Submitted:
02 March 2026
Posted:
04 March 2026
Read the latest preprint version here
Abstract
Keywords:
1. Introduction
2. Materials and Methods
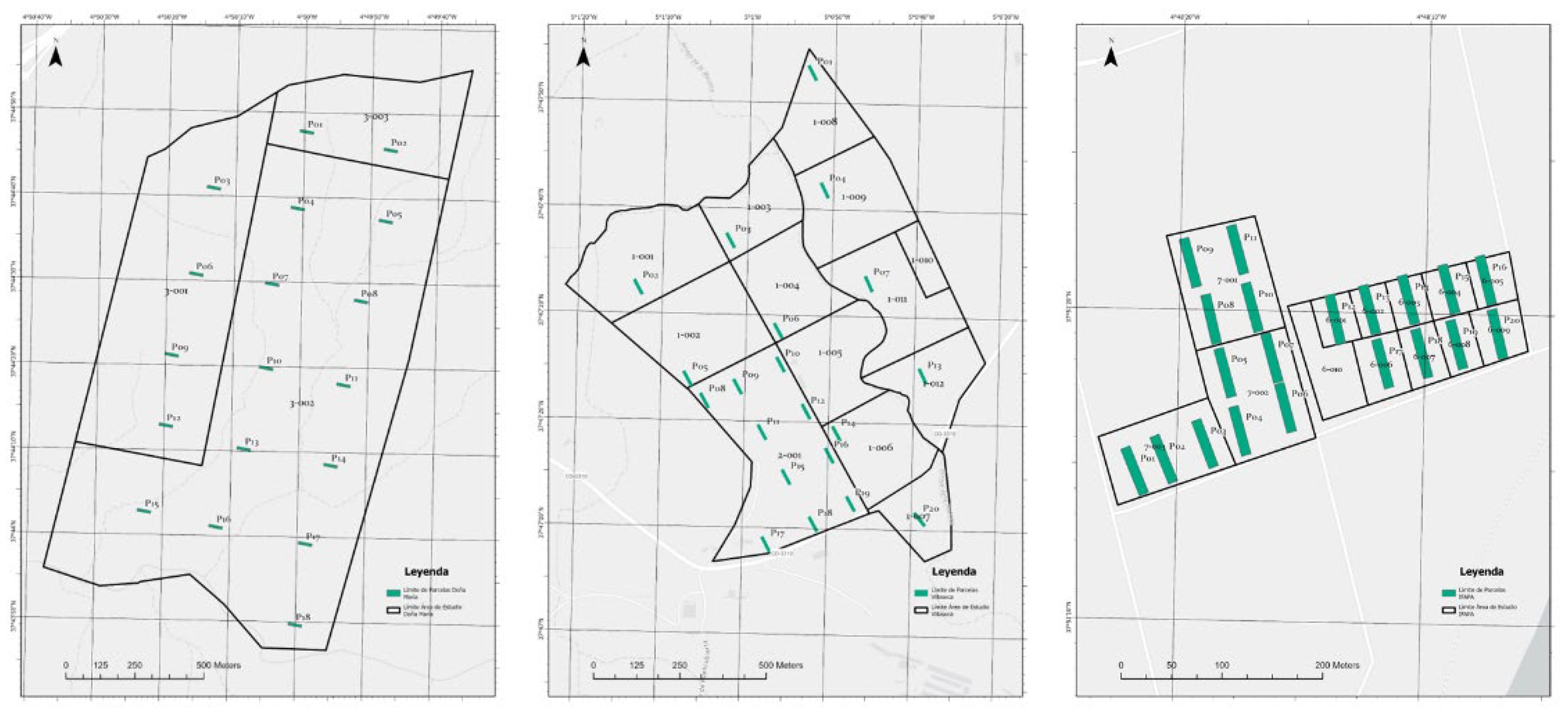
2.1. Study Sites
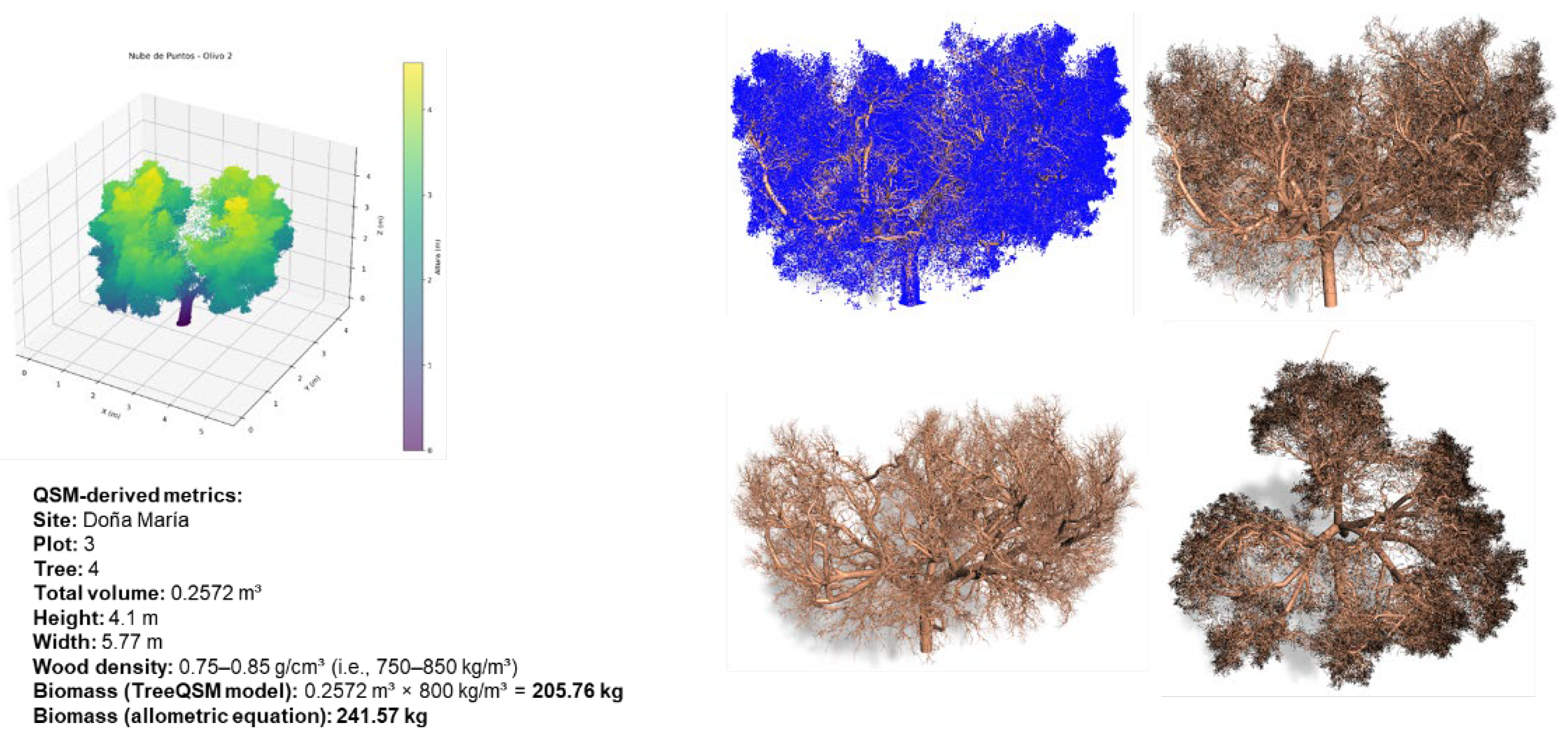
2.2. Field Inventory, Plot Design, and Biomass/Carbon Computation
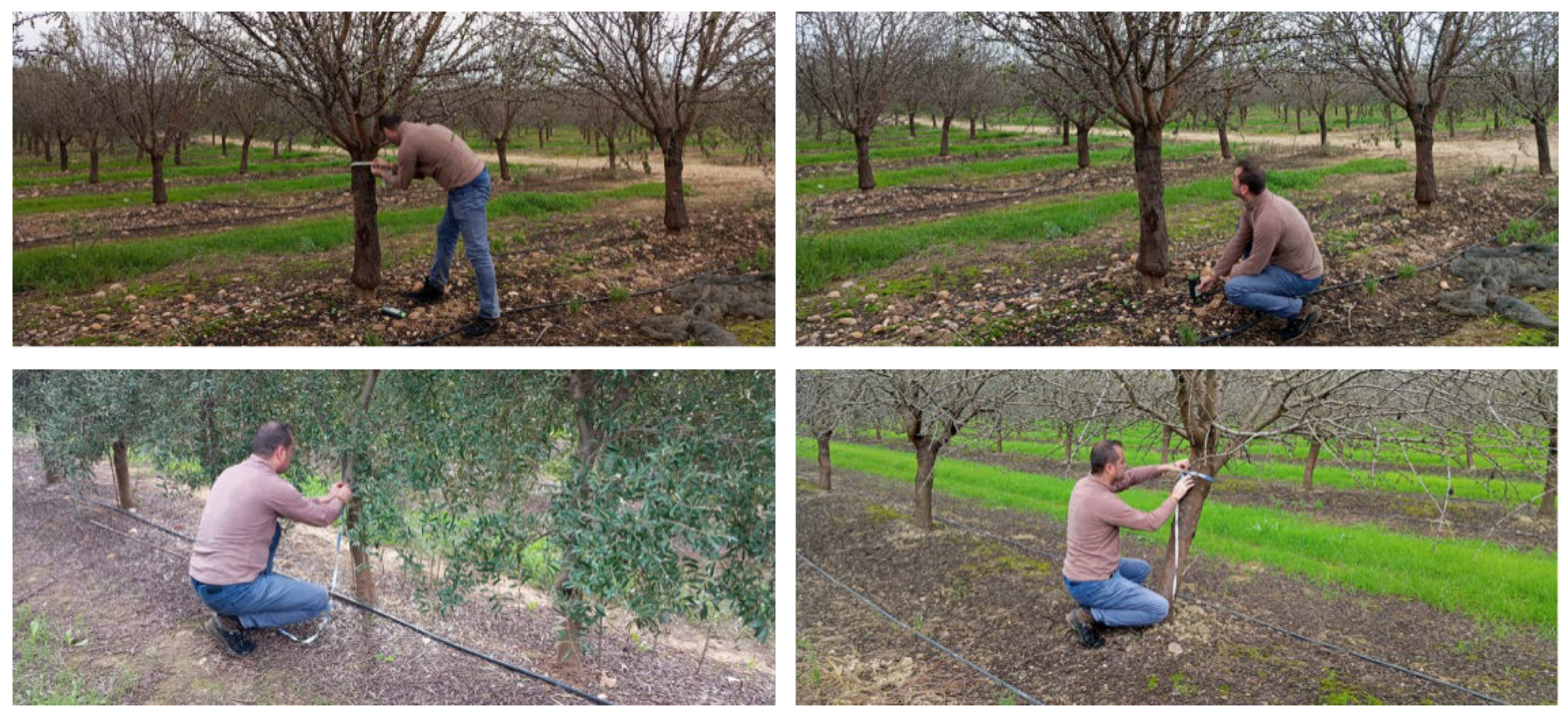
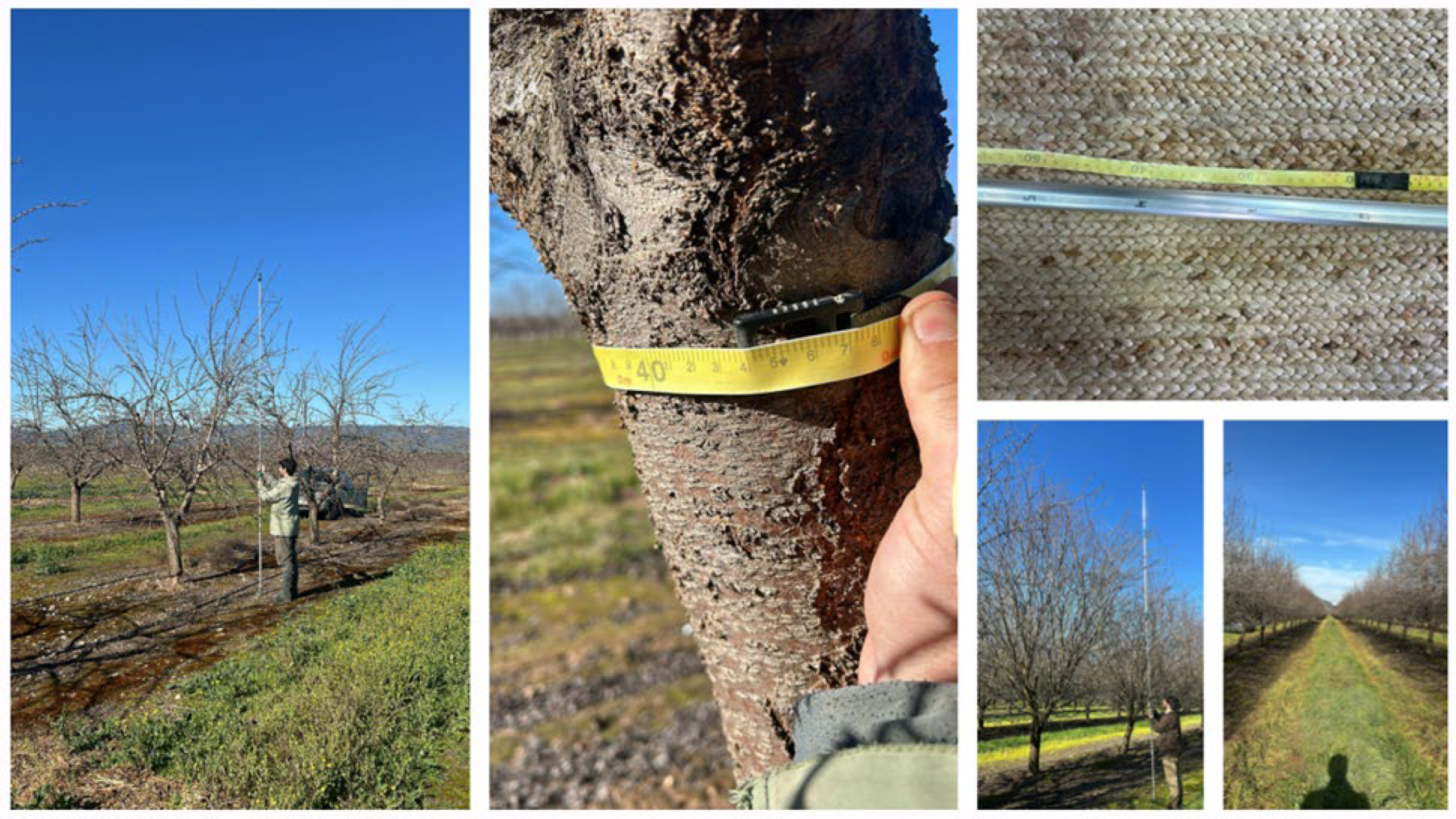


| Site | Statistic | N_trees_plot | D2r_cm | Height_m | AGB_Mg_ha | C_Mg_ha |
|---|---|---|---|---|---|---|
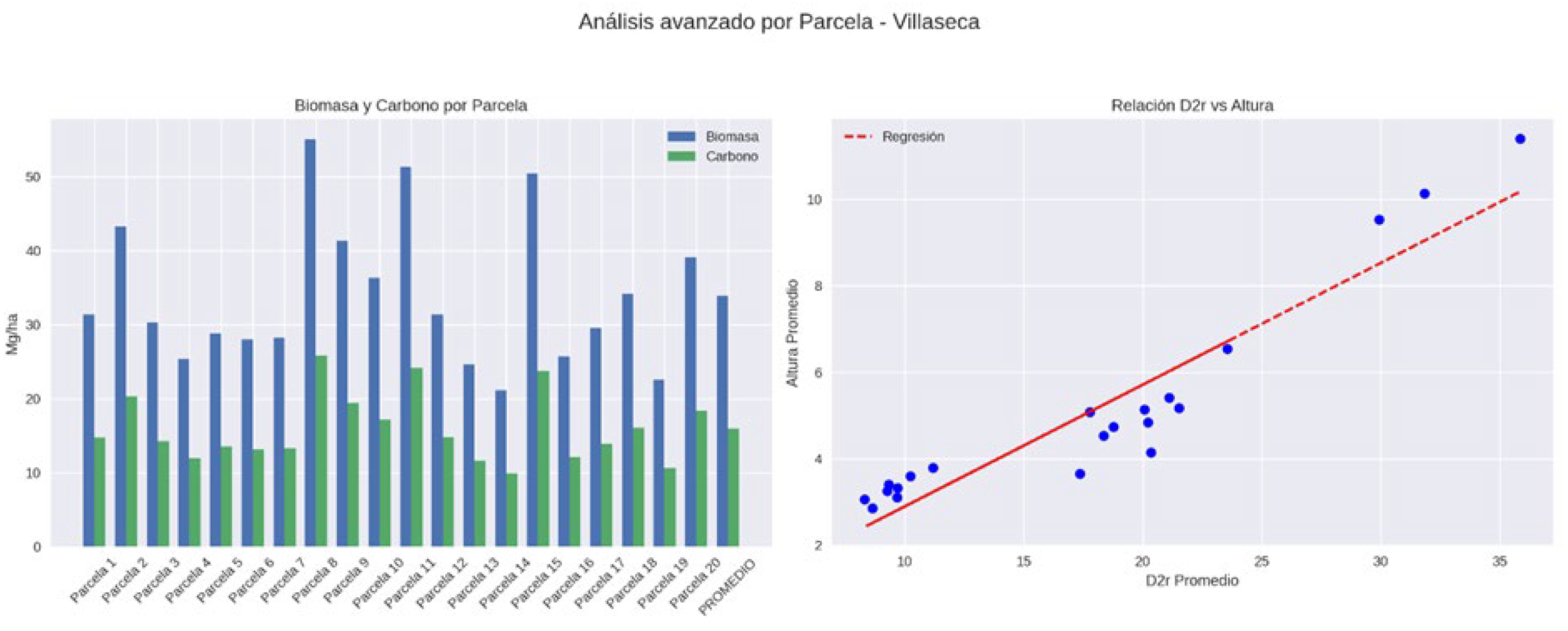
| Villaseca | Media | 55.1 | 17.76 | 5.09 | 33.89 | 15.93 |
| Villaseca | Desv. Est. | 27.48 | 8.06 | 2.42 | 9.64 | 4.53 |
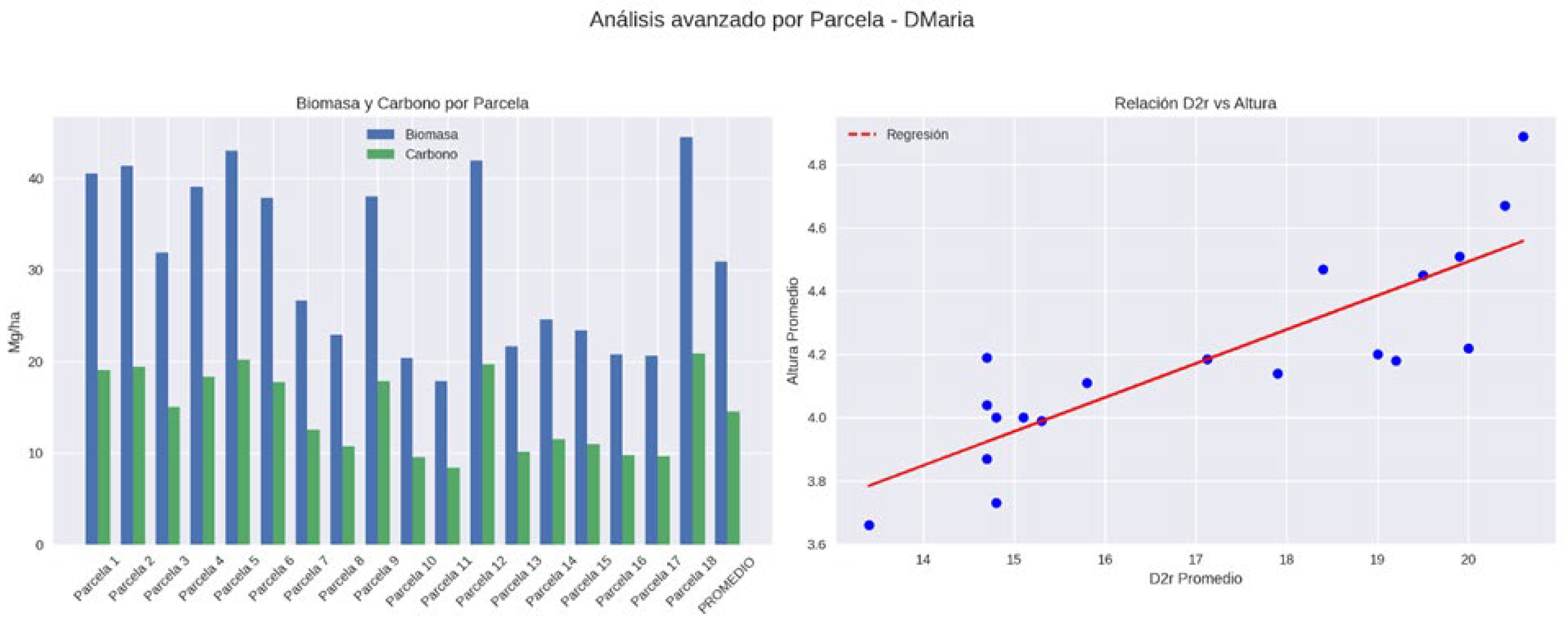
| Doña María | Media | 18.44 | 17.12 | 4.18 | 30.94 | 14.54 |
| Doña María | Desv. Est. | 0.83 | 2.43 | 0.31 | 9.35 | 4.39 |
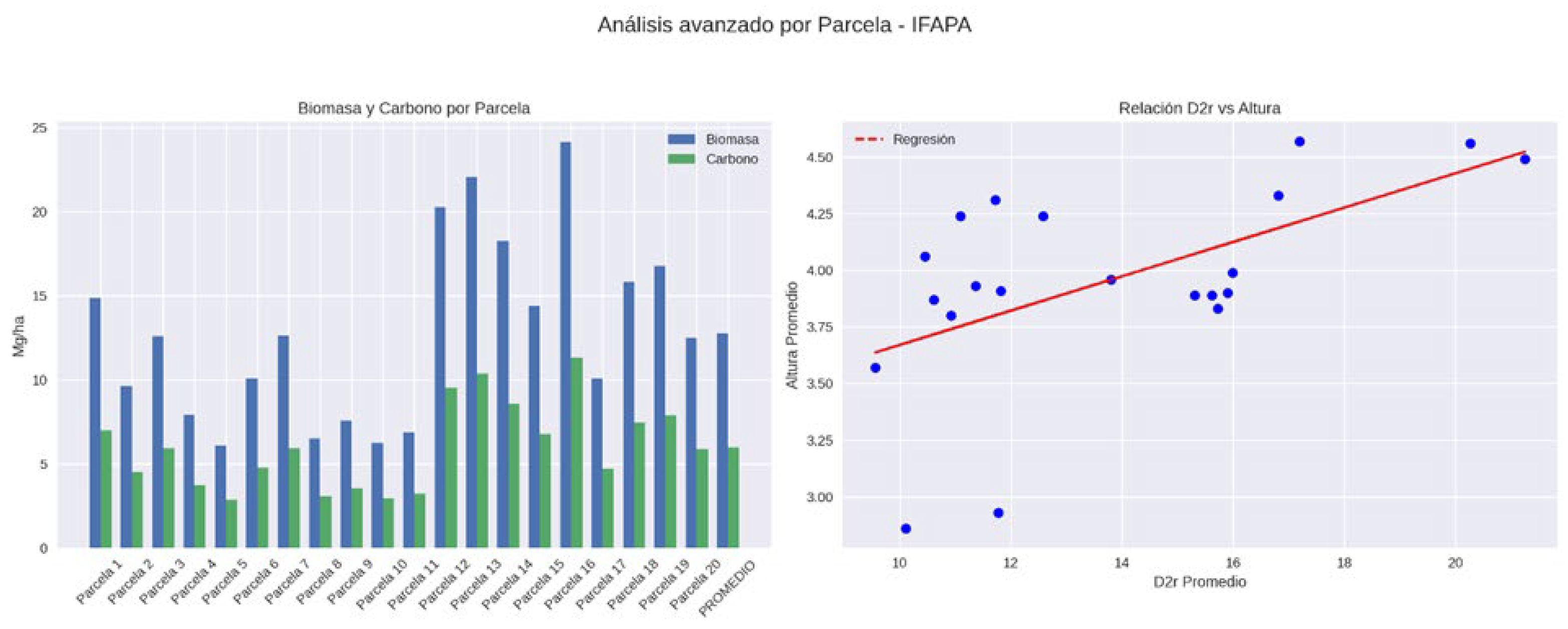
| IFAPA | Media | 21.65 | 13.8 | 3.96 | 12.76 | 6 |
| IFAPA | Desv. Est. | 12.04 | 3.35 | 0.44 | 5.34 | 2.51 |
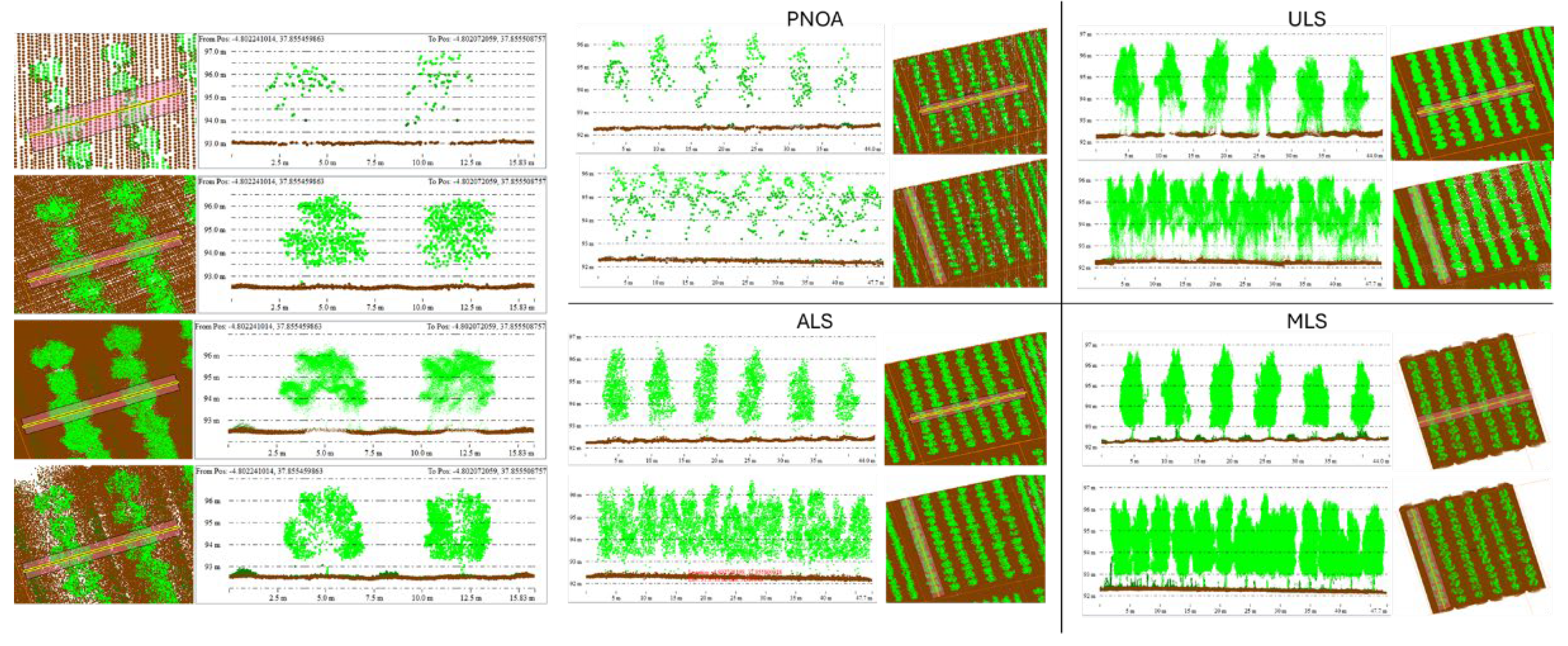
2.3. LiDAR Datasets
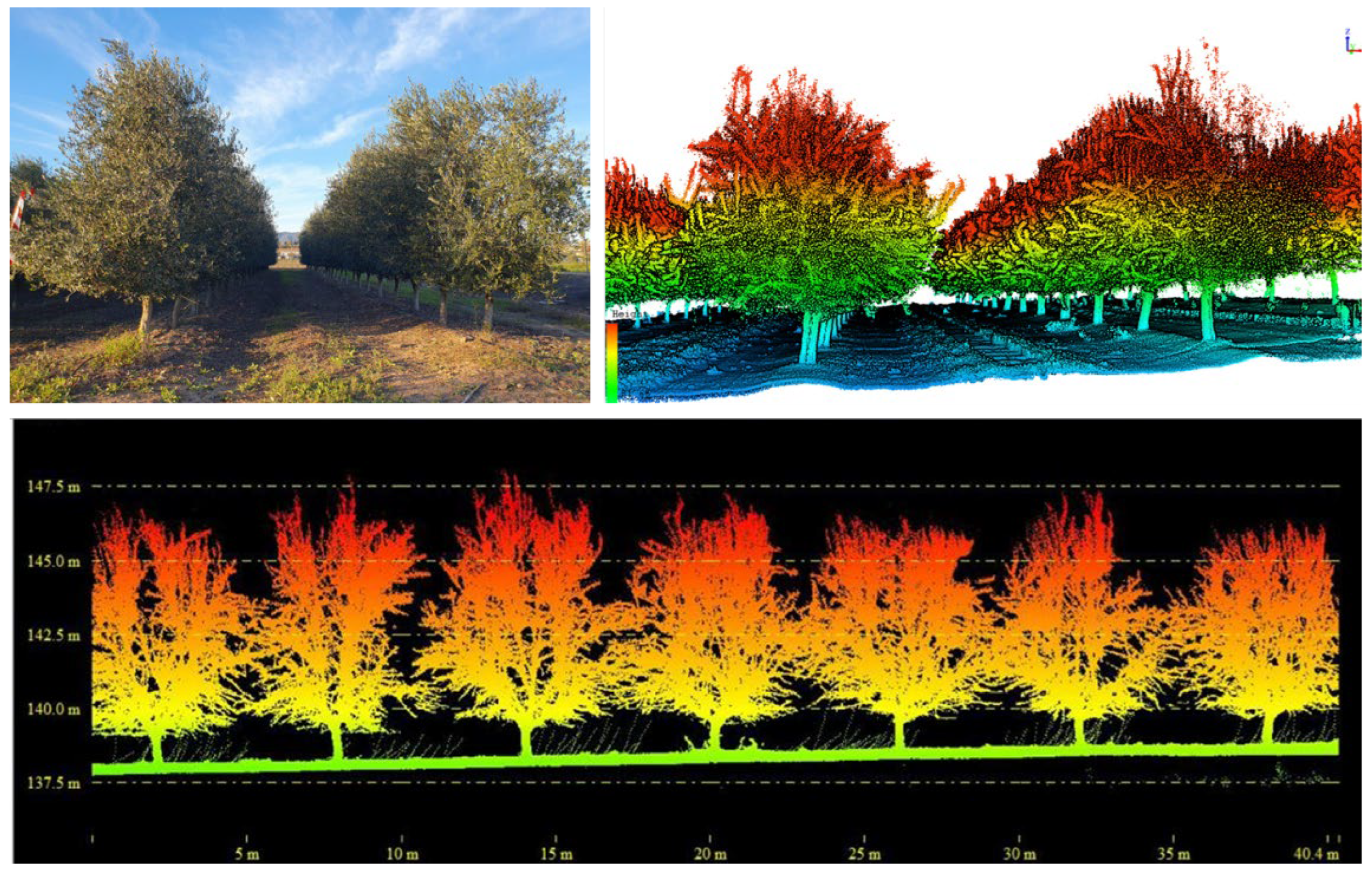
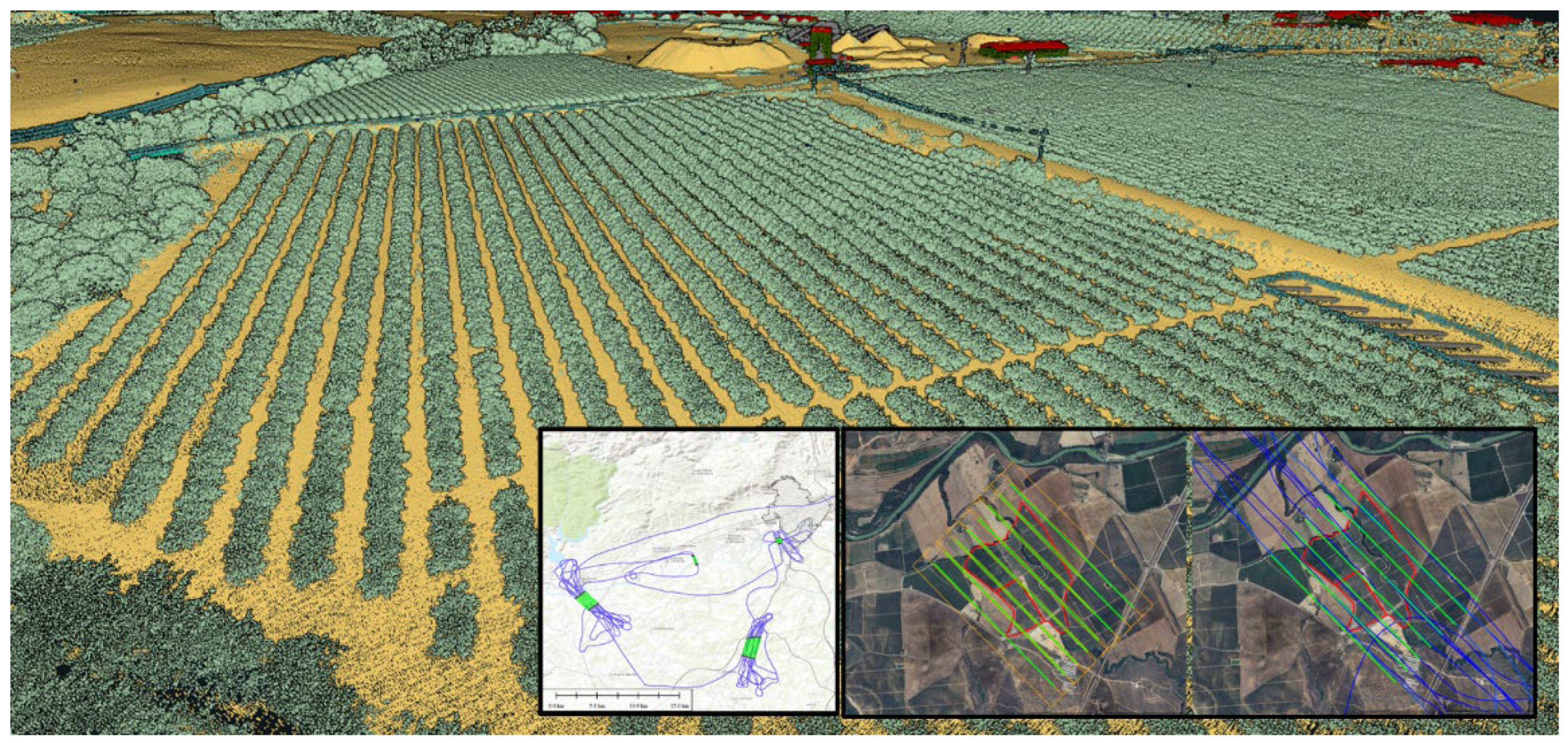
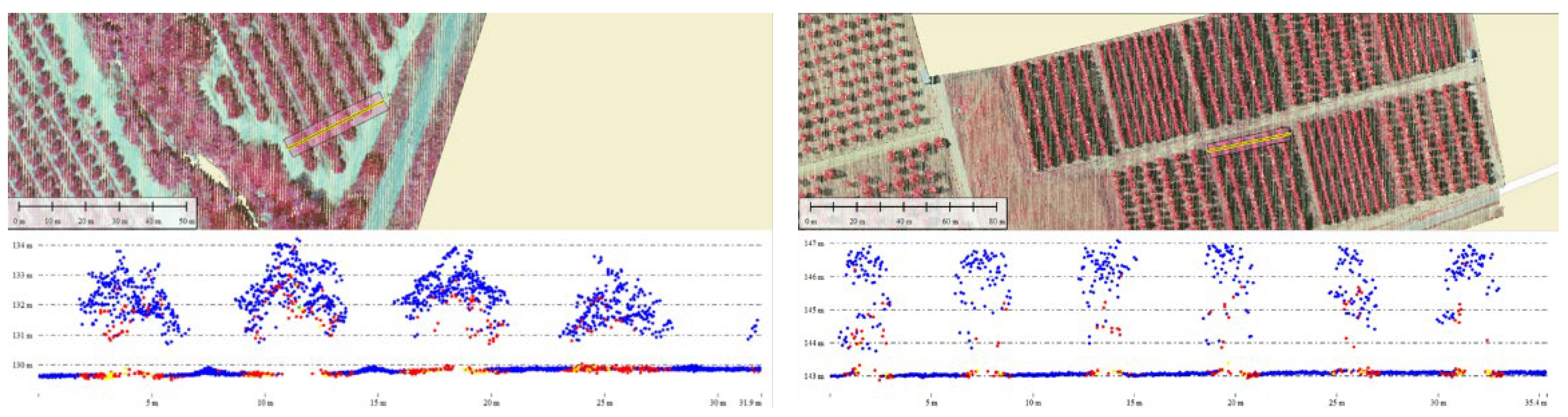
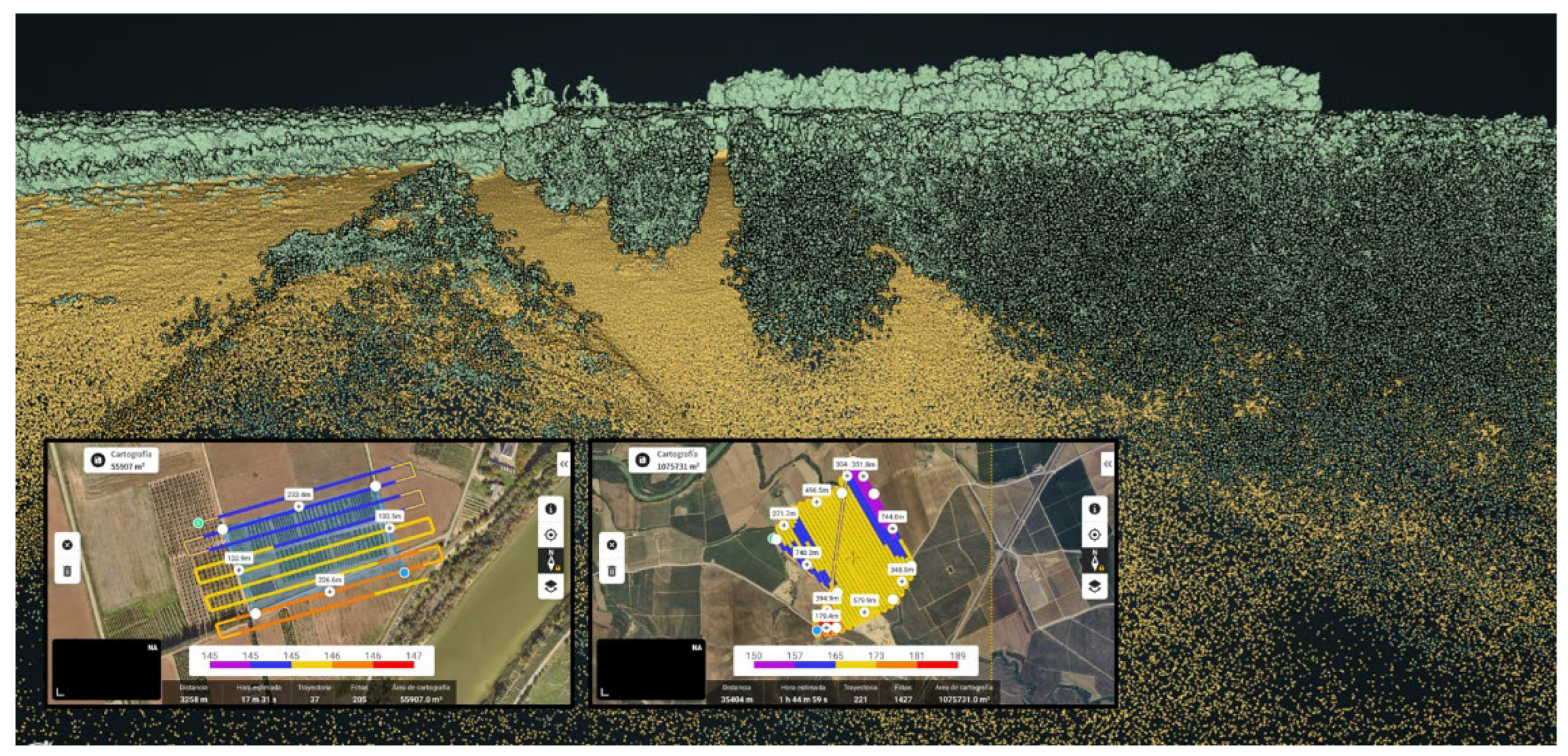


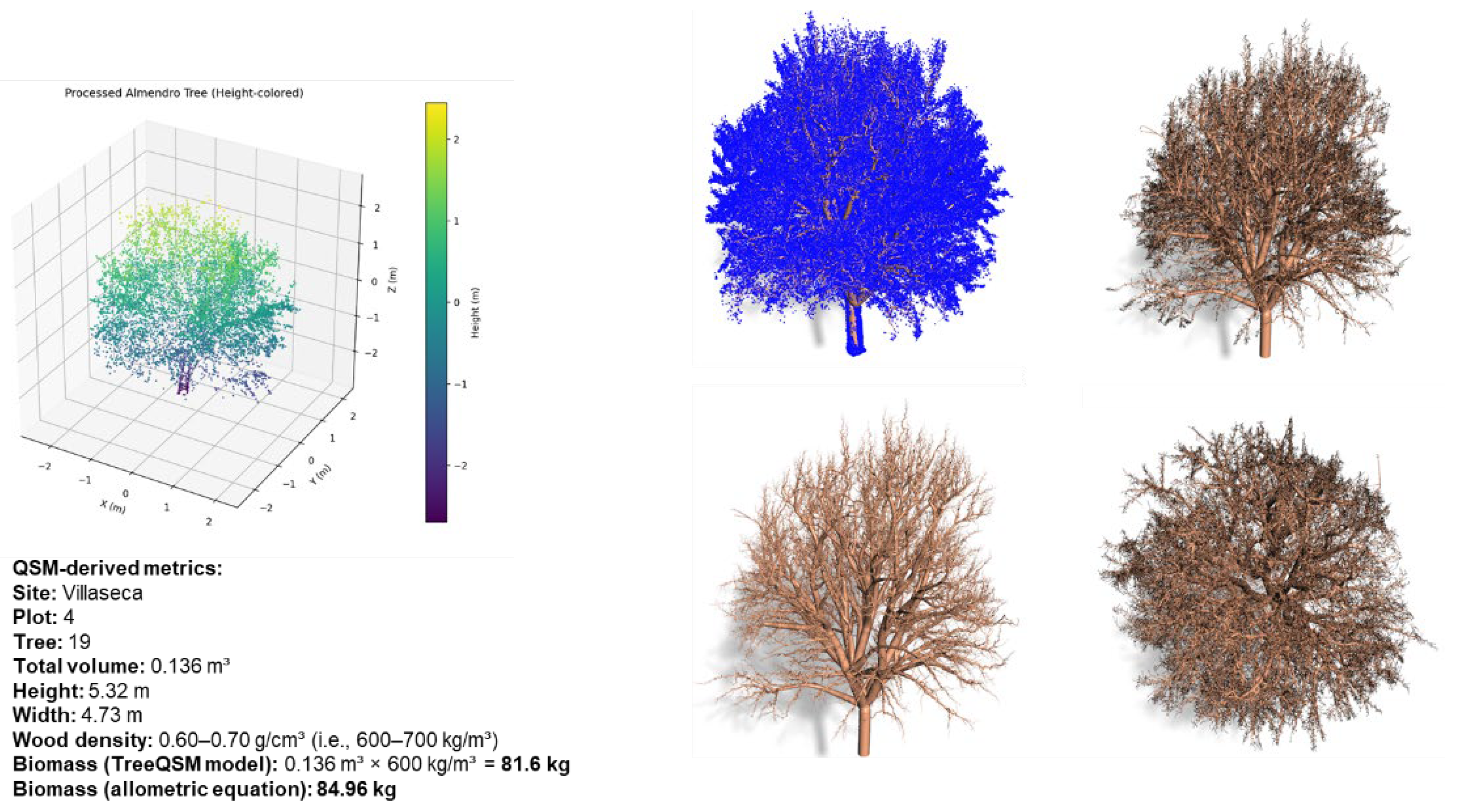
2.4. Pre-Processing and Metric Extraction
2.5. Modeling and Validation
3. Results
3.1. Field Biomass and Carbon Stocks
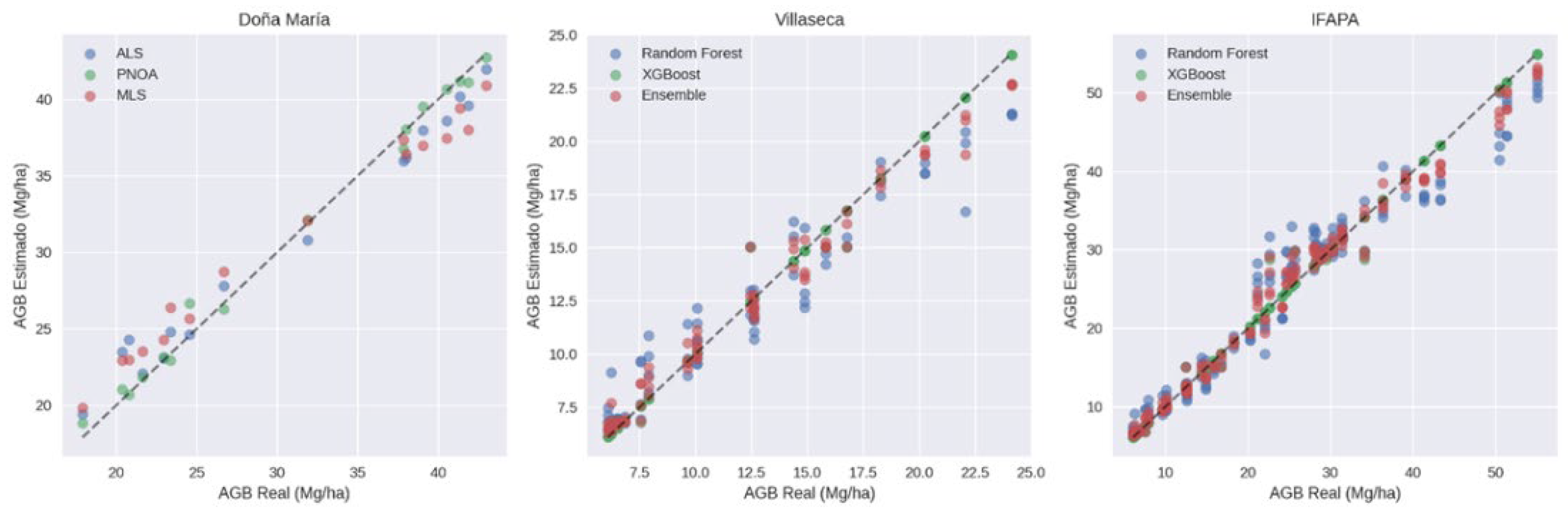
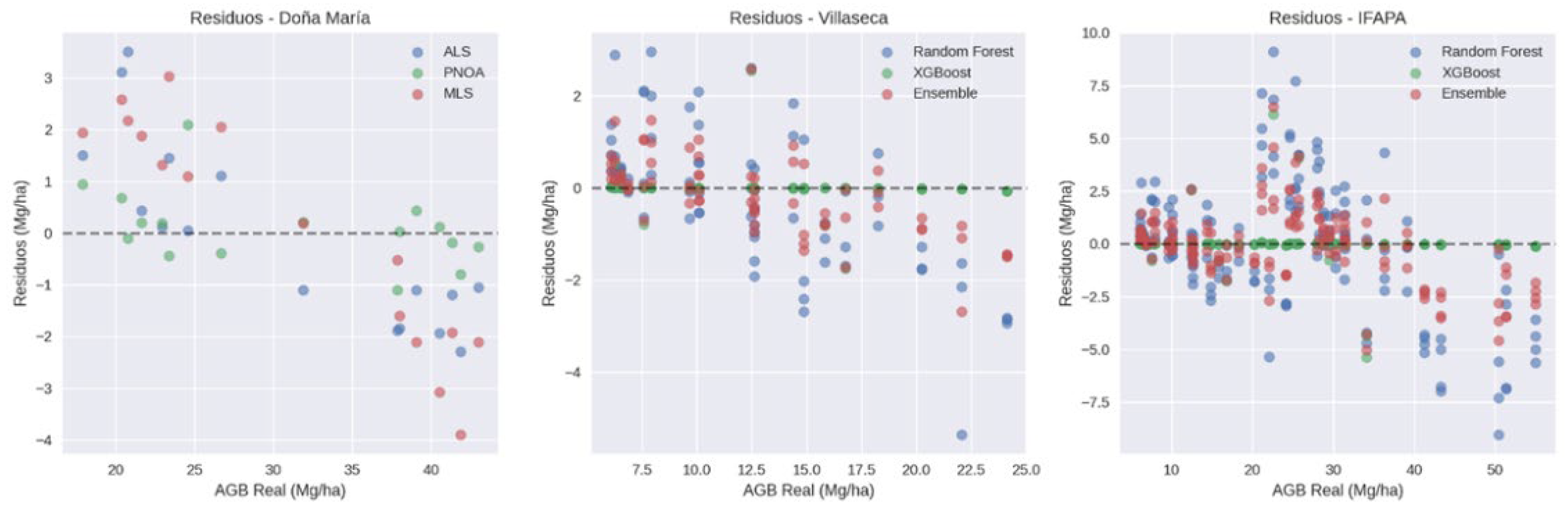
3.2. Model Performance
3.3. Cross-Site Comparison of Results
3.4. Statistical Comparison Across Sites (ANOVA and Kruskal–Wallis)
| Variable | ANOVA: F-statistic (p-value) | Kruskal-Wallis: H-statistic (p-value) |
|---|---|---|
| D2r | 3.35 (0.0421) | 5.34 (0.0694) |
| Tree Height | 3.51 (0.0365) | 2.34 (0.3104) |
| N_Tree | 27.28 (<0.0001) | 28.86 (<0.0001) |
| Biomass | 39.48 (<0.0001) | 37.97 (<0.0001) |
| Carbon | 39.48 (<0.0001) | 37.97 (<0.0001) |
| Site | Group | RMSE_Mg_ha | Rel_RMSE_% | Bias_Mg_ha | Rel_Bias_% | R² | Mean_Rel_Error_% | SD_Rel_Error_% |
|---|---|---|---|---|---|---|---|---|
| Doña María (platform comparison) | ALS | 1.743 | 5.671 | -0.0675 | -0.219 | 0.985 | 1.382 | 6.865 |
| Doña María (platform comparison) | PNOA | 0.725 | 2.359 | 0.1025 | 0.333 | 0.994 | 0.713 | 2.809 |
| Doña María (platform comparison) | MLS | 2.168 | 7.053 | 0.065 | 0.211 | 0.988 | 2.298 | 7.496 |
| IFAPA (model comparison) | Random Forest | 1.542 | 12.559 | -0.182 | -1.484 | 0.932 | 1.859 | 13.150 |
| IFAPA (model comparison) | XGBoost | 0.400 | 3.257 | -0.003 | -0.025 | 0.994 | 0.105 | 3.275 |
| IFAPA (model comparison) | Ensemble | 0.849 | 6.918 | -0.092 | -0.755 | 0.981 | 0.982 | 7.185 |
| Villaseca (model comparison) | Random Forest | 3.060 | 12.853 | -0.010 | -0.042 | 0.950 | 2.547 | 12.797 |
| Villaseca (model comparison) | XGBoost | 0.872 | 3.666 | -0.004 | -0.017 | 0.995 | 0.137 | 3.796 |
| Villaseca (model comparison) | Ensemble | 1.714 | 7.200 | -0.007 | -0.029 | 0.984 | 1.342 | 7.223 |
4. Discussion
4.1. Comparative Performance of LiDAR Modalities
4.2. Implications for AGB Modeling
4.3. Scalability and Hierarchical Integration Strategy
4.4. Limitations and Future Perspectives
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| AGB | Aboveground biomass |
| ALS | Airborne laser scanning |
| CHM | Canopy height model |
| CO2e | Carbon dioxide equivalent |
| DTM | Digital terrain model |
| GEDI | Global Ecosystem Dynamics Investigation |
| GNSS | Global Navigation Satellite System |
| IFAPA | Instituto de Investigación y Formación Agraria y Pesquera |
| IMU | Inertial measurement unit |
| MLS | Mobile laser scanning |
| MRV | Measurement–reporting–verification |
| PNOA | National Aerial Orthophotography Plan (Spain) |
| ULS | Unmanned laser scanning |
Appendix A
Appendix A.1. Plot Centroid Coordinates
| Site | Plot | Easting | Northing |
|---|---|---|---|
| Doña María | 1 | 338442.891 | 4177491.422 |
| Doña María | 2 | 338479.943 | 4177785.198 |
| Doña María | 3 | 338572.069 | 4178069.313 |
| Doña María | 4 | 338619.368 | 4178361.078 |
| Doña María | 5 | 338682.799 | 4178665.661 |
| Doña María | 6 | 338773.056 | 4178955.806 |
| Doña María | 7 | 338791.065 | 4179213.691 |
| Doña María | 8 | 338360.591 | 4178729.133 |
| Doña María | 9 | 338453.508 | 4179003.026 |
| Doña María | 10 | 338486.634 | 4179280.926 |
| Doña María | 11 | 338154.990 | 4177847.548 |
| Doña María | 12 | 338257.349 | 4178129.910 |
| Doña María | 13 | 338338.505 | 4178423.586 |
| Doña María | 14 | 337894.781 | 4177905.352 |
| Doña María | 15 | 337974.339 | 4178216.396 |
| Doña María | 16 | 338084.834 | 4178765.124 |
| Doña María | 17 | 338149.355 | 4179078.420 |
| Doña María | 18 | 337995.796 | 4178472.399 |
| Villaseca | 1 | 322488.703 | 4183949.082 |
| Villaseca | 2 | 322625.775 | 4184007.307 |
| Villaseca | 3 | 322734.451 | 4184066.151 |
| Villaseca | 4 | 322547.632 | 4184144.189 |
| Villaseca | 5 | 322673.036 | 4184206.089 |
| Villaseca | 6 | 322477.923 | 4184274.370 |
| Villaseca | 7 | 322407.454 | 4184406.609 |
| Villaseca | 8 | 322311.322 | 4184366.350 |
| Villaseca | 9 | 322605.783 | 4184333.879 |
| Villaseca | 10 | 322531.865 | 4184470.533 |
| Villaseca | 11 | 322525.107 | 4184568.767 |
| Villaseca | 12 | 322119.568 | 4184696.117 |
| Villaseca | 13 | 322787.640 | 4184703.639 |
| Villaseca | 14 | 322660.101 | 4184975.439 |
| Villaseca | 15 | 322943.758 | 4184435.829 |
| Villaseca | 16 | 322934.346 | 4184020.871 |
| Villaseca | 17 | 322693.976 | 4184269.419 |
| Villaseca | 18 | 322262.459 | 4184430.594 |
| Villaseca | 19 | 322625.344 | 4185315.724 |
| Villaseca | 20 | 322387.154 | 4184830.815 |
| IFAPA | 1 | 341170.695 | 4191371.022 |
| IFAPA | 2 | 341232.111 | 4191326.678 |
| IFAPA | 3 | 341205.203 | 4191261.724 |
| IFAPA | 4 | 341217.546 | 4191383.992 |
| IFAPA | 5 | 341191.305 | 4191315.048 |
| IFAPA | 6 | 341251.737 | 4191276.441 |
| IFAPA | 7 | 341314.457 | 4191314.919 |
| IFAPA | 8 | 341348.624 | 4191324.491 |
| IFAPA | 9 | 341387.266 | 4191334.331 |
| IFAPA | 10 | 341427.348 | 4191344.763 |
| IFAPA | 11 | 341464.099 | 4191353.589 |
| IFAPA | 12 | 341475.656 | 4191300.190 |
| IFAPA | 13 | 341435.214 | 4191289.675 |
| IFAPA | 14 | 341400.434 | 4191280.931 |
| IFAPA | 15 | 341361.690 | 4191270.790 |
| IFAPA | 16 | 341219.843 | 4191204.017 |
| IFAPA | 17 | 341265.128 | 4191226.961 |
| IFAPA | 18 | 341185.157 | 4191191.251 |
| IFAPA | 19 | 341144.323 | 4191175.878 |
| IFAPA | 20 | 341115.363 | 4191164.295 |
References
- Asner, G.P.; Mascaro, J. Mapping tropical forest carbon: Calibrating plot estimates to a simple LiDAR metric. Remote Sens. Environ. 2014, 140, 614–624. [CrossRef]
- Silva, C.A.; Hudak, A.T.; Vierling, L.A.; Klauberg, C.; Garcia, M.; Ferraz, A.; Keller, M.; Eitel, J.U.H.Saatchi, S. Impacts of airborne lidar pulse density on estimating biomass stocks and changes in a selectively logged tropical forest. Remote Sens. 2017, 9, 1068. [CrossRef]
- Estornell, J.; Velázquez-Martí, B.; López-Cortés, I.; Salazar, D.; Fernández-Sarría, A. Estimation of wood volume and height of olive tree plantations using airborne discrete-return LiDAR data. GISci. Remote Sens. 2014, 51, 17–29. [CrossRef]
- Velázquez-Martí, B.; López-Cortés, I.; Salazar-Hernández, D.M. Dendrometric analysis of olive trees for wood biomass quantification in Mediterranean orchards. Agrofor. Syst. 2014, 88, 755–765. [CrossRef]
- López-Bellido, P.J.; López-Bellido, R.J.; García-Morillo, J.; Fernández-García, P. Assessment of carbon sequestration and the carbon footprint in olive groves in Southern Spain. Int. J. Sustain. Dev. World Ecol. 2016, 23, 1–13. [CrossRef]
- Dubayah, R.; et al. The Global Ecosystem Dynamics Investigation: High-resolution laser ranging of the Earth’s forests and topography. Remote Sens. Environ. 2020, 240, 111779. [CrossRef]
- Duncanson, L.; et al. Aboveground biomass density models for NASA’s GEDI mission. Remote Sens. Environ. 2022, 270, 112845. [CrossRef]
- Khosravipour, A.; Skidmore, A.K.; Isenburg, M.; Wang, T.; Hussin, Y.A. Generating pit-free canopy height models from airborne lidar. Photogramm. Eng. Remote Sens. 2014, 80, 863–872. [CrossRef]
- Kukko, A.; Kaartinen, H.; Hyyppä, J.; Chen, Y. Multiplatform mobile laser scanning: Usability and performance. Sensors 2012, 12, 11712–11733. [CrossRef]
- Breiman, L. Random forests. Mach. Learn. 2001, 45, 5–32. [CrossRef]
- Chen, T.; Guestrin, C. XGBoost: A scalable tree boosting system. In Proceedings of the 22nd ACM SIGKDD International Conference on Knowledge Discovery and Data Mining, San Francisco, CA, USA, 13–17 August 2016; pp. 785–794. [CrossRef]
- IPCC. 2006 IPCC Guidelines for National Greenhouse Gas Inventories; Intergovernmental Panel on Climate Change: Geneva, Switzerland, 2006.
- Kebede, B.; Soromessa, T. Allometric equations for aboveground biomass estimation of Olea europaea L. subsp. cuspidata in Mana Angetu Forest. Ecosyst. Health Sustain. 2018, 4, 1–12. [CrossRef]
- Henry, M.; Bombelli, A.; Trotta, C.; Alessandrini, A.; Birigazzi, L.; Sola, G.; Vieilledent, G.; Santenoise, P.; Longuetaud, F.; Valentini, R.; Picard, N.; Saint-André, L. GlobAllomeTree: International platform for tree allometric equations to support volume, biomass and carbon assessment. iForest 2013, 6, 326–330. [CrossRef]
- Frank, B.; Mauro, F.; Allensworth, E. allometric: Structured Allometric Models for Trees. R package version 3.0.0, 2024. Available online: https://allometric.org/ (accessed on 26 February 2026).
- Proietti, S.; et al. Carbon footprint of an olive tree grove. Appl. Energy 2014, 127, 115–124. [CrossRef]
- Roussel, J.-R.; et al. lidR: An R package for analysis of airborne laser scanning (ALS) data. Remote Sens. Environ. 2020, 251, 112061. [CrossRef]
- LiDAR, Especificaciones técnicas. Plan Nacional de Ortofotografía Aérea (PNOA), Instituto Geográfico Nacional (IGN). Available online: https://pnoa.ign.es/pnoa-lidar/especificaciones-tecnicas (accessed on 26 February 2026).
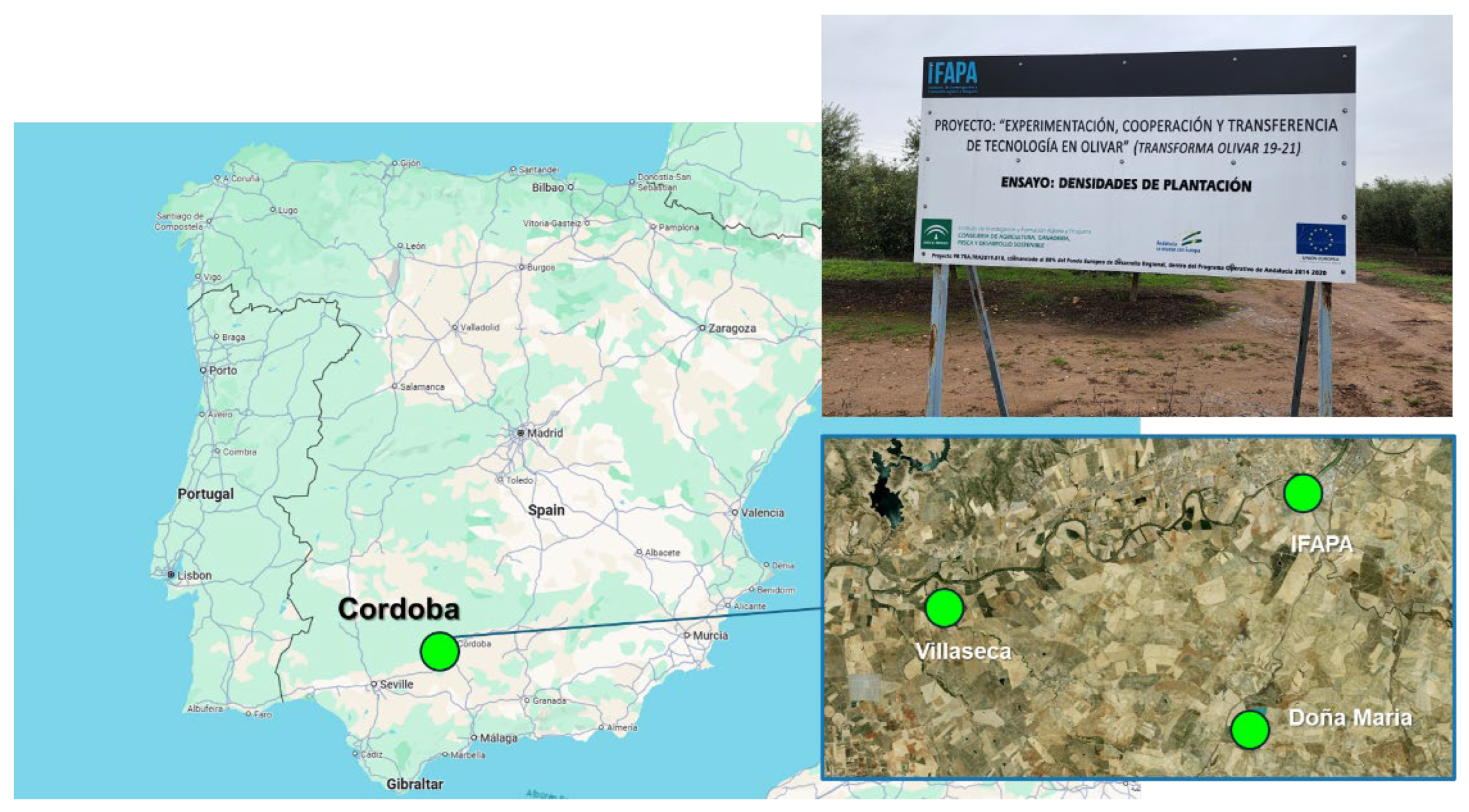

| System | Point spacing (m) | Density (pts/m²) | Intensity (bits) | Color | Multiple returns | Scan angle (deg) |
Flight/ Survey date |
EPSG | Orthometric heights | Stated Accuracy XY/Z |
|---|---|---|---|---|---|---|---|---|---|---|
| PNOA ALS | 0.447 | 5.0 | 16 | RGBI | Yes (4) | -19.62 – 3.17° | 27/09/2024 | 25830 | EGM2008-REDNAP | 0.30 / 0.10 m |
| Riegl ALS | 0.146 | 46.8 | 8 | RGB | Yes (7) | -30 – 30° | 21/11/2024 | 32630 | EVRS | 0.06 / 0.06 m |
| ULS | 0.02 | 2470.0 | 8 | RGB | No | -35 – 35° | 18/12/2024 | 32630 | EVRS | 0.10 / 0.05 m |
| MLS | 0.005 | 36545.0 | 8 | N/A | No | -90 – 65° | 10/12/2024 | 32630 | EVRS | 0.02 / 0.02 m |
| PNOA/ALS (national) | Riegl ALS | ULS | MLS | |
|---|---|---|---|---|
| Sistema LiDAR | Optech T2000 | Riegl LMS-Q680i | DJI Zenmuse L2 | LiDAR USA Velodyne hdl-32e |
| Sistema GNSS/IMU | POS AV™ AP60 (OEM) | IGI IMU LIE | DJI 200 Hz | Snoopy INS (OEM) |
| Sw Plan/Naveg./Post | Optech FMS y LMS (LiDAR Mapping Suite) | IGI Plan v1.5.5 /Aerocontrol/ Inertial Explorer 8.9 y AeroOffice 5.58 | DJI Terra | Scan look PC LiDAR 360 Inertial explorer |
| Hv (m) | 2.200 | 420 | 80 | 2 |
| FOV | 50 | 60 | 70 | 40ºV – 360ºHZ |
| Frec.- Escaneo (Hz) | 400 | |||
| Frec. Pulso (Kz) | 700-1.100 | 266 | 240 | 1.200 |
| Velocidad | 160 Knot | 85 Knot | 6 m/s | 1 m/s |
| Densidad nominal (ps/m2) | 5 | 11 | 300 | 33.000 |
| FWF | Sí, no disponible | Sí | No | No |
| Retornos | 4 | 7 | 3 | 1 |
| Intensidad | 16 | 8 | 8 | 8 |
| Color | RGBI | ND | RGB | ND |
| Rec. Transversal | >15% | >90% | >90% | N/A |
| EERR | IGN/RAP | IGN/RAP (CRBD) | IGN/RAP | IGN/RAP |
| Precisión (XY/Z m) | 0,30/0,10 | 0,06/0,06 | 0,10/0,05 | 0,02/0,02 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




