Submitted:
04 March 2026
Posted:
05 March 2026
Read the latest preprint version here
Abstract
Keywords:
1. Introduction
- This study used a hybrid combination ensemble approach using feature-level fusion and weighted ensemble methods.
- This study applied a transfer learning approach to transfer and combine the feature-level fusion ensemble result with the weighted ensemble method to increase the performance result.
- This study experimented and found the best combination of algorithms to combine in the ensemble method to improve the performance result.
- This study achieved better performance results compared to the existing studies.
2. Related Works
2.1. Gaps and Contributions
3. Research Methods
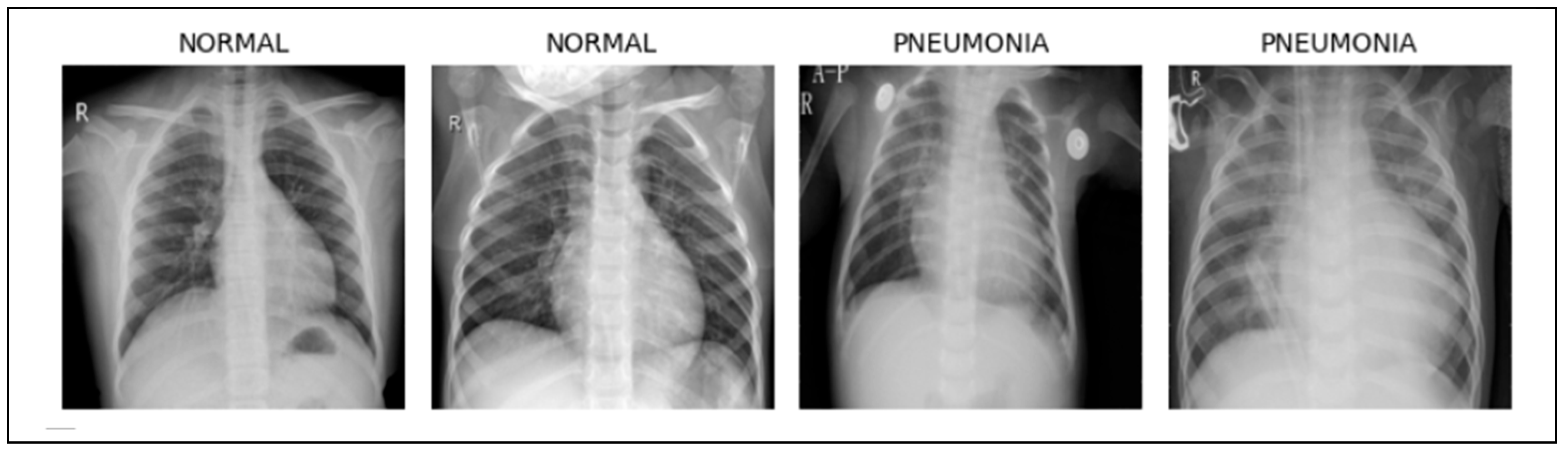
3.1. Dataset Description
3.2. Data Preprocessing
- A.
- Image Resizing
- B.
- Image Pixel Intensity Normalization
- -
- Mathematical Explanation
- -
- Enhanced Training Stability and Gradient Control
- -
- Acceleration of Convergence via Loss Landscape Reshaping
- -
- Mitigating Activation Function Saturation
- C.
- Label Encoding
- D.
- Data Structure Optimization
3.3. Data Augmentation
- -
- Horizontal flipping (T_flip) applied with a probability of P = 0.5.
- -
- Random rotation (T_rotate) within the range of ±10°.
- -
- Random scaling (zoom) (T_Zoom) by up to ±10%.
- -
- Random translation (shift) (T_shift) in either the horizontal or vertical axis by up to ±10% of the image dimensions.
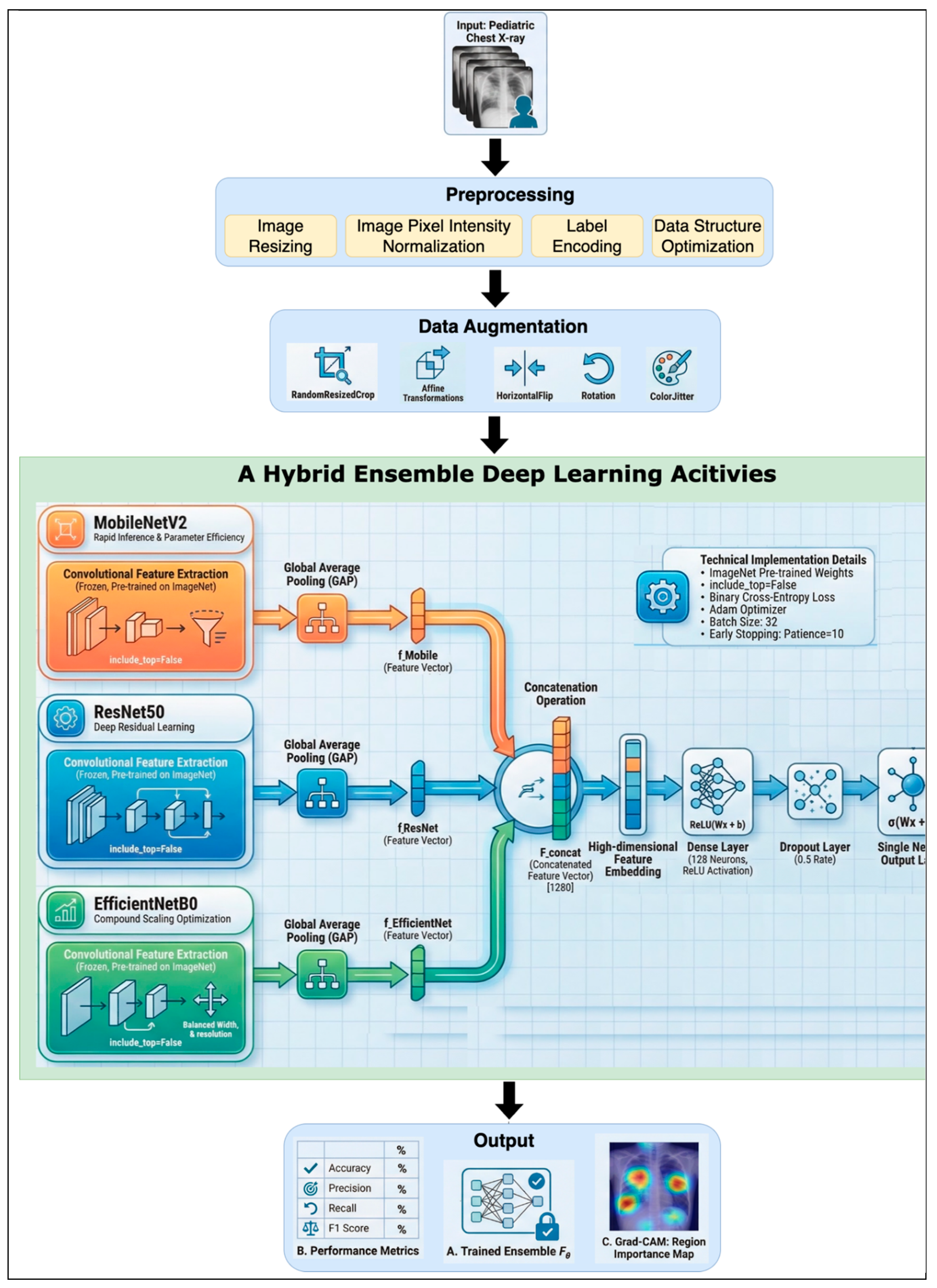
3.4. Model Architecture of Hybrid Convolutional Neural Network (CNN) Ensemble
3.5. Performance Evaluation and Clinical Significance of The Proposed Ensemble Model
- It introduces a novel hybrid ensemble framework that leverages the complementary strengths of lightweight (MobileNetV2) and deep semantic (ResNet50, EfficientNetB0) networks to balance computational efficiency with high-level feature representation.
- The model is uniquely tailored to a pediatric diagnostic setting, focusing on a sensitive and underrepresented population often overlooked in mainstream AI medical research.
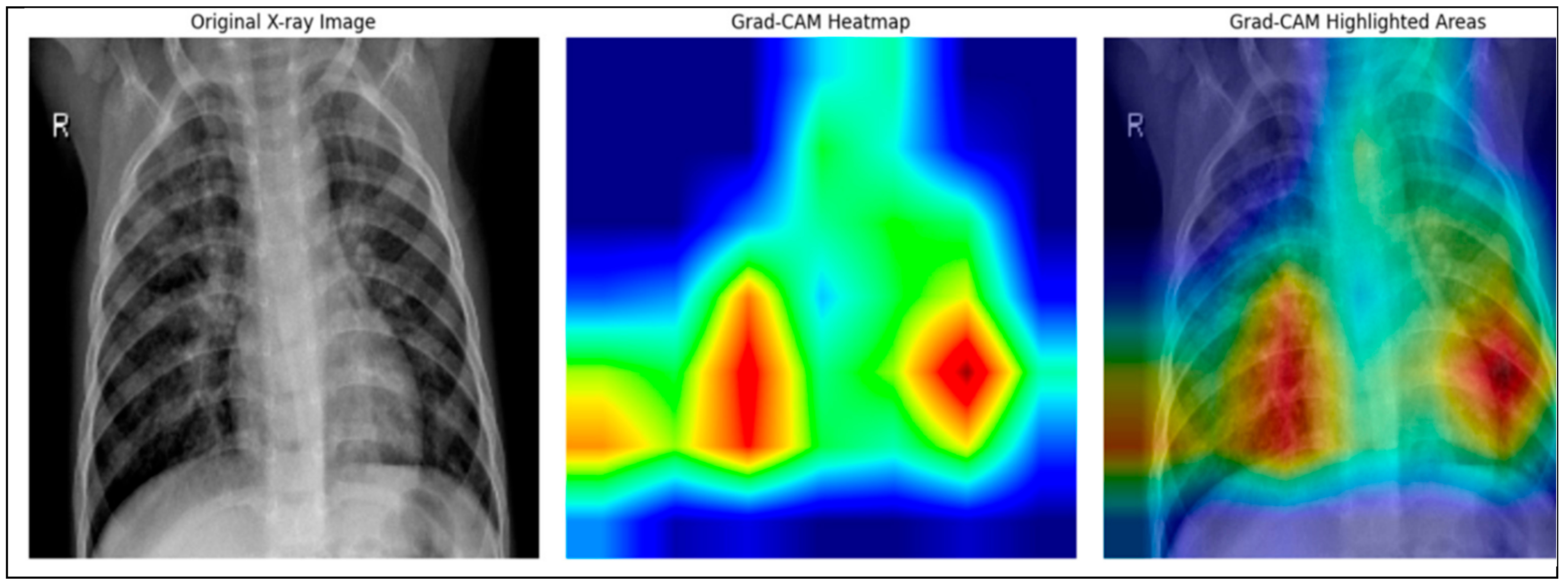
- To enhance transparency and foster trust in clinical environments, the model incorporates explainable AI (XAI) techniques via Grad-CAM, allowing practitioners to visualize and interpret decision regions within chest X-rays.
- A fully reproducible and well-documented pipeline has been developed, covering every stage from data preprocessing and augmentation to model training and evaluation, ensuring scientific rigor and practical deployment readiness.
- The exceptional F1-score of 94.97% confirms the model’s potential for real-world application in automated pneumonia screening tools, especially in resource-constrained healthcare environments.
4. Results and Discussions
4.1. Experimental Setup
4.2. Performance Metrics
4.3. Classification and Explanation
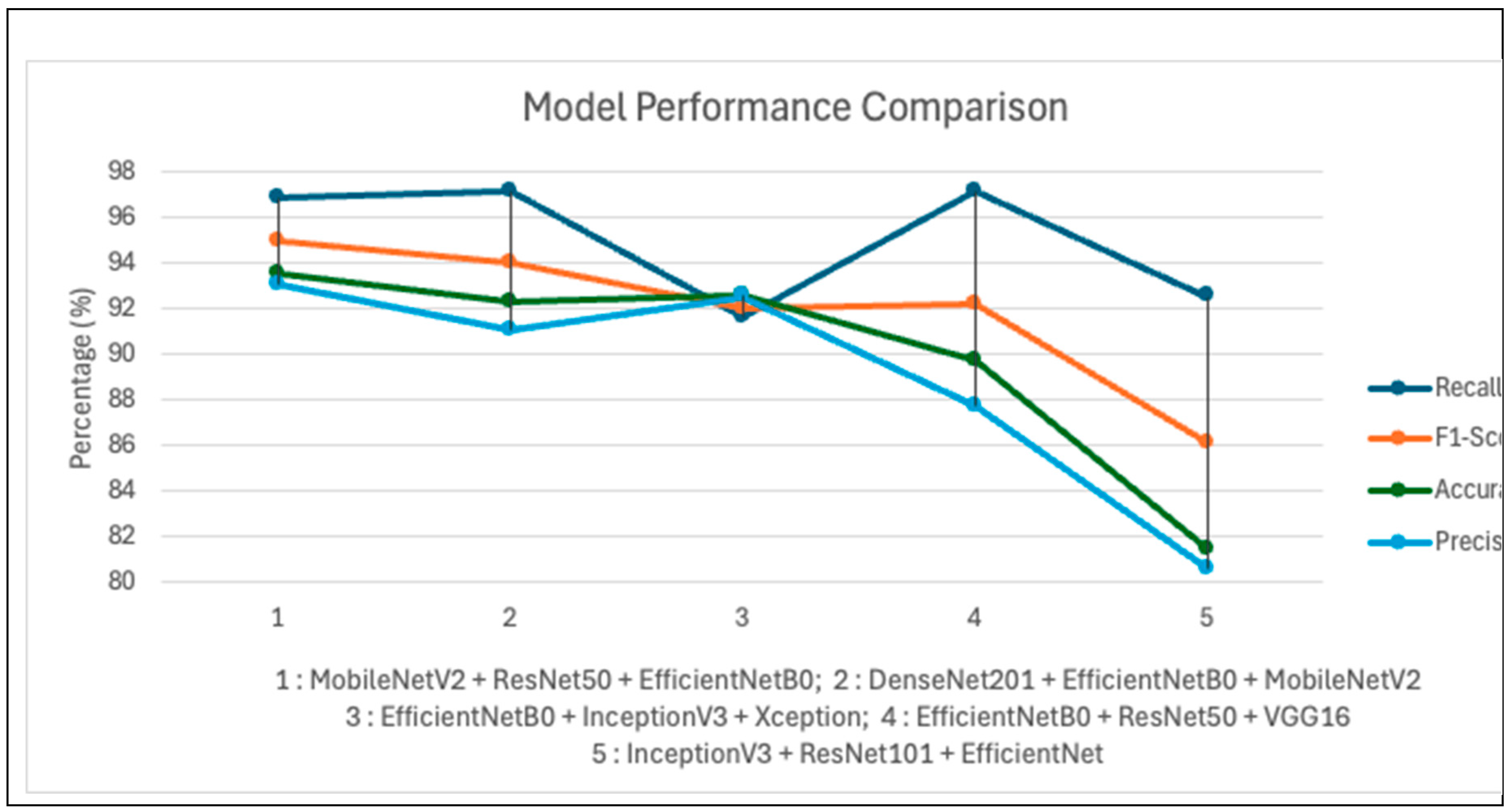
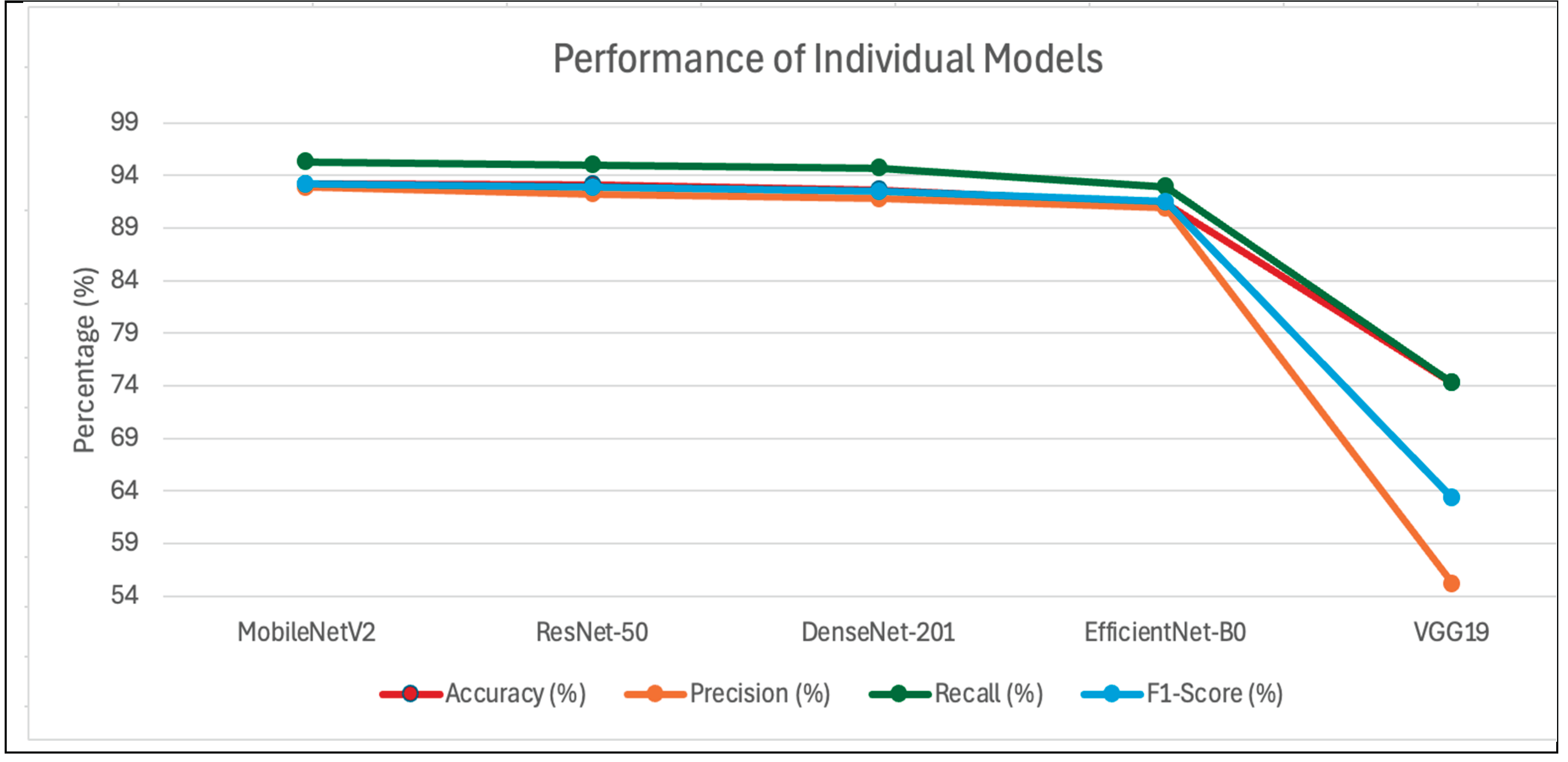
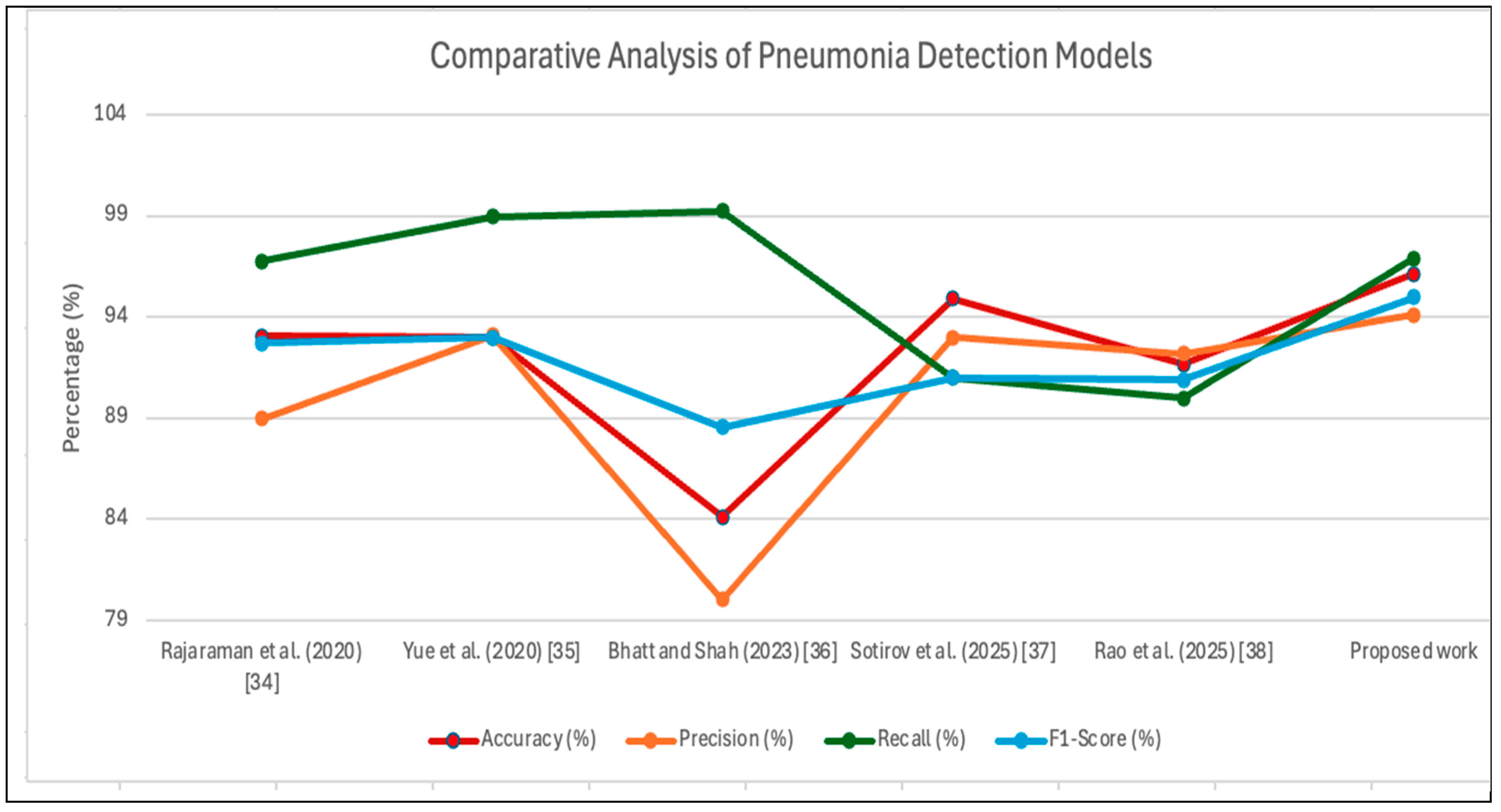
4.4. Comparative Analysis
- MobileNetV2: Employs depth-wise separable convolutions and linear bottlenecks. It is highly parameter-efficient (3.4M parameters) and fast, making it suitable for edge deployment. Its lower-level features capture local textures and edges, useful for detecting small consolidations.
- ResNet50: Introduces residual connections that enable training of very deep networks. Its 25.6M parameters allow learning of hierarchical, semantically rich features, particularly effective for identifying diffuse interstitial patterns characteristic of viral pneumonia.
- EfficientNetB0: Achieves state-of-the-art accuracy with compound scaling (depth, width, resolution). Its 5.3M parameters and balanced receptive field provide a complementary middle ground between the lightweight MobileNetV2 and the deeper ResNet50.
4.5. Clinical Relevance
5. Conclusion and Future Work
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- P. Radočaj, and G. Martinović, “Interpretable Deep Learning for Pediatric Pneumonia Diagnosis Through Multi-Phase Feature Learning and Activation Patterns,” Electronics, vol. 14, no. 9, pp. 1899, 2025. [CrossRef]
- Rudan, K. L. O'Brien, H. Nair, L. Liu, E. Theodoratou, S. Qazi, I. Lukšić, C. L. Fischer Walker, R. E. Black, and H. Campbell, “Epidemiology and etiology of childhood pneumonia in 2010: estimates of incidence, severe morbidity, mortality, underlying risk factors and causative pathogens for 192 countries,” J Glob Health, vol. 3, no. 1, pp. 010401, Jun, 2013. [CrossRef]
- L. P. Tavares, I. Galvão, and M. R. Ferrero, "5.30 - Novel Immunomodulatory Therapies for Respiratory Pathologies," Comprehensive Pharmacology, T. Kenakin, ed., pp. 554-594, Oxford: Elsevier, 2022.
- W. H. Organization, "Pneumonia in children," https://www.who.int/news-room/fact-sheets/detail/pneumonia, [13 May 2025, 2022].
- Z. X. Zhang, Y. Yong, W. C. Tan, L. Shen, H. S. Ng, and K. Y. Fong, “Prognostic factors for mortality due to pneumonia among adults from different age groups in Singapore and mortality predictions based on PSI and CURB-65,” Singapore Med J, vol. 59, no. 4, pp. 190-198, Apr, 2018. [CrossRef]
- T. Eurich, T. J. Marrie, J. K. Minhas-Sandhu, and S. R. Majumdar, “Risk of heart failure after community acquired pneumonia: prospective controlled study with 10 years of follow-up,” Bmj, vol. 356, pp. j413, Feb 13, 2017. [CrossRef]
- J. P. Metlay, and M. J. Fine, “Testing strategies in the initial management of patients with community-acquired pneumonia,” Ann Intern Med, vol. 138, no. 2, pp. 109-18, Jan 21, 2003. [CrossRef]
- M. Khalifa, and M. Albadawy, “Artificial Intelligence for Clinical Prediction: Exploring Key Domains and Essential Functions,” Computer Methods and Programs in Biomedicine Update, vol. 5, pp. 100148, 2024/01/01/, 2024. [CrossRef]
- D. Panteli, K. Adib, S. Buttigieg, F. Goiana-da-Silva, K. Ladewig, N. Azzopardi-Muscat, J. Figueras, D. Novillo-Ortiz, and M. McKee, “Artificial intelligence in public health: promises, challenges, and an agenda for policy makers and public health institutions,” The Lancet Public Health, vol. 10, no. 5, pp. e428-e432, 2025/05/01/, 2025. [CrossRef]
- Yunianta, “A Novel Advanced Performance Ensemble-Based Model (APEM) Framework: A Case Study on Diabetes Prediction,” Journal of Advances in Information Technology, vol. 15, no. 10, pp. 1193-1204, 2024. [CrossRef]
- J. Bajwa, U. Munir, A. Nori, and B. Williams, “Artificial intelligence in healthcare: transforming the practice of medicine,” Future Healthcare Journal, vol. 8, no. 2, pp. e188-e194, 2021/07/01/, 2021. [CrossRef]
- M. Tsuneki, “Deep learning models in medical image analysis,” Journal of Oral Biosciences, vol. 64, no. 3, pp. 312-320, 2022/09/01/, 2022. [CrossRef]
- M. Kaya, and Y. Çetin-Kaya, “A novel ensemble learning framework based on a genetic algorithm for the classification of pneumonia,” Engineering Applications of Artificial Intelligence, vol. 133, pp. 108494, 2024/07/01/, 2024. [CrossRef]
- S. Sotirov, D. Orozova, B. Angelov, E. Sotirova, and M. Vylcheva, “Transforming Pediatric Healthcare with Generative AI: A Hybrid CNN Approach for Pneumonia Detection,” Electronics, vol. 14, no. 9, pp. 1878, 2025. [CrossRef]
- S. Rao, Z. Zeng, and J. Zhang, “Robust Multiclass Pneumonia Classification via Multi-Head Attention and Transfer Learning Ensemble,” Applied Sciences, vol. 15, no. 21, pp. 11426, 2025. [CrossRef]
- H. Bhatt, and M. Shah, “A Convolutional Neural Network ensemble model for Pneumonia Detection using chest X-ray images,” Healthcare Analytics, vol. 3, pp. 100176, 2023/11/01/, 2023. [CrossRef]
- Z. Yue, L. Ma, and R. Zhang, “Comparison and Validation of Deep Learning Models for the Diagnosis of Pneumonia,” Computational Intelligence and Neuroscience, vol. 2020, no. 1, pp. 8876798, 2020. [CrossRef]
- S. Rajaraman, I. Kim, and S. K. Antani, “Detection and visualization of abnormality in chest radiographs using modality-specific convolutional neural network ensembles,” PeerJ, vol. 8, pp. e8693, 2020. [CrossRef]
- M. N. Islam, "Classification of pediatric pneumonia using chest X-rays by functional regression," https://arxiv.org/abs/2005.03243, 2020].
- R. Alsharif, Y. Al-Issa, A. M. Alqudah, I. A. Qasmieh, W. A. Mustafa, and H. Alquran, “PneumoniaNet: Automated Detection and Classification of Pediatric Pneumonia Using Chest X-ray Images and CNN Approach,” Electronics, vol. 10, no. 23, pp. 2949, 2021. [CrossRef]
- V. Ravi, H. Narasimhan, and T. D. Pham, “A cost-sensitive deep learning-based meta-classifier for pediatric pneumonia classification using chest X-rays,” Expert Systems, vol. 39, no. 7, pp. e12966, 2022.
- Mohammed, and R. Kora, “A comprehensive review on ensemble deep learning: Opportunities and challenges,” Journal of King Saud University - Computer and Information Sciences, vol. 35, no. 2, pp. 757-774, 2023/02/01/, 2023. [CrossRef]
- J. Arun Prakash, C. R. Asswin, V. Ravi, V. Sowmya, and K. P. Soman, “Pediatric pneumonia diagnosis using stacked ensemble learning on multi-model deep CNN architectures,” Multimedia Tools and Applications, vol. 82, no. 14, pp. 21311-21351, 2023/06/01, 2023. [CrossRef]
- T. S. Arulananth, S. W. Prakash, R. K. Ayyasamy, V. P. Kavitha, P. G. Kuppusamy, and P. Chinnasamy, “Classification of Paediatric Pneumonia Using Modified DenseNet-121 Deep-Learning Model,” IEEE Access, vol. 12, pp. 35716-35727, 2024. [CrossRef]
- Z. Pan, H. Wang, J. Wan, L. Zhang, J. Huang, and Y. Shen, “Efficient federated learning for pediatric pneumonia on chest X-ray classification,” Scientific Reports, vol. 14, no. 1, pp. 23272, 2024/10/07, 2024. [CrossRef]
- T. Yoon, and D. Kang, “Enhancing pediatric pneumonia diagnosis through masked autoencoders,” Scientific Reports, vol. 14, no. 1, pp. 6150, 2024/03/14, 2024. [CrossRef]
- E. Galvis Ruiz, J. Benavides-Cruz, D. M. Corredor, E. Morales-Mendoza, H. D. A. Cotrino Palma, and A. Cely-Jiménez, “Development of deep learning-based classification models for opacity differentiation in pediatric chest radiography,” Informatics in Medicine Unlocked, vol. 52, pp. 101605, 2025/01/01/, 2025. [CrossRef]
- P. R, G. Gajendran, S. Boulaaras, and S. S. Tantawy, “PediaPulmoDx: Harnessing cutting edge preprocessing and explainable AI for pediatric chest X-ray classification with DenseNet121,” Results in Engineering, vol. 25, pp. 104320, 2025/03/01/, 2025. [CrossRef]
- S. Katreddi, A. Midatani, A. P. Roy, U. Velpuri, and S. Kasani, “Pediatric pneumonia X-ray image classification: predictive model development with DenseNet-169 transfer learning,” Journal of Medical Artificial Intelligence, 2025. [CrossRef]
- S. Nazir, D. M. Dickson, and M. U. Akram, “Survey of explainable artificial intelligence techniques for biomedical imaging with deep neural networks,” Computers in Biology and Medicine, vol. 156, pp. 106668, 2023/04/01/, 2023. [CrossRef]
- R. R. Selvaraju, M. Cogswell, A. Das, R. Vedantam, D. Parikh, and D. Batra, "Grad-CAM: Visual Explanations from Deep Networks via Gradient-Based Localization." pp. 618-626. [CrossRef]
- D. Kermany, K. Zhang, and M. Goldbaum, "Labeled Optical Coherence Tomography (OCT) and Chest X-Ray Images for Classification," 2018.
- M. Mujahid, F. Rustam, R. Álvarez, J. Luis Vidal Mazón, I. T. Díez, and I. Ashraf, “Pneumonia Classification from X-ray Images with Inception-V3 and Convolutional Neural Network,” Diagnostics (Basel), vol. 12, no. 5, May 21, 2022. [CrossRef]
- Ke, W. Ellsworth, O. Banerjee, A. Y. Ng, and P. Rajpurkar, “CheXtransfer: performance and parameter efficiency of ImageNet models for chest X-Ray interpretation,” in Proceedings of the Conference on Health, Inference, and Learning, Virtual Event, USA, 2021, pp. 116–124.
- Goodfellow, Y. Bengio, and A. Courville, Deep Learning: The MIT Press, 2016.
- C. Aggarwal, Neural Networks and Deep Learning: Springer Cham, 2023.
- S. Santurkar, D. Tsipras, A. Ilyas, and A. Madry, “How Does Batch Normalization Help Optimization?,” in 32nd Conference on Neural Information Processing Systems (NIPS 2018), Montréal, Canada, 2018.
- S. Ioffe, and C. Szegedy, “Batch normalization: accelerating deep network training by reducing internal covariate shift,” in Proceedings of the 32nd International Conference on International Conference on Machine Learning - Volume 37, Lille, France, 2015, pp. 448–456.
- Mumuni, and F. Mumuni, “Data augmentation: A comprehensive survey of modern approaches,” Array, vol. 16, pp. 100258, 2022/12/01/, 2022. [CrossRef]
| Component | Configuration / Description |
|---|---|
| Optimizer | Adam (Adaptive Moment Estimation) with decoupled weight decay for stable and efficient updates |
| Learning Rate | 1×10−41 \times 10^(-4)1×10−4 — fine-tuned to ensure steady convergence without overshooting minima |
| Loss Function | Binary Cross-Entropy — appropriate for probabilistic outputs in binary classification |
| Epochs | 100 — capped with early stopping (patience = 10) to prevent overfitting |
| Batch Size | 32 — balanced for computational efficiency and learning stability |
| Regularization | Dropout with p=0.5 p = 0.5 p=0.5 applied in the fully connected layers to mitigate overfitting |
| Hardware | NVIDIA Tesla T4 GPU via Google Colab Pro for accelerated parallel training |
| Training Parameters | Values/Types |
|---|---|
| Model Architecture | MobileNetV2, VGG19, ResNet-50, DenseNet-201, EfficientNet-B0 (Pre-trained) |
| Optimizer | Adam (Learning Rate: 1e-4) |
| Loss Function | Categorical Crossentropy |
| Batch Size | 32 |
| Epochs | 100 |
| Dropout Rate (Layer 1) | 0.5 |
| Dropout Rate (Layer 2) | 0.3 |
| Learning Rate | 1e-4 |
| Weight Initialization | He Initialization |
| Activation Function | ReLU |
| Final Activation Function | Softmax |
| Input Size | 224 × 224 × 3 |
| Model Combination | Accuracy (%) | Precision (%) | Recall (%) | F1-Score (%) |
|---|---|---|---|---|
| MobileNetV2 + ResNet50 + EfficientNetB0 | 96.14 | 94.10 | 96.92 | 94.97 |
| DenseNet201 + EfficientNetB0 + MobileNetV2 | 92.31 | 91.11 | 97.18 | 94.04 |
| EfficientNetB0 + InceptionV3 + Xception | 92.63 | 92.56 | 91.62 | 92.05 |
| EfficientNetB0 + ResNet50 + VGG16 | 89.74 | 87.73 | 97.18 | 92.21 |
| InceptionV3 + ResNet101 + EfficientNet | 81.41 | 80.58 | 92.56 | 86.16 |
| Model | Accuracy (%) | Precision (%) | Recall (%) | F1-Score (%) |
|---|---|---|---|---|
| MobileNetV2 | 93.18 | 92.86 | 95.26 | 93.18 |
| ResNet-50 | 93.11 | 92.24 | 94.97 | 92.87 |
| DenseNet-201 | 92.64 | 91.76 | 94.68 | 92.47 |
| EfficientNet-B0 | 91.36 | 90.89 | 92.93 | 91.48 |
| VGG19 | 74.29 | 55.19 | 74.29 | 63.33 |
| Model | Accuracy (%) | Precision (%) | Recall (%) | F1-Score (%) |
|---|---|---|---|---|
| MobileNetV2 | 86.16 | 84.43 | 88.17 | 85.47 |
| ResNet-50 | 88.38 | 86.47 | 89.72 | 87.62 |
| EfficientNet-B0 | 87.85 | 85.79 | 88.39 | 85.96 |
| Static Ensemble | 89.14 | 87.28 | 90.48 | 88.35 |
| Proposed Work | 94.73 | 91.03 | 96.12 | 93.47 |
| Study | Dataset | Model | Accuracy (%) | Precision (%) | Recall (%) | F1-Score (%) | Notes |
|---|---|---|---|---|---|---|---|
| Rajaraman et al. (2020) [18] | 1000 Chest X-rays | ResNet50 | 93.06 | 88.97 | 96.78 | 92.71 | High performance with deep residual learning. |
| Yue et al. (2020) [17] | 5863 Chest X-rays | MobileNet | 92.98 | 93.10 | 98.98 | 93.00 | Balanced metrics suitable for clinical applications. |
| Bhatt and Shah (2023) [16] | 5863 Chest X-rays | ensemble network of 3 CNN models | 84.12 | 80.04 | 99.23 | 88.56 | Combines CNN feature extraction with machine learning classifier. |
| Sotirov et al. (2025) [14] | 5863 Chest X-rays | (CNN) with intuitionistic fuzzy estimation (IFE) | 94.93 | 93.00 | 91.00 | 91.00 | Combines convolutional neural networks with intuitionistic fuzzy estimators. |
| Rao et al. (2025) [15] | 5863 Chest X-rays | Ensemble DenseNet-121, ResNet-50, and VGG-19 | 91.67 | 92.19 | 90.00 | 90.89 | Proposes multimodel ensemble learning framework based on multi-head attention mechanism. |
| Proposed Work |
5863 Chest X-rays | MobileNetV2 + ResNet50 + EfficientNetB0 | 96.14 | 94.10 | 96.92 | 94.97 | Achieve superior accuracy and recall, ensuring robust and balanced. |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




