Submitted:
03 March 2026
Posted:
04 March 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
3. Theory
4. Results
- (i)
- Single lifetime and single correlation time.
- (ii)
- Single lifetime and two correlation times
- (iii)
- Two lifetimes and one correlation time
- (iv) Two lifetimes and two correlation times (un-associated)
- (v) Two lifetimes and two correlation times (associated)
4. Discussion
5. Conclusions
Funding
Acknowledgments
Conflicts of Interest
Abbreviations
| FLIM | Fluorescence lifetime imaging microscopy |
| ICS | Image Correlation Spectroscopy |
| CD | Cluster Density |
| B | Brightness |
Appendix A
Appendix A.1
References
- Magde, D.; Elson, E. L.; Webb, W. W. Thermodynamic fluctuations in a reacting system: measurement by fluorescence correlation spectroscopy. Phys. Rev. Lett. 1972, 29, 705–708. [Google Scholar] [CrossRef]
- Petersen, N. O. Scanning fluorescence correlation spectroscopy. I. Theory and simulation of aggregation measurements. Biophys. J. 1986, 49, 809–815. [Google Scholar] [CrossRef]
- St-Pierre, P. R.; Petersen, N. O. Relative ligand binding to small or large aggregates measured by scanning correlation spectroscopy. Biophys. J. 1990, 58, 503–511. [Google Scholar] [CrossRef]
- Petersen, N. O.; Hoddelius, P. L.; Wiseman, P.W.; Seger, O.; Magnusson, K. E. Quantitation of membrane receptor distributions by image correlation spectroscopy: concept and application. Biophys. J. 1993, 65, 1135–1146. [Google Scholar] [CrossRef]
- Wiseman, P. W.; Petersen, N. O. Image correlation spectroscopy. II. Optimization for ultrasensitive detection of preexisting platelet-derived growth factor-β receptor oligomers on intact cells. Biophys. J. 1999, 76, 963–977. [Google Scholar] [CrossRef]
- Petersen, N. O.; Brown, C.; Wiseman, P. W. Analysis of membrane protein cluster densities and sizes in situ by image correlation spectroscopy. Faraday Discuss. 1998, 111, 289–305. [Google Scholar] [CrossRef]
- Costantino, S.; Comeau, J. W.; Wiseman, P. W. Accuracy and dynamic range of spatial image correlation and cross-correlation spectroscopy. Biophys. J. 2005, 89, 1251–1260. [Google Scholar] [CrossRef] [PubMed]
- Comeau, J. W.; Kolin, D. L.; Wiseman, P. W. Accurate measurements of protein interactions in cells via improved spatial image cross-correlation spectroscopy. Mol. Biosyst. 2008, 4, 672–685. [Google Scholar] [CrossRef] [PubMed]
- Hebert, B.; Costantino, S.; Wiseman, P. W. Spatiotemporal image correlation spectroscopy (STICS) theory, verification, and application to protein velocity mapping in living CHO cells. Biophys. J. 2005, 88, 3601–3614. [Google Scholar] [CrossRef] [PubMed]
- Lakowicz, J.R.; Berndt, K.W. Lifetime selective fluorescence imaging using an rf phase sensitive camera. Rev. Sci. Instrum. 1991, 62, 1727. [Google Scholar] [CrossRef]
- Datta, R; Heaster, TM; Sharick, JT; Gillette, AA; Skala, MC. Fluorescence lifetime imaging microscopy: fundamentals and advances in instrumentation, analysis, and applications. J Biomed Opt. 2020, 25(7), 1–43. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Clayton, AH; Hanley, QS; Verveer, PJ. Graphical representation and multicomponent analysis of single-frequency fluorescence lifetime imaging microscopy data. J Microsc. 2004, 213(1), 1–5. [Google Scholar] [CrossRef] [PubMed]
- Redford, GI; Clegg, RM. Polar plot representation for frequency-domain analysis of fluorescence lifetimes. J Fluoresc. 2005, 15(5), 805–15. [Google Scholar] [CrossRef] [PubMed]
- Digman, MA; Caiolfa, VR; Zamai, M; Gratton, E. The phasor approach to fluorescence lifetime imaging analysis. Biophys J 2008, 94(2), L14–6. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Vallmitjana, A; Torrado, B; Dvornikov, A; Ranjit, S; Gratton, E. Blind Resolution of Lifetime Components in Individual Pixels of Fluorescence Lifetime Images Using the Phasor Approach. J Phys Chem B 2020, 124(45), 10126–10137. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Oleksiievets, N; Thiele, JC; Weber, A; Gregor, I; Nevskyi, O; Isbaner, S; Tsukanov, R; Enderlein, J. Wide-Field Fluorescence Lifetime Imaging of Single Molecules. J Phys Chem A 2020, 124(17), 3494–3500. [Google Scholar] [CrossRef] [PubMed]
- Weber, G. Polarization of the fluorescence of macromolecules. I. Theory and experimental method Biochem. J. 1952, 51, 145–55. [Google Scholar] [PubMed]
- Weber, G. Polarization of the fluorescence of macromolecules. II. Fluorescent conjugates of ovalbumin and bovine serum albumin. Biochem. J. 1952, 51, 155–67. [Google Scholar] [CrossRef] [PubMed]
- Porter, G; Sadkowski, P J; Treadle, C J. Picosecond rotational diffusion in kinetic and steady-state fluorescence spectroscopy. Chem. Phys. Lett. 1977, 49, 416–20. [Google Scholar] [CrossRef]
- Spencer, R D; Weber, G. Influence of Brownian rotations and energy transfer upon the measurements of fluorescence lifetime. J. Chem. Phys. 1970, 52, 1654. [Google Scholar] [CrossRef]
- Cross, A J; Fleming, G R. Analysis of time-resolved fluorescence anisotropy decays. Biophys J. 1984, 46, 45–56. [Google Scholar] [CrossRef] [PubMed]
- Lakowicz, J R; Cherek, H; Maliwal, B P; Gratton, E. Time resolved fluorescence anisotropies of diphenylhexatriene and perylene in solvents and lipid bilayers obtained from multifrequency phase-modulation fluorometry. Biochemistry 1985, 24, 376–83. [Google Scholar] [CrossRef]
- Siegel, J; Suhling, K; French, P M W; Phillips, D; Davis, D M; Webb, S E D; Sabharwal, Y; Levequefort, S. Wide-field time-resolved fluorescence anisotropy imaging (TR-FAIM): imaging the rotational mobility of a fluorophore. Rev. Sci. Instrum. 2003, 74, 182–92. [Google Scholar] [CrossRef]
- Clayton, A H A; Hanley, Q S; Arndt-Jovin, D J; Subramaniam, V; Jovin, T M. 2002 Dynamic fluorescence anisotropy imaging microscopy in the frequency domain (rFLIM). Biophys. J. 83, 1631–49. [CrossRef] [PubMed]
- Weber, G. Theory of differential phase fluorometry: detection of anisotropic molecular rotations. J. Chem. Phys. 1977, 66, 4081–91. [Google Scholar] [CrossRef]
- Lakowicz, J R; Prendergast, F G. Quantitation of hindered rotations of diphenylhexatriene in lipid bilayers by differential polarized phase fluorometry. Science 1978, 200, 1399–401. [Google Scholar] [CrossRef]
- Weber, G; Helgerson, S L; Cramer, W A; Mitchell, G W. Changes in rotational motion of a cell-bound fluorophore caused by colicin E1: a study by fluorescence polarization and differential polarized phase fluorometry. Biochemistry 1976, 15, 4429–32. [Google Scholar] [CrossRef]
- Kozer, N; Clayton, AHA. Analysis of complex anisotropy decays from single-frequency polarized-phasor ellipse plots. Methods Appl. Fluoresc. 2016, 4, 024005. [Google Scholar] [CrossRef]
- Kozer, N; Clayton, AHA. In-cell structural dynamics of an EGF receptor during ligand-induced dimer-oligomer transition. Eur Biophys 2020, 49(1), 21–37. [Google Scholar] [CrossRef]
- Clayton, AHA. Phase-Sensitive Fluorescence Image Correlation Spectroscopy. Int J Mol Sci. 2024, 25(20), 11165. [Google Scholar] [CrossRef]
- Smith, TA; Ghiggino, KP. A review of the analysis of complex time-resolved fluorescence anisotropy data. Methods Appl Fluoresc. 2015, 3(2), 022001. [Google Scholar] [CrossRef]
- O.Ciccotosto, GD; Kozer, N; Chow, TT; Chon, JW; Clayton, AH. Aggregation distributions on cells determined by photobleaching image correlation spectroscopy. Biophys J. 2013, 104(5), 1056–64. [Google Scholar] [CrossRef]
- Klymchenko, AS. Fluorescent Probes for Lipid Membranes: From the Cell Surface to Organelles. Acc Chem Res. 2023, 56(1), 1–12. [Google Scholar] [CrossRef]
- Steinmark, IE; James, AL; Chung, PH; Morton, PE; Parsons, M; Dreiss, CA; Lorenz, CD; Yahioglu, G; Suhling, K. Targeted fluorescence lifetime probes reveal responsive organelle viscosity and membrane fluidity. PLoS One 2019, 14(2), e0211165. [Google Scholar] [CrossRef]
- Lira, RB; Dillingh, LS; Schuringa, JJ; Yahioglu, G; Suhling, K; Roos, WH. Fluorescence lifetime imaging microscopy of flexible and rigid dyes probes the biophysical properties of synthetic and biological membranes. Biophys J 2024, 123(12), 1592–1609. [Google Scholar] [CrossRef]
- Lim, C; Seah, D; Vendrell, M. Chemical fluorophores for fluorescence lifetime imaging 2026. Chem Soc Rev. 2026, 55(3), 1352–1370. [Google Scholar] [CrossRef] [PubMed]
- Long, Y.; Stahl, Y.; Weidtkamp-Peters, S.; et al. In vivo FRET–FLIM reveals cell-type-specific protein interactions in Arabidopsis roots. Nature 2017, 548, 97–102. [Google Scholar] [CrossRef]
- Jares-Erijman, E.; Jovin, T. FRET imaging. Nat Biotechnol 2003, 21, 1387–1395. [Google Scholar] [CrossRef]
- Weber, A.; Hartig, R.; Zuschratter, W. FRET-analysis in living cells by fluorescence lifetime imaging microscopy: experimental workflow and methodology. Methods in Microscopy 2025, 2(1), 73–84. [Google Scholar] [CrossRef]
- Berezin, MY; Achilefu, S. Fluorescence lifetime measurements and biological imaging. Chem Rev. 2010, 110(5), 2641–84. [Google Scholar] [CrossRef] [PubMed]
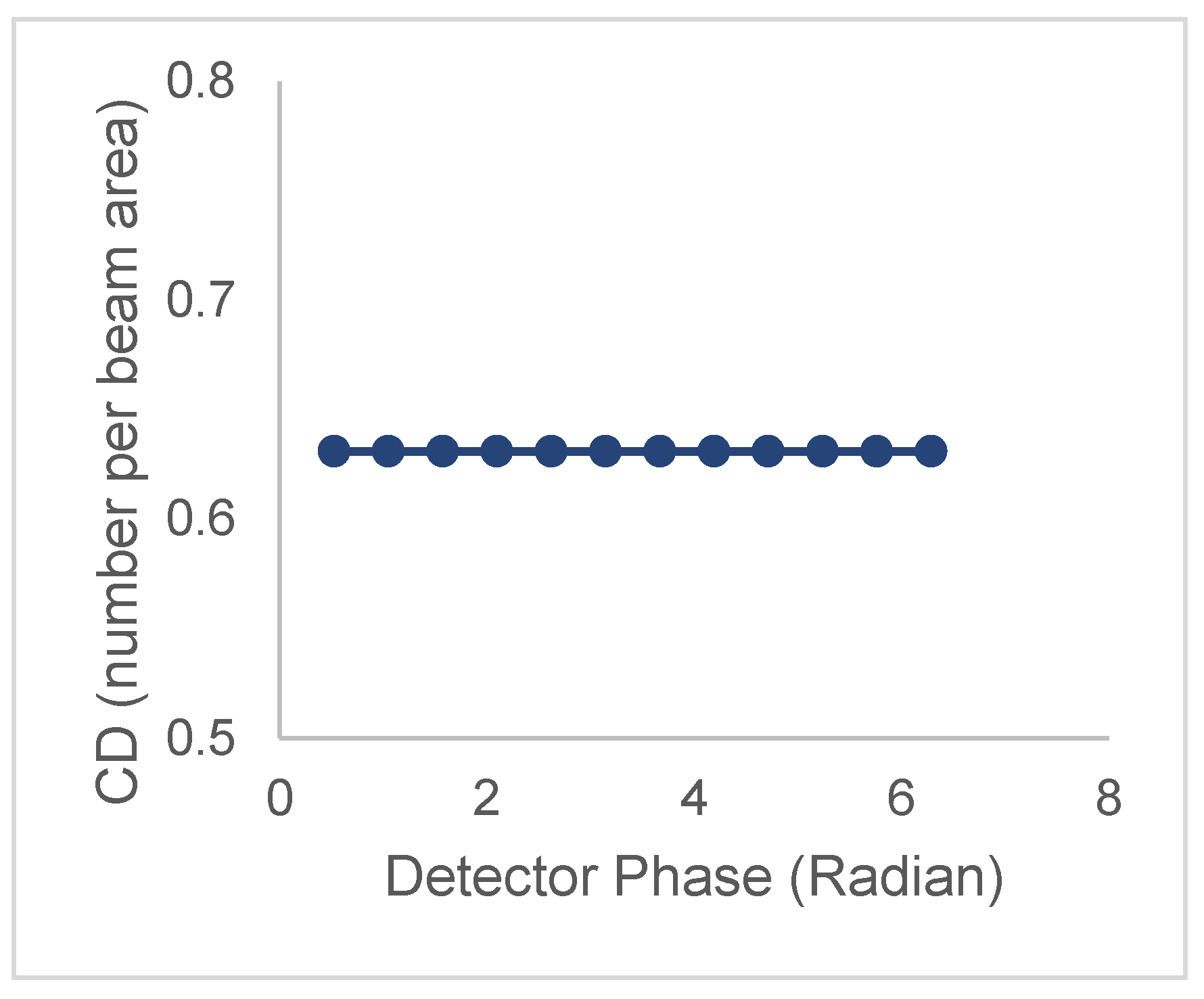
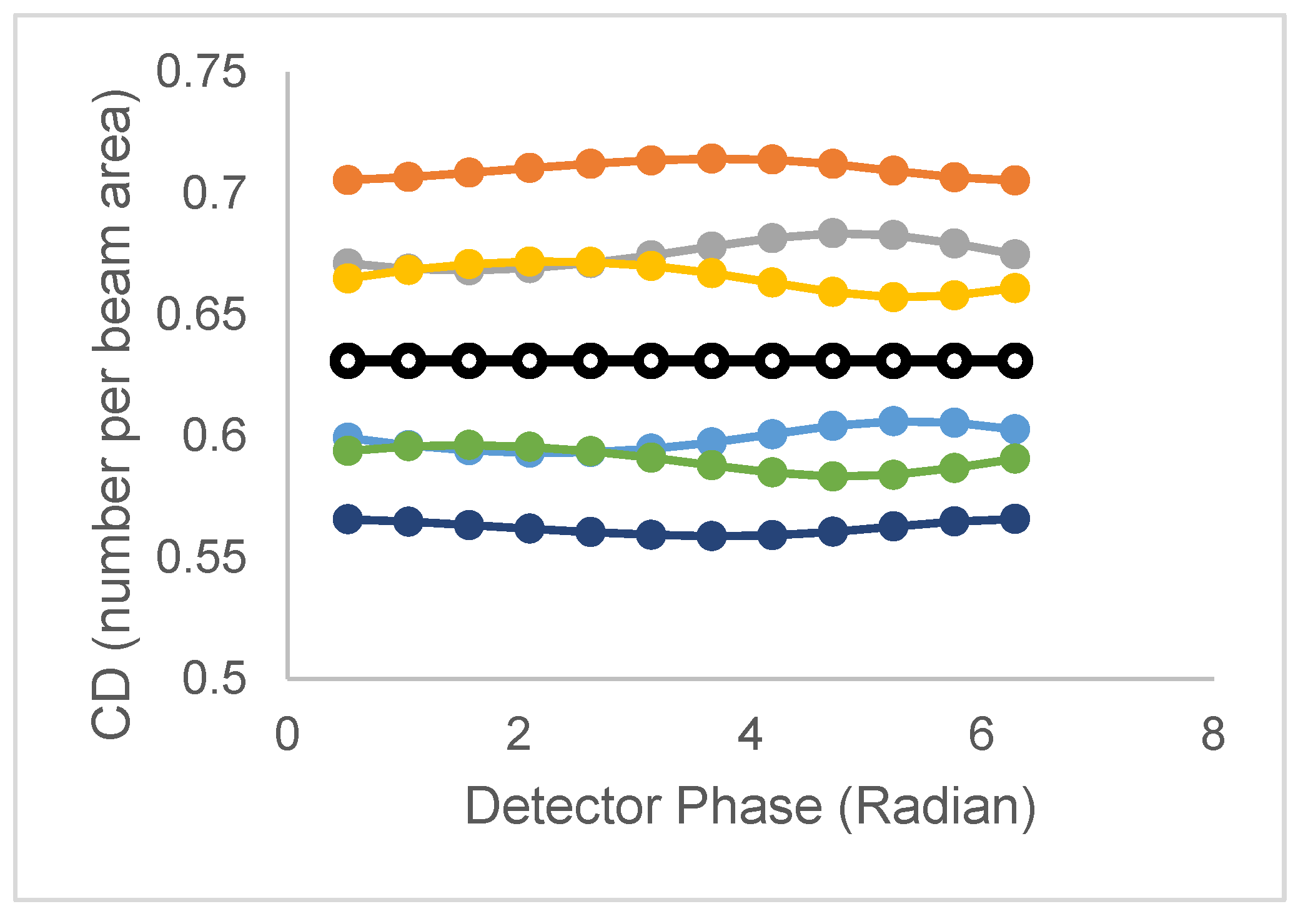
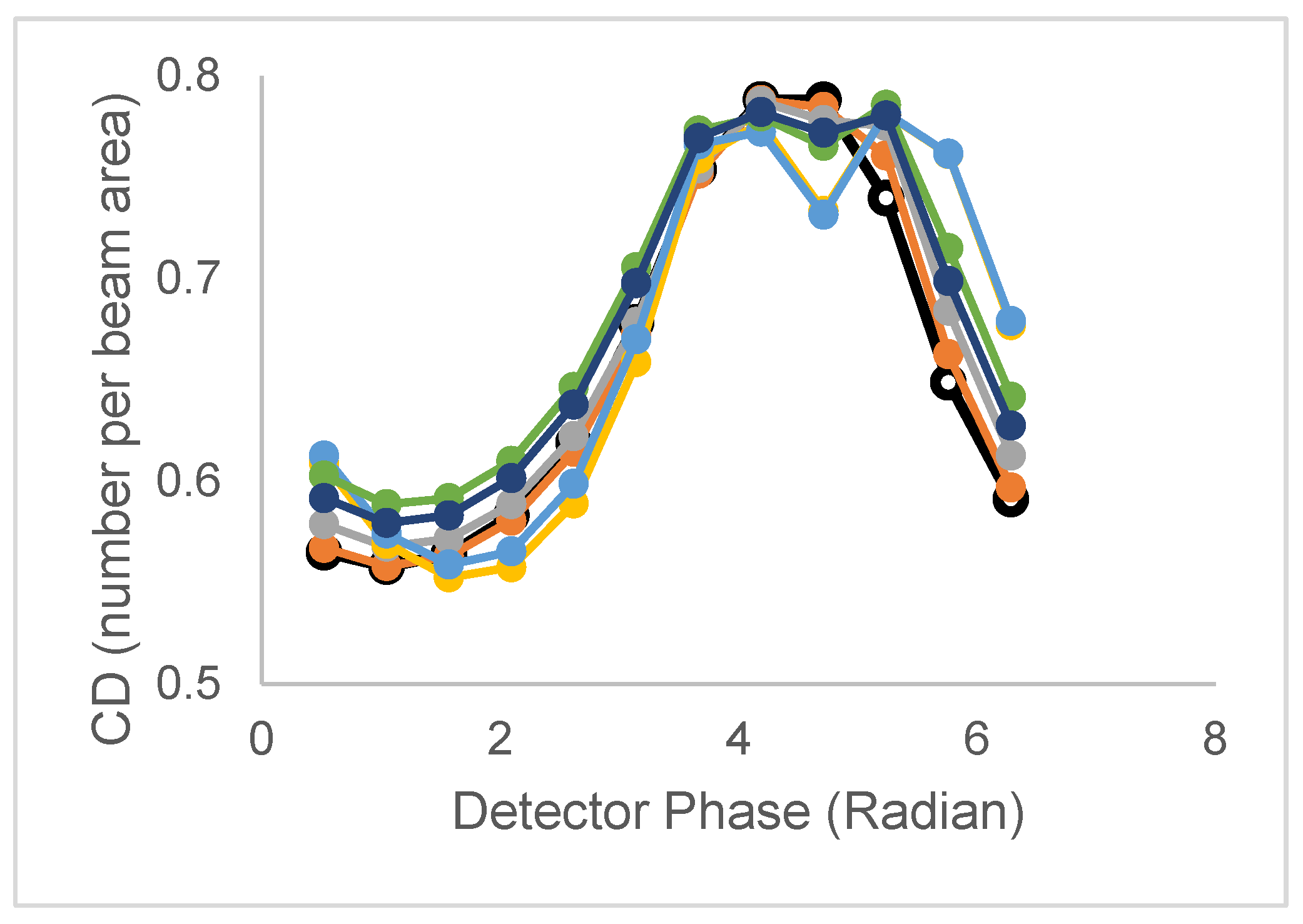
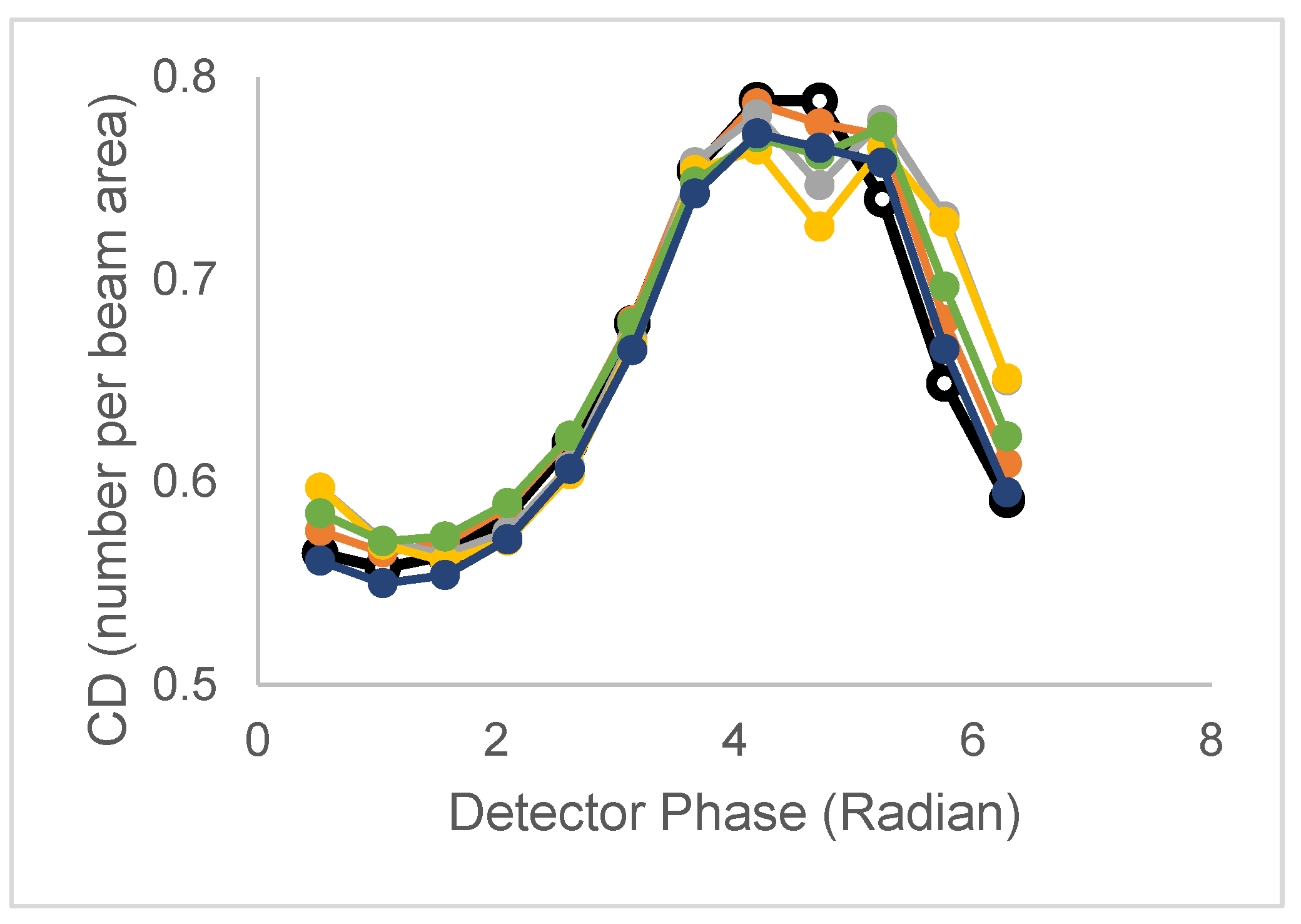
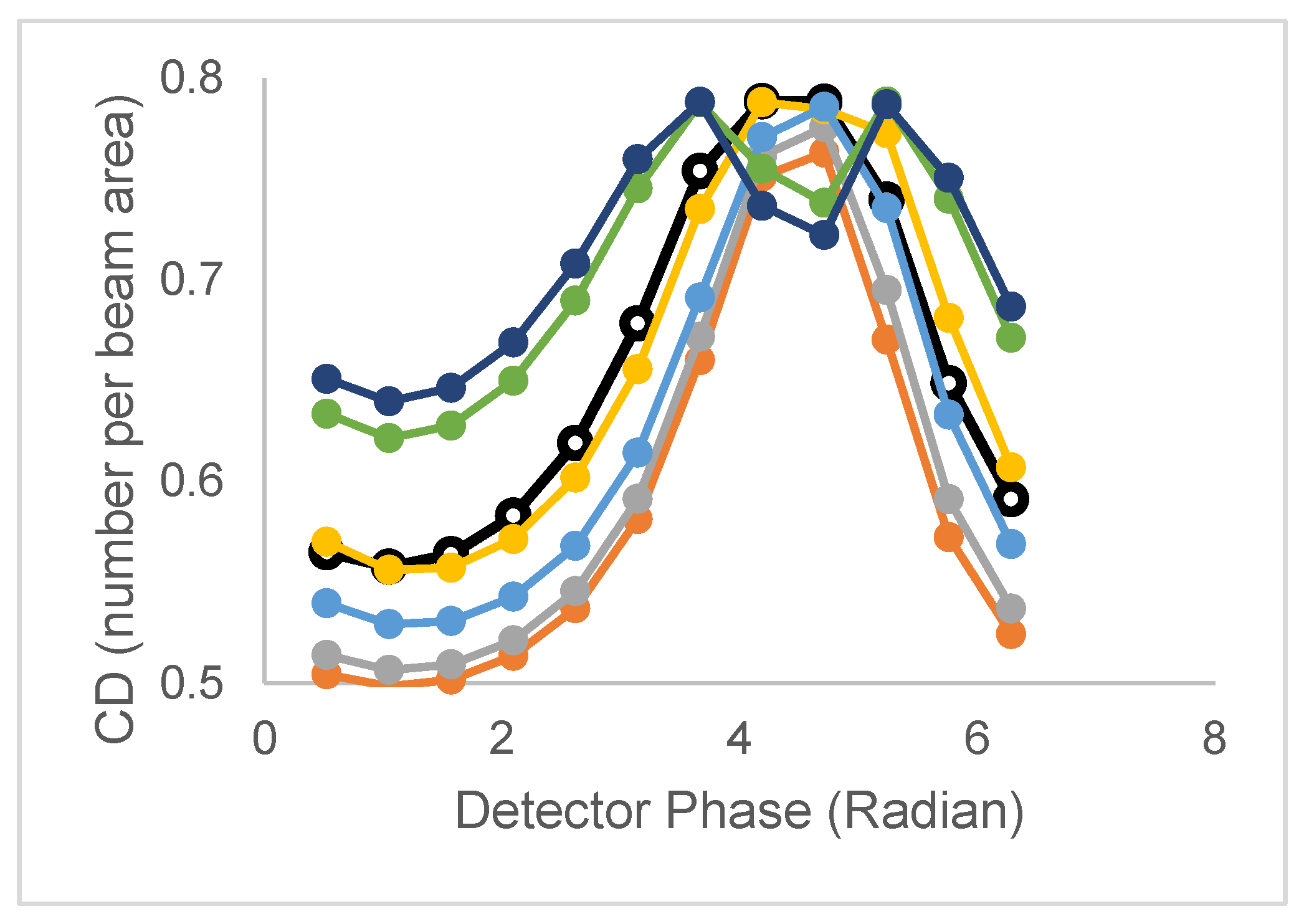
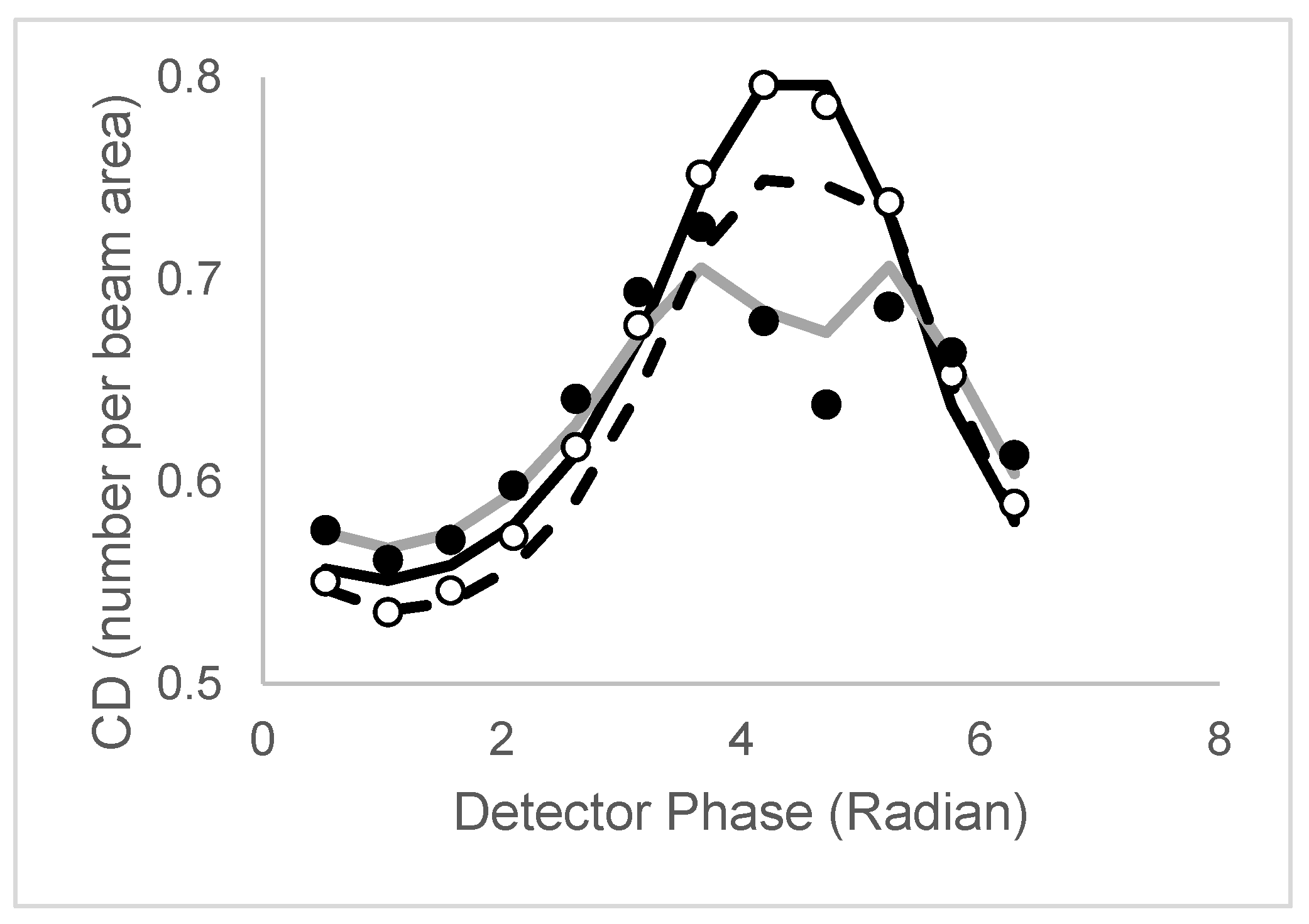
| Model | Lifetime | Correlation time | CD/CDperp |
|---|---|---|---|
| 1 τ and 1 ϕ | CD is constant | ||
| τ | ϕ | CDperp is constant | |
| 1 τ and 2 ϕ | τ1=τ2 | ϕ1,ϕ2 | CD is constant |
| CDperp =F(ϕ1,ϕ2) | |||
| 2 τ and 1 ϕ | τ1,τ2 | ϕ | CD has peak CDperp broad,split |
| 2 τ and 2 ϕ (non-assoc) | τ1,τ2 | ϕ1,ϕ2 | CD has peak CDperp broad,split |
| 2 τ and 2 ϕ (assoc) | τ1,τ2 | ϕ1,ϕ2 | CD has peak CDperp broad, split, and amplitude |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




