Submitted:
02 March 2026
Posted:
03 March 2026
You are already at the latest version
Abstract
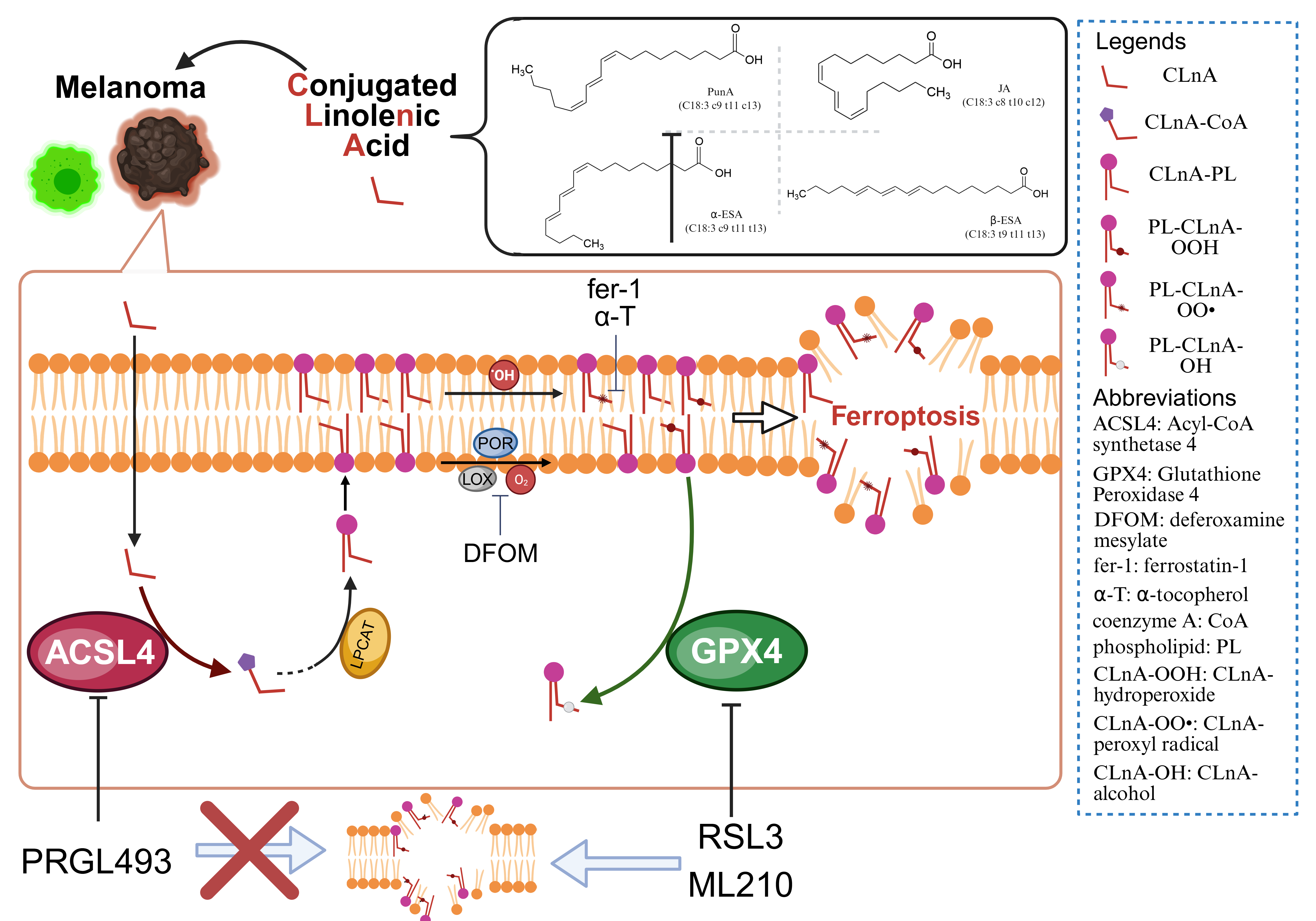
Keywords:
1. Introduction
2. Materials and Methods
2.1 Cell culture
2.2 Cell viability test
2.3 Kinetics of ZMEL1 cell viability loss
2.4 Fatty acid uptake and metabolic processing in ZMEL1 cells
2.5 Impact of CLnAs on acsl4 and gpx4 expression in ZMEL1 cells
2.6 Statistical analysis
3. Results
3.1 CLnAs are cytotoxic to melanoma cells
3.2 Ferroptosis inhibitors mitigate CLnA-induced toxicity in melanoma cells
3.3 GPX4 or ACSL4 inhibition influences PunA cytotoxicity in melanoma cells
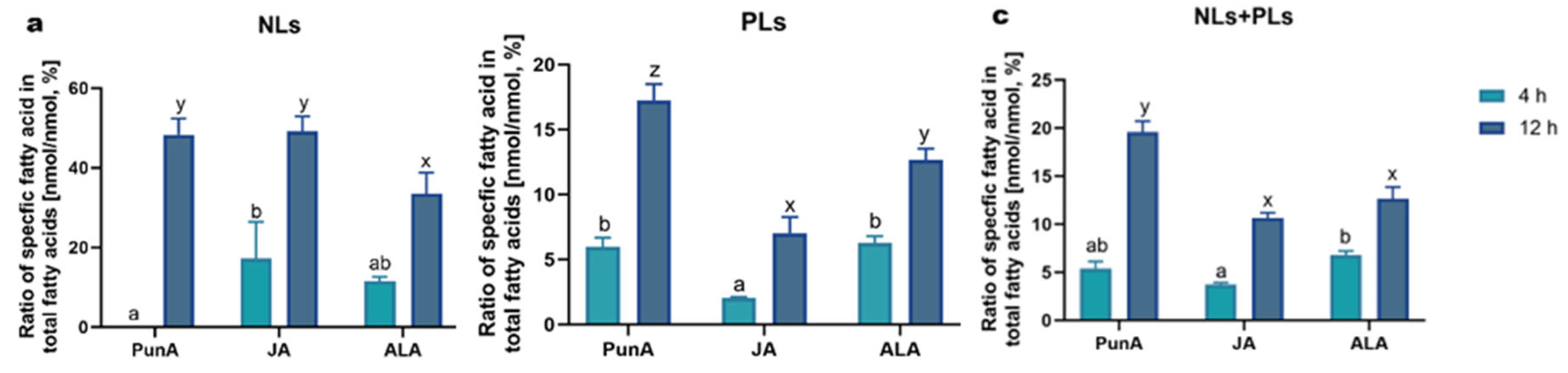
3.4 CLnAs show efficient cellular uptake and differential distribution in different fractions
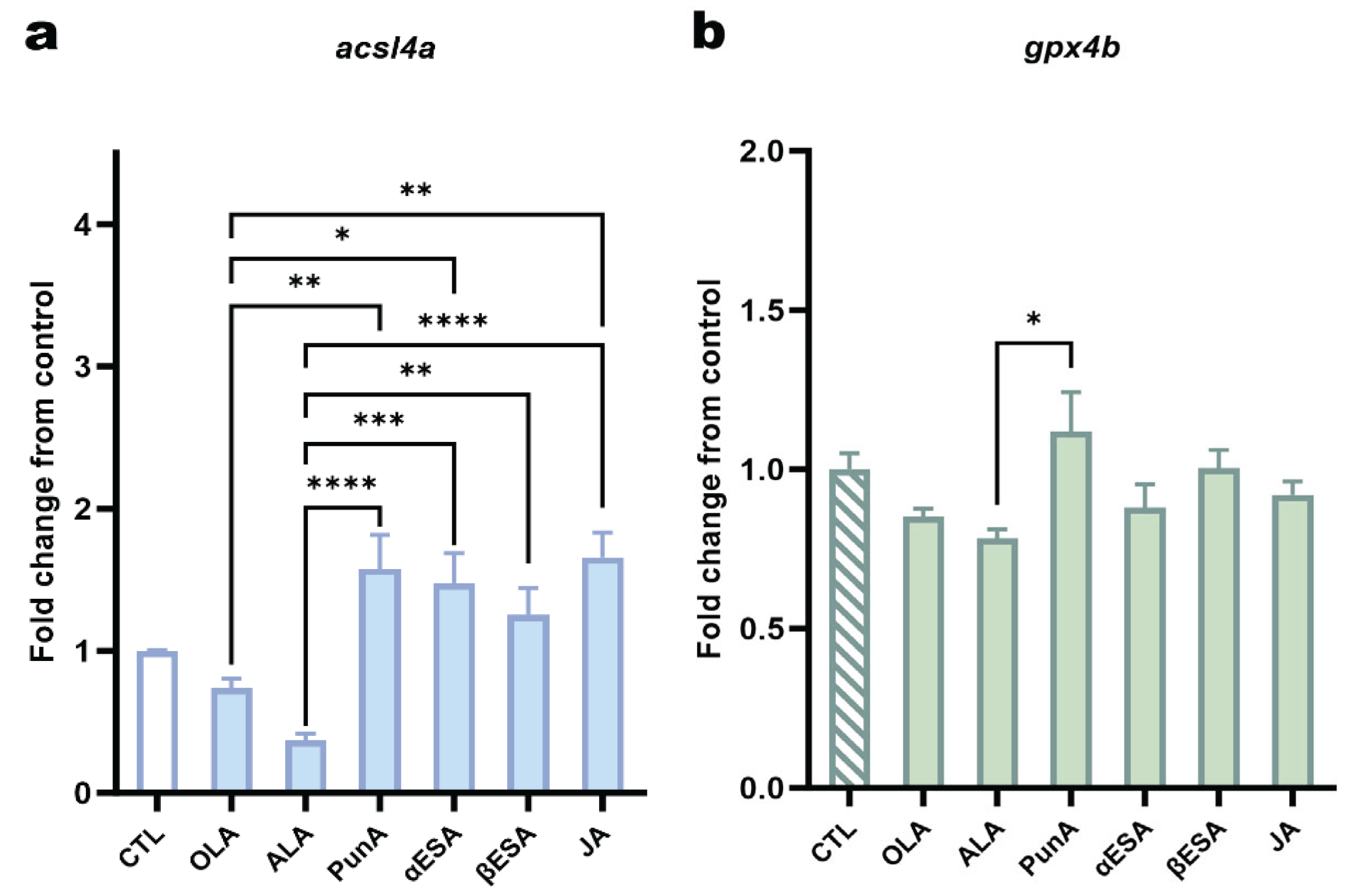
3.5 CLnA isomers differentially modulate ACSL4 and GPX4 expression in ZMEL1 cells
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
Abbreviations
| α-ESA | α-eleostearic acid |
| ALA | α-linolenic acid |
| α-T | α-tocopherol |
| β-ESA | β-eleostearic acid |
| ACSL4 | Acyl-CoA synthetase long-chain family member 4 |
| CLnA | Multidisciplinary Digital Publishing Institute |
| PunA | Directory of open access journals |
| DFOM | deferoxamine mesylate |
| fer-1 | ferrostatin-1 |
| GPX4 | glutathione peroxidase 4 |
| JA | jacaric acid |
| LPCAT | lysophosphatidylcholine acyltransferase |
| nec-1 | necrostatin-1 |
| OLA | oleic acid |
| PL | phospholipid |
| PUFA | polyunsaturated fatty acid |
| PunA | punicic acid |
| RQ | relative quantity |
References
- Mirza, F.N.; Yumeen, S.; Walter, F.M. The Epidemiology of Malignant Melanoma, Squamous Cell Carcinoma and Basal Cell Carcinoma in the UK from 2004 to 2014: A Population-based Cohort Analysis Using the Clinical Practice Research Datalink. Br. J. Dermatol. 2021, 184, 365–367. [Google Scholar] [CrossRef]
- Van Rooijen, E.; Fazio, M.; Zon, L.I. From Fish Bowl to Bedside: The Power of Zebrafish to Unravel Melanoma Pathogenesis and Discover New Therapeutics. Pigm. Cell Melanoma Res. 2017, 30, 402–412. [Google Scholar] [CrossRef]
- Wagle, N.; Emery, C.; Berger, M.F.; Davis, M.J.; Sawyer, A.; Pochanard, P.; Kehoe, S.M.; Johannessen, C.M.; MacConaill, L.E.; Hahn, W.C.; et al. Dissecting Therapeutic Resistance to RAF Inhibition in Melanoma by Tumor Genomic Profiling. J. Clin. Oncol. 2011, 29, 3085–3096. [Google Scholar] [CrossRef] [PubMed]
- Arnold, M.; Singh, D.; Laversanne, M.; Vignat, J.; Vaccarella, S.; Meheus, F.; Cust, A.E.; De Vries, E.; Whiteman, D.C.; Bray, F. Global Burden of Cutaneous Melanoma in 2020 and Projections to 2040. JAMA Dermatol. 2022, 158, 495. [Google Scholar] [CrossRef]
- Pérez, C.N.; Falcón, C.R.; Mons, J.D.; Orlandi, F.C.; Sangiacomo, M.; Fernandez-Muñoz, J.M.; Guerrero, M.; Benito, P.G.; Colombo, M.I.; Zoppino, F.C.M..; et al. Melanoma Cells with Acquired Resistance to Vemurafenib Have Decreased Autophagic Flux and Display Enhanced Ability to Transfer Resistance. Biochim. Biophys. Acta (BBA) - Mol. Basis Dis. 2023, 1869, 166801. [Google Scholar] [CrossRef] [PubMed]
- Boz Er, A.B.; Sheldrake, H.M.; Sutherland, M. Overcoming Vemurafenib Resistance in Metastatic Melanoma: Targeting Integrins to Improve Treatment Efficacy. Int. J. Mol. Sci. 2024, 25, 7946. [Google Scholar] [CrossRef]
- Natarelli, N.; Aleman, S.J.; Mark, I.M.; Tran, J.T.; Kwak, S.; Botto, E.; Aflatooni, S.; Diaz, M.J.; Lipner, S.R. A Review of Current and Pipeline Drugs for Treatment of Melanoma. Pharmaceuticals 2024, 17, 214. [Google Scholar] [CrossRef]
- Tanaka, T.; Hosokawa, M.; Yasui, Y.; Ishigamori, R.; Miyashita, K. Cancer Chemopreventive Ability of Conjugated Linolenic Acids. Int. J. Mol. Sci. 2011, 12, 7495–7509. [Google Scholar] [CrossRef]
- Ngo Njembe, M.T.; Dormal, E.; Gardin, C.; Mignolet, E.; Debier, C.; Larondelle, Y. Effect of the Dietary Combination of Flaxseed and Ricinodendron Heudelotii or Punica Granatum Seed Oil on the Fatty Acid Profile of Eggs. Food Chem. 2021, 344, 128668. [Google Scholar] [CrossRef]
- Gasmi, J.; Sanderson, J.T. Jacaric Acid and Its Octadecatrienoic Acid Geoisomers Induce Apoptosis Selectively in Cancerous Human Prostate Cells: A Mechanistic and 3-D Structure–Activity Study. Phytomedicine 2013, 20, 734–742. [Google Scholar] [CrossRef] [PubMed]
- Dhar Dubey, K.K.; Sharma, G.; Kumar, A. Conjugated Linolenic Acids: Implication in Cancer. J. Agric. Food. Chem. 2019, 67, 6091–6101. [Google Scholar] [CrossRef] [PubMed]
- Igarashi, M.; Miyazawa, T. Newly Recognized Cytotoxic Effect of Conjugated Trienoic Fatty Acids on Cultured Human Tumor Cells. Cancer Lett. 2000, 148, 173–179. [Google Scholar] [CrossRef]
- Yasui, Y.; Hosokawa, M.; Kohno, H.; Tanaka, T.; Miyashita, K. Troglitazone and 9Cis,11Trans,13Trans-Conjugated Linolenic Acid: Comparison of Their Antiproliferative and Apoptosis-Inducing Effects on Different Colon Cancer Cell Lines. Chemotherapy 2006, 52, 220–225. [Google Scholar] [CrossRef] [PubMed]
- Liu, W.-N.; Leung, K.-N. Apoptosis- and Differentiation-Inducing Activities of Jacaric Acid, a Conjugated Linolenic Acid Isomer, on Human Eosinophilic Leukemia EoL-1 Cells. Oncol. Rep. 2014, 32, 1881–1888. [Google Scholar] [CrossRef]
- Cuvelier, G.; Vermonden, P.; Debisschop, P.; Martin, M.; Derouane, F.; Liebisch, G.; Ecker, J.; Hoering, M.; Berlière, M.; Van Bockstal, M.; et al. Jacaric Acid Empowers RSL3-Induced Ferroptotic Cell Death in Two- and Three-Dimensional Breast Cancer Cell Models. Int. J. Mol. Sci. 2025, 26, 3375. [Google Scholar] [CrossRef]
- Vermonden, P.; Vancoppenolle, M.; Dierge, E.; Mignolet, E.; Cuvelier, G.; Knoops, B.; Page, M.; Debier, C.; Feron, O.; Larondelle, Y. Punicic Acid Triggers Ferroptotic Cell Death in Carcinoma Cells. Nutrients 2021, 13, 2751. [Google Scholar] [CrossRef]
- Vermonden, P.; Martin, M.; Glowacka, K.; Neefs, I.; Ecker, J.; Höring, M.; Liebisch, G.; Debier, C.; Feron, O.; Larondelle, Y. Phospholipase PLA2G7 Is Complementary to GPX4 in Mitigating Punicic-Acid-Induced Ferroptosis in Prostate Cancer Cells. iScience 2024, 27, 109774. [Google Scholar] [CrossRef]
- Cuvelier, G.; Vermonden, P.; Rousseau, J.; Feron, O.; Rezsohazy, R.; Larondelle, Y. Resistance to CLnA-Induced Ferroptosis Is Acquired in Caco-2 Cells upon Differentiation. Front. Cell Death 2023, 2, 1219672. [Google Scholar] [CrossRef]
- Zhang, T.; Gao, Y.; Mao, Y.; Zhang, Q.; Lin, C.; Lin, P.; Zhang, J.; Wang, X. Growth Inhibition and Apoptotic Effect of Alpha-Eleostearic Acid on Human Breast Cancer Cells. J. Nat. Med. 2012, 66, 77–84. [Google Scholar] [CrossRef]
- Yasui, Y.; Hosokawa, M.; Sahara, T.; Suzuki, R.; Ohgiya, S.; Kohno, H.; Tanaka, T.; Miyashita, K. Bitter Gourd Seed Fatty Acid Rich in 9c,11t,13t-Conjugated Linolenic Acid Induces Apoptosis and up-Regulates the GADD45, P53 and PPARγ in Human Colon Cancer Caco-2 Cells. Prostaglandins, Leukotrienes Essent. Fatty Acids 2005, 73, 113–119. [Google Scholar] [CrossRef] [PubMed]
- Dixon, S.J.; Lemberg, K.M.; Lamprecht, M.R.; Skouta, R.; Zaitsev, E.M.; Gleason, C.E.; Patel, D.N.; Bauer, A.J.; Cantley, A.M.; Yang, W.S.; et al. Ferroptosis: An Iron-Dependent Form of Nonapoptotic Cell Death. Cell 2012, 149, 1060–1072. [Google Scholar] [CrossRef]
- Suzuki, R.; Noguchi, R.; Ota, T.; Abe, M.; Miyashita, K.; Kawada, T. Cytotoxic Effect of Conjugated Trienoic Fatty Acids on Mouse Tumor and Human Monocytic Leukemia Cells. Lipids 2001, 36, 477–482. [Google Scholar] [CrossRef] [PubMed]
- Beatty, A.; Singh, T.; Tyurina, Y.Y.; Tyurin, V.A.; Samovich, S.; Nicolas, E.; Maslar, K.; Zhou, Y.; Cai, K.Q.; Tan, Y.; et al. Ferroptotic Cell Death Triggered by Conjugated Linolenic Acids Is Mediated by ACSL1. Nat. Commun. 2021, 12, 2244. [Google Scholar] [CrossRef]
- Do, Q.; Zhang, R.; Hooper, G.; Xu, L. Differential Contributions of Distinct Free Radical Peroxidation Mechanisms to the Induction of Ferroptosis. JACS Au 2023, 3, 1100–1117. [Google Scholar] [CrossRef]
- Dierge, E.; Larondelle, Y.; Feron, O. Cancer Diets for Cancer Patients: Lessons from Mouse Studies and New Insights from the Study of Fatty Acid Metabolism in Tumors. Biochimie 2020, 178, 56–68. [Google Scholar] [CrossRef] [PubMed]
- Snaebjornsson, M.T.; Janaki-Raman, S.; Schulze, A. Greasing the Wheels of the Cancer Machine: The Role of Lipid Metabolism in Cancer. Cell Metab. 2020, 31, 62–76. [Google Scholar] [CrossRef] [PubMed]
- Lee, J.; Shin, D.; Roh, J.-L. Lipid Metabolism Alterations and Ferroptosis in Cancer: Paving the Way for Solving Cancer Resistance. Eur. J. Pharmacol. 2023, 941, 175497. [Google Scholar] [CrossRef]
- Doll, S.; Proneth, B.; Tyurina, Y.Y.; Panzilius, E.; Kobayashi, S.; Ingold, I.; Irmler, M.; Beckers, J.; Aichler, M.; Walch, A.; et al. ACSL4 Dictates Ferroptosis Sensitivity by Shaping Cellular Lipid Composition. Nat. Chem. Biol. 2017, 13, 91–98. [Google Scholar] [CrossRef]
- Kagan, V.E.; Mao, G.; Qu, F.; Angeli, J.P.F.; Doll, S.; Croix, C.S.; Dar, H.H.; Liu, B.; Tyurin, V.A.; Ritov, V.B.; et al. Oxidized Arachidonic and Adrenic PEs Navigate Cells to Ferroptosis. Nat. Chem. Biol. 2017, 13, 81–90. [Google Scholar] [CrossRef]
- Jiang, X.; Stockwell, B.R.; Conrad, M. Ferroptosis: Mechanisms, Biology and Role in Disease. Nat. Rev. Mol. Cell Biol. 2021, 22, 266–282. [Google Scholar] [CrossRef]
- Friedmann Angeli, J.P.; Schneider, M.; Proneth, B.; Tyurina, Y.Y.; Tyurin, V.A.; Hammond, V.J.; Herbach, N.; Aichler, M.; Walch, A.; Eggenhofer, E.; et al. Inactivation of the Ferroptosis Regulator Gpx4 Triggers Acute Renal Failure in Mice. Nat. Cell Biol. 2014, 16, 1180–1191. [Google Scholar] [CrossRef]
- Hangauer, M.J.; Viswanathan, V.S.; Ryan, M.J.; Bole, D.; Eaton, J.K.; Matov, A.; Galeas, J.; Dhruv, H.D.; Berens, M.E.; Schreiber, S.L.; et al. Drug-Tolerant Persister Cancer Cells Are Vulnerable to GPX4 Inhibition. Nat 2017, 551, 247–250. [Google Scholar] [CrossRef]
- Hou, J.; Jiang, C.; Wen, X.; Li, C.; Xiong, S.; Yue, T.; Long, P.; Shi, J.; Zhang, Z. ACSL4 as a Potential Target and Biomarker for Anticancer: From Molecular Mechanisms to Clinical Therapeutics. Front. Pharmacol. 2022, 13, 949863. [Google Scholar] [CrossRef] [PubMed]
- Beatty, A.; Singh, T.; Tyurina, Y.Y.; Nicolas, E.; Maslar, K.; Zhou, Y.; Cai, K.Q.; Tan, Y.; Doll, S.; Conrad, M.; et al. Conjugated Linolenic Fatty Acids Trigger Ferroptosis in Triple-Negative Breast Cancer 2019.
- Avram, S.; Coricovac, D.-E.; Pavel, I.Z.; Pinzaru, I.; Ghiulai, R.; Baderca, F.; Soica, C.; Muntean, D.; Branisteanu, D.E.; Spandidos, D.A.; et al. Standardization of A375 Human Melanoma Models on Chicken Embryo Chorioallantoic Membrane and Balb/c Nude Mice. Oncol. Rep. 2017, 38, 89–99. [Google Scholar] [CrossRef]
- Campbell, N.R.; Rao, A.; Zhang, M.; Baron, M.; Heilmann, S.; Deforet, M.; Kenny, C.; Ferretti, L.; Huang, T.-H.; Garg, M.; et al. Cell State Diversity Promotes Metastasis through Heterotypic Cluster Formation in Melanoma. Cancer Biology 2020. [Google Scholar]
- Dang, M.; Henderson, R.E.; Garraway, L.A.; Zon, L.I. Long-Term Drug Administration in the Adult Zebrafish Using Oral Gavage for Cancer Preclinical Studies. Dis. Models Mech. 2016, dmm.024166. [Google Scholar] [CrossRef] [PubMed]
- Bligh, E.G.; Dyer, W.J. A Rapid Method of Total Lipid Extraction and Purification. Canadian Journal of Biochemistry and Physiology 1959, 37, 911–917. [Google Scholar]
- Kaluzny, M.A.; Duncan, L.A.; Merritt, M.V.; Epps, D.E. Rapid Separation of Lipid Classes in High Yield and Purity Using Bonded Phase Columns. J. Lipid Res. 1985, 26, 135–140. [Google Scholar] [CrossRef]
- Dierge, E.; Debock, E.; Guilbaud, C.; Corbet, C.; Mignolet, E.; Mignard, L.; Bastien, E.; Dessy, C.; Larondelle, Y.; Feron, O. Peroxidation of N-3 and n-6 Polyunsaturated Fatty Acids in the Acidic Tumor Environment Leads to Ferroptosis-Mediated Anticancer Effects. Cell Metab. 2021, 33, 1701–1715.e5. [Google Scholar] [CrossRef]
- Bajbouj, K.; Shafarin, J.; Hamad, M. High-Dose Deferoxamine Treatment Disrupts Intracellular Iron Homeostasis, Reduces Growth, and Induces Apoptosis in Metastatic and Nonmetastatic Breast Cancer Cell Lines. Technol. Cancer Res. Treat. 2018, 17, 153303381876447. [Google Scholar] [CrossRef]
- Eaton, J.K.; Furst, L.; Ruberto, R.A.; Moosmayer, D.; Hilpmann, A.; Ryan, M.J.; Zimmermann, K.; Cai, L.L.; Niehues, M.; Badock, V.; et al. Selective Covalent Targeting of GPX4 Using Masked Nitrile-Oxide Electrophiles. Nat. Chem. Biol. 2020, 16, 497–506. [Google Scholar] [CrossRef] [PubMed]
- Lopes-Marques, M.; Cunha, I.; Reis-Henriques, M.A.; Santos, M.M.; Castro, L.F.C. Diversity and History of the Long-Chain Acyl-CoA Synthetase (Acsl) Gene Family in Vertebrates. BMC Evol. Biol. 2013, 13, 271. [Google Scholar] [CrossRef] [PubMed]
- Mendieta-Serrano, M.A.; Schnabel, D.; Lomelí, H.; Salas-Vidal, E. Spatial and Temporal Expression of Zebrafish Glutathione Peroxidase 4 a and b Genes during Early Embryo Development. Gene Expression Patterns 2015, 19, 98–107. [Google Scholar] [CrossRef]
- Rochette, L.; Dogon, G.; Rigal, E.; Zeller, M.; Cottin, Y.; Vergely, C. Lipid Peroxidation and Iron Metabolism: Two Corner Stones in the Homeostasis Control of Ferroptosis. Int. J. Mol. Sci. 2022, 24, 449. [Google Scholar] [CrossRef]
- Grignano, E.; Birsen, R.; Chapuis, N.; Bouscary, D. From Iron Chelation to Overload as a Therapeutic Strategy to Induce Ferroptosis in Leukemic Cells. Front. Oncol. 2020, 10, 586530. [Google Scholar] [CrossRef]
- Miotto, G.; Rossetto, M.; Di Paolo, M.L.; Orian, L.; Venerando, R.; Roveri, A.; Vučković, A.-M.; Bosello Travain, V.; Zaccarin, M.; Zennaro, L.; et al. Insight into the Mechanism of Ferroptosis Inhibition by Ferrostatin-1. Redox Biol. 2020, 28, 101328. [Google Scholar] [CrossRef]
- Shinohara, N.; Tsuduki, T.; Ito, J.; Honma, T.; Kijima, R.; Sugawara, S.; Arai, T.; Yamasaki, M.; Ikezaki, A.; Yokoyama, M.; et al. Jacaric Acid, a Linolenic Acid Isomer with a Conjugated Triene System, Has a Strong Antitumor Effect in Vitro and in Vivo. Biochim. Biophys. Acta (BBA) - Mol. Cell Biol. Lipids 2012, 1821, 980–988. [Google Scholar] [CrossRef]
- Tang, Y.; Zhou, J.; Hooi, S.; Jiang, Y.; Lu, G. Fatty Acid Activation in Carcinogenesis and Cancer Development: Essential Roles of Long-chain acyl-CoA Synthetases (Review). Oncol Lett 2018. [Google Scholar] [CrossRef]
- Sheraj, I.; Guray, N.T.; Banerjee, S. A Pan-Cancer Transcriptomic Study Showing Tumor Specific Alterations in Central Metabolism. Sci Rep 2021, 11, 13637. [Google Scholar] [CrossRef] [PubMed]
- Hassannia, B.; Vandenabeele, P.; Vanden Berghe, T. Targeting Ferroptosis to Iron out Cancer. Cancer Cell 2019, 35, 830–849. [Google Scholar] [CrossRef]
- Danielli, M.; Perne, L.; Jarc Jovičić, E.; Petan, T. Lipid Droplets and Polyunsaturated Fatty Acid Trafficking: Balancing Life and Death. Front. Cell Dev. Biol. 2023, 11, 1104725. [Google Scholar] [CrossRef]
- Jarc, E.; Kump, A.; Malavašič, P.; Eichmann, T.O.; Zimmermann, R.; Petan, T. Lipid Droplets Induced by Secreted Phospholipase A2 and Unsaturated Fatty Acids Protect Breast Cancer Cells from Nutrient and Lipotoxic Stress. Biochim. Biophys. Acta (BBA) - Mol. Cell Biol. Lipids 2018, 1863, 247–265. [Google Scholar] [CrossRef]
- Lange, M.; Wölk, M.; Li, V.W.; Doubravsky, C.E.; Hendricks, J.M.; Kato, S.; Otoki, Y.; Styler, B.; Johnson, S.L.; Harris, C.A.; et al. FSP1-Mediated Lipid Droplet Quality Control Prevents Neutral Lipid Peroxidation and Ferroptosis. Nature Cell Biology 2025, 27, 1902–1913. [Google Scholar] [CrossRef]
- Li, F.-J.; Long, H.-Z.; Zhou, Z.-W.; Luo, H.-Y.; Xu, S.-G.; Gao, L.-C. System Xc−/GSH/GPX4 Axis: An Important Antioxidant System for the Ferroptosis in Drug-Resistant Solid Tumor Therapy. Front. Pharmacol. 2022, 13, 910292. [Google Scholar] [CrossRef] [PubMed]
- Doll, S.; Freitas, F.P.; Shah, R.; Aldrovandi, M.; Da Silva, M.C.; Ingold, I.; Goya Grocin, A.; Xavier Da Silva, T.N.; Panzilius, E.; Scheel, C.H.; et al. FSP1 Is a Glutathione-Independent Ferroptosis Suppressor. Nat 2019, 575, 693–698. [Google Scholar] [CrossRef] [PubMed]
- White, R.; Rose, K.; Zon, L. Zebrafish Cancer: The State of the Art and the Path Forward. Nat. Rev. Cancer 2013, 13, 624–636. [Google Scholar] [CrossRef]
- Heilmann, S.; Ratnakumar, K.; Langdon, E.M.; Kansler, E.R.; Kim, I.S.; Campbell, N.R.; Perry, E.B.; McMahon, A.J.; Kaufman, C.K.; Van Rooijen, E.; et al. A Quantitative System for Studying Metastasis Using Transparent Zebrafish. Cancer Res. 2015, 75, 4272–4282. [Google Scholar] [CrossRef] [PubMed]
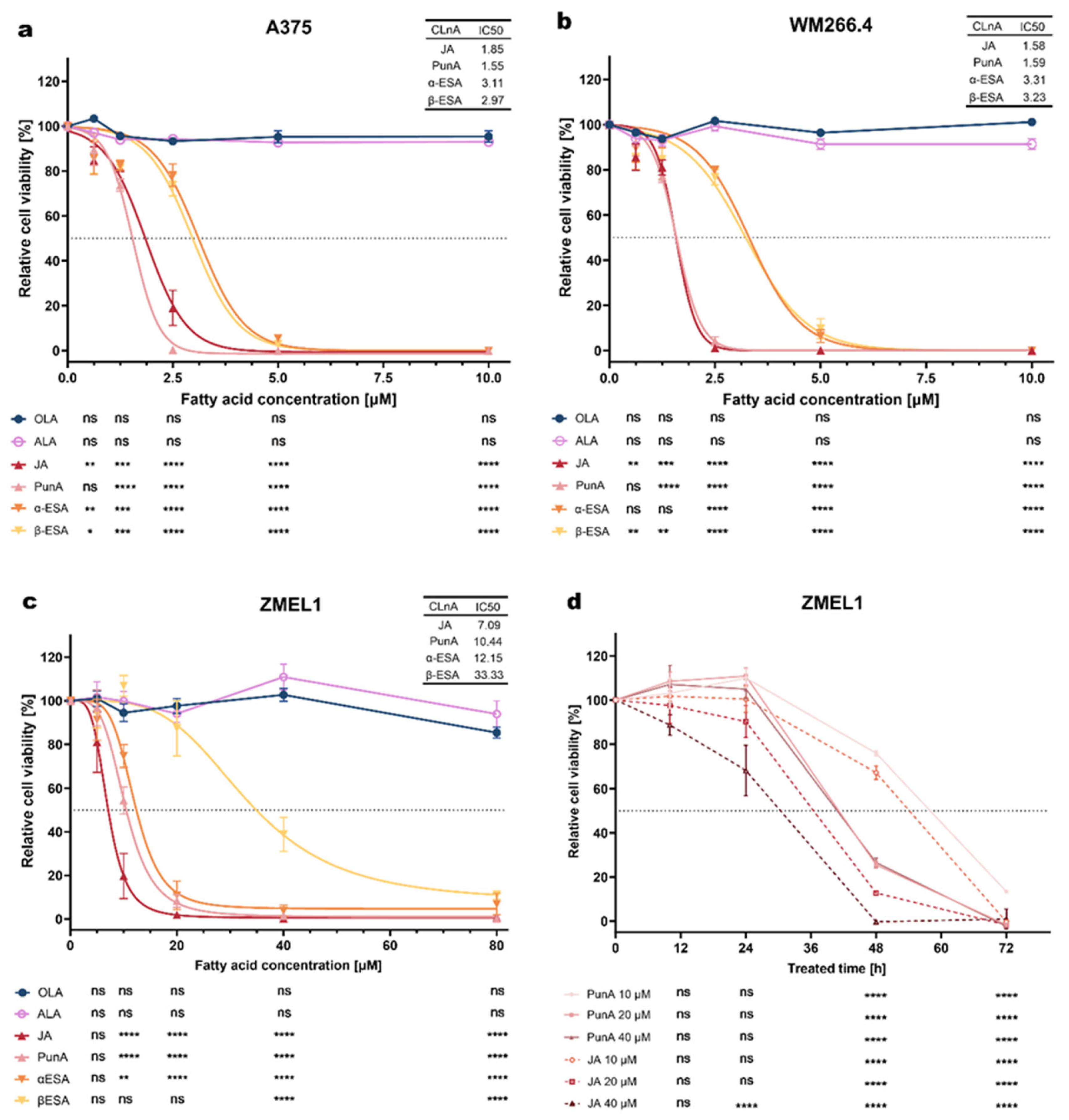
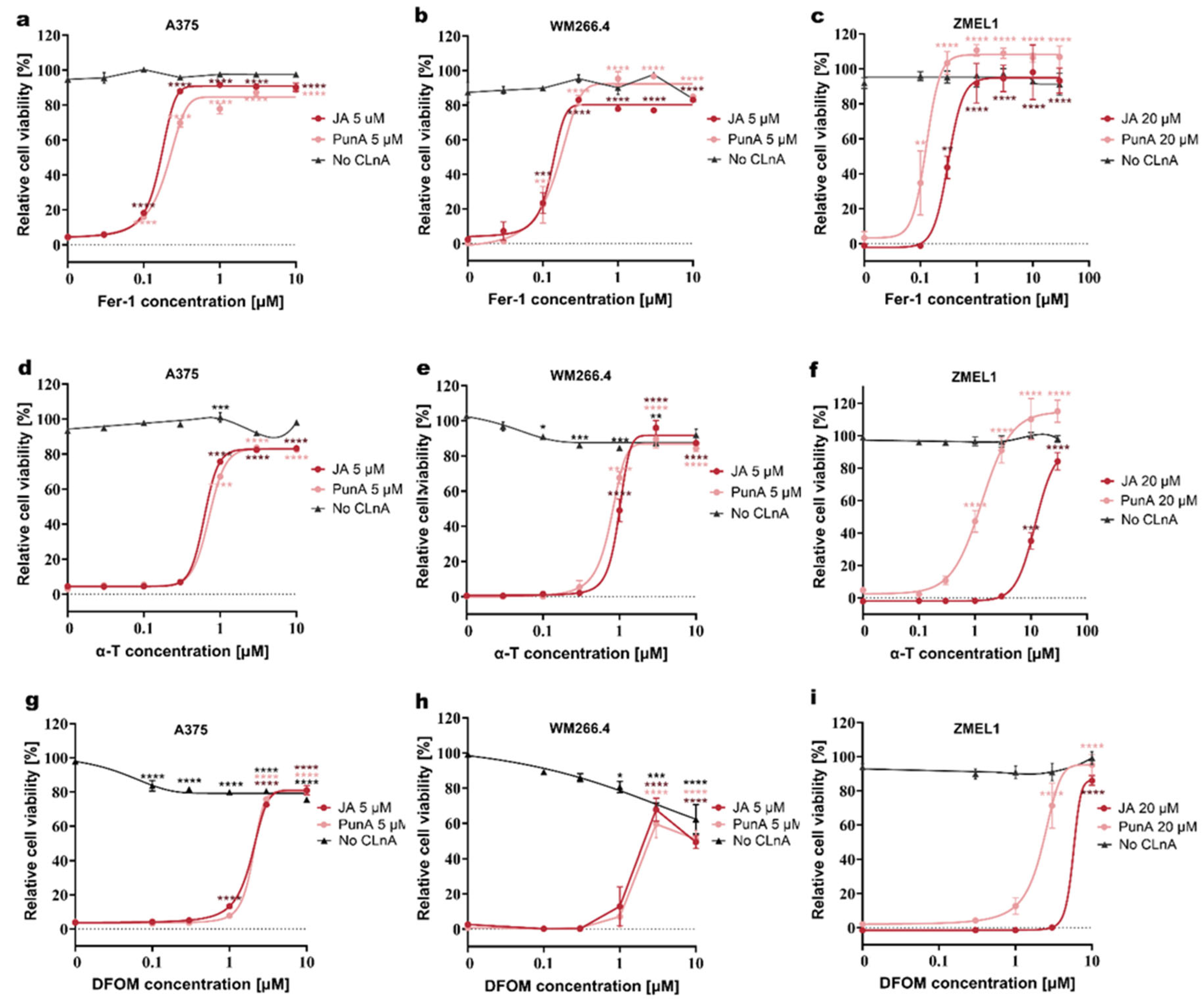
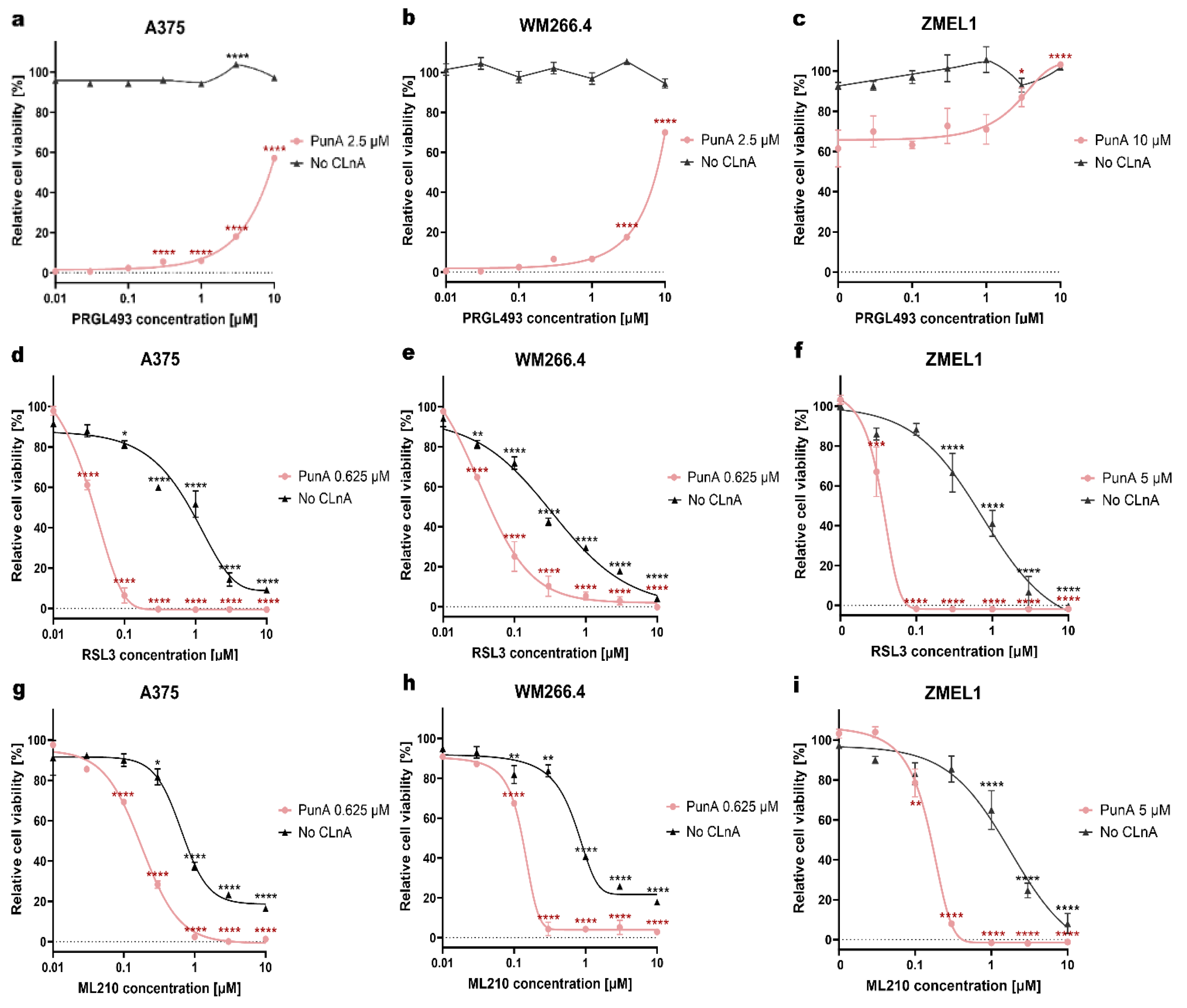
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).



