Submitted:
05 March 2026
Posted:
05 March 2026
You are already at the latest version
Abstract
Keywords:
Introduction
1. Definition of Core Terms
- Multi-track Cognition (MTC): The original core theory proposed in this study, which refers to a cognitive framework that constructs independent and interoperable data processing channels for different medical systems, to preserve the native theoretical logic of each system while realizing cross-system semantic alignment.
- Cognitive Track (CT): A standardized data processing channel constructed corresponding to the native logic of a single medical system, which completely retains its core feature dimensions, evaluation rules, and application logic without forced cross-paradigm transformation, providing an independent and complete data processing and logic mapping link for each independent medical system, with the formal definition: an independent data processing pathway preserving the native logic of a medical system.
- Homeostasis: The core basic concept of this study, which refers to the dynamic equilibrium state maintained by the human physiological system through self-regulation under the changes of internal and external environments, and is the core quantitative representation of health status, rather than the index stability under disease state.
- Homeostatic Representation Network (HRN): A structured data network with human system homeostasis as the core, composed of quantifiable homeostatic dimensions with predefined association relationships and target ranges. It is the only general quantitative mediation benchmark and semantic anchor for cross-system data interoperability, with the formal definition: a structured network of quantifiable homeostatic dimensions with predefined target ranges.
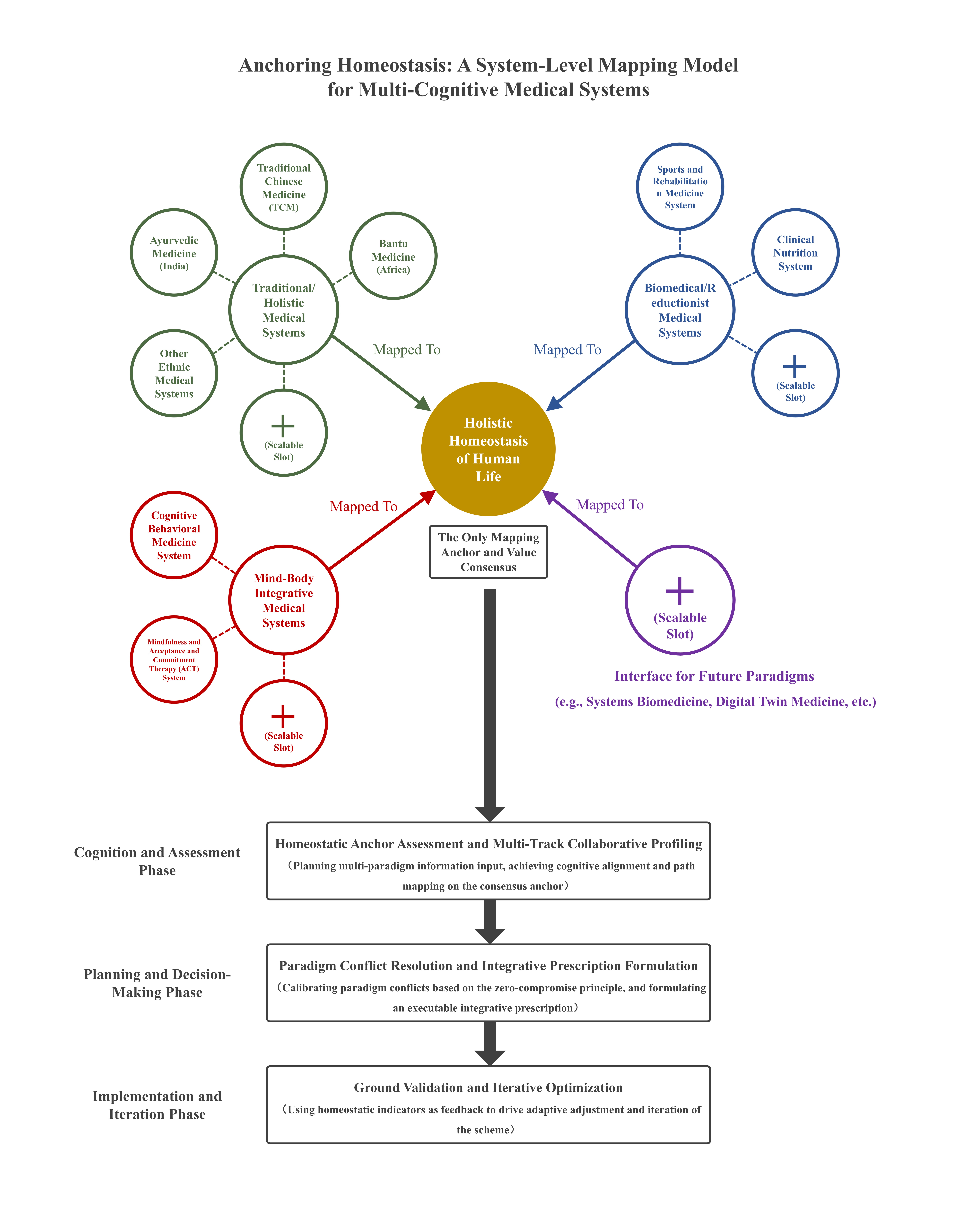
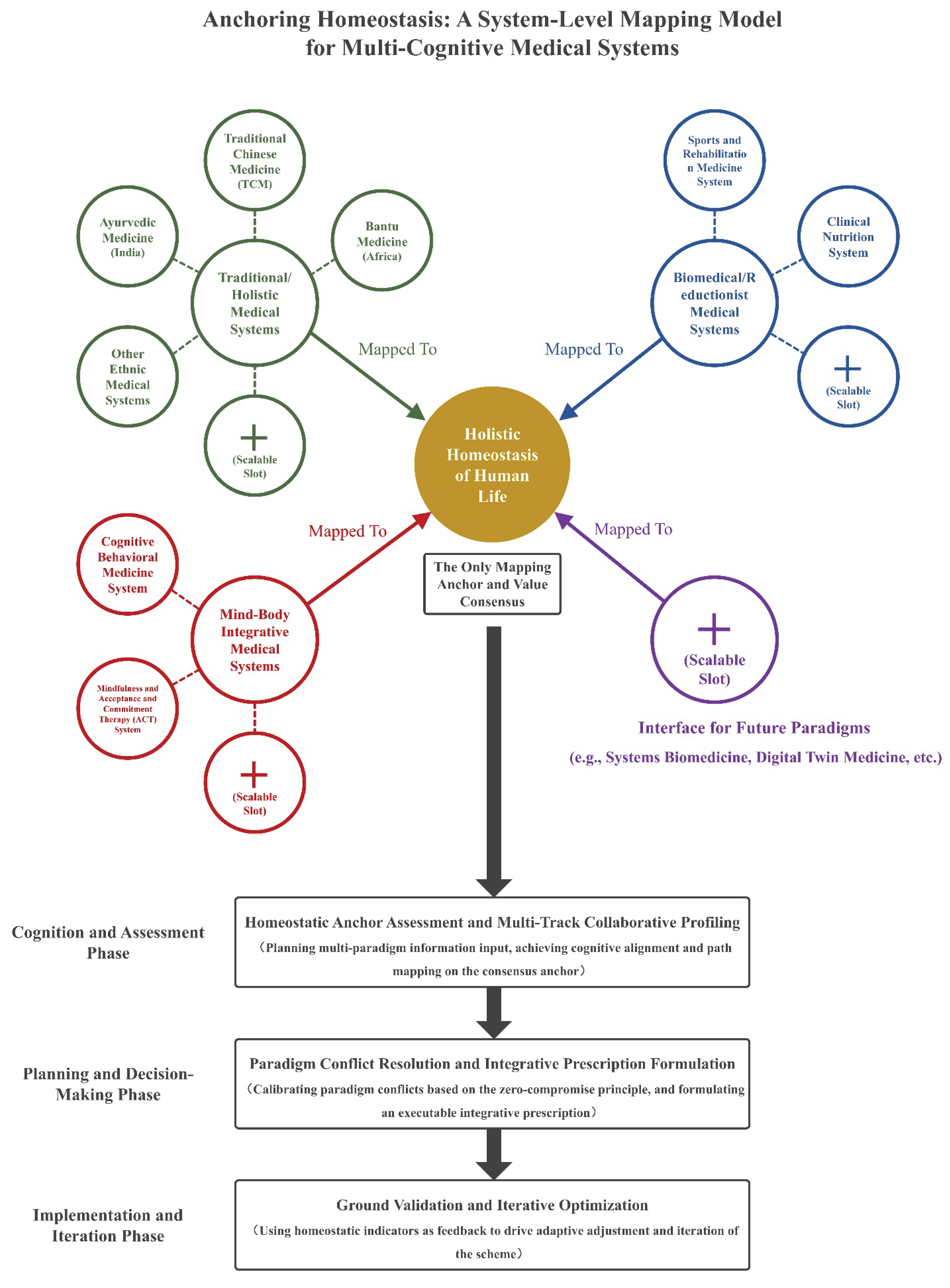
- System-Level Mapping Topology: A full-link mapping architecture defined by three core rules, which realizes the complete mapping from the native feature dimensions of the medical system to the homeostatic representation network, avoids the defects of fragmented analysis, and fits the holism logic of the medical system.
- Semantic Alignment Accuracy (SAA): The core evaluation index of this study, which is used to quantify the semantic consistency of cross-system data fusion. The specific calculation method and determination basis are shown in Section 2.4.
2. Construction of Multi-Track Cognition System-Level Mapping Framework
2.1. Multi-track Cognitive Dimension Construction Module
- Standardized feature dimension set: is derived entirely from the core theoretical framework of the respective medical system. For example, the TCM cognitive track includes dimensions such as nature and flavor, meridian tropism, efficacy and indication, syndrome adaptation, and compatibility taboo; the Western medicine cognitive track includes dimensions such as active ingredients, action targets, pharmacological effects, safe dose, and toxicological characteristics; the Tibetan medicine cognitive track includes dimensions such as five-source attribute, six tastes, eight properties, seventeen effects, and disease adaptation.
- Data normalization rules: Comply with the clinical application specifications of the corresponding medical system. For example, TCM adopts the grading standard of herbal medicine property intensity and the quantitative scoring standard of TCM syndromes; Western medicine adopts the standardization specification of clinical test indicators. All indicators are finally mapped to the [0,1] interval to eliminate dimensional differences.
- Systemic effect evaluation index: Adopt the native evaluation system of the corresponding medical system. For example, TCM takes the harmony degree of viscera function and the balance state of yin-yang qi and blood as the core; Western medicine takes the change range of physiological indicators and the effective rate of target regulation as the core, which is used as the optimization objective of mapping model training.
2.2. General Anchor Mediation Layer Construction Module
2.2.1. Setting of Homeostatic Dimensions
2.2.2. Network Topology and Weight Construction
- Topology structure determination stage: Through authoritative physiological literature and expert consultation with 3 chief physicians of integrated TCM and Western medicine, determine the existence of association relationships between dimensions (i.e., edge connection), and clarify the basic topology of the network, to ensure that the association relationships conform to classical physiological theories and clinical consensus.
- Weight optimization stage: In clinical practice, the physiological indicators of the human body are not independent of each other, but have inherent synergistic and restrictive relationships that conform to the laws of human physiology. Subjective weight assignment by experts is prone to personal bias, which will affect the objectivity and clinical reproducibility of the homeostatic representation network. The Bayesian network can objectively quantify the strength of the association between homeostatic dimensions based on real clinical data, avoiding subjective bias to the greatest extent, and ensuring that the network structure fully conforms to the objective laws of human physiology.Using the data of patients with stable vital signs and test indicators within 24 hours of admission in the Medical Information Mart for Intensive Care III (MIMIC-III) dataset, the Bayesian network is used to complete parameter learning, and optimize the specific value and direction of the weight. The conditional probability distribution formula of the Bayesian network is:Where Pa() is the parent node set of node , that is, other homeostatic dimensions that have a direct impact on the node; P(∣Pa()) is the conditional probability distribution of node . The parameter learning of the Bayesian network is completed by the maximum likelihood estimation method, and the quantitative association relationship between homeostatic dimensions is finally determined.
2.3. System-Level Mapping Relationship Construction Module
2.3.1. Three Core Mapping Rules
- Cluster Correspondence Constraint: A systemic effect corresponds to the synergistic action of a cluster of features/components, rather than a one-to-one linear mapping. In the algorithm implementation, multiple feature data corresponding to the same systemic effect are input into the model as a feature group, rather than splitting a single feature for independent modeling. This rule fits the core theory of multi-component synergistic effect of TCM, and is consistent with the research paradigm of multi-component and multi-target of network pharmacology [4,7], with the supplementary definition: rule governing synergistic effects of feature clusters.
- Network Emergence Constraint: The mapping must capture the emergent properties arising from multi-feature, multi-target, and multi-pathway interactions, moving beyond simple additive effects. In the algorithm implementation, the high-order interaction terms between features are automatically learned through the model to completely restore the multi-factor synergistic logic of complex biological systems. This rule is based on the emergence theory of complex system science, conforms to the inherent law of the human body as a complex physiological system, and is highly consistent with the holism of integrative medicine [1].
- Context Dependence Constraint: The mapping is context-aware, dynamically adapting to individual patient factors such as physical constitution and syndrome patterns, to fit the individualized intervention principle of different medical systems. In the algorithm implementation, a hierarchical training mechanism is adopted: first, the data set is stratified based on the patient's physical constitution/syndrome labels, and mapping sub-models are trained separately for different stratifications; for new samples, the corresponding syndrome/physical constitution stratification is matched first, and then the corresponding sub-model is called to complete the mapping, realizing the dynamic adjustment of association weights with the human body state.
2.3.2. Implementation of Mapping Model Algorithm
- Feature vectorization processing: For the standardized feature dimensions of each cognitive track, the combination of ordinal encoding and one-hot encoding is used to complete vectorization conversion: the cold/hot attribute of TCM nature is graded into 7 levels of "great cold, cold, slight cold, neutral, slight hot, hot, great hot", which are mapped to ordinal values from -3 to +3; classification features such as TCM meridian tropism, efficacy and indication, and Western medicine disease classification are processed by one-hot encoding and converted into binary feature vectors. Finally, all features are integrated into a unified input feature matrix X with consistent dimensions.
- Input, output and loss function: The input of the model is the standardized feature matrix X of each cognitive track, and the output is the prediction matrix Y of the corresponding dimension of the homeostatic representation network (i.e., the change of human homeostatic indicators after intervention). The core clinical goal of this model is to accurately predict the real changes of human homeostatic state after intervention with different medical systems. Therefore, the model training must take the deviation between the predicted value and the real clinical observed value as the core optimization objective, to ensure that the model output has clear clinical guiding significance. The optimization objective of model training is to minimize the mean squared error (MSE) between the predicted value and the real value. The loss function formula is:Where n is the number of training samples, is the real value of the change of homeostatic indicators, and is the predicted value of the model.
- Model training process: 5-fold cross-validation is used to complete model training and parameter optimization. The data set is randomly divided into 5 equal subsets. Each time, 4 subsets are taken as the training set and 1 subset as the verification set, and the cycle is repeated 5 times. The average performance is taken as the final performance of the model. The core hyperparameters of the model are optimized by grid search, and the optimal parameters are finally determined: number of decision trees n_estimators=100, maximum tree depth max_depth=10, minimum number of samples for node splitting min_samples_split=5. The clinical data of integrative medicine has strong heterogeneity due to the differences in patients' physical constitution, syndrome patterns and underlying diseases. To avoid model overfitting and ensure the stability and generalizability of the model in real clinical scenarios, we use 5-fold cross-validation to complete model training and hyperparameter optimization. The complete algorithm flow is as follows: The algorithm pseudo-code is as follows:
2.4. Definition of Core Evaluation Indicators
- Semantic Alignment Accuracy (SAA): The core evaluation index, used to quantify the semantic consistency of cross-system data fusion, The biggest clinical pain point in the field of integrative medicine is the lack of objective, quantifiable and reproducible indicators to evaluate whether the interventions of different medical systems have achieved clinically meaningful consensus. Semantic Alignment Accuracy (SAA) takes human homeostasis as the unified anchor, and quantifies the semantic consistency of intervention data from different medical systems at the clinical level, providing an objective evaluation tool for the effect of multi-medical system integration. with the calculation formula:
- 2.
- Goodness of Fit (: Used to evaluate the prediction accuracy of the model for the change of human homeostatic indicators, This indicator is used to evaluate the interpretability of the model for the changes of human homeostatic state. The closer the value is to 1, the more accurately the model can restore the real impact of interventions from different medical systems on human homeostasis, which directly determines the clinical reference value of the model output. with the calculation formula:
- 3.
-
Mean Absolute Error (MAE): Used to evaluate the absolute deviation between the predicted value and the real value of the model, This indicator quantifies the average absolute deviation between the predicted value of the model and the real clinical observed value, which directly reflects the prediction accuracy of the model for a single homeostatic dimension, and ensures that the prediction of each physiological indicator is consistent with the real clinical situation without systematic deviation. with the calculation formula:The smaller the MAE value, the higher the prediction accuracy of the model.
- 4.
-
Root Mean Square Error (RMSE): Used to evaluate the deviation degree between the predicted value and the real value of the model, This indicator is highly sensitive to the abnormal predicted values of the model, which can effectively test the stability of the model in extreme clinical scenarios (such as patients with severe homeostatic imbalance), and ensure that the model can maintain reliable prediction effect in patient groups with different health states, adapting to the complex scenarios of real clinical practice. with the calculation formula:The smaller the RMSE value, the better the prediction stability of the model.
3. Model Verification and Result Analysis
3.1. Experimental Data and Preprocessing
- Data cleaning: Invalid samples with missing values > 30% were eliminated, and the remaining missing values were filled by multiple imputation method.
- Standardization processing: Aiming at the format differences of data from different medical systems, a unified feature vectorization framework was adopted. One-hot encoding was used for all classification features, ordinal encoding for ordinal grade features, and min-max standardization for continuous numerical features. Finally, the features of all cognitive tracks were uniformly mapped to the [0,1] interval to eliminate the format and dimensional differences of data from different systems.
- Dataset division: The preprocessed dataset was randomly divided into training set (70%) and test set (30%) according to the ratio of 7:3. The training set was used for model training and parameter optimization, and the test set was used for model performance evaluation.
3.2. Selection of Baseline Methods
- Baseline 1: Single-point Linear Correlation Analysis: The mainstream traditional method in the current field of TCM-Western medicine data fusion, which realizes the linear correspondence between a single TCM feature and a single Western medicine indicator through Pearson correlation analysis.
- Baseline 2: LASSO Regression Feature Association Method: A commonly used method for feature screening and association of high-dimensional medical data, which realizes feature selection through L1 regularization and constructs a linear association model between TCM features and Western medicine indicators.
- Baseline 3: Knowledge Graph Semantic Alignment Method: The current mainstream technical method for medical term alignment, which realizes the semantic alignment between TCM syndromes and Western medicine diseases, traditional Chinese medicine and pharmacological effects by constructing a medical knowledge graph [12].
3.3. Basic Verification Results of TCM-Western Medicine Dual-Track
- The semantic alignment accuracy reaches 91.27%, which is 14.69 percentage points higher than the best-performing Baseline 3, and 32.14 percentage points higher than the traditional single-point linear correlation method. The Cohen's d effect sizes are all >0.8, among which the Cohen's d value for the performance difference between the proposed framework and the best-performing baseline (Baseline 3) reaches 1.94, far exceeding the 0.8 threshold for a large effect size, further confirming the clinical practical value of the performance improvement..
- The goodness of fit of the model reaches 0.852, which is much higher than the baseline methods, proving that the proposed framework has excellent prediction accuracy for the changes of human homeostatic indicators.
- Both MAE and RMSE are significantly lower than the baseline methods, proving that the prediction accuracy and stability of the proposed framework are better.
3.4. Extended Adaptation Verification Results of Multi-Medical System
3.5. Case Demonstration
- Multi-track cognitive data input: Dual-track data were collected simultaneously for the patient, fully retaining the native logic of each medical system without cross-paradigm transformation:
- 2.
- Model mapping and fusion: The dual-track data of TCM and Western medicine were input into the trained model, which were respectively mapped to the homeostatic representation network to obtain unified benchmark data, and cross-track data fusion was completed. The mapping process strictly follows the three core constraints of the framework: the cluster correspondence constraint is reflected in the synergistic effect of the feature component group corresponding to the qi-tonifying systemic effect; the network emergence constraint is reflected in the fitting of the overall systemic effect of multi-component synergistic intervention; the context dependence constraint is reflected in the dynamic adjustment of the mapping weight based on the patient's Lung-Spleen Qi Deficiency Syndrome label.
- 3.
- Homeostatic assessment and output: The model outputs the systematic homeostatic assessment results of the patient, and the core abnormal dimensions are the homeostatic imbalance of the immune system, endocrine system, and digestive system. At the same time, based on the fusion results, the intervention means corresponding to the TCM and Western medicine cognitive tracks are matched, and the integrated intervention scheme recommendation of TCM and Western medicine is output, including the recommendation of modified Sijunzi Tang, nutritional intervention scheme, and lifestyle guidance, and the safety assessment is completed simultaneously.
4. Discussion
4.1. Core Innovation Value and Theoretical Connection
4.2. Research Limitations
4.3. Compatibility with Existing Medical Standards
5. Conclusion
References
- Fan, DM. Theory and practice of integrative medicine[J]. National Medical Journal of China 2022, 102(12), 865–868. [Google Scholar] [CrossRef]
- Zhang, BL; Zhang, JH. Historical achievements and future direction of the development of integrated traditional Chinese and Western medicine[J]. Chinese Journal of Integrated Traditional and Western Medicine 2023, 43(1), 17–21. [Google Scholar] [CrossRef]
- Li, YB; Cui, M; Yang, Y. Research on the correlation between TCM syndromes and Western medicine diseases based on knowledge graph[J]. Chinese Journal of Information on Traditional Chinese Medicine 2021, 28(3), 1–5. [Google Scholar] [CrossRef]
- Li, S. Network pharmacology and innovation of integrated traditional Chinese and Western medicine[J]. Chinese Journal of Integrated Traditional and Western Medicine 2021, 41(2), 141–144. [Google Scholar] [CrossRef]
- Liu, BY; He, LY; Zhang, RS. Thoughts and methods of quantitative research on TCM syndromes[J]. Chinese Journal of Basic Medicine in Traditional Chinese Medicine 2021, 27(3), 364–367. [Google Scholar]
- World Health Organization. WHO global strategy on traditional and complementary medicine 2023-2032[R]; World Health Organization: Geneva, 2022. [Google Scholar] [CrossRef]
- Chen, J; Li, Y; Zhang, Y. Machine learning in traditional Chinese medicine research: advances and prospects[J]. Journal of Ethnopharmacology 2021, 276, 114186. [Google Scholar] [CrossRef]
- Breiman, L. Random Forests[J]. Machine Learning 2001, 45(1), 5–32. [Google Scholar] [CrossRef]
- Chinese Medical Association. National Clinical Laboratory Operation Procedures (4th Edition)[M]; People's Medical Publishing House: Beijing, 2015. [Google Scholar]
- Lv, AP; He, XJ; Ma, C. Core scientific issues and methodological exploration of big data research in integrated traditional Chinese and Western medicine[J]. Chinese Journal of Integrated Traditional and Western Medicine 2022, 42(6), 655–658. [Google Scholar] [CrossRef]
- Kuhn, TS. The Structure of Scientific Revolutions[M]; University of Chicago Press: Chicago, 1962. [Google Scholar]
- Hogan, A; Blomqvist, E; Cochez, M; et al. Knowledge graphs[J]. ACM Computing Surveys 2021, 54(4), 1–37. [Google Scholar] [CrossRef]
| Dataset Name | Data Source | Core Purpose |
| TCMSP Traditional Chinese Medicine System Pharmacology Database | Northwest A&F University | Feature extraction of TCM cognitive track |
| MIMIC-III Intensive Care Medical Dataset | Massachusetts Institute of Technology | Feature extraction of Western medicine cognitive track and extraction of real homeostatic values |
| Clinical Diagnosis and Treatment Dataset of China Academy of Chinese Medical Sciences | China Academy of Chinese Medical Sciences | Feature extraction of TCM cognitive track and model training verification |
| Public Dataset of Clinical Research on Tibetan Medicine in China | Chinese Journal of Tibetan Medicine | Feature extraction of Tibetan medicine cognitive track and multi-system extended verification |
| Evaluation Indicator | Proposed Framework | Baseline 1: Single-point Linear Correlation | Baseline 2: LASSO Regression | Baseline 3: Knowledge Graph Alignment |
| Semantic Alignment Accuracy (SAA) | 91.27% | 59.13% | 68.42% | 76.58% |
| Goodness of Fit () | 0.852 | 0.426 | 0.583 | 0.617 |
| Mean Absolute Error (MAE) | 0.076 | 0.213 | 0.168 | 0.152 |
| Root Mean Square Error (RMSE) | 0.098 | 0.257 | 0.204 | 0.189 |
| Cohen's d Effect Size | - | 2.87 | 2.31 | 1.94 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




