Submitted:
01 March 2026
Posted:
03 March 2026
You are already at the latest version
Abstract
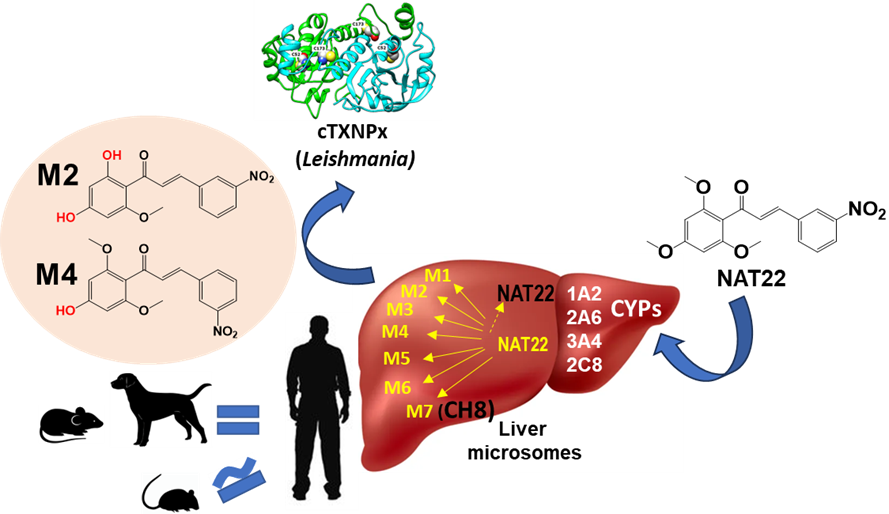
Keywords:
1. Introduction
2. Materials and Methods
2.1. Materials
2.2. In Silico Prediction
2.3. Liver Microsomal Assay
2.4. LC-MS/MS Analysis
2.5. NMR Spectroscopy
2.6. Protein Modeling and Molecular Docking
3. Results
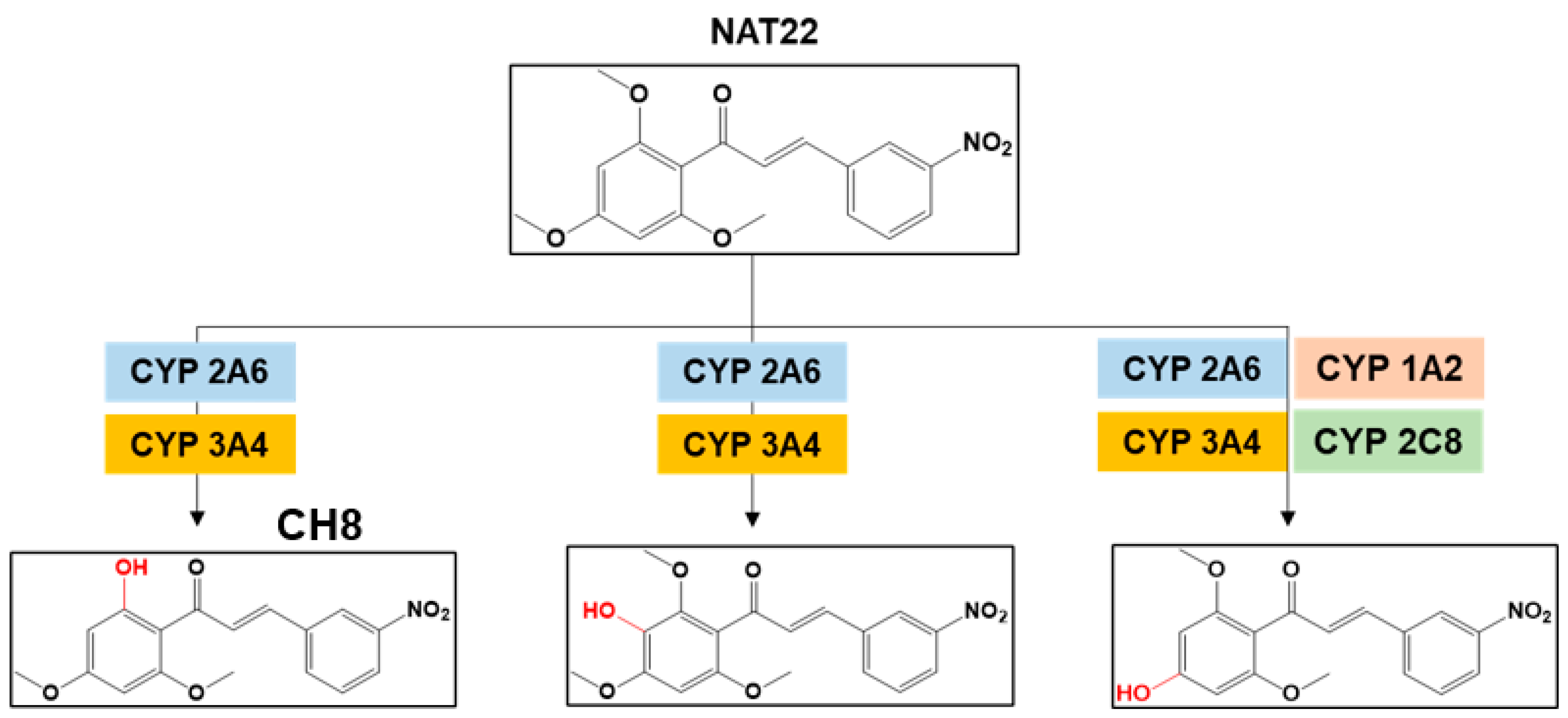
3.1. In Silico Prediction of Hepatic Metabolites of NAT22
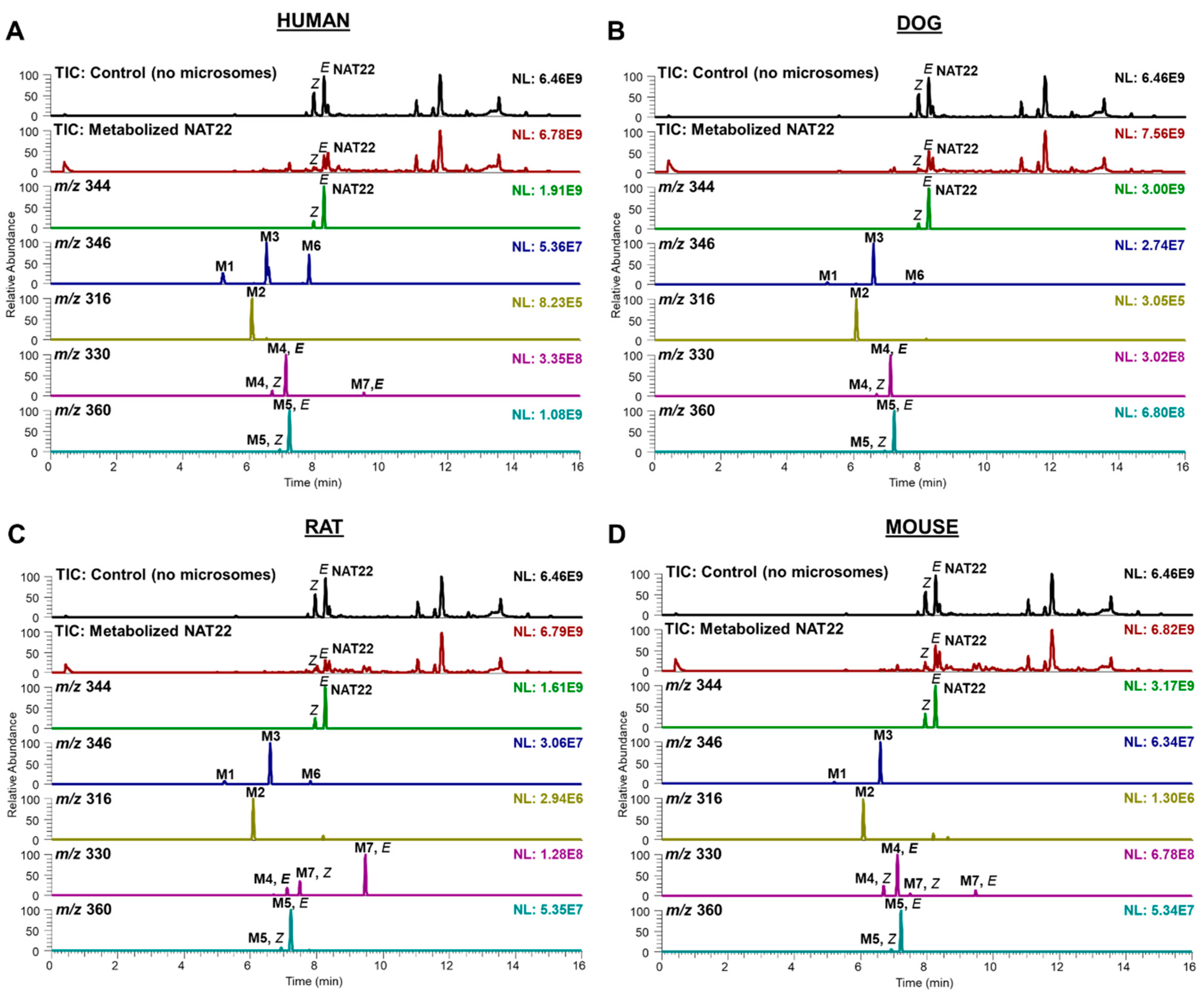
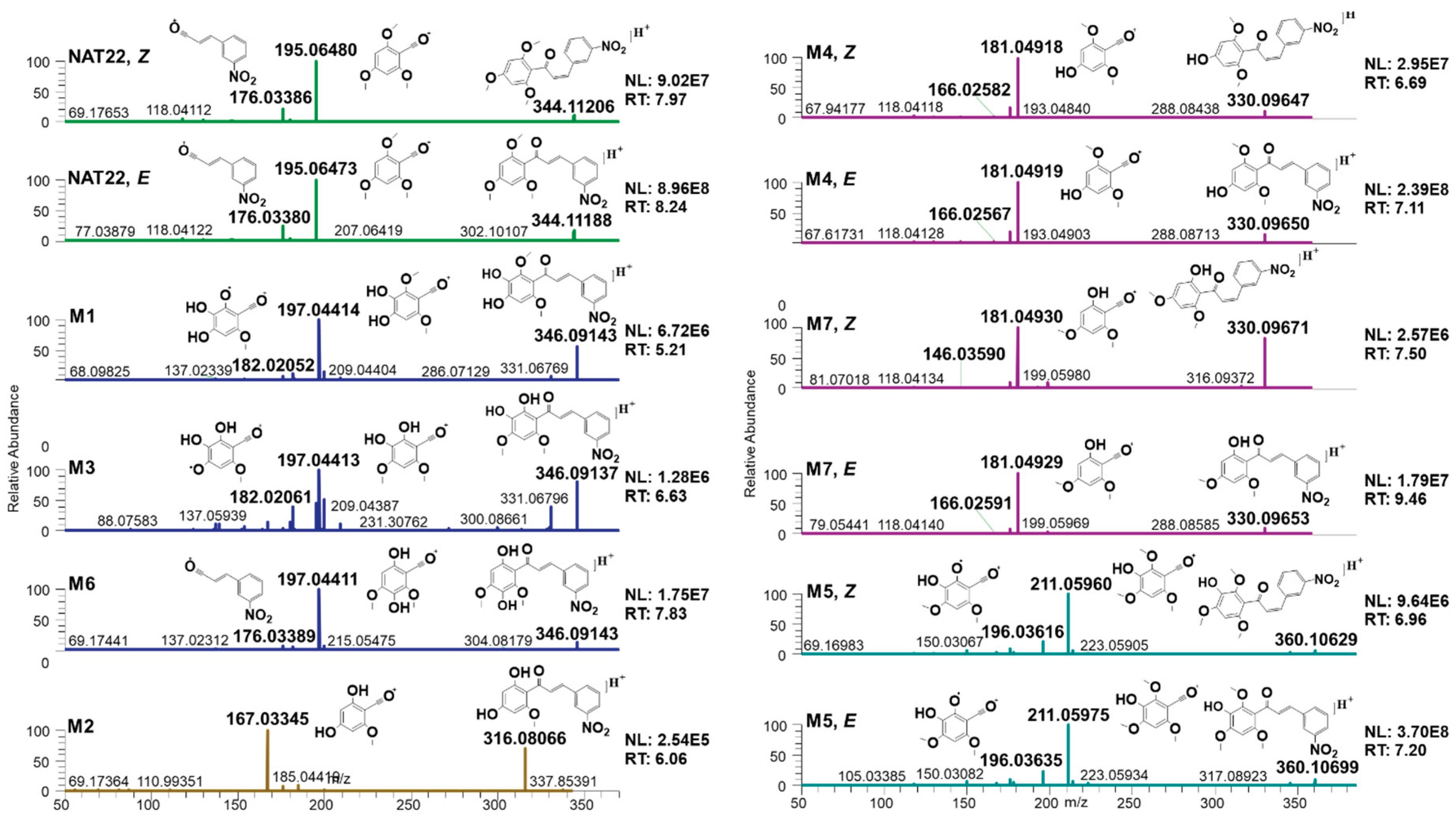
3.2. Identification of NAT22 Metabolites by LC-MS/MS and NMR
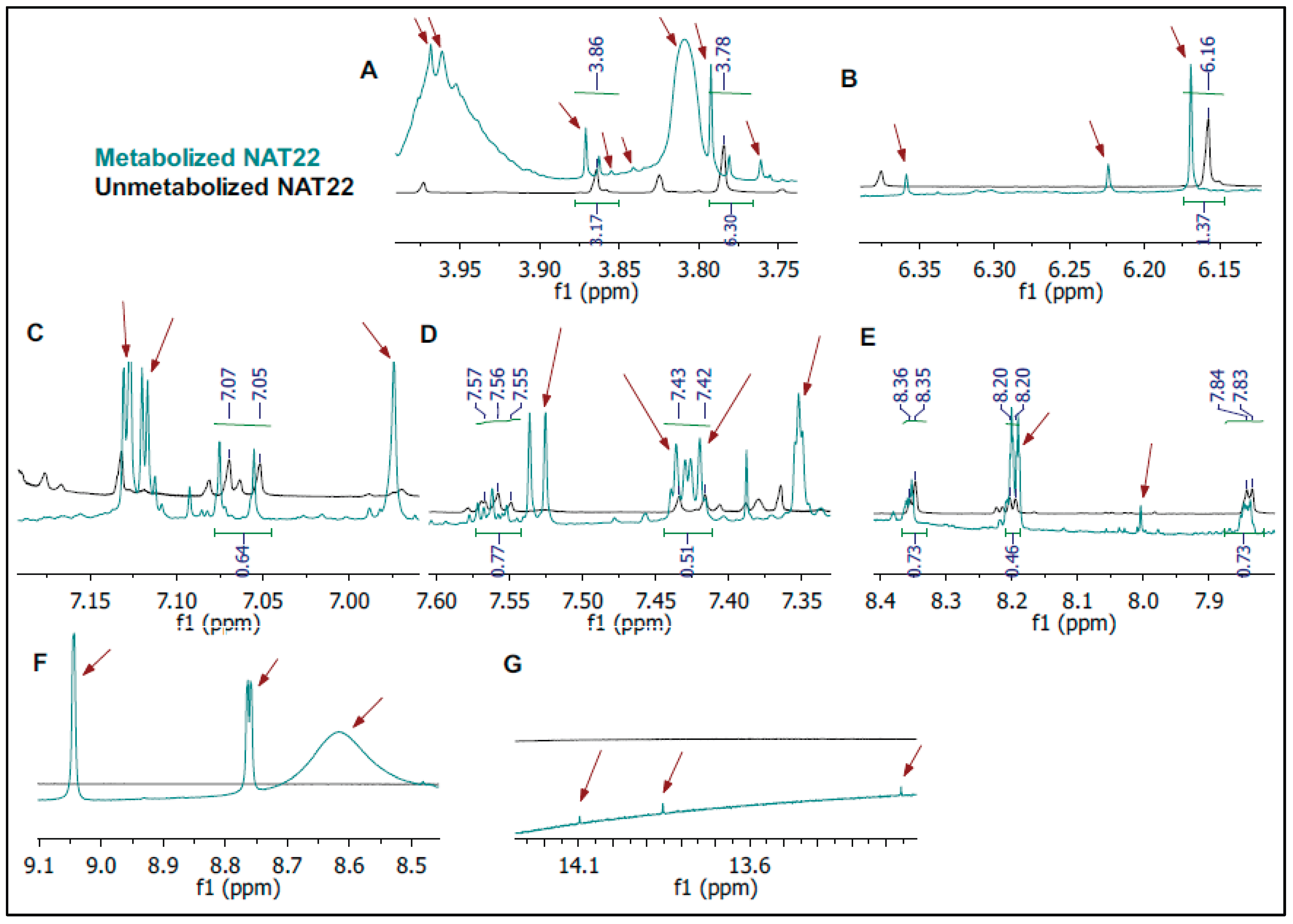
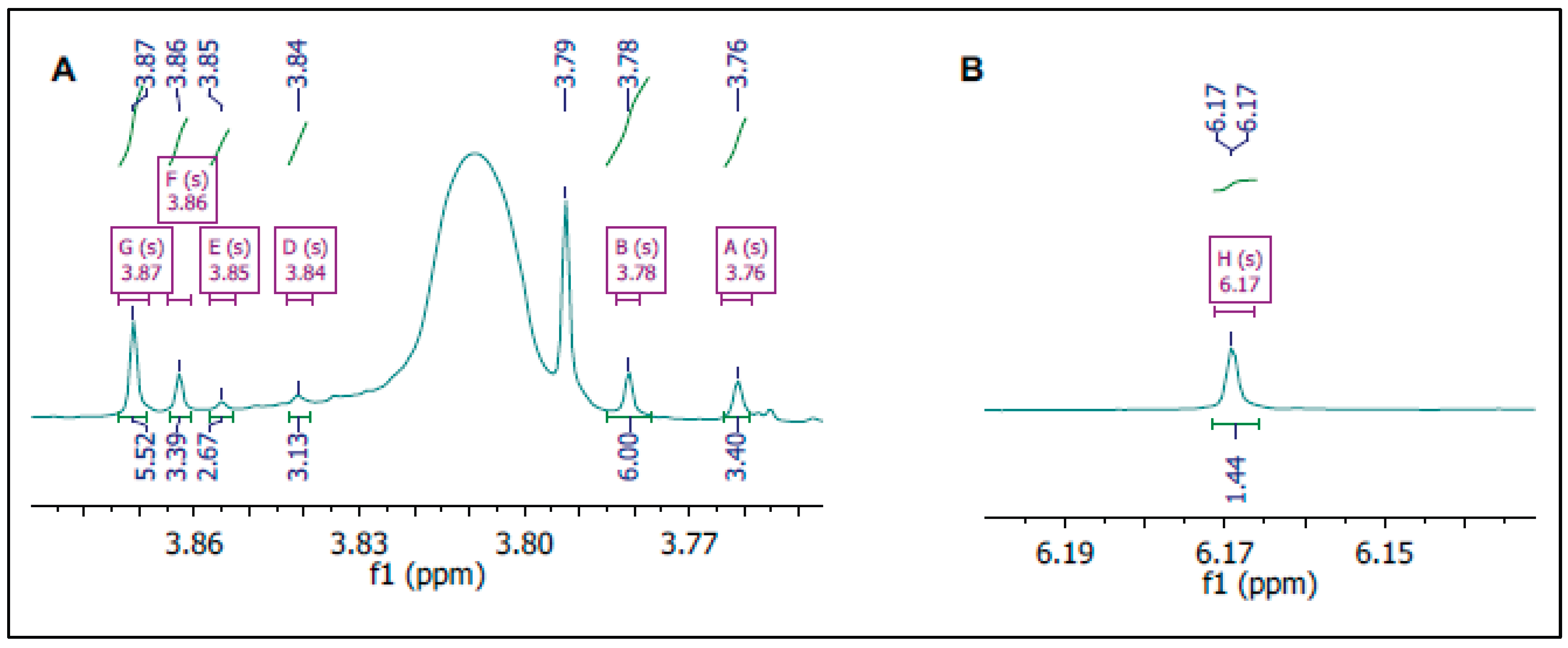
3.3. NMR Analysis
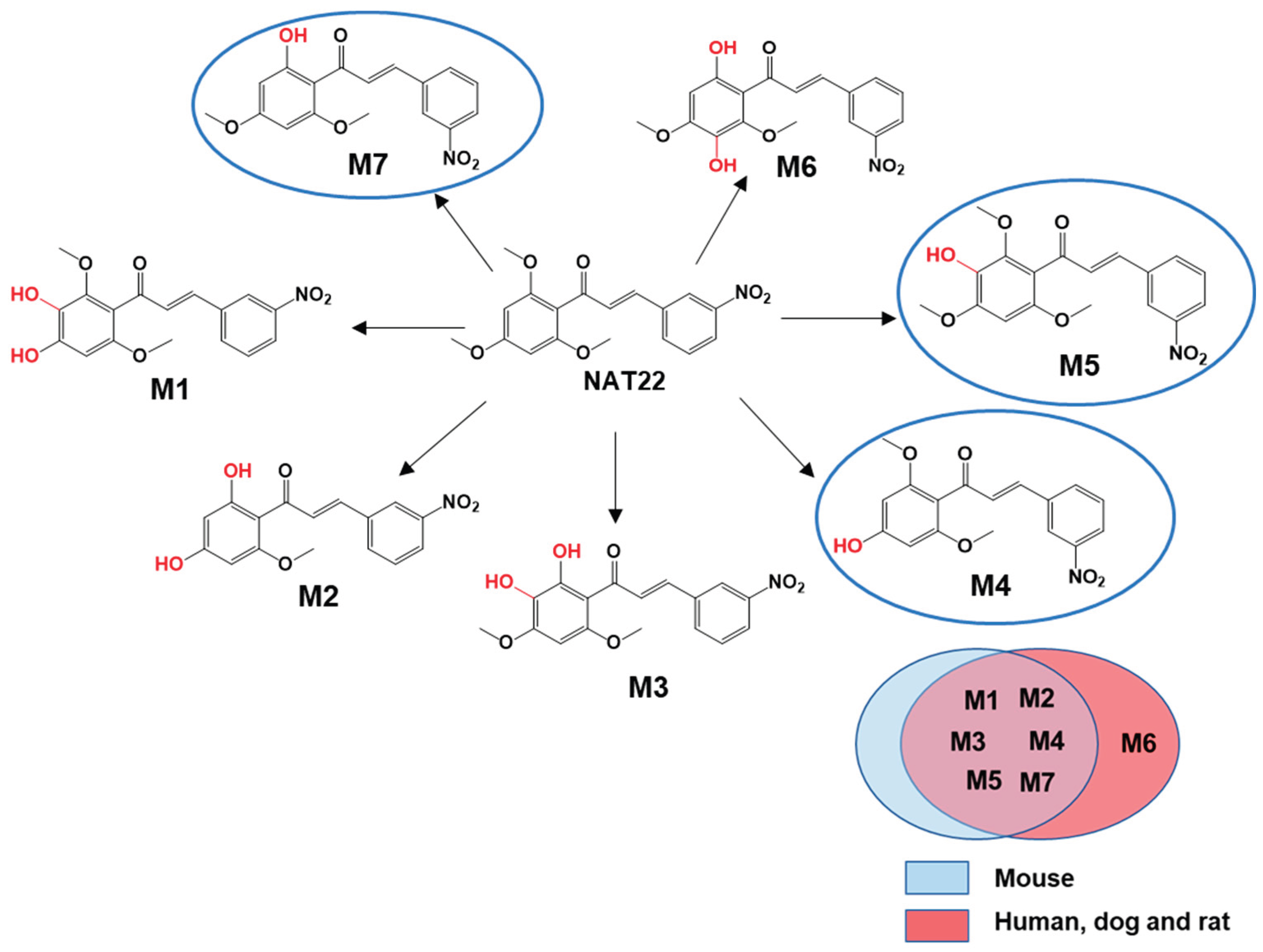
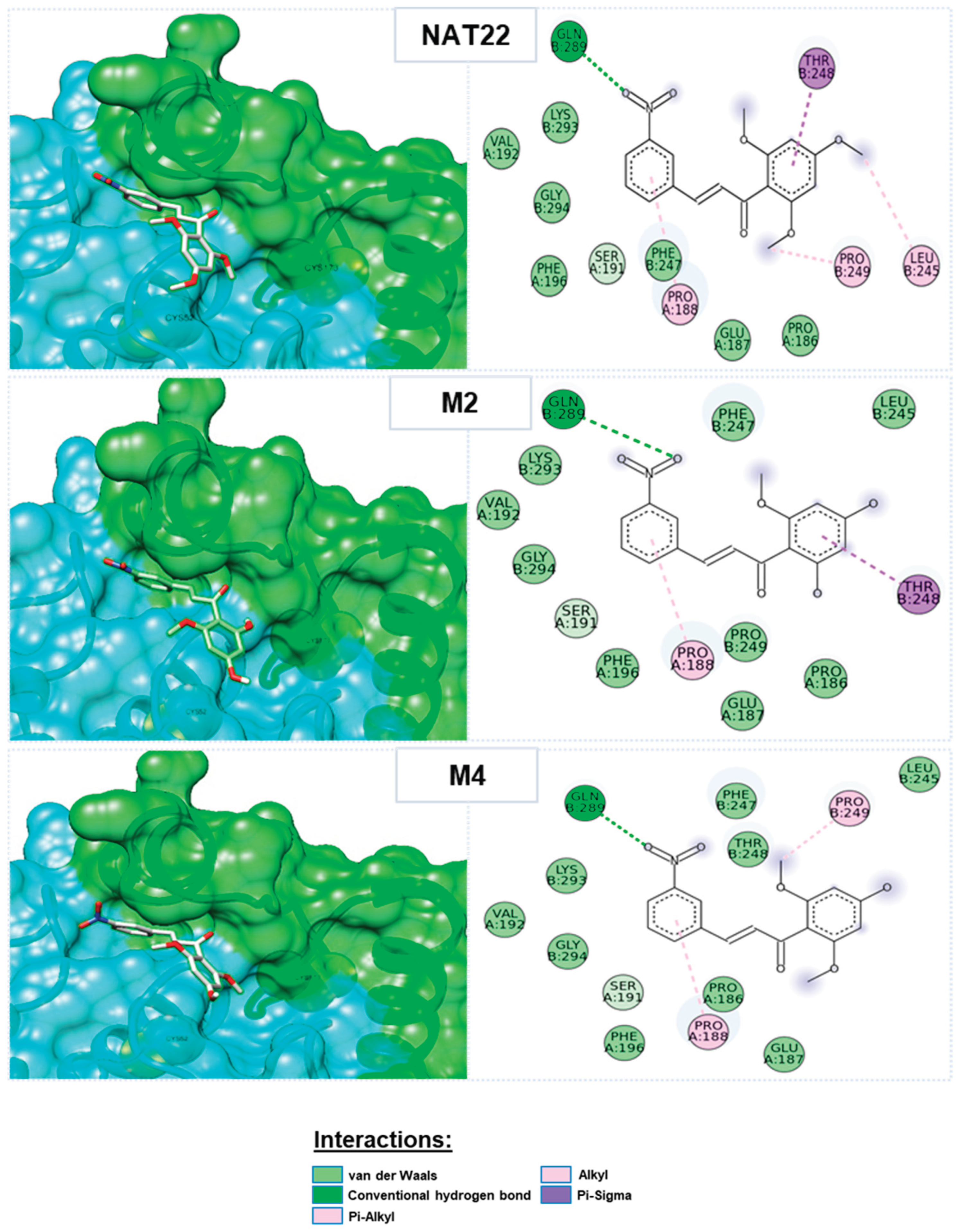
3.4. Molecular Docking of NAT22 Metabolites
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
Abbreviations
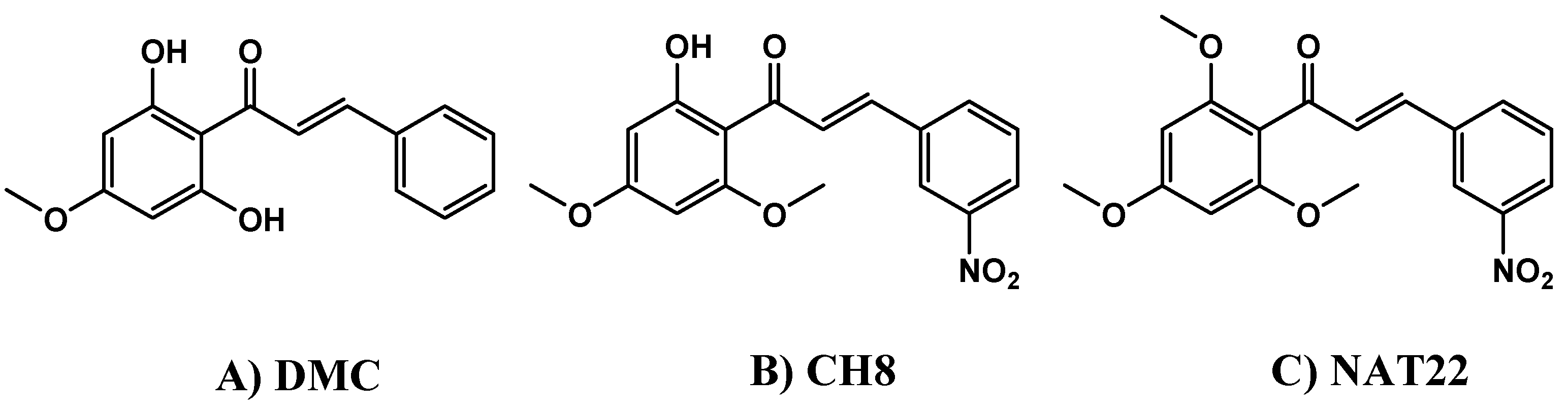
- NAT22 — 3-nitro-2’-hydroxy-4’,6’-dimethoxychalcone
- cTXNPx — Tryparedoxin peroxidase
- CH8 — 3-nitro-2’-hydroxy-4’,6’-dimethoxychalcone
- DMC — 2’,6’-dihydroxy-4’-methoxychalcone
- CYP — Cytochrome P450
- LC-MS/MS — Liquid Chromatography coupled to Tandem Mass Spectrometry
- NMR — Nuclear Magnetic Resonance
- PBS — Phosphate-Buffered Saline
- MTBE — Methyl tert-butyl ether
- HPLC — High Performance Liquid Chromatography
- NADP⁺ — Nicotinamide Adenine Dinucleotide Phosphate
- Tris-HCl — Tris(hydroxymethyl)aminomethane hydrochloride buffer
- ADMET — Absorption, Distribution, Metabolism, Excretion, and Toxicity
- BLAST — Basic Local Alignment Search Tool
- PDB — Protein Data Bank
- TXI probe — Probe type for NMR spectroscopy (1H NMR)
Appendix A
| Analyte | Accurate mass (m/z) |
Human | Dog | Rat | Mouse | Molecular Formula | ||||
|---|---|---|---|---|---|---|---|---|---|---|
| Exact mass (m/z) |
Error (ppm) |
Exact mass (m/z) |
Error (ppm) |
Exact mass (m/z) |
Error (ppm) |
Exact mass (m/z) |
Error (ppm) |
|||
| NAT22 (Z) | 344,11286 | 344,11246 | 1,16 | 344,11209 | 2,24 | 344,11209 | 2,24 | 344,11215 | 2,06 | C18H17NO6 |
| NAT22 (E) | 344,11230 | 1,63 | 344,11206 | 2,32 | 344,11191 | 2,76 | 344,11218 | 1,98 | ||
| M1 (E) | 346,09213 | 346,09186 | 2,92 | 346,09149 | 1,85 | 346,09186 | 0,78 | 346,09171 | 1,21 | C17H15NO7 |
| M2 (E) | 316,08156 | 316,08112 | 1,39 | 316,08099 | 1,80 | 316,08081 | 2,37 | 316,08145 | 0,35 | C16H13NO6 |
| M3 (E) | 346,09213 | 346,09189 | 0,69 | 346,09143 | 2,02 | 346,09140 | 2,11 | 346,09164 | 1,42 | C17H15NO7 |
| M4 (Z) | 330,09721 | 330,09702 | 0,58 | 330,09653 | 2,06 | 330,09631 | 2,73 | 330,09665 | 1,70 | C17H15NO6 |
| M4 (E) | 330,09680 | 1,24 | 330,09641 | 2,42 | 330,09637 | 2,54 | 330,09650 | 2,15 | ||
| M5 (E) | 360,10778 | 360,10745 | 0,92 | 360,10690 | 2,44 | 360,10687 | 2,53 | 360,10709 | 1,92 | C18H17NO7 |
| M6 (E) | 346,09213 | 346,09180 | 0,95 | 346,09140 | 2,11 | 346,09131 | 2,37 | - | - | C17H15NO7 |
| M7 (Z) | 330,09721 | 330,09702 | 0,58 | 330,09647 | 2,24 | 330,09647 | 2,24 | 330,09653 | 2,06 | C17H15NO6 |
| M7 (E) | 330,09708 | 0,39 | 330,09644 | 2,33 | 330,09659 | 1,88 | 330,09677 | 1,33 | ||
| Ligands | Docking energy (kcal/mol) | H-bond | Pi-sigma | Pi-alkyl | Alkyl | Van der Waals |
|---|---|---|---|---|---|---|
| M2 | -6,045 | Gln-289 | - | Pro-188 | - | Pro-186, Glu-187, Ser-191, Val-192, Phe-196, Phe-247, Leu-245, Pro-249, Lys-293, Gly-294 |
| M4 | -5,970 | Gln-289 | - | Pro-188 | Pro-249 | Pro-186, Glu-187, Ser-191, Val-192, Phe-196, Phe-247, Leu-245, Thr-248, Lys-293, Gly-294 |
| NAT22 | -5,964 | Gln-289 | Thr-248 | Pro-188 | Leu-245, Pro-249 | Pro-186, Glu-187, Ser-191, Val-192, Phe-196, Phe-247, Lys-293, Gly-294 |
| M7 | -5,927 | Gln-289 | - | Pro-188 | Leu-245 | Pro-186, Glu-187, Ser-191, Val-192, Phe-196, Phe-247, Thr-248, Lys-293, Pro-249, Gly-294 |
| M3 | -5,886 | Gln-289 | - | Pro-188 | Pro-188, Pro-249 | Glu-187, Ser-191, Val-192, Phe-196, Leu-245, Phe-247, Thr-248, Lys-293, Gly-294 |
| M1 | -5,851 | Gln-289 | Thr-248 | Pro-188 | - | Pro-186, Glu-187, Ser-191, Val-192, Phe-196, Phe-247, Pro-249, Lys-293, Gly-294 |
| M6 | -5,841 | Gln-289 | - | Pro-188 | Leu-245 | Pro-186, Glu-187, Ser-191, Val-192, Phe-196, Phe-247, Thr-248, Lys-293, Pro-249, Gly-294 |
| M5 | -5,831 | Gln-289 | - | Pro-188 | Pro-186, Leu-245, Pro-249 | Glu-187, Ser-191, Val-197, Phe-196, Phe-247, Thr-248, Lys-293, Gly-294 |
References
- Adhikari, S.; Nath, P.; Deb, V. K.; Das, N.; Banerjee, A.; Pathak, S.; Duttaroy, A. K. Pharmacological potential of natural chalcones: a recent studies and future perspective. Front Pharmacol. 2025, 16, 1570385. [Google Scholar] [CrossRef]
- Nematollahi, M. H.; Mehrabani, M.; Hozhabri, Y.; Mirtajaddini, M.; Iravani, S. Antiviral and antimicrobial applications of chalcones and their derivatives: from nature to greener synthesis. Heliyon 2023, 9, e20428. [Google Scholar] [CrossRef] [PubMed]
- Chen, M.; Christensen, S. B.; Blom, J.; Lemmich, E.; Nadelmann, L.; Fich, K.; Theander, A.; Kharazmi, T. G. Licochalcone A, a novel antiparasitic agent with potent activity against human pathogenic protozoan species of Leishmania. Antimicrob Agents and Chemother. 1993, 37, 2550–2556. [Google Scholar] [CrossRef] [PubMed]
- Torres-Santos, E. C.; Moreira, D. L.; Kaplan, M. A. C.; Meirelles, M. N.; Rossi-Bergmann, B. Selective effect of 2',6'-dihydroxy-4'-methoxychalcone isolated from Piper aduncum on Leishmania amazonensis. Antimicrob Agents Chemother. 1999, 43, 1234–1241. [Google Scholar] [CrossRef]
- Boeck, P.; Falcão, C. A. B.; Leal, P. C.; Yunes, R. A.; Cechinel-Filho, V.; Torres-Santos, E. C.; Rossi-Bergmann, B. Synthesis of chalcone analogues with increased antileishmanial activity. Bioorg Med Chem. 2006, 14, 1538–1545. [Google Scholar] [CrossRef] [PubMed]
- Andrighetti-Fröhner, C. R.; de Oliveira, K. N.; Gaspar-Silva, D.; Pacheco, L. K.; Joussef, A. C.; Steindel, M.; Simões, C. M. O.; de Souza, A. M. T.; Magalhães, U. O.; Afonso, I. F.; Rodrigues, C. R.; Nunes, R. J.; Castro, H. C. Synthesis, biological evaluation and SAR of sulfonamide 4-methoxychalcone derivatives with potential antileishmanial activity. Eur J Med Chem 2009, 44, 755–763. [Google Scholar] [CrossRef] [PubMed]
- Aponte, J. C.; Castillo, D.; Estevez, Y.; Gonzalez, G.; Arevalo, J.; Hammond, G. B.; Sauvain, M. In vitro and in vivo anti-Leishmania activity of polysubstituted synthetic chalcones. ACS Med Chem Lett. 2010, 20, 100–103. [Google Scholar] [CrossRef]
- Gupta, S.; Shivahare, R.; Korthikunta, V.; Singh, R.; Gupta, S.; Tadigoppula, N. Synthesis and biological evaluation of chalcones as potential antileishmanial agents. Eur J Med Chem. 2014, 81, 359–366. [Google Scholar] [CrossRef] [PubMed]
- De Mello, M. V. P.; Abrahim-Vieira, B. D. A.; Domingos, T. F. S.; de Jesus, J. B.; de Sousa, A. C. C.; Rodrigues, C. R.; Souza, A. M. T. D. A comprehensive review of chalcone derivatives as antileishmanial agents. Eur J Med Chem 2018, 150, 920–929. [Google Scholar] [CrossRef] [PubMed]
- Sousa-Batista, A. J.; Arruda-Costa, N.; Escrivani, D. O.; Reynaud, F.; Steel, P. G.; Rossi-Bergmann, B. Single-dose treatment for cutaneous leishmaniasis with an easily synthesized chalcone entrapped in polymeric microparticles. Parasitol 2020, 147, 1032–1037. [Google Scholar] [CrossRef]
- Garcia, A. R.; Oliveira, D. M. P.; Jesus, J. B.; Souza, A. M. T.; Sodero, A. C. R.; Vermelho, A. B.; Leal, I. C. R.; Souza, R. O. M. A.; Miranda, L. S. M.; Pinheiro, A. S.; Rodrigues, I. A. Identification of chalcone derivatives as inhibitors of Leishmania infantum arginase and promising antileishmanial agents. Front Chem 2021, 8, 624678. [Google Scholar] [CrossRef] [PubMed]
- De Santiago-Silva, K. M.; Bortoleti, B. T. da S.; Oliveira, L. do N.; Maia, F. L. de A.; Castro, J. C.; Costa, I. C.; Lazarin, D. B.; Wardell, J. L.; Wardell, S. M. S. V.; Albuquerque, M. G.; Lima, C. H. da S.; Pavanelli, W. R.; Bispo, M. de L. F.; Gonçalves, R. S. B. Antileishmanial Activity of 4,8-Dimethoxynaphthalenyl chalcones on Leishmania amazonensis. Antibiotics 2022, 11, 1402. [Google Scholar] [CrossRef] [PubMed]
- Alvar, J.; Velez, I. D.; Bern, C.; Herrero, M.; Desjeux, P.; Cano, J.; Jannin, J.; den Boer, M. Leishmaniasis worldwide and global estimates of its incidence. PLoS One 2012, 7, e35671. [Google Scholar] [CrossRef] [PubMed]
- Ribeiro, L. R.; Silva, S. N.; Saliba, M. F.; Carvalho, J. de P.; Cota, G. PloS One 2024, 19, e0315710.
- Esfandiarpour, I.; Farajzadeh, S.; Rahnama, Z.; Fathabadi, E. A.; Heshmatkhah, A. Adverse effects of intralesional meglumine antimoniate and its influence on clinical laboratory parameters in the treatment of cutaneous leishmaniasis. Int J Dermatol 2012, 51, 1221–1225. [Google Scholar] [CrossRef] [PubMed]
- Escrivani, D. O.; Charlton, R. L.; Borsodi, M. P. G.; Zingali, R. B.; Souza, A. M. T.; Abrahim-Vieira, B.; Freitag-Pohl, S.; Pohl, E.; Denny, P. W.; Rossi-Bergmann, B.; Steel, P. G. Chalcones identify cTXNPx as a potential antileishmanial drug target. PLoS Negl Trop Dis 2021, 15, e0009951. [Google Scholar] [CrossRef]
- Sousa-Batista, A. J.; Escrivani-Oliveira, D.; Falcão, C. A. B.; Da S. Philipon, C. I. M.; Rossi-Bergmann, B. Broad spectrum and safety of oral treatment with a promising nitrosylated chalcone in murine leishmaniasis. Antimicrob Agents Chemother. 2018a, 62, e00792. [Google Scholar] [CrossRef] [PubMed]
- Sousa-Batista, A. J.; Pacienza-Lima, W.; Arruda-Costa, N.; Falcão, C. A. B.; Ré, M. I.; Rossi-Bergmann, B. Depot subcutaneous injection with chalcone CH8-loaded poly (Lactic-Co-Glycolic Acid) microspheres as a single-dose treatment of cutaneous leishmaniasis. Antimicrob Agents Chemother 2018b, 62, e01822-17. [Google Scholar] [CrossRef]
- Sousa-Batista, A. J.; Arruda-Costa, N.; Rossi-Bergmann, B.; Ré, M. I. Improved drug loading via spray drying of a chalcone implant for local treatment of cutaneous leishmaniasis. Drug Dev Ind Pharm 2018c, 44, 1473–1480. [Google Scholar] [CrossRef] [PubMed]
- Caridha, D.; Vesely, B.; van Bocxlaer, K.; Arana, B.; Mowbray, C. E.; Rafati, S.; Uliana, S.; Reguera, R.; Kreishman-Deitrick, M.; Sciotti, R.; Buffet, P.; Croft, S. L. Route map for the discovery and pre-clinical development of new drugs and treatments for cutaneous leishmaniasis. Int J Parasitol Drugs Drug Resist 2019, 11, 106–117. [Google Scholar] [CrossRef]
- Turpeinen, M.; Ghiciuc, C.; Opritoui, M.; Tursas, L.; Pelkonen, O.; Pasanen, M. Predictive value of animal models for human cytochrome P450 (CYP)-mediated metabolism: a comparative study in vitro. Xenobiotica 2007, 37, 1367–1377. [Google Scholar] [CrossRef]
- Guengerich, F. P.; Waterman, M. R.; Egli, M. Recent structural insights into cytochrome P450 function. Trends Pharmacol Sci 2016, 37, 625–640. [Google Scholar] [CrossRef]
- Esteves, F.; Rueff, J.; Karanendonk, M. The central role of cytochrome P450 in xenobiotic metabolism—A brief review on a fascinating enzyme family. J Xenobiot. 2021, 11, 94–114. [Google Scholar] [CrossRef] [PubMed]
- Mukherjee, P.; Roy, S.; Ghosh, D.; Nandi, S. K. Role of animal models in biomedical research: a review. Lab Anim Res 2022, 38, 18. [Google Scholar] [CrossRef] [PubMed]
- Marques, L. M. M.; Da Silva-Júnior, E. A.; Gouvea, D. R.; Vessecchi, R.; Pupo, M. T.; Lopes, N. P.; Kato, M. J.; De Oliveira, A. R. M. In vitro metabolism of the alkaloid piplartine by rat liver microsomes. J Pharm Biomed Anal 2014, 95, 113–120. [Google Scholar] [CrossRef] [PubMed]
- Barth, T.; Habenschus, M. D.; Moreira, F. L.; Ferreira, L. de S.; Lopes, N. P.; De Oliveira, A. R. M. In vitro metabolism of the lignan (-)-grandisin, an anticancer drug candidate, by human liver microsomes. Drug Test Anal 2015, 7, 780–786. [Google Scholar] [CrossRef]
- Nielsen, S. E.; Breinholt, V.; Justesen, U.; Cornett, C.; Dragsted, L. O. In vitro biotransformation of flavonoids by rat liver microsomes. Xenobiot 1998, 28, 389–401. [Google Scholar] [CrossRef] [PubMed]
- Zenger, K.; Agnolet, S.; Schneider, B.; Kraus, B. Biotransformation of flavokawains A, B, and C, chalcones from Kava (Piper methysticum), by human liver microsomes. J Agric Food Chem 2015, 63, 6376–6385. [Google Scholar] [CrossRef]
- Guo, J.; Liu, D.; Nikolic, D.; Zhu, D.; Pezzuto, J. M.; van Breemen, R. B. In vitro metabolism of isoliquiritigenin by human liver microsomes. Drug Metab Dispos. 2008, 36, 461–468. [Google Scholar] [CrossRef] [PubMed]
- Qin, Z.; Wang, P.; Duan, S.; Wan, X.; Xing, H.; Yang, J.; Zhang, X.; Yao, Z.; Yao, X. Potential determinants for metabolic fates and inhibitory effects of isobavachalcone involving in human cytochrome P450, UDP-glucuronosyltransferase enzymes, and efflux transporters. J Pharma Sci. 2021, 110, 2285–2294. [Google Scholar] [CrossRef]
- Altschul, S. F.; Gish, W.; Miller, W.; Myers, E. W.; Lipman, D. J. Basic local alignment search tool. J Mol Biol. 1990, 215, 403–410. [Google Scholar] [CrossRef] [PubMed]
- Bateman, A.; et al. UniProt: the universal protein knowledgebase in 2021. Nucleic Acids Res 2021, 49, D480–D489. [Google Scholar]
- Brindisi, M.; Brogi, S.; Relitti, N.; Vallone, V.; Butini, S.; Gemma, S.; Novellino, E.; Colotti, G.; Angiulli, G.; Di Chiaro, F.; Fiorillo, A.; Ilari, A.; Campiani, G. Structure-based discovery of the first non-covalent inhibitors of Leishmania major tryparedoxin peroxidase by high throughput docking. Sci Rep. 2015, 5, 9705. [Google Scholar] [CrossRef]
- De Oliveira, N. S.; De Souza, L. G.; De Almeida, V. M.; Barreto, A. R. R.; Carvalho-Gondim, F.; Schaeffer, E.; Santos-Filho, O. A; Rossi-Bergmann, B.; Da Silva, A. J. M. Synthesis and evaluation of hybrid sulfonamide-chalcones with potential antileishmanial activity. Arch Pharm (Weinheim) 2024, 357, e2300440. [Google Scholar] [CrossRef] [PubMed]
- Almazroo, O. A.; Miah, M. K.; Venkataramanan, R. Drug Metabolism in the liver. Clin Liver Dis. 2017, 21, 1–20. [Google Scholar] [CrossRef] [PubMed]
- Issa, N. T.; Wathieu, H.; Ojo, A.; Byers, S. W.; Dakshanamurthy, S. Drug metabolism in preclinical drug development: A survey of the discovery process, toxicology, and computational tools. Curr Drug Metab. 2017, 18, 556–565. [Google Scholar] [CrossRef]
- Zhang, B.; Duan, D.; Ge, C.; Yao, J.; Liu, Y.; Li, X.; Fang, J. Synthesis of xanthohumol analogues and discovery of potent thioredoxin reductase inhibitor as potential anticancer agent. J Med Chem. 2015, 58, 1795–1805. [Google Scholar] [CrossRef] [PubMed]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




