T. T. Seppälä, I. Bernstein, E. Holinski-Feder, P. Sala, D. G. Evans, et al. 2018. “Cancer risk and survival in path_MMR carriers by gene and gender up to 75 years of age: Prospective Lynch Syndrome Database report.” Gut 67: 1306–1316. [CrossRef]
- Blair, V. R., M. McLeod, F. Carneiro, et al. 2020. “Hereditary diffuse gastric cancer: Updated clinical practice guidelines.” Lancet Oncology 21: e386–e397. [CrossRef]
- Wiese, J., S. Sirigireddy, A. Elghezewi, et al. 2025. “Hereditary cancer syndromes in familial breast cancer: Insights into proactive gastric cancer surveillance.” Journal of Investigative Medicine High Impact Case Reports 13: 23247096251358660. [CrossRef]
- Ishida, H., K. Kumamoto, O. Suzuki, et al. 2022. “JSCCR guidelines 2020 for the clinical practice of hereditary colorectal cancer: Peutz–Jeghers syndrome.” International Journal of Clinical Oncology 27: 14–30.
- Dorling, S., S. Carvalho, J. Allen, et al. 2021. “Breast cancer risk genes—association analysis in more than 113,000 women.” New England Journal of Medicine 384: 428–439. [CrossRef]
- Antoniou, A. C., S. Casadei, T. Heikkinen, et al. 2014. “Breast-cancer risk in families with mutations in PALB2.” New England Journal of Medicine 371: 497–506. [CrossRef]
- Slaughter, D. P., H. W. Southwick, and W. Smejkal. 1953. “Field cancerization in oral stratified squamous epithelium: Clinical implications of multicentric origin.” Cancer 6: 963–968.
- Martincorena, I., J. C. Fowler, A. Wabik, et al. 2018. “Somatic mutant clones colonize the human esophagus with age.” Science 362: 911–917. [CrossRef]
- Yizhak, K., F. Aguet, J. Buchanan, et al. 2019. “RNA sequence analysis reveals macroscopic somatic clonal expansion across normal tissues.” Science 364: eaaw0726. [CrossRef]
- The GTEx Consortium. 2013. “The Genotype-Tissue Expression (GTEx) project.” Nature Genetics 45: 580–585. [CrossRef]
- Alexandrov, L. B., J. Kim, N. J. Haradhvala, M. N. Huang, A. W. Tian Ng, Y. Wu, et al. 2020. “The repertoire of mutational signatures in human cancer.” Nature 578: 94–101. [CrossRef]
- Martincorena, I., A. Roshan, M. Gerstung, P. Ellis, P. Van Loo, S. McLaren, et al. 2015. “High burden and pervasive positive selection of somatic mutations in normal human skin.” Science 348: 880–886. [CrossRef]
- Lee-Six, H., N. F. Øbro, M. S. Shepherd, S. Grossmann, K. Dawson, M. Belmonte, et al. 2018. “Population dynamics of normal human blood inferred from somatic mutations.” Nature 561: 473–478. [CrossRef]
- Yokoyama, A., N. Kakiuchi, T. Yoshizato, Y. Nannya, H. Suzuki, Y. Takeuchi, et al. 2019. “Age-related remodelling of oesophageal epithelia by mutated cancer drivers.” Nature 565: 312–317. [CrossRef]
- de Bono, J., J. Mateo, K. Fizazi, F. Saad, N. Shore, S. Sandhu, et al. 2020. “Olaparib for Metastatic Castration-Resistant Prostate Cancer.” New England Journal of Medicine 382: 2091–2102. [CrossRef]
- Le, D. T., J. N. Durham, K. N. Smith, H. Wang, B. R. Bartlett, L. K. Aulakh, et al. 2017. “Mismatch repair deficiency predicts response of solid tumors to PD-1 blockade.” Science 357: 409–413. [CrossRef]
- Vousden, K. H., and C. Prives. 2009. “Blinded by the light: The growing complexity of p53.” Cell 137: 413–431. [CrossRef]
- Kastenhuber, E. R., and S. W. Lowe. 2017. “Putting p53 in Context.” Cell 170: 1062–1078. [CrossRef]
- Brennan, J. A., L. Mao, R. H. Hruban, J. O. Boyle, Y. J. Eby, W. M. Koch, S. N. Goodman, and D. Sidransky. 1995. “Molecular assessment of histopathological staging in squamous-cell carcinoma of the head and neck.” New England Journal of Medicine 332: 429–435. [CrossRef]
- Ross-Innes, C. S., J. Becq, A. Warren, R. K. Cheetham, H. Northen, M. O’Donovan, et al. 2015. “Whole-genome sequencing provides new insights into the clonal architecture of Barrett’s esophagus and esophageal adenocarcinoma.” Nature Genetics 47: 1038–1046. [CrossRef]
- Jaiswal, S., P. Fontanillas, J. Flannick, A. Manning, P. V. Grauman, B. G. Mar, et al. 2014. “Age-related clonal hematopoiesis associated with adverse outcomes.” New England Journal of Medicine 371: 2488–2498. [CrossRef]
- Yazaki, S., M. Tokura, H. Aiba, Y. Kojima, and K. Shiraishi. 2025. “Clinical applications of cell-free DNA-based liquid biopsy analysis.” Translational Oncology 61: 102519. [CrossRef]
- Turabi, K., K. Klute, and P. Radhakrishnan. 2024. “Decoding the dynamics of circulating tumor DNA in liquid biopsies.” Cancers 16: 2432. [CrossRef]
- Tie, J., Y. Wang, S. N. Lo, K. Lahouel, J. D. Cohen, R. Wong, et al. 2025. “Circulating tumor DNA analysis guiding adjuvant therapy in stage II colon cancer: 5-year outcomes of the randomized DYNAMIC trial.” Nature Medicine 31: 1509–1518. [CrossRef]
- Parikh, A. R., E. E. Van Seventer, G. Siravegna, A. V. Hartwig, A. Jaimovich, Y. He, et al. 2021. “Minimal residual disease detection using a plasma-only circulating tumor DNA assay in patients with colorectal cancer.” Clinical Cancer Research 27: 5586–5594. [CrossRef]
- Negro, S., A. Pulvirenti, C. Trento, S. Indraccolo, S. Ferrari, M. Scarpa, et al. 2025. “Circulating tumor DNA as a real-time biomarker for minimal residual disease and recurrence prediction in stage II colorectal cancer: A systematic review and meta-analysis.” International Journal of Molecular Sciences 26: 2486. [CrossRef]
- Abbosh, C., N. J. Birkbak, G. A. Wilson, M. Jamal-Hanjani, T. Constantin, R. Salari, et al. 2017. “Phylogenetic ctDNA analysis depicts early-stage lung cancer evolution.” Nature 545: 446–451. [CrossRef]
- Chaudhuri, A. A., J. J. Chabon, A. F. Lovejoy, A. M. Newman, H. Stehr, T. D. Azad, et al. 2017. “Early Detection of Molecular Residual Disease in Localized Lung Cancer by Circulating Tumor DNA Profiling.” Cancer Discovery 7: 1394–1403. [CrossRef]
- Hassan, W. A. 2025. “Molecular advances in early-stage and locally advanced non-small cell lung carcinoma: Shaping the future of precision oncology—systematic review.” Science Progress 108: 368504251383055. [CrossRef]
- Genovese, G., A. K. Kähler, R. E. Handsaker, J. Lindberg, S. A. Rose, S. F. Bakhoum, et al. 2014. “Clonal hematopoiesis and blood-cancer risk inferred from blood DNA sequence.” New England Journal of Medicine 371: 2477–2487. [CrossRef]
- Evans, M. A., and K. Walsh. 2026. “The Immunological Consequences of Clonal Hematopoiesis in Heart Failure.” Immunological Reviews 337: e70094. [CrossRef]
- Razavi, P., B. T. Li, D. N. Brown, B. Jung, E. Hubbell, R. Shen, et al. 2019. “High-intensity sequencing reveals the sources of plasma circulating cell-free DNA variants.” Nature Medicine 25: 1928–1937. [CrossRef]
- Kostic, A. D., D. Gevers, C. S. Pedamallu, M. Michaud, F. Duke, A. M. Earl, et al. 2012. “Genomic analysis identifies association of Fusobacterium with colorectal carcinoma.” Genome Research 22: 292–298. [CrossRef]
- Thomas, A. M., P. Manghi, F. Asnicar, E. Pasolli, F. Armanini, M. Zolfo, et al. 2019. “Metagenomic analysis of colorectal cancer datasets identifies cross-cohort microbial diagnostic signatures and a link with choline degradation.” Nature Medicine 25: 667–678. [CrossRef]
- Flemer, B., D. B. Lynch, J. M. Brown, I. B. Jeffery, F. J. Ryan, M. J. Claesson, M. O’Riordain, F. Shanahan, and P. W. O’Toole. 2017. “Tumour-associated and non-tumour-associated microbiota in colorectal cancer.” Gut 66: 633–643. [CrossRef]
- Huning, N. C., M. H. Buhaya, V. V. Nguyen, A. Khazi-Syed, H. A. Ali, A. Khan, et al. 2026. “Biomechanical Phenotyping Reveals Unique Mechanobiological Signatures of Early-Onset Colorectal Cancer.” Advanced Science 13: e14693. [CrossRef]
- Yachida, S., S. Mizutani, H. Shiroma, S. Shiba, T. Nakajima, T. Sakamoto, et al. 2019. “Metagenomic and metabolomic analyses reveal distinct stage-specific phenotypes of the gut microbiota in colorectal cancer.” Nature Medicine 25: 968–976. [CrossRef]
- Louis, P., G. L. Hold, and H. J. Flint. 2014. “The gut microbiota, bacterial metabolites and colorectal cancer.” Nature Reviews Microbiology 12: 661–672. [CrossRef]
- O’Keefe, S. J. 2016. “Diet, microorganisms and their metabolites, and colon cancer.” Nature Reviews Gastroenterology & Hepatology 13: 691–706. [CrossRef]
- Janney, A., F. Powrie, and E. H. Mann. 2020. “Host-microbiota maladaptation in colorectal cancer.” Nature 585: 509–517. [CrossRef]
- Li, R., T. Ugai, L. Xu, D. Zucker, S. Ogino, and M. Wang. 2022. “Utility of continuous disease subtyping systems for improved evaluation of etiologic heterogeneity.” Cancers 14: 1811. [CrossRef]
- Bosch, S., D. J. Berkhout, I. Ben Larbi, T. G. de Meij, and N. K. de Boer. 2019. “Fecal volatile organic compounds for early detection of colorectal cancer: Where are we now?” Journal of Cancer Research and Clinical Oncology 145 (1): 223–234. [CrossRef]
- Arasaradnam, R. P., M. McFarlane, C. Ryan-Fisher, et al. 2014. “Detection of colorectal cancer (CRC) by urinary volatile organic compound analysis.” PLOS ONE 9: e108750. [CrossRef]
- de Meij, T. G., I. B. Larbi, M. P. van der Schee, Y. E. Lentferink, T. Paff, J. S. Terhaar Sive Droste, C. J. Mulder, A. A. van Bodegraven, and N. K. de Boer. 2014. “Electronic nose can discriminate colorectal carcinoma and advanced adenomas by fecal volatile biomarker analysis: Proof of principle study.” International Journal of Cancer 134: 1132–1138. [CrossRef]
- Bhattacharyya, N., Z. Sastimoglu, D. Armstrong, P. R. Selvaganapathy, and M. J. Deen. 2025. “Fecal volatile organic compounds for colorectal cancer detection: A systematic review and meta-analysis.” Computers in Biology and Medicine 195: 110561. [CrossRef]
- Ji, Y., W. Wang, S. Dong, Y. Huang, Z. Yu, Y. Zhao, and X. Li. 2025. “Discriminating Colorectal Cancer via Exhaled Volatile Organic Compounds.” QJM (in press): hcaf317. [CrossRef]
- Attene-Ramos, M. S., E. D. Wagner, M. J. Plewa, and H. R. Gaskins. 2006. “Evidence that hydrogen sulfide is a genotoxic agent.” Molecular Cancer Research 4: 9–14. [CrossRef]
- Blachier, F., M. Beaumont, and E. Kim. 2019. “Cysteine-derived hydrogen sulfide and gut health: A matter of endogenous or bacterial origin.” Current Opinion in Clinical Nutrition and Metabolic Care 22: 68–75. [CrossRef]
- Carbonero, F., A. C. Benefiel, A. H. Alizadeh-Ghamsari, and H. R. Gaskins. 2012. “Microbial pathways in colonic sulfur metabolism and links with health and disease.” Frontiers in Physiology 3: 448. [CrossRef]
- Wong, S. H., and J. Yu. 2019. “Gut microbiota in colorectal cancer: Mechanisms of action and clinical applications.” Nature Reviews Gastroenterology & Hepatology 16: 690–704. [CrossRef]
- Ala, M. 2022. “Tryptophan metabolites modulate inflammatory bowel disease and colorectal cancer by affecting immune system.” International Reviews of Immunology 41: 326–345. [CrossRef]
- Liu, Y., Z. Pei, T. Pan, H. Wang, W. Chen, and W. Lu. 2023. “Indole metabolites and colorectal cancer: Gut microbial tryptophan metabolism, host gut microbiome biomarkers, and potential intervention mechanisms.” Microbiological Research 272: 127392. [CrossRef]
- Botasini, S., D. Zhana, N. Fischera, et al. 2025. “Smart underwear: A novel wearable for long-term monitoring of gut microbial gas production via flatus.” Biosensors and Bioelectronics: X 13: 100699. [CrossRef]
- Kalantar-Zadeh, K., N. Ha, J. Z. Ou, and K. J. Berean. 2017. “Ingestible Sensors.” ACS Sensors 2: 468–483. [CrossRef]
- Rehan, M., I. Al-Bahadly, D. G. Thomas, W. Young, L. K. Cheng, and E. Avci. 2024. “Smart capsules for sensing and sampling the gut: Status, challenges and prospects.” Gut 73: 186–202. [CrossRef]
- Wang, J. 2008. “Electrochemical glucose biosensors.” Chemical Reviews 108: 814–825. [CrossRef]
- Röck, F., N. Barsan, and U. Weimar. 2008. “Electronic nose: Current status and future trends.” Chemical Reviews 108: 705–725. [CrossRef]
- Grate, J. W., S. J. Martin, and R. M. White. 1993. “Acoustic wave microsensors.” Analytical Chemistry 65: 940A–948A. [CrossRef]
- Fraser, C. G. 2012. “Reference change values.” Clinical Chemistry and Laboratory Medicine 50: 807–812. [CrossRef]
- Røys, EÅ., K. Viste, R. Kellmann, N. A. Guldhaug, B. Alaour, M. S. Sylte, et al. 2025. “Estimating reference change values using routine patient data: A novel pathology database approach.” Clinical Chemistry 71: 307–318. [CrossRef]
- Coskun, A., S. Sandberg, I. Unsal, M. Serteser, and A. K. Aarsand. 2022a. “Personalized reference intervals: From theory to practice.” Critical Reviews in Clinical Laboratory Sciences 59: 501–516. [CrossRef]
- Coskun, A., S. Sandberg, I. Unsal, F. G. Yavuz, C. Cavusoglu, M. Serteser, M. Kilercik, and A. K. Aarsand. 2022b. “Personalized reference intervals—statistical approaches and considerations.” Clinical Chemistry and Laboratory Medicine 60: 629–635. [CrossRef]
- Tebani, A., A. Gummesson, W. Zhong, I. S. Koistinen, T. Lakshmikanth, L. M. Olsson, et al. 2020. “Integration of molecular profiles in a longitudinal wellness profiling cohort.” Nature Communications 11: 4487. [CrossRef]
- Babu, M., and M. Snyder. 2023. “Multi-Omics Profiling for Health.” Molecular & Cellular Proteomics 22: 100561. [CrossRef]
- McHugh, J. K., E. K. Bancroft, E. Saunders, M. N. Brook, E. McGrowder, S. Wakerell, et al. 2025. “Assessment of a polygenic risk score in screening for prostate cancer.” New England Journal of Medicine 392: 1406–1417. [CrossRef]
- Vassy, J. L., C. A. Brunette, M. S. Lebo, K. MacIsaac, T. Yi, M. E. Danowski, et al. 2023. “The GenoVA study: Equitable implementation of a pragmatic randomized trial of polygenic-risk scoring in primary care.” American Journal of Human Genetics 110: 1841–1852. [CrossRef]
- Croitoru, V. M., I. M. Cazacu, I. Popescu, D. Paul, S. O. Dima, A. E. Croitoru, and A. D. Tanase. 2022. “Clonal Hematopoiesis and Liquid Biopsy in Gastrointestinal Cancers.” Frontiers in Medicine (Lausanne) 8: 772166. [CrossRef]
- Cai, Z., R. C. Poulos, J. Liu, and Q. Zhong. 2022. “Machine learning for multi-omics data integration in cancer.” iScience 25: 103798. [CrossRef]
- Nicora, G., F. Vitali, A. Dagliati, N. Geifman, and R. Bellazzi. 2020. “Integrated multi-omics analyses in oncology: A review of machine learning methods and tools.” Frontiers in Oncology 10: 1030. [CrossRef]
- Srivastava, S., E. J. Koay, A. D. Borowsky, A. M. De Marzo, S. Ghosh, P. D. Wagner, and B. S. Kramer. 2019. “Cancer overdiagnosis: A biological challenge and clinical dilemma.” Nature Reviews Cancer 19: 349–358. [CrossRef]
- Dunn, B. K., S. Woloshin, H. Xie, and B. S. Kramer. 2022. “Cancer overdiagnosis: A challenge in the era of screening.” Journal of the National Cancer Center 2: 235–242. [CrossRef]
- Kim, A., K. C. Chung, C. Keir, and D. L. Patrick. 2022. “Patient-reported outcomes associated with cancer screening: A systematic review.” BMC Cancer 22: 223. [CrossRef]
- Jansen, S. N. G., B. A. Kamphorst, B. C. Mulder, et al. 2024. “Ethics of early detection of disease risk factors: A scoping review.” BMC Medical Ethics 25: 25. [CrossRef]
- Kim, H., and K. U. Park. 2023. “Clinical Circulating Tumor DNA Testing for Precision Oncology.” Cancer Research and Treatment 55: 351–366. [CrossRef]
- Sempere, L. F. 2025. “Ethical considerations and implications of multi-cancer early detection screening: Reliability, access and cost to test and treat.” Cambridge Quarterly of Healthcare Ethics 34: 489–498. [CrossRef]
- Imai, M., Y. Nakamura, and T. Yoshino. 2025. “Transforming cancer screening: The potential of multi-cancer early detection (MCED) technologies.” International Journal of Clinical Oncology 30: 180–193. [CrossRef]
- Kansal, A. R., A. Tafazzoli, W. Ye, W. Morris, D. Zou, B. Quinn, and A. M. Fendrick. 2025. “Potential Economic Value of Multi-cancer Early Detection Testing Under Differential Cost Trends for Cancer Screening and Management.” PharmacoEconomics Open (in press). [CrossRef]
- Lewis, D., W. W. L. Wong, J. Lipscomb, and S. Horton. 2024. “An exploratory analysis of the cost-effectiveness of a multi-cancer early detection blood test compared with standard of care screening in Ontario, Canada.” PharmacoEconomics 42: 393–407. [CrossRef]
- Doherty, C., M. Baldwin, R. Lambe, M. Altini, and B. Caulfield. 2025. “Privacy in consumer wearable technologies: A living systematic analysis of data policies across leading manufacturers.” NPJ Digital Medicine 8: 363. [CrossRef]
- Sui, A., W. Sui, S. Liu, and R. Rhodes. 2023. “Ethical considerations for the use of consumer wearables in health research.” Digital Health 9: 20552076231153740. [CrossRef]
- Canali, S., V. Schiaffonati, and A. Aliverti. 2022. “Challenges and recommendations for wearable devices in digital health: Data quality, interoperability, health equity, fairness.” PLOS Digital Health 1: e0000104. [CrossRef]
Figure 1.
Threshold vs. Trajectory Monitoring. Traditional screening relies on a population-average threshold (red dashed line). In contrast, personalized medicine monitors an individual's unique biological baseline (blue line). Early detection is triggered by a statistically significant "drift" (trajectory) rather than the absolute crossing of a population threshold.
Figure 1.
Threshold vs. Trajectory Monitoring. Traditional screening relies on a population-average threshold (red dashed line). In contrast, personalized medicine monitors an individual's unique biological baseline (blue line). Early detection is triggered by a statistically significant "drift" (trajectory) rather than the absolute crossing of a population threshold.
Figure 2.
The Molecular Field of Cancerization. The evolutionary landscape of field cancerization. Tissues harbor "molecular field memory"—clones with driver mutations (e.g., TP53) and mutational signatures (e.g., UV, tobacco) that precede and surround the overt malignancy.
Figure 2.
The Molecular Field of Cancerization. The evolutionary landscape of field cancerization. Tissues harbor "molecular field memory"—clones with driver mutations (e.g., TP53) and mutational signatures (e.g., UV, tobacco) that precede and surround the overt malignancy.
Figure 3.
Liquid Biopsy and Molecular Residual Disease (MRD). Timeline backbone of post-treatment monitoring. ctDNA acts as a "molecular echo," detecting molecular residual disease (MRD) months before clinical or radiologic recurrence becomes visible.
Figure 3.
Liquid Biopsy and Molecular Residual Disease (MRD). Timeline backbone of post-treatment monitoring. ctDNA acts as a "molecular echo," detecting molecular residual disease (MRD) months before clinical or radiologic recurrence becomes visible.
Figure 4.
Concept of Wearable "Volatomics" for CRC. Proposed extension toward wearable multi-gas volatomics. Continuous metabolic surveillance allows for baseline-adjusted triage, capturing dynamic tumor–microbiome interactions that single-point sampling may miss.
Figure 4.
Concept of Wearable "Volatomics" for CRC. Proposed extension toward wearable multi-gas volatomics. Continuous metabolic surveillance allows for baseline-adjusted triage, capturing dynamic tumor–microbiome interactions that single-point sampling may miss.
Figure 5.
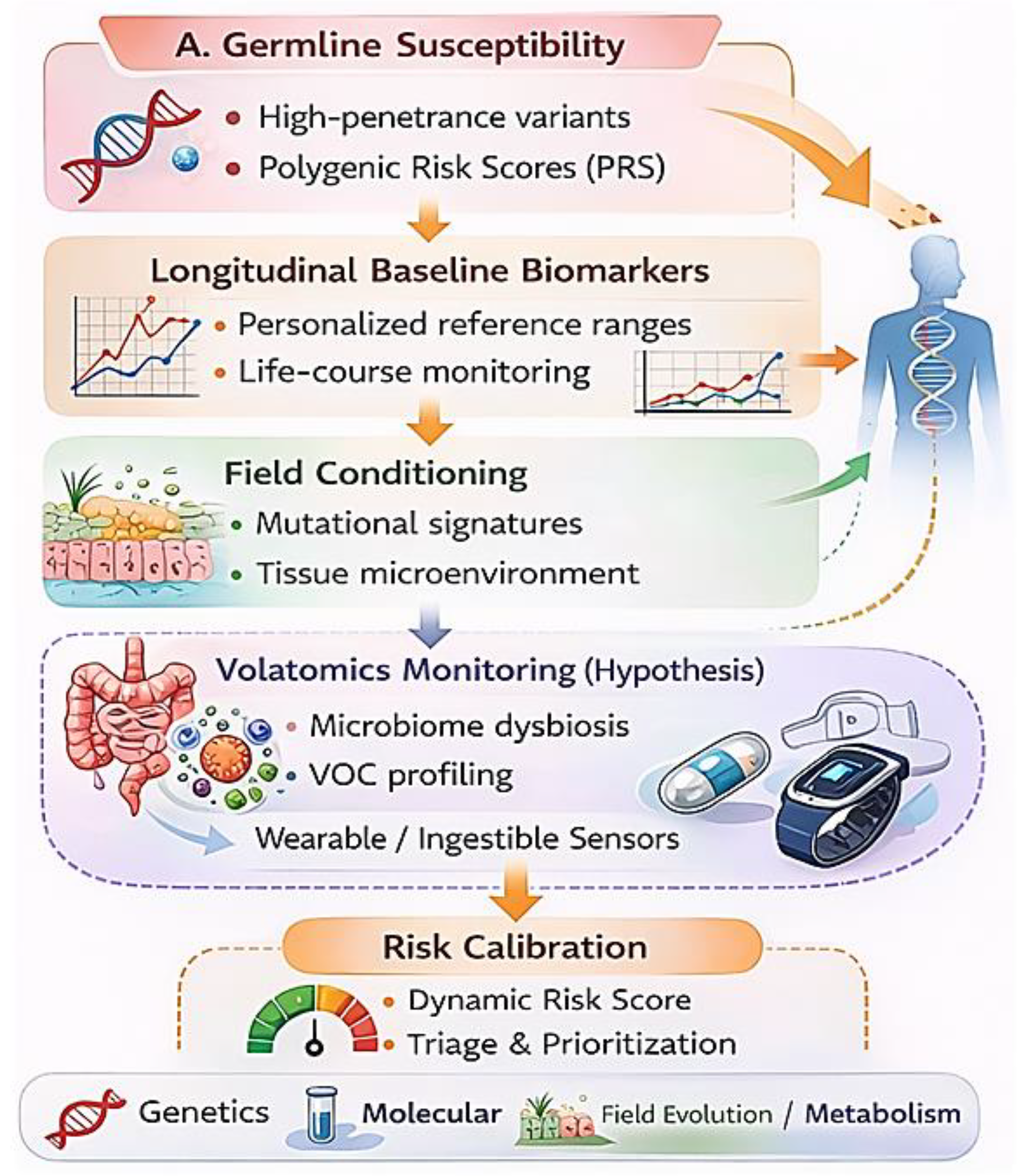
Integrated framework for primary cancer risk assessment in cancer-naïve individuals. In individuals without a prior cancer diagnosis, early detection is structured as longitudinal risk calibration rather than single time-point screening. Germline susceptibility (high-penetrance variants and polygenic risk scores) defines inherited risk. Longitudinal baseline biomarkers establish personalized reference ranges over time. Field conditioning (mutational signatures and tissue microenvironment changes) reflects preclinical biological evolution. The proposed still hypothetic volatomics monitoring layer—based on microbiome dysbiosis and volatile organic compound (VOC) profiling—aims to detect sustained metabolic deviation from baseline. These inputs converge into dynamic risk calibration, guiding triage toward imaging, endoscopy, or biopsy.
Figure 5.
Integrated framework for primary cancer risk assessment in cancer-naïve individuals. In individuals without a prior cancer diagnosis, early detection is structured as longitudinal risk calibration rather than single time-point screening. Germline susceptibility (high-penetrance variants and polygenic risk scores) defines inherited risk. Longitudinal baseline biomarkers establish personalized reference ranges over time. Field conditioning (mutational signatures and tissue microenvironment changes) reflects preclinical biological evolution. The proposed still hypothetic volatomics monitoring layer—based on microbiome dysbiosis and volatile organic compound (VOC) profiling—aims to detect sustained metabolic deviation from baseline. These inputs converge into dynamic risk calibration, guiding triage toward imaging, endoscopy, or biopsy.
Figure 6.
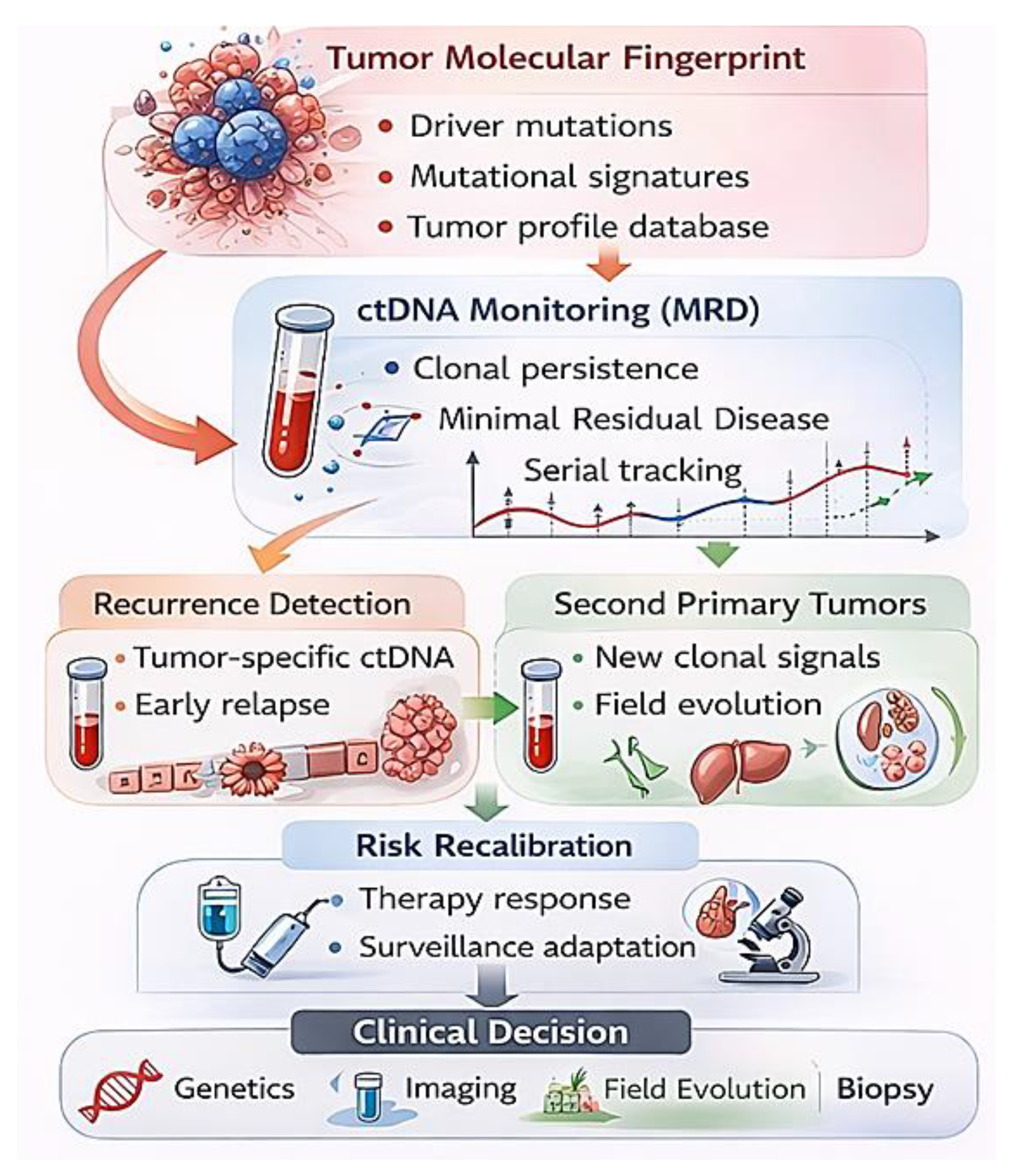
Molecular surveillance framework for post-primary cancer recurrence and second primary detection. In patients with a history of prior cancer, surveillance focuses on recurrence and second primary detection. Molecular fingerprinting of the primary tumor establishes a patient-specific genomic template. Circulating tumor DNA (ctDNA) monitoring enables serial assessment of minimal residual disease and clonal persistence. Concordant signals suggest recurrence, whereas divergent clonal profiles may indicate second primary tumors arising within a conditioned field. Ongoing molecular monitoring supports risk recalibration and informs clinical decision-making, including imaging, biopsy, or therapeutic intervention.
Figure 6.
Molecular surveillance framework for post-primary cancer recurrence and second primary detection. In patients with a history of prior cancer, surveillance focuses on recurrence and second primary detection. Molecular fingerprinting of the primary tumor establishes a patient-specific genomic template. Circulating tumor DNA (ctDNA) monitoring enables serial assessment of minimal residual disease and clonal persistence. Concordant signals suggest recurrence, whereas divergent clonal profiles may indicate second primary tumors arising within a conditioned field. Ongoing molecular monitoring supports risk recalibration and informs clinical decision-making, including imaging, biopsy, or therapeutic intervention.
Table 1.
The Paradigm Shift in Cancer Screening The table summarizes the core transition from traditional methods to a personalized baseline approach.
Table 1.
The Paradigm Shift in Cancer Screening The table summarizes the core transition from traditional methods to a personalized baseline approach.
| Feature |
Traditional Screening |
Personalized Baseline Medicine |
| Conceptual Foundation |
Population-level risk (age, sex, limited history) |
Individual biological "set point" |
| Diagnostic Logic |
Threshold-based (static cut-offs) |
Trajectory-based (dynamic drift from baseline) |
| Testing Frequency |
Episodic (e.g., every 2–5 years) |
Longitudinal / Integrated surveillance |
| Intervention Goal |
Early clinical/anatomic detection |
Molecular interception at inception |
Table 2.
Key Germline Susceptibility Syndromes and Management Strategies. The table details the genetic vulnerabilities that transform early detection into a life-course strategy. HBOC: Hereditary Breast and Ovarian Cancer; HDGC: Hereditary Diffuse Gastric Cancer. MMR: Mismatch Repair; MSI: Microsatellite Instability.
Table 2.
Key Germline Susceptibility Syndromes and Management Strategies. The table details the genetic vulnerabilities that transform early detection into a life-course strategy. HBOC: Hereditary Breast and Ovarian Cancer; HDGC: Hereditary Diffuse Gastric Cancer. MMR: Mismatch Repair; MSI: Microsatellite Instability.
| Syndrome |
Primary Genes |
Associated Cancer Risks |
Sentinel/Clinical Action |
| HBOC |
BRCA1, BRCA2 |
Breast, Ovarian, Pancreatic, Prostate |
Prophylactic surgery; PARP inhibitor eligibility |
| Lynch Syndrome |
MLH1, MSH2, MSH6, PMS2 |
Colorectal, Endometrial, Gastric, Urothelial |
Universal MMR screening; MSI-high immunotherapy |
| HDGC |
CDH1 |
Diffuse Gastric, Lobular Breast |
Prophylactic total gastrectomy |
| Peutz–Jeghers |
STK11 |
Pancreatic, GI, Breast, Gynecologic |
Specialized pancreatic imaging in early adulthood |
Table 3.
Mechanistic Classes of VOCs in Colorectal Cancer The table links metabolic output to the underlying tumor-microbiome interaction.
Table 3.
Mechanistic Classes of VOCs in Colorectal Cancer The table links metabolic output to the underlying tumor-microbiome interaction.
| VOC Class |
Example Compounds |
Mechanistic Pathway |
| Sulfur-Containing |
Methyl mercaptan, Dimethyl trisulfide |
Altered sulfur amino acid metabolism; bacterial H2S production |
| Short-Chain Fatty Acids |
Butyrate (often reduced) |
Microbiome dysbiosis; loss of anti-inflammatory signaling |
| Aldehydes & Ketones |
Volatile aldehydes |
Lipid peroxidation; mitochondrial dysfunction under oxidative stress |
| Nitrogenous |
Indoles |
Microbial tryptophan metabolism; modulation of AhR signaling |
|
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).