Submitted:
22 March 2026
Posted:
26 March 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
Scope and Limitations
2. Methods
2.1. AI-Assisted Workflow
2.2. Custom SMILES Descriptor Calculator
2.3. Training Sets and Ensemble ML Models
2.4. Bioisosteric Modifications
2.5. External In Silico Validation
2.6. Iterative Lead Optimization
2.7. Property-Space MIC Estimation
3. Results
3.1. Model Performance and Screening
3.2. External Validation of the Initial Lead
3.3. Iterative Lead Optimization (Table 1)
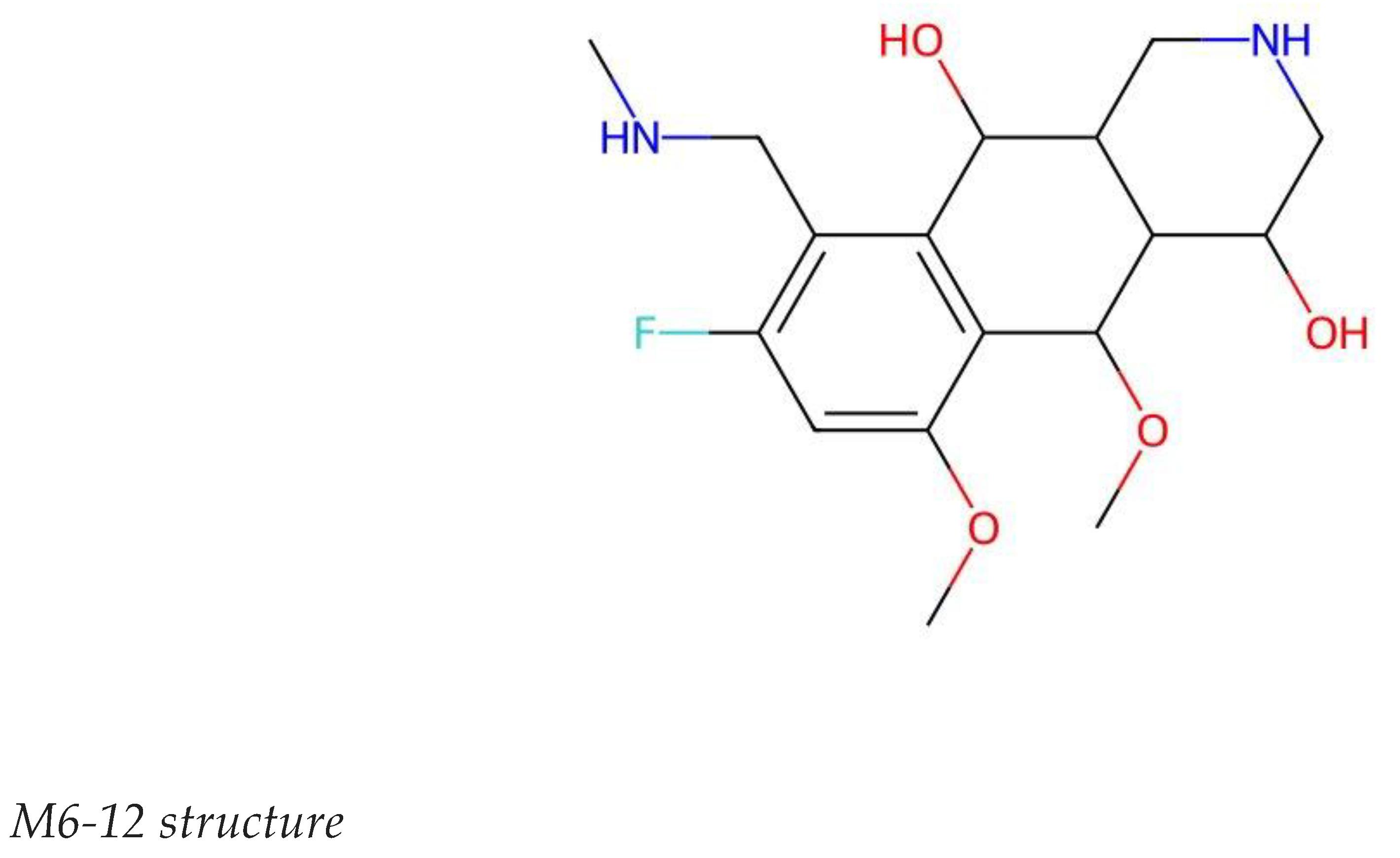
3.4. Comprehensive Profile of M6-12
3.4.1. ML Scoring
3.4.2. SwissADME
3.4.3. RDKit Cross-Validation (Table 2)
3.4.4. pkCSM ADMET
3.4.5. ChEMBL
3.5. Property-Space MIC Estimation
4. Discussion
4.1. The “Signaling First, Killing Later” Paradigm
4.2. Comparison with Existing AI-Driven Antibiotic Discovery Approaches
4.3. Contextualizing the pkCSM Hepatotoxicity Prediction
4.4. Scaffold Simplification and the Piperidine Pharmacophore
4.5. Why M6-12 Passes Both ML Models: A Structural Analysis
4.6. Cross-Platform Concordance
4.7. Interpretive Limitations of the Antibiotic-Likeness Model
5. Limitations
6. Conclusions
Supplementary Materials
Author Contributions
Funding
Declaration of generative AI and AI-assisted technologies in the research process
Data Availability Statement
Conflicts of Interest
Abbreviations
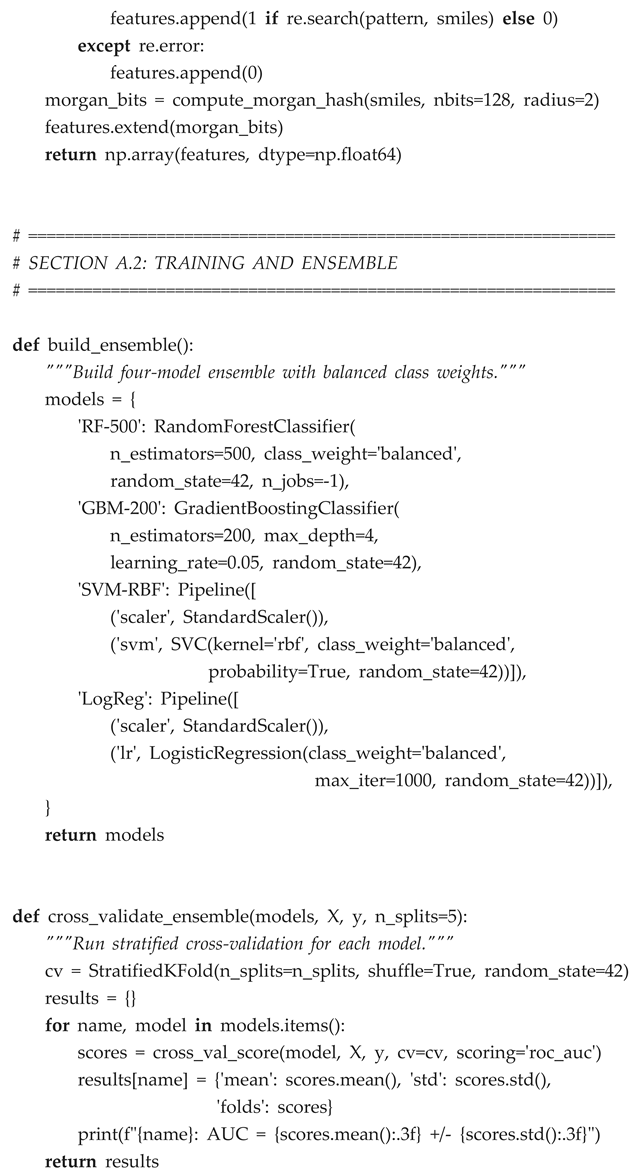
Appendix A. ML Pipeline Code










References
- Abegg, M. A. Exploratory AI-assisted in silico evidence that bacterial signaling molecules may occupy a drug-like pharmacokinetic space. Preprints 2026, 2026020854. [Google Scholar] [CrossRef]
- Andersson, D. I.; Hughes, D. Microbiological effects of sublethal levels of antibiotics. Nat Rev Microbiol 2014, 12, 465–478. [Google Scholar] [CrossRef] [PubMed]
- Baell, J. B.; Holloway, G. A. New Substructure Filters for Removal of Pan Assay Interference Compounds (PAINS) from Screening Libraries and for Their Exclusion in Bioassays. J Med Chem 2010, 53, 2719–2740. [Google Scholar] [CrossRef] [PubMed]
- Beyersmann, P. G.; Tomasch, J.; Son, K.; Stocker, R.; Göker, M.; Wagner-Döbler, I.; Simon, M.; Brinkhoff, T. Dual function of tropodithietic acid as antibiotic and signaling molecule in global gene regulation of the probiotic bacterium Phaeobacter inhibens. Scientific Reports 2017, 7(1), Article 730. [Google Scholar] [CrossRef]
- Brown, E. D.; Wright, G. D. Antibacterial drug discovery in the resistance era. Nature 2016, 529, 336–343. [Google Scholar] [CrossRef]
- Cavasotto, C. N.; Scardino, V. Machine learning toxicity prediction: Latest advances by toxicity end point. ACS Omega 2022, 7(51), 47536–47546. [Google Scholar] [CrossRef]
- Cesaro, A.; Bagheri, M.; Torres, M.; Wan, F.; de la Fuente-Nunez, C. Deep learning tools to accelerate antibiotic discovery. Expert Opin Drug Discov 2023, 18(11), 1245–1257. [Google Scholar] [CrossRef]
- Chen, M.; Tung, C.-W.; Shi, Q.; Guo, L.; Shi, L.; Fang, H.; Borlak, J.; Tong, W. A testing strategy to predict risk for drug-induced liver injury in humans using high-content screen assays and the ‘rule-of-two’ model. Archives of Toxicology 2014, 88(7), 1439–1449. [Google Scholar] [CrossRef]
- Daina, A.; Michielin, O.; Zoete, V. SwissADME: a free web tool to evaluate pharmacokinetics, drug-likeness and medicinal chemistry friendliness of small molecules. Sci Rep 2017, 7, 42717. [Google Scholar] [CrossRef]
- Davies, J.; Davies, D. Origins and evolution of antibiotic resistance. Microbiol Mol Biol Rev 2010, 74, 417–433. [Google Scholar] [CrossRef]
- Desai, J. V.; Mitchell, A. P. Candida albicans biofilm development and its genetic control. Microbiology Spectrum 2015, 3(3), Article MB-0005-2014. [Google Scholar] [CrossRef] [PubMed]
- Fajardo, A.; Martínez, J. L. Antibiotics as signals that trigger specific bacterial responses. Curr Opin Microbiol 2008, 11(2), 161–167. [Google Scholar] [CrossRef] [PubMed]
- Fernández-Delgado, M.; Cernadas, E.; Barro, S.; Amorim, D. Do we need hundreds of classifiers to solve real world classification problems? Journal of Machine Learning Research 2014, 15(90), 3133–3181. [Google Scholar] [CrossRef]
- Feng, X.; Marcon, V.; Pisula, W.; Hansen, M. R.; Kirkpatrick, J.; Grozema, F.; Andrienko, D.; Kremer, K.; Müllen, K. Towards high charge-carrier mobilities by rational design of the shape and periphery of discotics. Nat Mater 2009, 8(5), 421–426. [Google Scholar] [CrossRef]
- Gaulton, A.; Hersey, A.; Nowotka, M.; Bento, A. P.; Chambers, J.; Mendez, D.; Mutowo, P.; Atkinson, F.; Bellis, L. J.; Cibrián-Uhalte, E.; et al. The ChEMBL database in 2017. Nucleic Acids Res 2017, 45(D1), D945–D954. [Google Scholar] [CrossRef]
- Gfeller, D.; Grosdidier, A.; Wirth, M.; Daina, A.; Michielin, O.; Zoete, V. SwissTargetPrediction: a web server for target prediction of bioactive small molecules. Nucleic Acids Res 2014, 42(W1), W32–W38. [Google Scholar] [CrossRef]
- Goh, E.-B.; Yim, G.; Tsui, W.; McClure, J.; Surette, M. G.; Davies, J. Transcriptional modulation of bacterial gene expression by subinhibitory concentrations of antibiotics. Proc Natl Acad Sci USA 2002, 99(26), 17025–17030. [Google Scholar] [CrossRef]
- Guengerich, F. P. Mechanisms of drug toxicity and relevance to pharmaceutical development. Drug Metab Pharmacokinet 2011, 26, 167–176. [Google Scholar] [CrossRef]
- Hagmann, W. K. The many roles for fluorine in medicinal chemistry. J Med Chem 2008, 51, 4359–4369. [Google Scholar] [CrossRef]
- Henriksen, N. N. S. E.; Lindqvist, L. L.; Wibowo, M.; Sonnenschein, E. C.; Bentzon-Tilia, M.; Gram, L. Role is in the eye of the beholder—the multiple functions of the antibacterial compound tropodithietic acid produced by marine Rhodobacteraceae. FEMS Microbiol Rev 2022, 46, fuac007. [Google Scholar] [CrossRef]
- Hentzer, M.; Riedel, K.; Rasmussen, T. B.; Heydorn, A.; Andersen, J. B.; Parsek, M. R.; Rice, S. A.; Eberl, L.; Molin, S.; Høiby, N.; Kjelleberg, S.; Givskov, M. Inhibition of quorum sensing in Pseudomonas aeruginosa biofilm bacteria by a halogenated furanone compound. Microbiology 2002, 148 Pt 1, 87–102. [Google Scholar] [CrossRef]
- Hoffman, L. R.; D’Argenio, D. A.; MacCoss, M. J.; Zhang, Z.; Jones, R. A.; Miller, S. I. Aminoglycoside antibiotics induce bacterial biofilm formation. Nature 2005, 436, 1171–1175. [Google Scholar] [CrossRef] [PubMed]
- Irwin, J. J.; Sterling, T.; Mysinger, M. M.; Bolstad, E. S.; Coleman, R. G. ZINC: a free tool to discover chemistry for biology. J Chem Inf Model 2012, 52(7), 1757–1768. [Google Scholar] [CrossRef] [PubMed]
- Kalia, V. C. Quorum sensing inhibitors: an overview. Biotechnol Adv 2013, 31, 224–245. [Google Scholar] [CrossRef] [PubMed]
- Landrum, G. RDKit: Open-source cheminformatics. 2016. Available online: https://www.rdkit.org.
- Linares, J. F.; Gustafsson, I.; Baquero, F.; Martinez, J. L. Antibiotics as intermicrobial signaling agents instead of weapons. Proc Natl Acad Sci USA 2006, 103, 19484–19489. [Google Scholar] [CrossRef]
- Liu, G.-Y.; Yu, D.; Fan, M.-M.; Zhang, X.; Jin, Z.-Y.; Tang, C.; Liu, X.-F. Antimicrobial resistance crisis: Could artificial intelligence be the solution? Military Medical Research 2024, 11(1), Article 7. [Google Scholar] [CrossRef]
- Lovering, F.; Bikker, J.; Humblet, C. Escape from Flatland: increasing saturation as an approach to improving clinical success. J Med Chem 2009, 52(21), 6752–6756. [Google Scholar] [CrossRef]
- McGovern, S. L.; Helfand, B. T.; Feng, B.; Shoichet, B. K. A specific mechanism of nonspecific inhibition. J Med Chem 2003, 46(20), 4265–4272. [Google Scholar] [CrossRef]
- Meanwell, N. A. Synopsis of bioisosteres in drug design. J Med Chem 2011, 54, 2529–2591. [Google Scholar] [CrossRef]
- Mugumbate, G.; Overington, J. P. The relationship between target-class and the physicochemical properties of antibacterial drugs. Bioorganic & Medicinal Chemistry 2015, 23(16), 5218–5224. [Google Scholar] [CrossRef]
- Mulliner, D.; Schmidt, F.; Stolte, M.; Spirkl, H.-P.; Czich, A.; Amberg, A. Computational models for human and animal hepatotoxicity with a global application scope. Chemical Research in Toxicology 2016, 29(5), 757–767. [Google Scholar] [CrossRef]
- Murray, C. J. L.; Ikuta, K. S.; Sharara, F.; Swetschinski, L.; Robles Aguilar, G.; Gray, A.; et al. Global burden of bacterial antimicrobial resistance in 2019: a systematic analysis. Lancet 2022, 399(10325), 629–655. [Google Scholar] [CrossRef] [PubMed]
- O’Shea, R.; Moser, H. E. Physicochemical properties of antibacterial compounds. J Med Chem 2008, 51, 2871–2878. [Google Scholar] [CrossRef] [PubMed]
- Patani, G. A.; LaVoie, E. J. Bioisosterism: a rational approach in drug design. Chem Rev 1996, 96, 3147–3176. [Google Scholar] [CrossRef] [PubMed]
- Payne, D. J.; Gwynn, M. N.; Holmes, D. J.; Pompliano, D. L. Drugs for bad bugs: confronting the challenges of antibacterial discovery. Nat Rev Drug Discov 2007, 6(1), 29–40. [Google Scholar] [CrossRef]
- Pedregosa, F.; Varoquaux, G.; Gramfort, A.; Michel, V.; Thirion, B.; Grisel, O.; Blondel, M.; Prettenhofer, P.; Weiss, R.; Dubourg, V.; Vanderplas, J.; Passos, A.; Cournapeau, D.; Brucher, M.; Perrot, M.; Duchesnay, É. Scikit-learn: Machine learning in Python. Journal of Machine Learning Research 2011, 12, 2825–2830. [Google Scholar]
- Pires, D. E. V.; Blundell, T. L.; Ascher, D. B. pkCSM: predicting small-molecule pharmacokinetic and toxicity properties using graph-based signatures. J Med Chem 2015, 58(9), 4066–4072. [Google Scholar] [CrossRef]
- Purser, S.; Moore, P. R.; Swallow, S.; Gouverneur, V. Fluorine in medicinal chemistry. Chem Soc Rev 2008, 37(2), 320–330. [Google Scholar] [CrossRef]
- Reisenbauer, J. C.; Green, O.; Franchino, A.; Finkelstein, P.; Morandi, B. Late-stage diversification of indole skeletons through nitrogen atom insertion. Science 2022, 377(6610), 1104–1109. [Google Scholar] [CrossRef]
- Richter, M. F.; Drown, B. S.; Riley, A. P.; Garcia, A.; Shirai, T.; Svec, R. L.; Hergenrother, P. J. Predictive compound accumulation rules yield a broad-spectrum antibiotic. Nature 2017, 545(7654), 299–304. [Google Scholar] [CrossRef]
- Ritchie, T. J.; Macdonald, S. J. F. The impact of aromatic ring count on compound developability – are too many aromatic rings a liability in drug design? Drug Discovery Today 2009, 14(21–22), 1011–1020. [Google Scholar] [CrossRef]
- Sengupta, S.; Chattopadhyay, M. K.; Grossart, H. P. Multifaceted roles of antibiotics and antibiotic resistance in nature. Front Microbiol 2013, 4, 47. [Google Scholar] [CrossRef] [PubMed]
- Skindersoe, M. E.; Alhede, M.; Phipps, R.; Yang, L.; Jensen, P. O.; Rasmussen, T. B.; Bjarnsholt, T.; Tolker-Nielsen, T.; Høiby, N.; Givskov, M. Effects of antibiotics on quorum sensing in Pseudomonas aeruginosa. Antimicrob Agents Chemother 2008, 52(10), 3648–3663. [Google Scholar] [CrossRef] [PubMed]
- Sterling, T.; Irwin, J. J. ZINC 15 – Ligand discovery for everyone. Journal of Chemical Information and Modeling 2015, 55(11), 2324–2337. [Google Scholar] [CrossRef] [PubMed]
- Stokes, J. M.; Yang, K.; Swanson, K.; Jin, W.; Cubillos-Ruiz, A.; Donghia, N. M.; MacNair, C. R.; French, S.; Carfrae, L. A.; Bloom-Ackermann, Z.; et al. A deep learning approach to antibiotic discovery. Cell 2020, 180(4), 688–702. [Google Scholar] [CrossRef]
- Suk, J. S.; Xu, Q.; Kim, N.; Hanes, J.; Ensign, L. M. PEGylation as a strategy for improving nanoparticle-based drug and gene delivery. Adv Drug Deliv Rev 2016, 99 Pt A, 28–51. [Google Scholar] [CrossRef]
- van Tilborg, D.; Brinkmann, H.; Criscuolo, E.; Rossen, L.; Özçelik, R.; Grisoni, F. Deep learning for low-data drug discovery: hurdles and opportunities. Curr Opin Struct Biol 2024, 86, 102818. [Google Scholar] [CrossRef]
- Vandeputte, O. M.; Kiendrebeogo, M.; Rajaonson, S.; Diallo, B.; Mol, A.; El Jaziri, M.; Baucher, M. Identification of catechin as one of the flavonoids from Combretum albiflorum bark extract that reduces the production of quorum-sensing-controlled virulence factors in Pseudomonas aeruginosa PAO1. Appl Environ Microbiol 2010, 76(1), 243–253. [Google Scholar] [CrossRef]
- Vitaku, E.; Smith, D. T.; Njardarson, J. T. Analysis of the structural diversity, substitution patterns, and frequency of nitrogen heterocycles among U.S. FDA approved pharmaceuticals. J Med Chem 2014, 57(24), 10257–10274. [Google Scholar] [CrossRef]
- Wöhrle, T.; Wurzbach, I.; Kirres, J.; Kostidou, A.; Kapernaum, N.; Litterscheidt, J.; Haenle, J. C.; Staffeld, P.; Baro, A.; Giesselmann, F.; Laschat, S. Discotic liquid crystals. Chemical Reviews 2016, 116(3), 1139–1241. [Google Scholar] [CrossRef]
- Wong, F.; Zheng, E. J.; Valeri, J. A.; Donghia, N. M.; Anahtar, M. N.; Omori, S.; Li, A.; Cubillos-Ruiz, A.; Krishnan, A.; Jin, W.; Manson, A. L.; Friedrichs, J.; Helbig, R.; Hajian, B.; Fiejtek, D. K.; Wagner, F. F.; Soutter, H. H.; Earl, A. M.; Stokes, J. M.; Renner, L. D.; Collins, J. J. Discovery of a structural class of antibiotics with explainable deep learning. Nature 2024, 626, 177–185. [Google Scholar] [CrossRef]
| Compound | Description | P(QS) | P(Abx) | MW | LogP | HBD | Lip. | PAINS | Brenk | SA | Issue |
|---|---|---|---|---|---|---|---|---|---|---|---|
| Parent | 6×OH | 0.969 | 0.813 | 274 | 1.60 | 6 | 1 | 0 | 2 | 1.44 | Catechol+HBD |
| Lead | 4×OAc | 0.849 | 0.876 | 442 | 2.72 | 2 | 0 | 0 | 2 | 2.66 | High TPSA |
| Mod 4c | CH₂NH₂+3OMe | 0.990 | 0.234 | 329 | 2.41 | 3 | 0 | 0 | 1† | 1.98 | PAH+AMES+ |
| Mod 5 | NHMe+sat.ring | 0.964 | 0.227 | ~305 | — | — | — | — | 0‡ | — | Low P_Abx |
| M6-12 | NHMe+F+pip.NH | 0.928 | 0.792 | 340 | 0.48* | 4 | 0 | 0 | 0 | 4.50 | ✓ Final |
| Property | Custom Desc. | SwissADME | RDKit | Concordance |
|---|---|---|---|---|
| MW (Da) | 340.4 | 340.39 | 340.3950 | ✔ |
| LogP | — | 0.63 (cons.) | 0.4847 (Crippen) | ≈¹ |
| TPSA (Ų) | — | 82.98 | 82.98 | ✔ |
| HBD | 4 | 4 | 4 | ✔ |
| HBA | 7 | 7 | 6² | ≈² |
| Fraction Csp3 | 0.65 | 0.65 | 0.6471 | ✔ |
| Rotatable bonds | 4 | 4 | 4 | ✔ |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




