Submitted:
08 March 2026
Posted:
12 March 2026
You are already at the latest version
Abstract
Keywords:
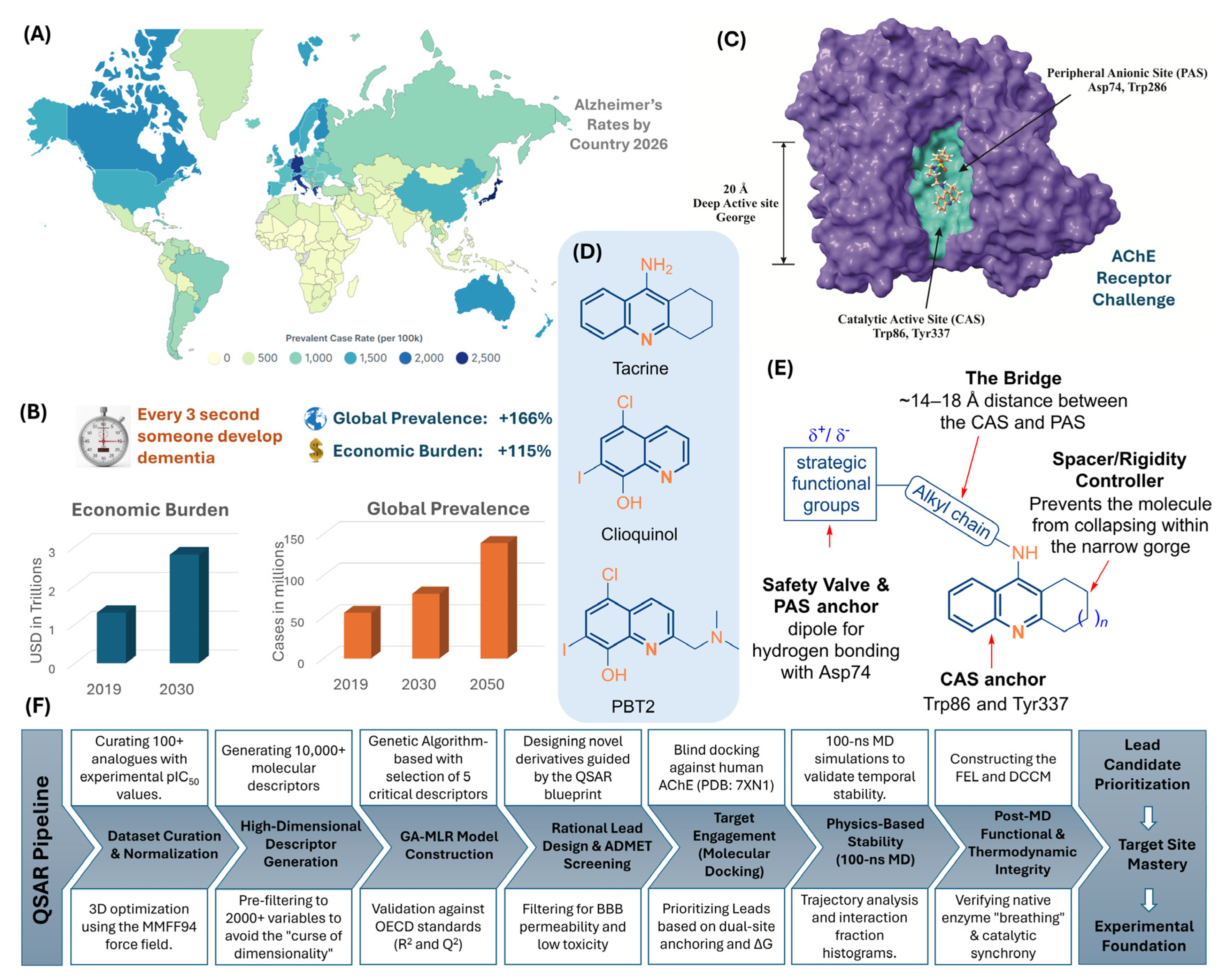
1. Introduction
2. Results and Discussion
2.1. QSAR Model Development and Statistical Robustness
2.2. Mechanistic Interpretation of Molecular Descriptors
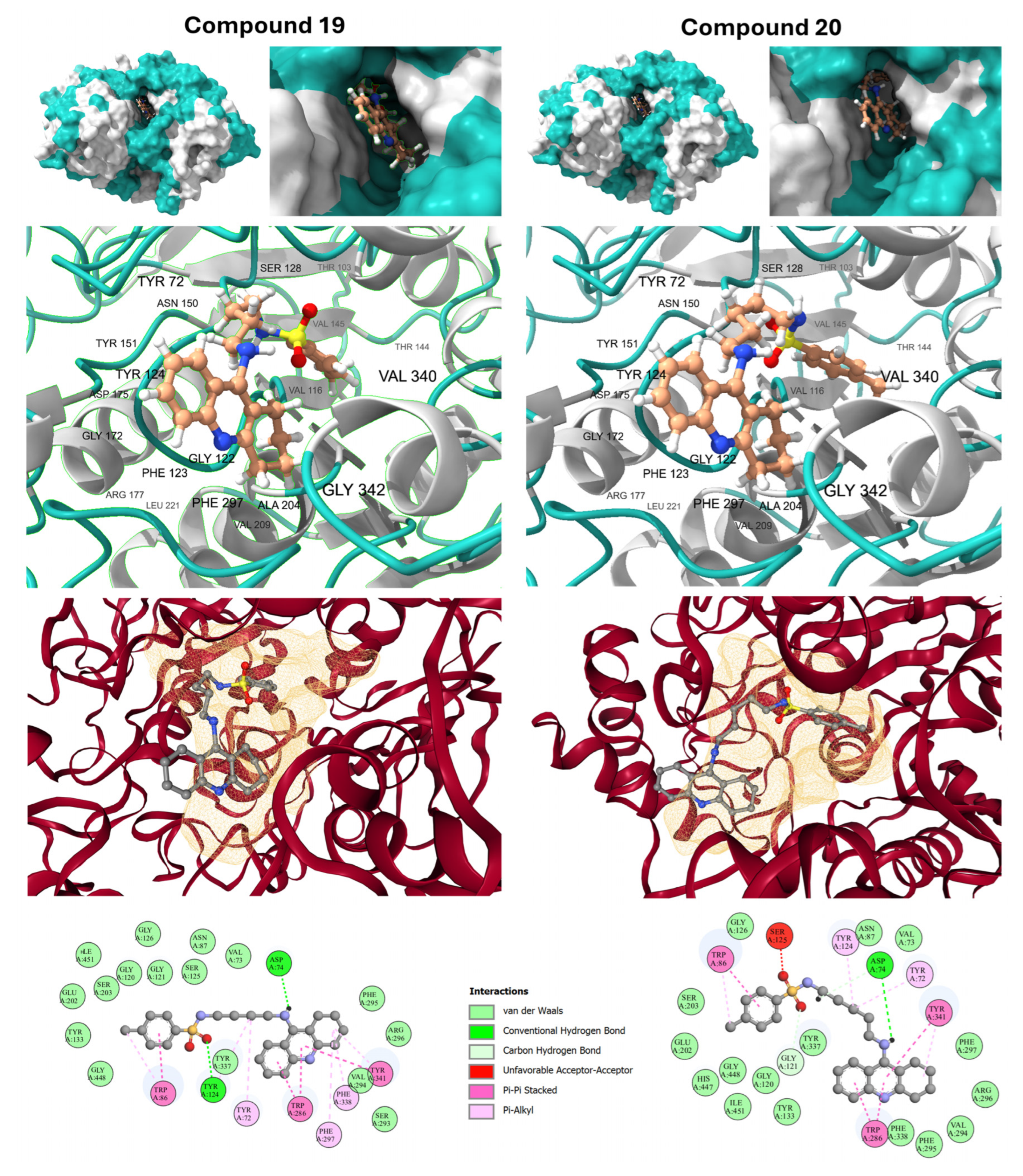
- max_conj_path (alvaDesc): This constitutional descriptor quantifies the length of the longest continuous π-conjugation path within the molecule. The positive coefficient identifies extended π-systems as primary potency drivers. Mechanistically, this facilitates high-occupancy π-π stacking interactions with the aromatic residues lining the 20-Å-deep enzyme gorge, specifically Trp86 in the CAS and Trp286/Tyr341 in the PAS.
- MATS3s (Dragon7): This 2D-autocorrelation index represents Moran autocorrelation of lag 3 weighted by the intrinsic state (I-state). It describes the electronic distribution and polarization across the molecular graph at a specific topological distance. In the context of AChE, electronic density distributed at a lag of 3 is critical for facilitating favorable electrostatic and π-cation interactions within the narrow gorge, particularly with Phe338.
- R8m (Dragon7): A GETAWAY (Geometry, Topology, and Atom Weights AssemblY) descriptor, R8m denotes R autocorrelation of lag 8 weighted by mass. This topological index captures the longitudinal reach and mass distribution of the scaffold. Its selection highlights the requirement for an inhibitor to effectively bridge the ~14–18 Å distance between the catalytic and peripheral binding sites within the human enzyme crystal structure (7XN1).
- C-N-C=O (Fragmentor): This fragment-based descriptor identifies the presence of carbamoyl or urea-like functional groups. Moieties identified by this descriptor act as essential hydrogen-bond donors or acceptors, facilitating polar contacts with the residues of the catalytic triad, such as Glu202 and Ser203, which are vital for competitive inhibition of acetylcholine hydrolysis.
- MNA (Multilevel Neighborhoods of Atoms): Multilevel Neighborhoods of Atoms (MNA) are 2D-substructural notations that describe the local environment of an atom. This specific MNA string accounts for subtle steric and hydrophobic effects in methyl-bearing local environments. Its inclusion suggests that the van der Waals fit within the narrow hydrophobic mid-gorge is highly sensitive to substitution patterns that modulate the local atomic neighborhood.
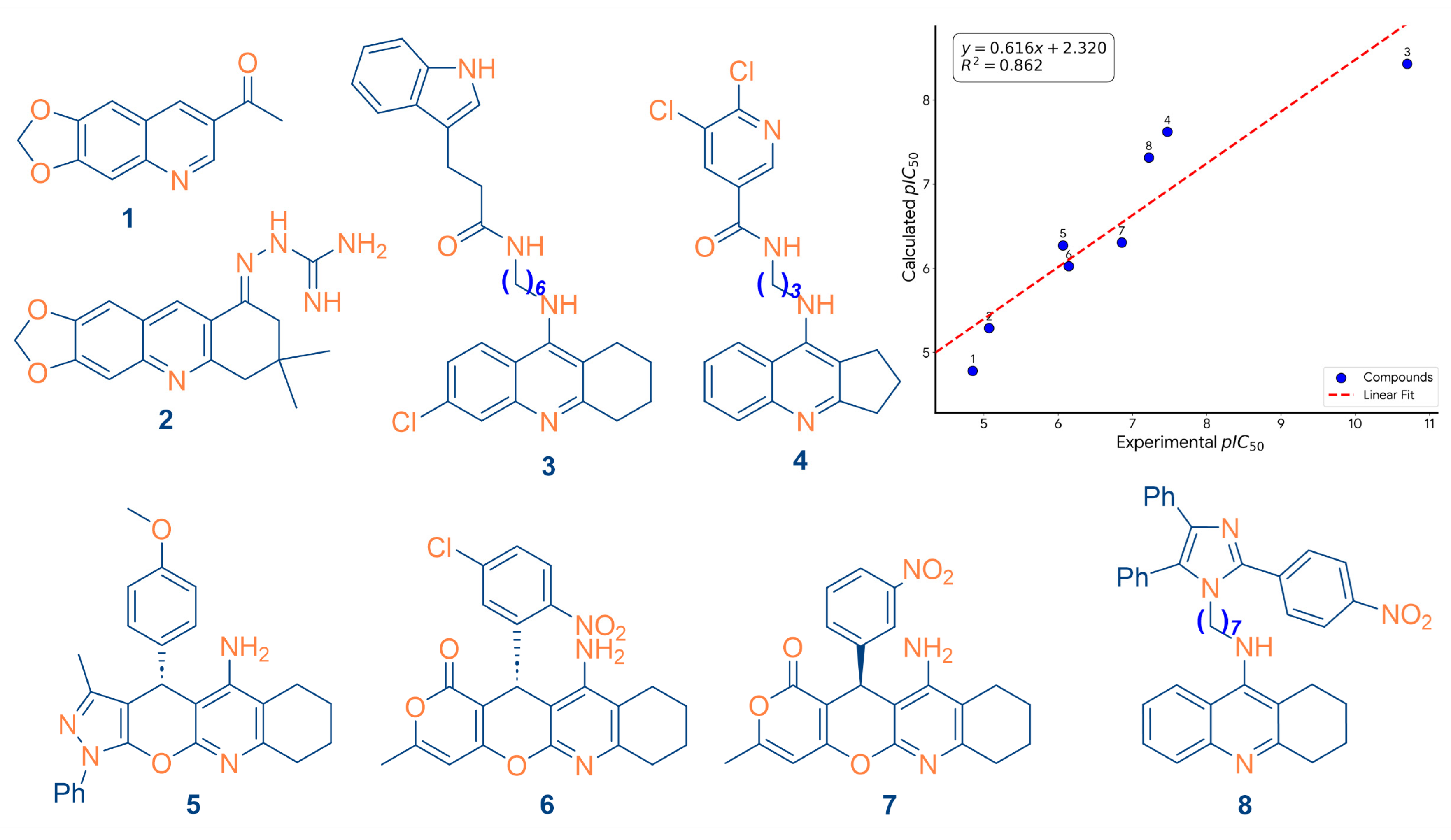
2.3. External Validation and Structural Generalizability
| Compound | 1 | 2 | 3 | 4 | 5 | 6 | 7 | 8 |
|---|---|---|---|---|---|---|---|---|
| Calculated pIC50 | 4.782 | 5.288 | 8.429 | 7.620 | 6.272 | 6.024 | 6.306 | 7.314 |
| Experimental pIC50 | 4.849 | 5.070 | 10.700 | 7.469 | 6.066 | 6.143 | 6.857 | 7.221 |
| Reference | [30] | [31] | [31] | [30] | [30] | [32] | [32] | [33] |
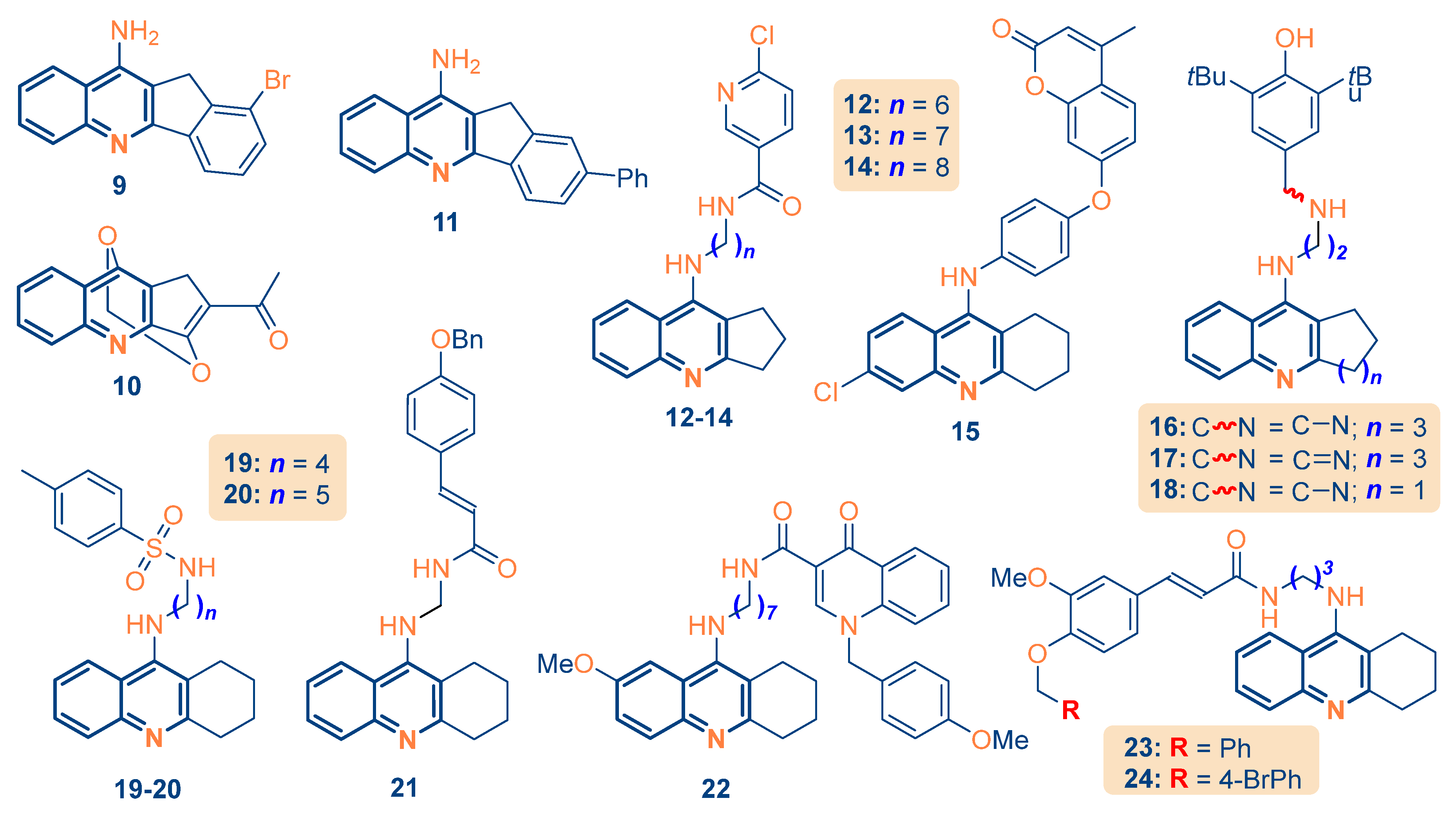
2.4. Design and Predictive Profiling of Novel Fused Quinolines
2.5. ADMET Screening and Toxicity Observations
2.6. Molecular Docking and Active-Site Gorge Interaction
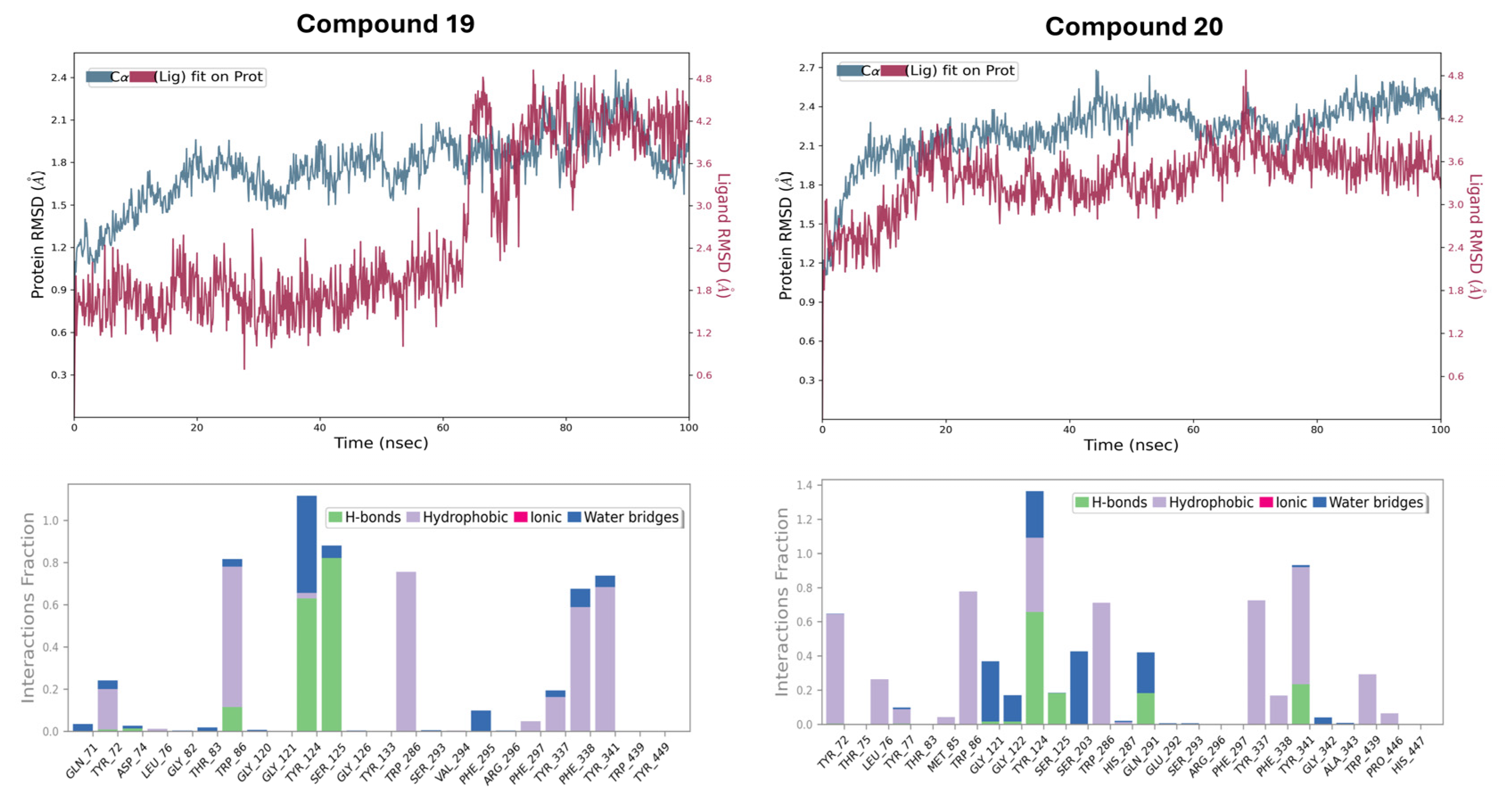
2.7. Molecular Dynamics (MD) Simulation and Dynamic Stability
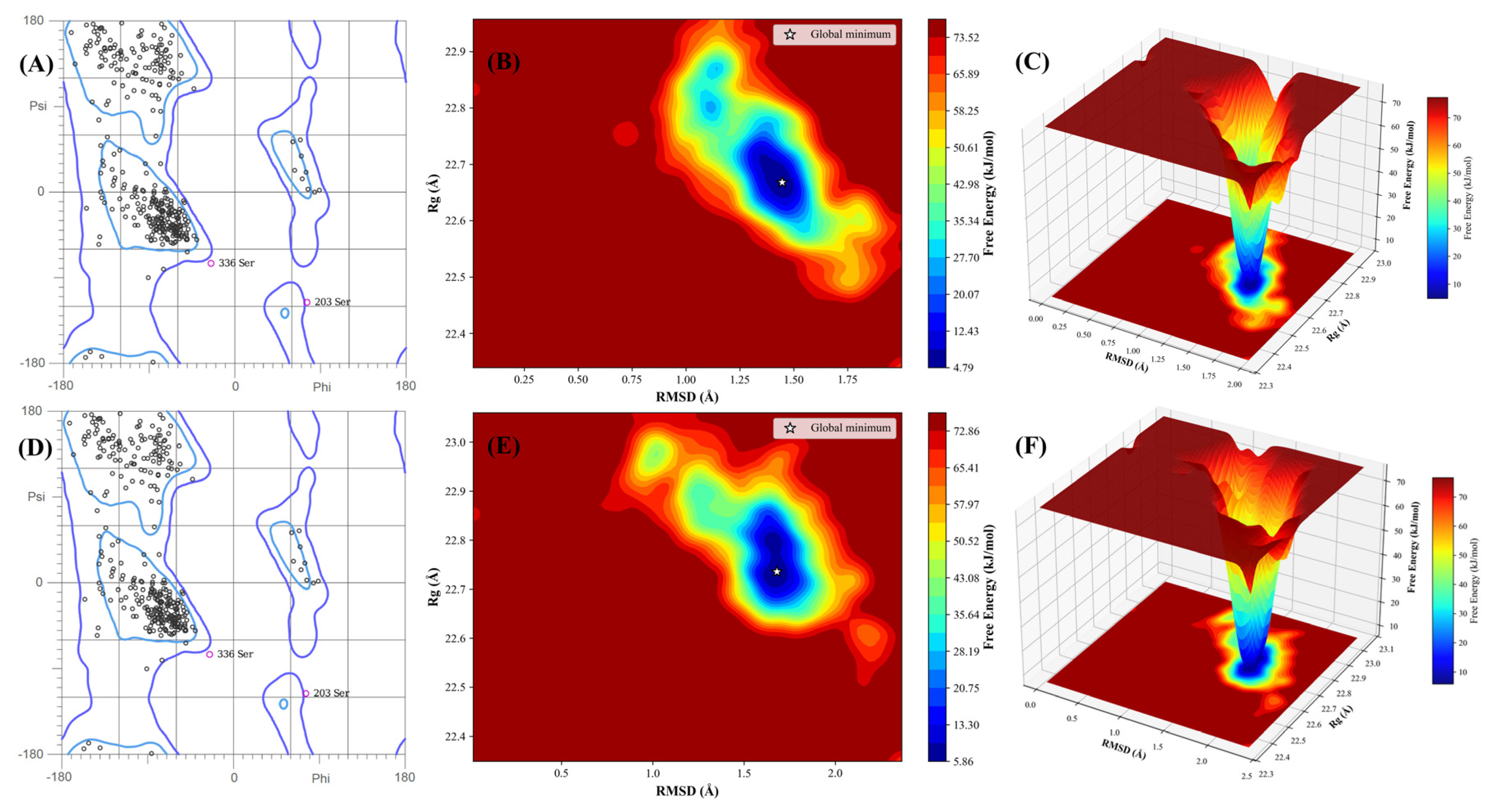
2.8. Post-MD Integrity and Energy Landscapes
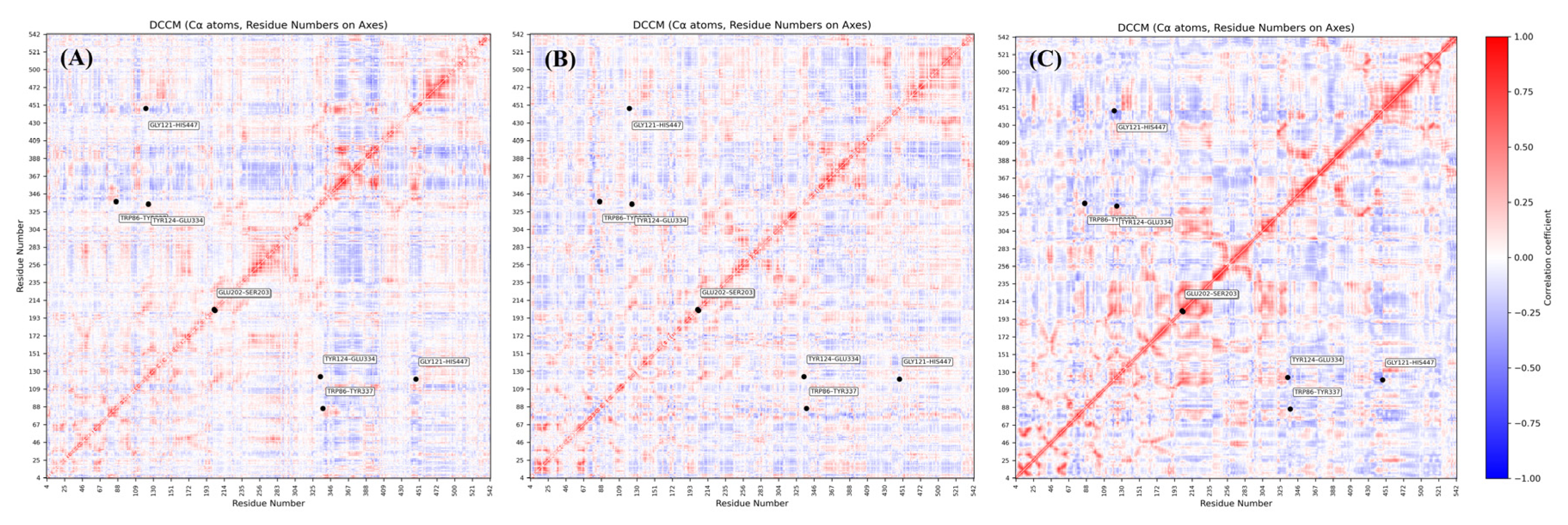
2.9. Functional Residue Synchrony and Catalytic Integrity
3. Materials and Methods
3.1. Dataset Preparation and Computational Framework
3.2. QSAR Model Construction and Regulatory Validation
3.3. Rational Design and ADMET Evaluation
3.4. Molecular Docking and Molecular Dynamics Protocols
3.5. Post-Simulation Integrity and Functional Dynamics Analysis
5. Conclusions
Supplementary Materials
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Cummings, J.; Zhou, Y.; Lee, G.; Zhong, K.; Fonseca, J.; Cheng, F. Alzheimer’s disease drug development pipeline: 2023. Alzheimer’s Dement. (N. Y.) 2023, 9, e12385. [Google Scholar] [CrossRef] [PubMed]
- Xiaopeng, Z.; Jing, Y.; Xia, L.; Xingsheng, W.; Juan, D.; Yan, L.; Baoshan, L. Global Burden of Alzheimer’s disease and other dementias in adults aged 65 years and older, 1991-2021: population-based study. Front. Public Health 2025, 13, 1585711. [Google Scholar] [CrossRef] [PubMed]
- World Population Review. Alzheimer’s Rates by Country 2026. Available online: https://worldpopulationreview.com/country-rankings/alzheimers-rates-by-country (accessed on 3 February 2026).
- Alzheimer’s Disease International. Dementia Statistics. Available online: https://www.alzint.org/about/dementia-facts-figures/dementia-statistics/ (accessed on 3 February 2026).
- Yiannopoulou, K.G.; Anastasiou, A.I.; Zachariou, V.; Pelidou, S.-H. Reasons for Failed Trials of Disease-Modifying Treatments for Alzheimer Disease and Their Contribution in Recent Research. Biomedicines 2019, 7, 97. [Google Scholar] [CrossRef]
- Kong, L.; Yang, X.; Xu, J. Comparison of safety of lecanemab and aducanumab: a real-world disproportionality analysis using the FDA adverse event reporting system. Front. Pharmacol. 2025, 16, 1593989. [Google Scholar] [CrossRef] [PubMed]
- Hampel, H.; Elhage, A.; Cho, M.; Apostolova, L.G.; Nicoll, J.A.R.; Atri, A. Amyloid-related imaging abnormalities (ARIA): radiological, biological and clinical characteristics. Brain 2023, 146, 4414–4424. [Google Scholar] [CrossRef]
- Wu, B.; Hu, Q.; Tian, F.; Wu, F.; Li, Y.; Xu, T. A pharmacovigilance study of association between proton pump inhibitor and dementia event based on FDA adverse event reporting system data. Sci. Rep. 2021, 11, 10709. [Google Scholar] [CrossRef]
- Saeed, R.; Tariq, H.Z.; Althobaiti, A.; Sadeghian, N.; Taslimi, P.; Al-Rashida, M.; Islam, T.; Thabet, H.K.; Aftab, H.; Şenol, H.; et al. Design, synthesis, and multi-target evaluation of 4-phenyl quinoline-8-sulfonate thiosemicarbazones as potential anti-Alzheimer agents. Sci. Rep. 2025, 15, 44212. [Google Scholar] [CrossRef]
- Grabowska, W.; Bijak, M.; Szelenberger, R.; Gorniak, L.; Podogrocki, M.; Harmata, P.; Cichon, N. Acetylcholinesterase as a Multifunctional Target in Amyloid-Driven Neurodegeneration: From Dual-Site Inhibitors to Anti-Agregation Strategies. Int. J. Mol. Sci. 2025, 26, 8726. [Google Scholar] [CrossRef]
- Cheng, S.; Song, W.; Yuan, X.; Xu, Y. Gorge Motions of Acetylcholinesterase Revealed by Microsecond Molecular Dynamics Simulations. Sci. Rep. 2017, 7, 3219. [Google Scholar] [CrossRef]
- Rawat, K.; Tewari, D.; Bisht, A.; Chandra, S.; Tiruneh, Y.K.; Hassan, H.M.; Al-Emam, A.; Sindi, E.R.; Al-Dies, A.-A.M. Identification of AChE targeted therapeutic compounds for Alzheimer’s disease: an in-silico study with DFT integration. Sci. Rep. 2024, 14, 30356. [Google Scholar] [CrossRef]
- Marucci, G.; Buccioni, M.; Dal Ben, D.; Lambertucci, C.; Volpini, R.; Amenta, F. Efficacy of acetylcholinesterase inhibitors in Alzheimer’s disease. Neuropharmacology 2021, 190, 108352. [Google Scholar] [CrossRef]
- Su, T.; Zhang, T.; Xie, S.; Yan, J.; Wu, Y.; Li, X.; Huang, L.; Luo, H.-B. Discovery of novel PDE9 inhibitors capable of inhibiting Aβ aggregation as potential candidates for the treatment of Alzheimer’s disease. Sci. Rep. 2016, 6, 21826. [Google Scholar] [CrossRef]
- Zhang, Y.; Chen, H.; Li, R.; Sterling, K.; Song, W. Amyloid β-based therapy for Alzheimer’s disease: challenges, successes and future. Sig. Transduct. Target Ther. 2023, 8, 248. [Google Scholar] [CrossRef] [PubMed]
- Hampel, H.; Hardy, J.; Blennow, K.; Chen, C.; Perry, G.; Kim, S.H.; Villemagne, V.L.; Aisen, P.; Vendruscolo, M.; Iwatsubo, T.; et al. The Amyloid-β Pathway in Alzheimer’s Disease. Mol. Psychiatry 2021, 26, 5481–5503. [Google Scholar] [CrossRef]
- Luo, J.; Wärmländer, S.K.; Gräslund, A.; Abrahams, J.P. Cross-interactions between the Alzheimer Disease Amyloid-β Peptide and Other Amyloid Proteins: A Further Aspect of the Amyloid Cascade Hypothesis. J. Biol. Chem. 2016, 291, 16485–16493. [Google Scholar] [CrossRef]
- Goyal, M.; Dhanjal, J.K.; Goyal, S.; Tyagi, C.; Hamid, R.; Grover, A. Development of dual inhibitors against Alzheimer’s disease using fragment-based QSAR and molecular docking. BioMed Res. Int. 2014, 2014, 979606. [Google Scholar] [CrossRef]
- Ferreira, J.P.S.; Albuquerque, H.M.T.; Cardoso, S.M.; Silva, A.M.S.; Silva, V.L.M. Dual-target compounds for Alzheimer’s disease: Natural and synthetic AChE and BACE-1 dual-inhibitors and their structure-activity relationship (SAR). Eur. J. Med. Chem. 2021, 221, 113492. [Google Scholar] [CrossRef]
- Grande, G.; Valletta, M.; Rizzuto, D.; Xia, X.; Qiu, C.; Orsini, N.; Dale, M.; Andersson, S.; Fredolini, C.; Laukka, E.J.; et al. Blood-based biomarkers of Alzheimer’s disease and incident dementia in the community. Nat. Med. 2025, 31, 2027–2035. [Google Scholar] [CrossRef]
- Qizilbash, N.; Birks, J.; Lopez Arrieta, J.; Lewington, S.; Szeto, S. WITHDRAWN: Tacrine for Alzheimer’s disease. Cochrane Database Syst. Rev. 2007, 3, CD000202. [Google Scholar] [CrossRef]
- Avram, S.; Mernea, M.; Limban, C.; Borcan, F.; Chifiriuc, C. Potential Therapeutic Approaches to Alzheimer’s Disease By Bioinformatics, Cheminformatics And Predicted ADMET Tools. Curr. Neuropharmacol. 2020, 18, 696–719. [Google Scholar] [CrossRef] [PubMed]
- Nilewar, S.S.; Chobe, S.; Dudhe, P.; Kumar, P.K.; Lodha, S.; Raut, A.D.; Fernández-Conde, D.; Farhan, M.; Muteeb, G.; Pawar, T.J. An Integrated QSAR-MD-DCCM Pipeline: A Predictive Computational Platform for the Rational Design and Dynamic Functional Validation of Dual-Target Directed Ligands. Pharmaceuticals 2026, 19, 249. [Google Scholar] [CrossRef]
- Moaddel, R.; Wainer, I.W. Development of immobilized membrane-based affinity columns for use in the online characterization of membrane bound proteins and for targeted affinity isolations. Anal. Chim. Acta 2006, 564, 97–105. [Google Scholar] [CrossRef]
- Bastikar, V.; Bastikar, A.; Gupta, P. Quantitative structure–activity relationship-based computational approaches. In Computational Approaches for Novel Therapeutic and Diagnostic Designing to Mitigate SARS-CoV-2 Infection; Elsevier: Amsterdam, The Netherlands, 2022; pp. 191–205. [Google Scholar] [CrossRef]
- Huang, J.; Fan, X. Why QSAR Fails: An Empirical Evaluation Using Conventional Computational Approach. Mol. Pharmaceutics 2011, 8, 600–608. [Google Scholar] [CrossRef] [PubMed]
- Leardi, R. Genetic Algorithms in Feature Selection. In Principles of QSAR and Drug Design; Rogers, J.P., Ed.; Academic Press: San Diego, CA, USA, 1996; pp. 67–86. [Google Scholar] [CrossRef]
- Reddy, A.S.; Kumar, S.; Garg, R. Hybrid-genetic algorithm based descriptor optimization and QSAR models for predicting the biological activity of Tipranavir analogs for HIV protease inhibition. J. Mol. Graph. Model. 2010, 28, 852–862. [Google Scholar] [CrossRef]
- Gupta, K.; Kumar, A.; Patel, R.; Ghode, P.; Kumar, H.; Murmu, A.; Sahu, N.; Verma, G.; Sahu, S.; Soni, S.; et al. Predictive QSAR Models Followed by Toxicity, Molecular Docking, and Molecular Dynamics Simulation in Search of Azole Derivatives as AChE Inhibitors for the Treatment of Alzheimer’s Disease. J. Chemom. 2025, 39, e70049. [Google Scholar] [CrossRef]
- Li, Z.H.; Yin, L.Q.; Zhao, D.H.; Jin, L.H.; Sun, Y.J.; Tan, C. SAR studies of quinoline and derivatives as potential treatments for Alzheimer’s disease. Arab. J. Chem. 2022, 16, 104502. [Google Scholar] [CrossRef]
- Muñoz-Ruiz, P.; Rubio, L.; García-Palomero, E.; Dorronsoro, I.; Monte-Millán, M.D.; Valenzuela, R.; Usán, P.; de Austria, C.; Bartolini, M.; Andrisano, V.; et al. Design, synthesis, and biological evaluation of dual binding site acetylcholinesterase inhibitors: new disease-modifying agents for Alzheimer’s disease. J. Med. Chem. 2005, 48, 7223–7233. [Google Scholar] [CrossRef] [PubMed]
- Hariri, R.; Afshar, Z.; Mahdavi, M.; Safavi, M.; Saeedi, M.; Najafi, Z.; Sabourian, R.; Karimpour-Razkenari, E.; Edraki, N.; Moghadam, F.H.; et al. Novel tacrine-based pyrano [3′,4′:5,6]pyrano [2,3-b]quinolinones: synthesis and cholinesterase inhibitory activity. Arch. Pharm. 2016, 349, 915–924. [Google Scholar] [CrossRef]
- da Costa, J.S.; Bizarro Lopes, J.P.; Russowsky, D.; Petzhold, C.L.; de Amorim Borges, A.C.; Ceschi, M.A.; Konrath, E.; Batassini, C.; Lunardi, P.S.; Saraiva Gonçalves, C.A. Synthesis of tacrine-lophine hybrids via one-pot four component reaction and biological evaluation as acetyl- and butyrylcholinesterase inhibitors. Eur. J. Med. Chem. 2013, 62, 556–563. [Google Scholar] [CrossRef]
- Sushko, I.; Pandey, A.K.; Novotarskyi, S.; Körner, R.; Rupp, M.; Teetz, W.; Brandmaier, S.; Abdelaziz, A.; Prokopenko, V.V.; Tanchuk, V.Y.; et al. Online chemical modeling environment (OCHEM): web platform for data storage, model development and publishing of chemical information. J. Cheminform. 2011, 3 (Suppl 1), P20. [Google Scholar] [CrossRef]
- Gramatica, P.; Chirico, N.; Papa, E.; Cassani, S.; Kovarich, S. QSARINS: A new software for the development, analysis, and validation of QSAR MLR models. J. Comput. Chem. 2013, 34, 2121–2132. [Google Scholar] [CrossRef]
- Gramatica, P.; Cassani, S.; Chirico, N. QSARINS-chem: Insubria datasets and new QSAR/QSPR models for environmental pollutants in QSARINS. J. Comput. Chem. 2014, 35, 1036–1044. [Google Scholar] [CrossRef]
- Fu, L.; Shi, S.; Yi, J.; Wang, N.; He, Y.; Wu, Z.; Peng, J.; Deng, Y.; Wang, W.; Wu, C.; et al. ADMETlab 3.0: an updated comprehensive online ADMET prediction platform enhanced with broader coverage, improved performance, API functionality and decision support. Nucleic Acids Res. 2024, 52, W422–W431. [Google Scholar] [CrossRef]
- Banerjee, P.; Kemmler, E.; Dunkel, M.; Preissner, R. ProTox 3.0: a webserver for the prediction of toxicity of chemicals. Nucleic Acids Res. 2024, 52, W513–W520. [Google Scholar] [CrossRef] [PubMed]
- Dileep, K.; Ihara, K.; Mishima-Tsumagari, C.; Kukimoto-Niino, M.; Yonemochi, M.; Hanada, K.; Shirouzu, M.; Zhang, K.Y. Crystal structure of human acetylcholinesterase in complex with tacrine: Implications for drug discovery. Int. J. Biol. Macromol. 2022, 210, 172–181. [Google Scholar] [CrossRef] [PubMed]
- Yang, X.; Liu, Y.; Gan, J.; Xiao, Z.-X.; Cao, Y. FitDock: protein–ligand docking by template fitting. Brief. Bioinform. 2022, 23, bbac087. [Google Scholar] [CrossRef]
- Liu, Y.; Yang, X.; Gan, J.; Chen, S.; Xiao, Z.-X.; Cao, Y. CB-Dock2: improved protein–ligand blind docking by integrating cavity detection, docking and homologous template fitting. Nucleic Acids Res. 2022, 50, W159–W164. [Google Scholar] [CrossRef]
- Meng, E.C.; Goddard, T.D.; Pettersen, E.F.; Ferrin, T.E.; ChimeraX, U.S. UCSF ChimeraX: Tools for structure building and analysis. Protein Sci. 2023, 32, e4792. [Google Scholar] [CrossRef]
- Schöning-Stierand, K.; Diedrich, K.; Ehrt, C.; Flachsenberg, F.; Graef, J.; Sieg, J.; Penner, P.; Poppinga, M.; Ungethüm, A.; Rarey, M. ProteinsPlus: a comprehensive collection of web-based molecular modeling tools. Nucleic Acids Res. 2022, 50, W611–W615. [Google Scholar] [CrossRef]
- Fährrolfes, R.; Bietz, S.; Flachsenberg, F.; Meyder, A.; Nittinger, E.; Rarey, M. ProteinsPlus: a web portal for structure analysis of macromolecules. Nucleic Acids Res. 2017, 45, W337–W343. [Google Scholar] [CrossRef] [PubMed]
- Schöning-Stierand, K.; Diedrich, K.; Fährrolfes, R.; Flachsenberg, F.; Meyder, A.; Nittinger, E.; Steinegger, R.; Rarey, M. ProteinsPlus: interactive analysis of protein–ligand binding interfaces. Nucleic Acids Res. 2020, 48, W48–W53. [Google Scholar] [CrossRef] [PubMed]
- Bowers, K.J.; Chow, D.E.; Xu, H.; Dror, R.O.; Eastwood, M.P.; Gregersen, B.A.; Klepeis, J.L.; Kolossvary, I.; Moraes, M.A.; Sacerdoti, F.D.; et al. Scalable Algorithms for Molecular Dynamics Simulations on Commodity Clusters. In Proceedings of the 2006 ACM/IEEE Conference on Supercomputing (SC’06); IEEE: Tampa, FL, USA, 2006; p. 43. [Google Scholar] [CrossRef]
- Nilewar, S.S.; Chobe, S.S.; Gurav, A.D.; Kureshi, S.B.; Palande, S.B.; Escobar-Cabrera, J.; Hernández-Rosas, F.; Pawar, T.J. Galloylation-Driven Anchoring of the Asp325-Asp336 Ridge: The Molecular Logic Behind the Superior Kinetic Stabilization of HMPV Fusion Protein by Green Tea Dimeric Catechins. Molecules 2026, 31, 821. [Google Scholar] [CrossRef]
- Michaud-Agrawal, N.; Denning, E.J.; Woolf, T.B.; Beckstein, O. MDAnalysis: A toolkit for the analysis of molecular dynamics simulations. J. Comput. Chem. 2011, 32, 2319–2327. [Google Scholar] [CrossRef] [PubMed]
- Williams, C.J.; Headd, J.J.; Moriarty, N.W.; Prisant, M.G.; Videau, L.L.; Deis, L.N.; Verma, V.; Keedy, D.A.; Hintze, B.J.; Chen, V.B.; et al. MolProbity: More and better reference data for improved all-atom structure validation. Protein Sci. 2018, 27, 293–315. [Google Scholar] [CrossRef] [PubMed]
- Hunter, J.D. Matplotlib: A 2D Graphics Environment. Comput. Sci. Eng. 2007, 9, 90–95. [Google Scholar] [CrossRef]
- Virtanen, P.; Gommers, R.; Oliphant, T.E.; Haberland, M.; Reddy, T.; Cournapeau, D.; Burovski, E.; Peterson, P.; Weckesser, W.; Bright, J.; van der Walt, S.J.; Brett, M.; Wilson, J.; Millman, K.J.; Mayorov, N.; Nelson, A.R.J.; Jones, E.; Kern, R.; Larson, E.; Carey, C.J.; Polat, İ.; Feng, Y.; Moore, E.W.; VanderPlas, J.; Laxalde, D.; Perktold, J.; Cimrman, R.; Henriksen, I.; Quintero, E.A.; Harris, C.R.; Archibald, A.M.; Ribeiro, A.H.; Pedregosa, F.; van Mulbregt, P. SciPy 1.0 Contributors. SciPy 1.0: fundamental algorithms for scientific computing in Python. Nat. Methods 2020, 17, 261–272. [Google Scholar] [CrossRef]
- Harris, C.R.; Millman, K.J.; van der Walt, S.J.; Gommers, R.; Virtanen, P.; Cournapeau, D.; Wieser, E.; Taylor, J.; Berg, S.; Smith, N.J.; Kern, R.; Picus, M.; Hoyer, S.; van Kerkwijk, M.H.; Brett, M.; Haldane, A.; del Río, J.F.; Wiebe, M.; Peterson, P.; Gérard-Marchant, P.; Sheppard, K.; Reddy, T.; Weckesser, W.; Abbasi, H.; Gohlke, C.; Oliphant, T.E. Array programming with NumPy. Nature 2020, 585, 357–362. [Google Scholar] [CrossRef]
- Ichiye, T.; Karplus, M. Collective motions in proteins: a covariance analysis of atomic fluctuations in molecular dynamics and normal mode simulations. Proteins 1991, 11, 205–217. [Google Scholar] [CrossRef] [PubMed]
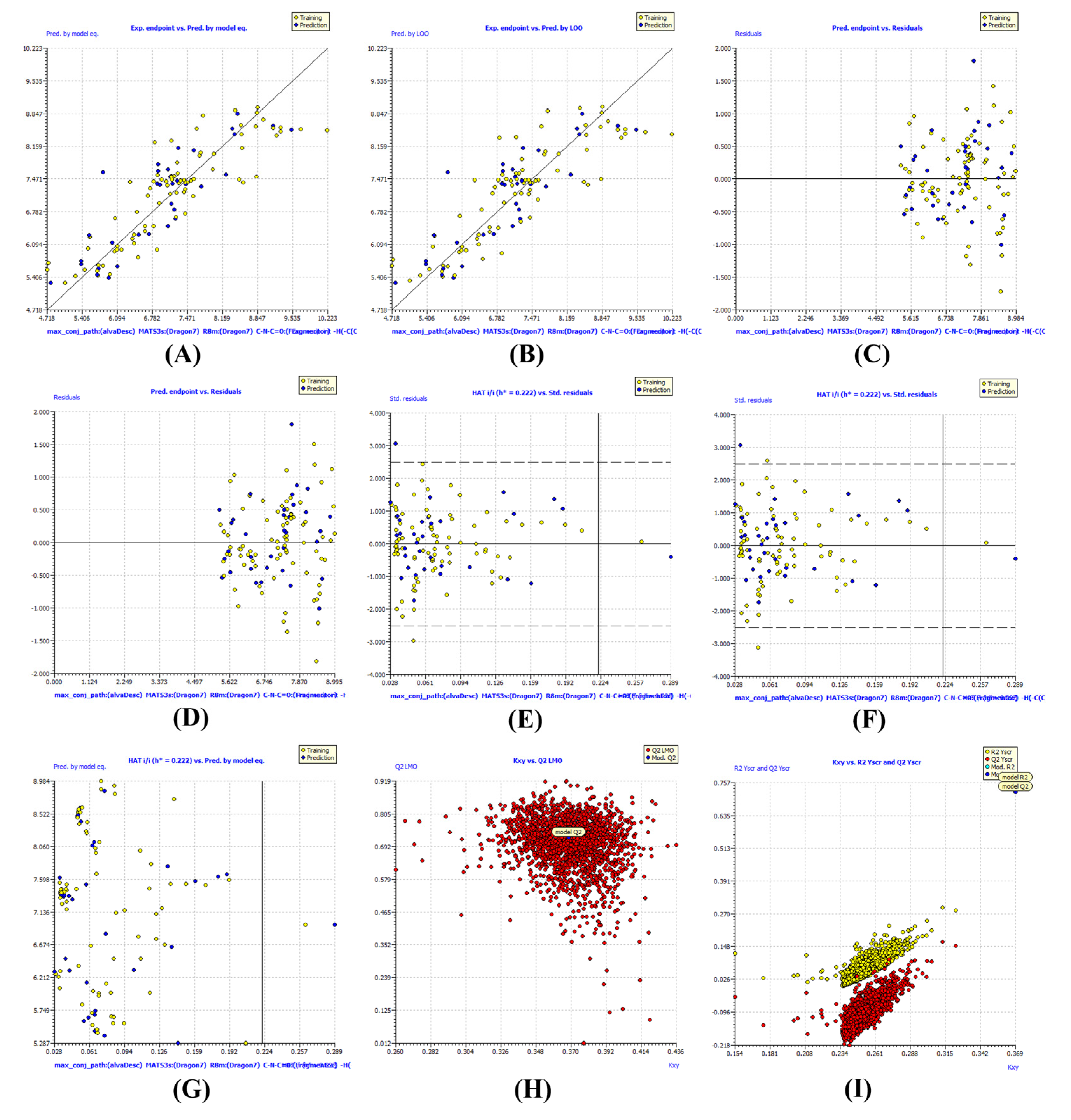
| Metric | QSAR Model | Threshold |
|---|---|---|
| R2 (training) | 0.7569 | >0.60 |
| Adjusted R2 | 0.7407 | Close to R2 |
| RMSE (training) | 0.5752 | Lower is better |
| MAE (training) | 0.4367 | Lower is better |
| Q2 (LOO) | 0.7244 | >0.50 |
| Q2 (LMO) | 0.7178 | >0.60 |
| CCC (training) | 0.8616 | >0.85 |
| (external, n = 8) | 0.737 | >0.60 |
| CCC (external) | 0.8509 | >0.85 |
| (average) | 0.6257 | >0.50 |
| 0.0996 | <0.20 |
| Compd | Modifications done | Descriptors Used | Rationale Behind Structure Design | pIC50 |
|---|---|---|---|---|
| 2 | Fused quinoline with alkyl substitutions and ester groups | max_conj_path, MAT53s, C-N-C=O | Alkyl and ester groups introduce flexibility and increase the complexity of molecular geometry, impacting polarity and size. | 4.3725 |
| 4 | Fused quinoline with amine, halogen, and ester groups | max_conj_path, MAT53s, R8m | Amine, halogen, and ester substitutions influence molecular shape, electronic properties, and size. | 8.0588 |
| 9 | Fused quinoline with alkyl and halogen substitutions | max_conj_path, MAT53s, R8m | Fused quinoline rings enhance conjugation, while alkyl and halogen substitutions influence lipophilicity and molecular shape. | 6.1265 |
| 10 | Fused quinoline attached to a benzene ring with ester groups |
max_conj_path, MAT53s, R8m, C-N-C=O |
Ester and benzene ring groups increase topological complexity, modifying polarity and molecular geometry. | 5.9286 |
| 11 | Fused quinoline with ester and amine substitutions | max_conj_path, MAT53s, R8m | Ester and amine substitutions modify molecular shape and increase flexibility, altering lipophilicity and geometry. | 6.3097 |
| 12 | Fused quinoline with extended amines and halogen substitutions | max_conj_path, MAT53s, C-N-C=O | Extended amines and halogen groups introduce greater flexibility and polarity, affecting molecular interactions. | 7.4803 |
| 13 | Fused quinoline with extended carbon chains and halogen | max_conj_path, MAT53s, R8m | Longer alkyl chains and halogen substitutions alter hydrophobicity and increase molecular complexity. | 7.5403 |
| 14 | Extended quinoline with additional carbon chains and halogen | max_conj_path, MAT53s, C-N-C=O | Additional carbon chains and halogen groups increase hydrophobicity and modify molecular geometry for better structural balance. | 7.5616 |
| 15 | Extended quinoline fused with a cyclic group, with ester substitutions |
max_conj_path, MAT53s, R8m, -H(-C(C-H-H-C)) |
Cyclic structures and ester substitutions modify the molecular geometry, improving structural complexity and hydrophobicity. | 5.6982 |
| 16 | Extended quinoline with multiple substitution groups |
max_conj_path, MAT53s, R8m, -H(-C(C-H-H-C)) |
Multiple substitutions alter the topological structure, increasing rigidity and adjusting molecular shape. | 5.3384 |
| 17 | Fused quinoline with additional nitrogen substitutions |
max_conj_path, MAT53s, R8m, C-N-C=O |
Nitrogen substitutions influence polarity and the complexity of the molecular structure, modifying its topology. | 5.3398 |
| 18 | Fused quinoline with nitrogen-containing functionalization |
max_conj_path, MAT53s, R8m, -H(-C(C-H-H-C)) |
Nitrogen-containing functional groups increase polarity and modify the shape of the molecule for better structural interaction. | 5.1299 |
| 19 | Fused quinoline with sulfonate group and amines |
max_conj_path, MAT53s, R8m, -H(-C(C-H-H-C)) |
Sulfonate groups add polarity, while amines introduce additional flexibility and complexity to the molecular structure. | 5.8935 |
| 20 | Sulfonate substituted quinoline fused with amine groups |
max_conj_path, MAT53s, R8m, -H(-C(C-H-H-C)) |
Sulfonate and amine substitutions affect polarity and increase the complexity of the molecular structure. | 5.9529 |
| 21 | Additional functional groups and fused rings | max_conj_path, MAT53s, C-N-C=O | Fused rings and additional functional groups introduce rigidity, modifying the shape and flexibility of the molecule. | 7.1775 |
| 22 | Fused quinoline with extended ring systems and ester functionalities | max_conj_path, MAT53s, R8m | Extended ring systems and ester groups provide additional rigidity and alter molecular flexibility. | 7.7726 |
| 23 | Extended quinoline with ester and oxygen-containing functional groups |
max_conj_path, MAT53s, R8m, C-N-C=O |
Ester and oxygen-containing groups enhance polarity and hydrophobicity, modifying the molecular geometry for better flexibility. | 7.1909 |
| 24 | Fused quinoline with halogen, ester, and amine groups |
max_conj_path, MAT53s, R8m, -H(-C(C-H-H-C)) |
Halogen, ester, and amine groups increase hydrophobicity and polarity, modifying the overall shape and topological complexity. | 7.0912 |
| Parameter | 19 | 20 | Tacrine |
|---|---|---|---|
| QED | 0.520 | 0.463 | 0.706 |
| logS (log mol/L) | -4.698 | -4.898 | -2.875 |
| Synth | 2.268 | 2.292 | 2.015 |
| caco2 (log cm/s) | -4.956 | -5.000 | -4.677 |
| logD | 3.703 | 3.748 | 2.101 |
| BBB | 0.647 | 0.768 | 0.977 |
| logP | 4.385 | 4.556 | 2.432 |
| ΔG (kcal/mol) | -11.1 | -10.6 | -9.0 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.




