Submitted:
04 March 2026
Posted:
05 March 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
2.1. Subjects
2.2. Ethics Statement
2.3. Fecal Sample Collection
2.4. DNA Extraction
2.5. Amplicon Library Preparation of ITS2 rRNA Region, Sequencing, and Fungal Taxonomic Identification
2.6. Classification of “Mycobiotypes”
2.7. Nested-PCR for Histoplasma Capsulatum
2.8. Statistical Analysis
3. Results
3.1. Demographic and Clinical Characteristics of the Study Participants
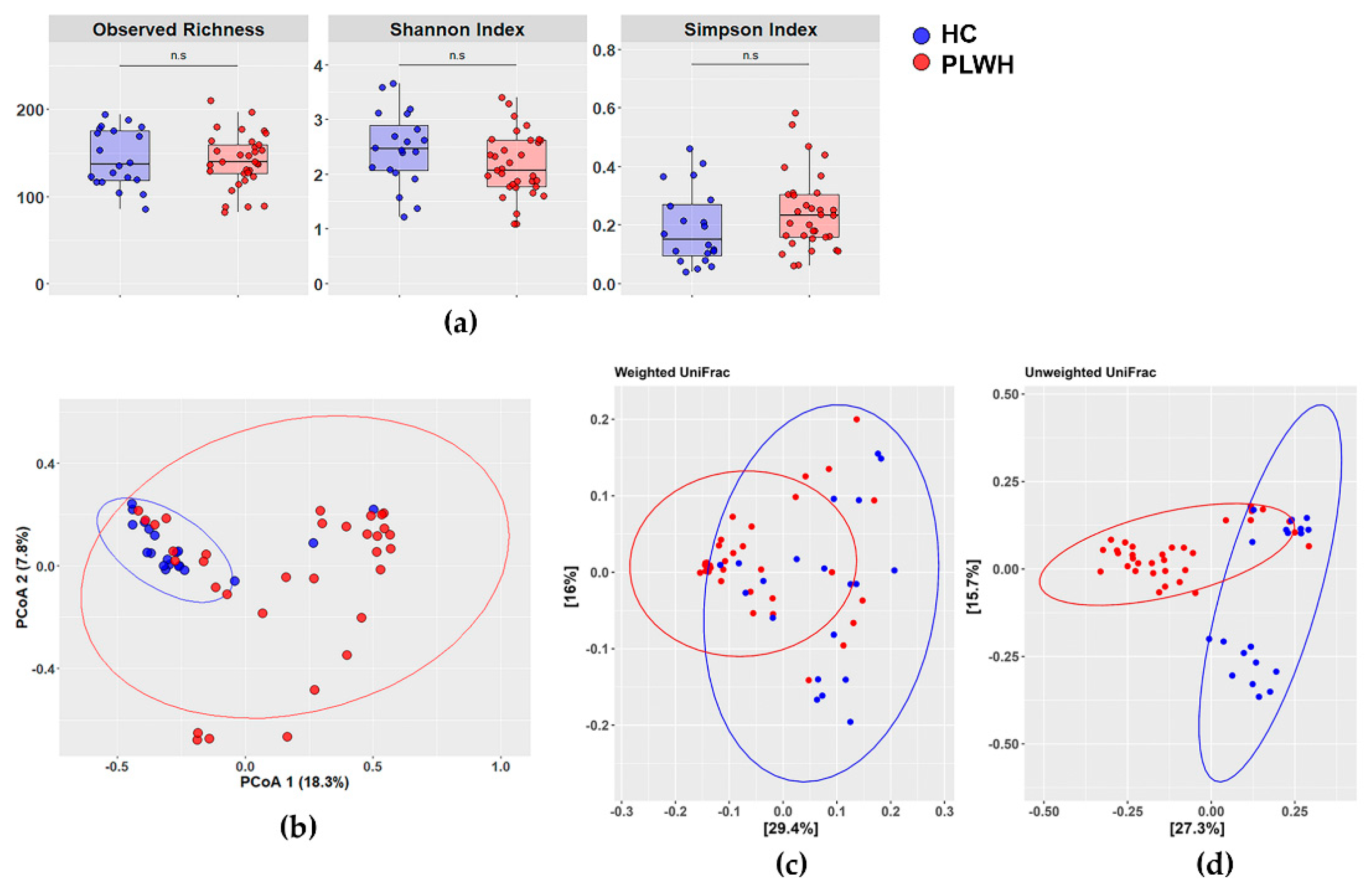
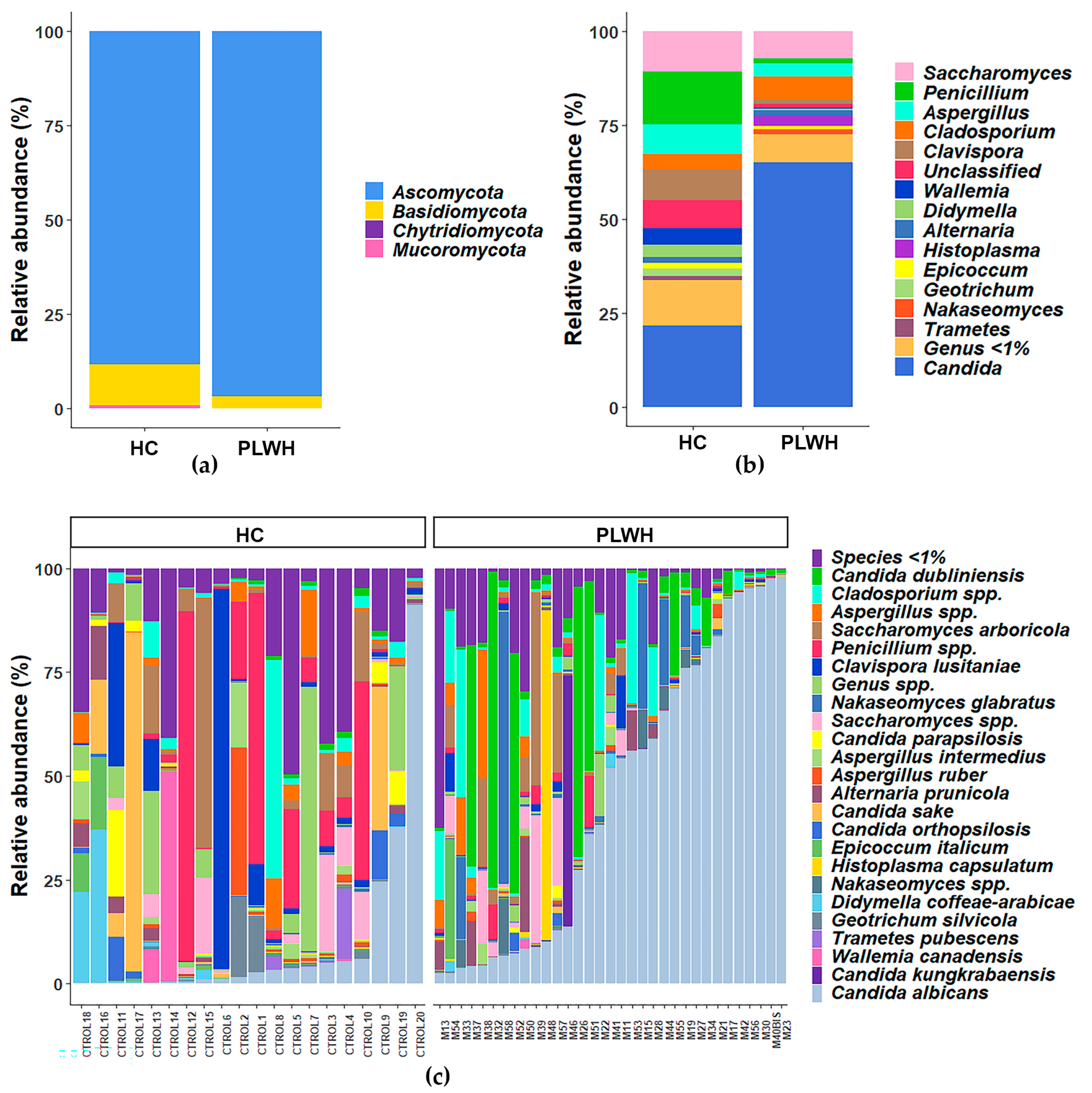
3.2. Gut Fungal Composition in PLWH Differs from That of HIV-Negative Individuals
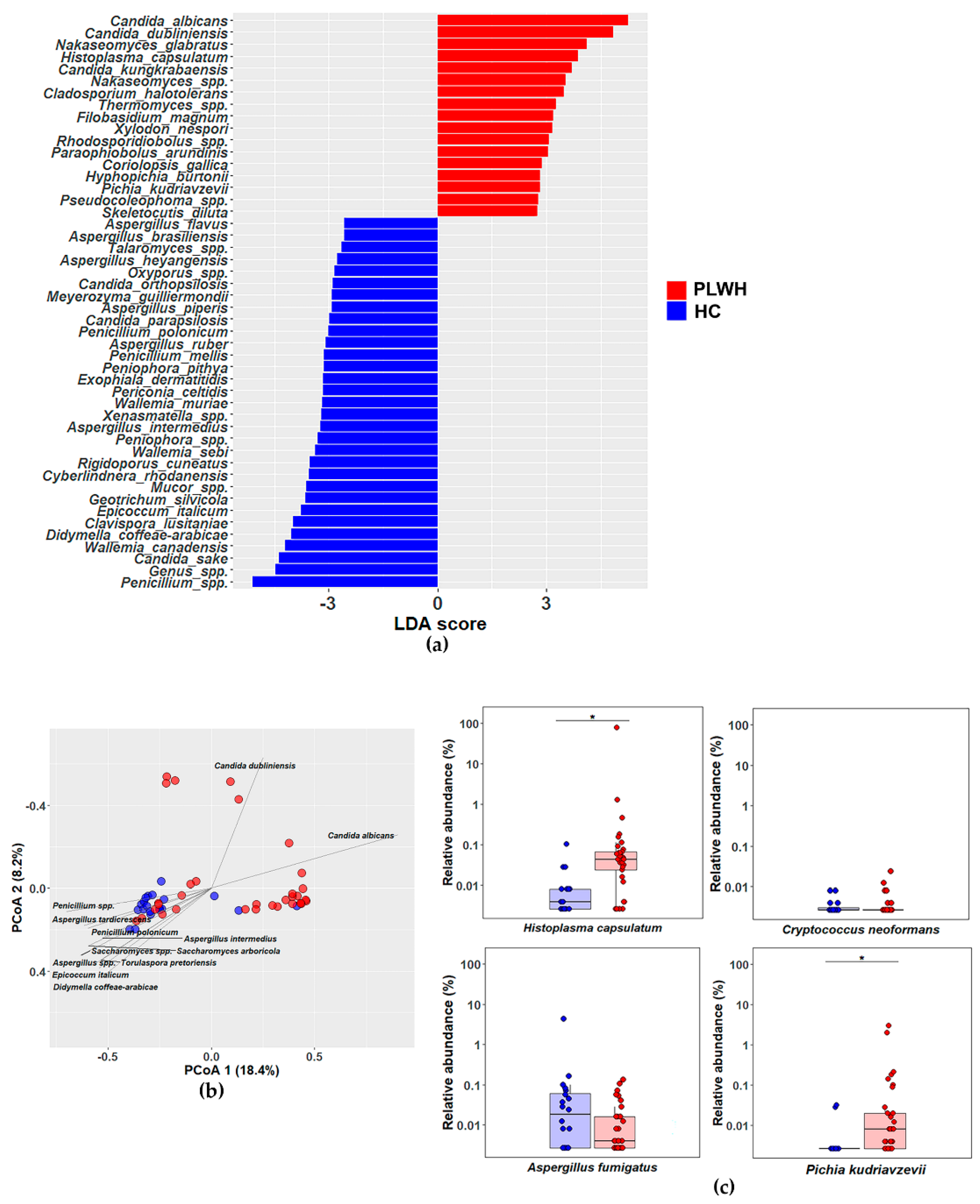
3.3. Distinct Mycobiota Signatures in People Living with HIV: Enrichment of Candida albicans, Candida dubliniensis, Nakaseomyces glabratus, and Histoplasma capsulatum
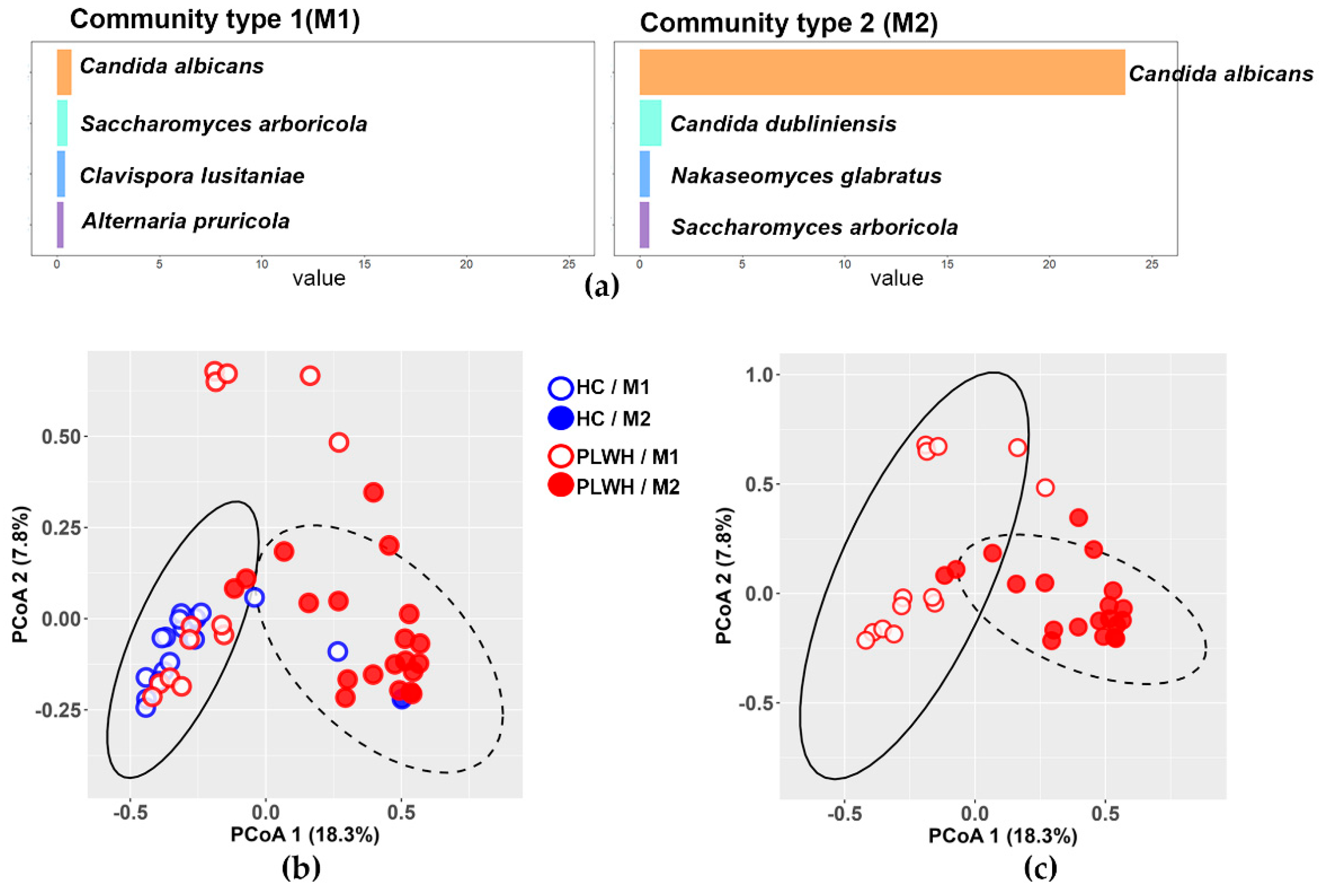
3.4. The Gut Mycobiota Clusters into Two Mycobiotypes, M1 and M2, with M1 Predominating in Healthy Controls
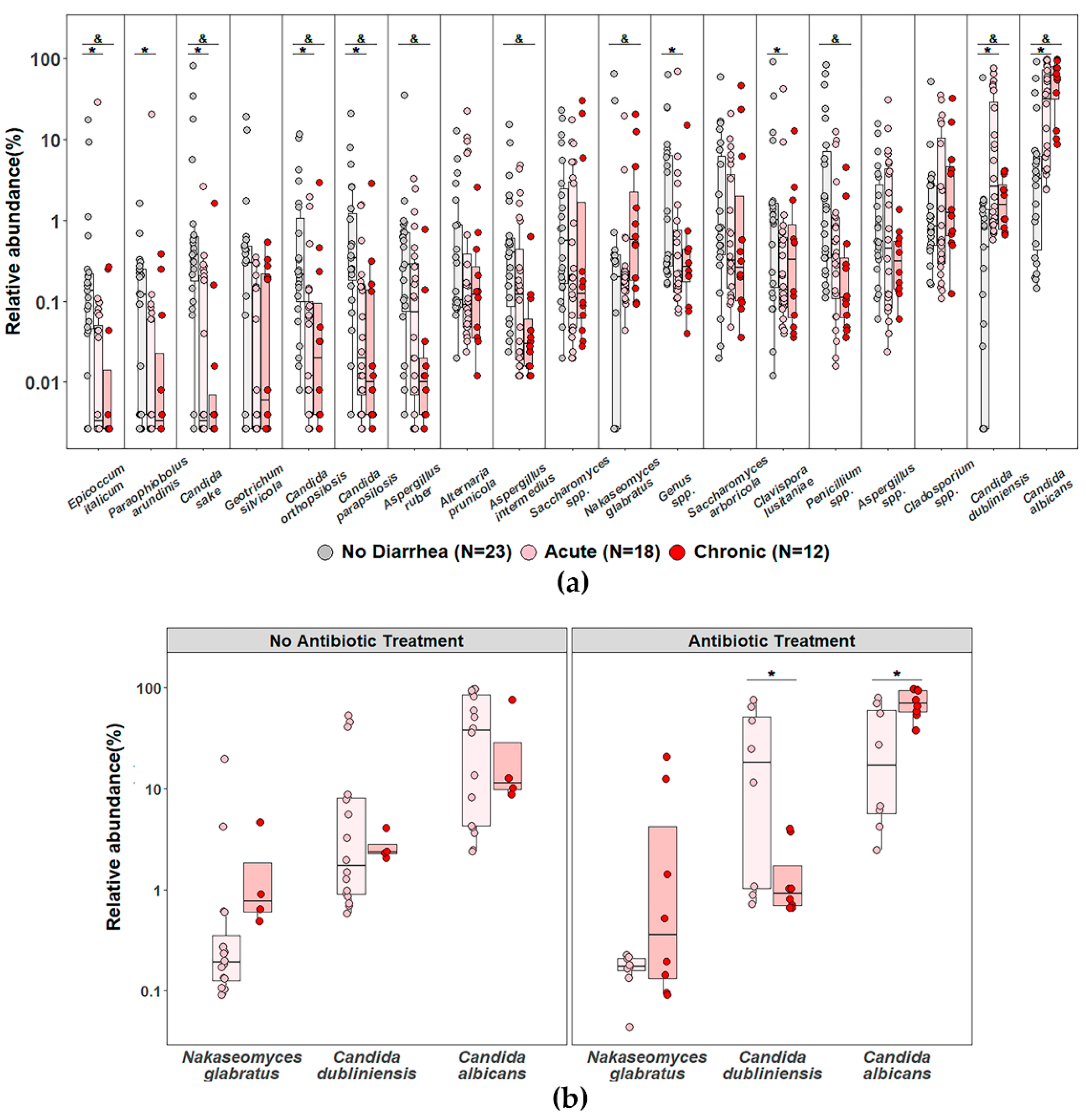
3.5. Impact of Diarrhea and Antibiotic Treatment on the Relative Abundance of Dominant Fungal Species in PLWH
4. Discussion
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| PLWH | People living with Human Immunodeficiency Virus |
| LEFSe | Linear Discriminant Analysis Effect Size |
| DMM analysis | Dirichlet multinomial mixture analysis |
| ART | antiretroviral therapy |
References
- Ministerio de Salud de La Nación (Argentina). Boletín No. 41: Respuesta al VIH y Las ITS En La Argentina. Año XXVII. Buenos Aires. 2024. Available online: Https://Www.Argentina.Gob.Ar/Sites/Default/Files/2017/11/Boletin_n_41-Respuesta_al_vih_y_las_its_en_la_argentina_2024.Pdf.
- Duarte, M.J.; Tien, P.C.; Somsouk, M.; Price, J.C. The Human Microbiome and Gut–Liver Axis in People Living with HIV. Curr HIV/AIDS Rep 2023, 20, 170–180. [Google Scholar] [CrossRef]
- Dolo, O.; Coulibaly, F.; Somboro, A.M.; Sun, S.; Diarra, M.; Maiga, A.; Bore, S.; Fofana, D.B.; Marcelin, A.-G.; Diakite, B.; et al. The Human Gut Microbiome and Its Metabolic Pathway Dynamics before and during HIV Antiretroviral Therapy. Microbiol Spectr 2025, 13, e02205-24. [Google Scholar] [CrossRef]
- Dillon, S.M.; Frank, D.N.; Wilson, C.C. The Gut Microbiome and HIV-1 Pathogenesis: A Two-Way Street. AIDS 2016, 30, 2737–2751. [Google Scholar] [CrossRef]
- Olaru, I.D.; Tacconelli, E.; Yeung, S.; Ferrand, R.A.; Stabler, R.A.; Hopkins, H.; Aiken, A.M.; Kranzer, K. The Association between Antimicrobial Resistance and HIV Infection: A Systematic Review and Meta-Analysis. Clinical Microbiology and Infection 2021, 27, 846–853. [Google Scholar] [CrossRef]
- Spragge, F.; Bakkeren, E.; Jahn, M.T.; B. N. Araujo, E.; Pearson, C.F.; Wang, X.; Pankhurst, L.; Cunrath, O.; Foster, K.R. Microbiome Diversity Protects against Pathogens by Nutrient Blocking. Science 2023, 382, eadj3502. [Google Scholar] [CrossRef]
- Ratiner, K.; Ciocan, D.; Abdeen, S.K.; Elinav, E. Utilization of the Microbiome in Personalized Medicine. Nat Rev Microbiol 2024, 22, 291–308. [Google Scholar] [CrossRef]
- Bandera, A.; De Benedetto, I.; Bozzi, G.; Gori, A. Altered Gut Microbiome Composition in HIV Infection: Causes, Effects and Potential Intervention. Current Opinion in HIV and AIDS 2018, 13, 73–80. [Google Scholar] [CrossRef]
- Li, X.V.; Leonardi, I.; Iliev, I.D. Gut Mycobiota in Immunity and Inflammatory Disease. Immunity 2019, 50, 1365–1379. [Google Scholar] [CrossRef] [PubMed]
- Richard, M.L.; Sokol, H. The Gut Mycobiota: Insights into Analysis, Environmental Interactions and Role in Gastrointestinal Diseases. Nat Rev Gastroenterol Hepatol 2019. [Google Scholar] [CrossRef]
- Iliev, I.D.; Brown, G.D.; Bacher, P.; Gaffen, S.L.; Heitman, J.; Klein, B.S.; Lionakis, M.S. Focus on Fungi. Cell 2024, 187, 5121–5127. [Google Scholar] [CrossRef] [PubMed]
- Dohlman, A.B.; Klug, J.; Mesko, M.; Gao, I.H.; Lipkin, S.M.; Shen, X.; Iliev, I.D. A Pan-Cancer Mycobiome Analysis Reveals Fungal Involvement in Gastrointestinal and Lung Tumors. Cell 2022, 185, 3807–3822.e12. [Google Scholar] [CrossRef]
- Leonardi, I.; Gao, I.H.; Lin, W.-Y.; Allen, M.; Li, X.V.; Fiers, W.D.; De Celie, M.B.; Putzel, G.G.; Yantiss, R.K.; Johncilla, M.; et al. Mucosal Fungi Promote Gut Barrier Function and Social Behavior via Type 17 Immunity. Cell 2022, 185, 831–846.e14. [Google Scholar] [CrossRef]
- Li, X.V.; Leonardi, I.; Putzel, G.G.; Semon, A.; Fiers, W.D.; Kusakabe, T.; Lin, W.-Y.; Gao, I.H.; Doron, I.; Gutierrez-Guerrero, A.; et al. Immune Regulation by Fungal Strain Diversity in Inflammatory Bowel Disease. Nature 2022, 603, 672–678. [Google Scholar] [CrossRef]
- Su, Q.; Wong, O.W.H.; Lu, W.; Wan, Y.; Zhang, L.; Xu, W.; Li, M.K.T.; Liu, C.; Cheung, C.P.; Ching, J.Y.L.; et al. Multikingdom and Functional Gut Microbiota Markers for Autism Spectrum Disorder. Nat Microbiol 2024. [Google Scholar] [CrossRef]
- Weiner, L.D.; Retuerto, M.; Hager, C.L.; El Kamari, V.; Shan, L.; Sattar, A.; Kulkarni, M.; Funderburg, N.; Ghannoum, M.A.; Dirajlal-Fargo, S.; et al. Fungal Translocation Is Associated with Immune Activation and Systemic Inflammation in Treated HIV. AIDS Research and Human Retroviruses 2019, 35, 461–472. [Google Scholar] [CrossRef]
- Narayanan, A.; Kieri, O.; Vesterbacka, J.; Manoharan, L.; Chen, P.; Ghorbani, M.; Ljunggren, H.-G.; Sällberg Chen, M.; Aleman, S.; Sönnerborg, A.; et al. Exploring the Interplay between Antiretroviral Therapy and the Gut-Oral Microbiome Axis in People Living with HIV. Sci Rep 2024, 14, 17820. [Google Scholar] [CrossRef]
- Rocafort, M.; Gootenberg, D.B.; Luévano, J.M.; Paer, J.M.; Hayward, M.R.; Bramante, J.T.; Ghebremichael, M.S.; Xu, J.; Rogers, Z.H.; Munoz, A.R.; et al. HIV-Associated Gut Microbial Alterations Are Dependent on Host and Geographic Context. Nat Commun 2024, 15, 1055. [Google Scholar] [CrossRef]
- Yin, Y.; Tuohutaerbieke, M.; Feng, C.; Li, X.; Zhang, Y.; Xu, Q.; Tu, J.; Yang, E.; Zou, Q.; Shen, T. Characterization of the Intestinal Fungal Microbiome in HIV and HCV Mono-Infected or Co-Infected Patients. Viruses 2022, 14, 1811. [Google Scholar] [CrossRef]
- Hamad, I.; Abou Abdallah, R.; Ravaux, I.; Mokhtari, S.; Tissot-Dupont, H.; Michelle, C.; Stein, A.; Lagier, J.-C.; Raoult, D.; Bittar, F. Metabarcoding Analysis of Eukaryotic Microbiota in the Gut of HIV-Infected Patients. PLoS ONE 2018, 13, e0191913. [Google Scholar] [CrossRef] [PubMed]
- Delavy, M.; Sertour, N.; Patin, E.; Le Chatelier, E.; Cole, N.; Dubois, F.; Xie, Z.; Saint-André, V.; Manichanh, C.; Walker, A.W.; et al. Unveiling Candida Albicans Intestinal Carriage in Healthy Volunteers: The Role of Micro- and Mycobiota, Diet, Host Genetics and Immune Response. Gut Microbes 2023, 15, 2287618. [Google Scholar] [CrossRef]
- Shankar, J. Food Habit Associated Mycobiota Composition and Their Impact on Human Health. Front. Nutr. 2021, 8, 773577. [Google Scholar] [CrossRef]
- Kabwe, M.H.; Vikram, S.; Mulaudzi, K.; Jansson, J.K.; Makhalanyane, T.P. The Gut Mycobiota of Rural and Urban Individuals Is Shaped by Geography. BMC Microbiol 2020, 20, 257. [Google Scholar] [CrossRef]
- Nash, A.K.; Auchtung, T.A.; Wong, M.C.; Smith, D.P.; Gesell, J.R.; Ross, M.C.; Stewart, C.J.; Metcalf, G.A.; Muzny, D.M.; Gibbs, R.A.; et al. The Gut Mycobiome of the Human Microbiome Project Healthy Cohort. Microbiome 2017, 5, 153. [Google Scholar] [CrossRef]
- Portela, N.D.; Mena, C.; Martín, M.G.; Burstein, V.L.; Chiapello, L.S.; Pesoa, S.A. Effect of DNA Extraction Method in Gut Fungal Community Assessment. Revista Argentina de Microbiología 2025, 57, 309–317. [Google Scholar] [CrossRef]
- Bialek, R.; Feucht, A.; Aepinus, C.; Just-Nübling, G.; Robertson, V.J.; Knobloch, J.; Hohle, R. Evaluation of Two Nested PCR Assays for Detection of Histoplasma Capsulatum DNA in Human Tissue. J Clin Microbiol 2002, 40, 1644–1647. [Google Scholar] [CrossRef]
- Dangarembizi, R.; Hoving, J.C.; Boulware, D.R.; Colombo, A.L.; Govender, N.P.; Oladele, R.; Dat, V.Q.; Schwartz, I.S.; Brown, G.D. AIDS-Related Mycoses: Advances, Challenges, and Future Directions. Trends in Microbiology 2025, 33, 141–144. [Google Scholar] [CrossRef]
- Borgognone, A.; Noguera-Julian, M.; Oriol, B.; Noël-Romas, L.; Ruiz-Riol, M.; Guillén, Y.; Parera, M.; Casadellà, M.; Duran, C.; Puertas, M.C.; et al. Gut Microbiome Signatures Linked to HIV-1 Reservoir Size and Viremia Control. Microbiome 2022, 10, 59. [Google Scholar] [CrossRef] [PubMed]
- Do Nascimento, W.M.; Machiavelli, A.; Ferreira, L.G.E.; Cruz Silveira, L.; De Azevedo, S.S.D.; Bello, G.; Smith, D.P.; Mezzari, M.P.; Petrosino, J.F.; Delgado Duarte, R.T.; et al. Gut Microbiome Profiles and Associated Metabolic Pathways in HIV-Infected Treatment-Naïve Patients. Cells 2021, 10, 385. [Google Scholar] [CrossRef]
- San-Juan-Vergara, H.; Zurek, E.; Ajami, N.J.; Mogollon, C.; Peña, M.; Portnoy, I.; Vélez, J.I.; Cadena-Cruz, C.; Diaz-Olmos, Y.; Hurtado-Gómez, L.; et al. A Lachnospiraceae-Dominated Bacterial Signature in the Fecal Microbiota of HIV-Infected Individuals from Colombia, South America. Sci Rep 2018, 8, 4479. [Google Scholar] [CrossRef] [PubMed]
- Rolling, T.; Hohl, T.M.; Zhai, B. Minority Report: The Intestinal Mycobiota in Systemic Infections. Current Opinion in Microbiology 2020, 56, 1–6. [Google Scholar] [CrossRef]
- Fiers, W.D.; Leonardi, I.; Iliev, I.D. From Birth and Throughout Life: Fungal Microbiota in Nutrition and Metabolic Health. Annu Rev Nutr 2020, 40, 323–343. [Google Scholar] [CrossRef]
- Loreto, É.S.; Scheid, L.A.; Nogueira, C.W.; Zeni, G.; Santurio, J.M.; Alves, S.H. Candida Dubliniensis: Epidemiology and Phenotypic Methods for Identification. Mycopathologia 2010, 169, 431–443. [Google Scholar] [CrossRef]
- Szóstak, N.; Handschuh, L.; Samelak-Czajka, A.; Tomela, K.; Pietrzak, B.; Schmidt, M.; Galus, Ł.; Mackiewicz, J.; Mackiewicz, A.; Kozlowski, P.; et al. Gut Mycobiota Dysbiosis Is Associated with Melanoma and Response to Anti–PD-1 Therapy. Cancer Immunology Research 2024, 12, 427–439. [Google Scholar] [CrossRef]
- Casalini, G.; Giacomelli, A.; Antinori, S. The WHO Fungal Priority Pathogens List: A Crucial Reappraisal to Review the Prioritisation. The Lancet Microbe 2024, 5, 717–724. [Google Scholar] [CrossRef]
- Huang, J.; Hu, P.; Ye, L.; Shen, Z.; Chen, X.; Liu, F.; Xie, Y.; Yu, J.; Fan, X.; Xiao, M.; et al. Pan-Drug Resistance and Hypervirulence in a Human Fungal Pathogen Are Enabled by Mutagenesis Induced by Mammalian Body Temperature. Nat Microbiol 2024, 9, 1686–1699. [Google Scholar] [CrossRef]
- Chamroensakchai, T.; Kanjanabuch, T.; Saikong, W.; Panya, W.; Thaweekote, S.; Eiam-Ong, S.; Hurdeal, V.G.; Hyde, K.D. The First Human Report of Hyphopichia Burtonii, Initially Misdiagnosed as Sterile Peritonitis in a Patient on Peritoneal Dialysis. Medical Mycology Case Reports 2021, 33, 26–29. [Google Scholar] [CrossRef]
- Cassone, A.; Cauda, R. Candida and Candidiasis in HIV-Infected Patients: Where Commensalism, Opportunistic Behavior and Frank Pathogenicity Lose Their Borders. AIDS 2012, 26, 1457–1472. [Google Scholar] [CrossRef]
- Kreulen, I.A.M.; De Jonge, W.J.; Van Den Wijngaard, R.M.; Van Thiel, I.A.M. Candida Spp. in Human Intestinal Health and Disease: More than a Gut Feeling. Mycopathologia 2023, 188, 845–862. [Google Scholar] [CrossRef]
- González-González, A.E.; Taylor, M.L.; Curiel-Quesada, E. Aspectos relevantes del marcador molecular Hcp100 de Histoplasma capsulatum y su potencial uso terapéutico en la histoplasmosis. Revista Iberoamericana de Micología 2012, 29, 115–119. [Google Scholar] [CrossRef]
- Li, W.; Qing, Y.; Yu, Q.; Zhang, H.; Rang, Z.; Li, S.; Cui, F. Alterations of the Intestinal Mucosal Barrier and Gut Fungal Microbiome in Asymptomatic HIV-Infected Patients. Canadian Journal of Infectious Diseases and Medical Microbiology 2024, 2024, 6995192. [Google Scholar] [CrossRef]
|
PLWH (n=33) |
HC (n=20) |
|
|
Age (years) median (min-max) |
42.0 (23-61) |
31.5 (25-65) |
|
Sex Male / Female (n) |
23 / 10 |
11 / 9 |
|
CD4+ T lymphocytes Absolute number/µl: median (min-max) |
50 (2-1886) |
RR: 410 - 1590 |
|
CD8+ T lymphocytes Absolute number/µl: median (min-max) |
470 (122 - 3518) |
RR: 190-1140 |
|
CD3+ T lymphocytes Absolute number/µl: median (min-max) |
559 (135-3693) |
RR: 690-2540 |
|
Viral Load Copies/ml: median (min-max) |
173.000 (0-10.000.000) |
Undetectable |
|
Diarrhea* Chronic / Acute / No diarrhea (n) |
12 /18 / 3 |
0 / 0 / 20 |
|
Hospitalization Outpatient /Inpatient / ICU (n) |
8 / 20 /5 |
20 / 0 / 0 |
|
Antiretroviral treatment (ART) Yes/ No (n) |
13 / 20 |
0 / 20 |
|
Antibacterial treatment (ATB) Yes / No (n) |
17 / 16 |
0 / 20 |
|
Antifungal treatment (ATF) Yes / No (n) |
9 / 24 |
0 / 20 |
|
Oral Candidiasis Yes / No (n) |
8 / 25 |
0 / 20 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




