Submitted:
01 March 2026
Posted:
02 March 2026
You are already at the latest version
Abstract
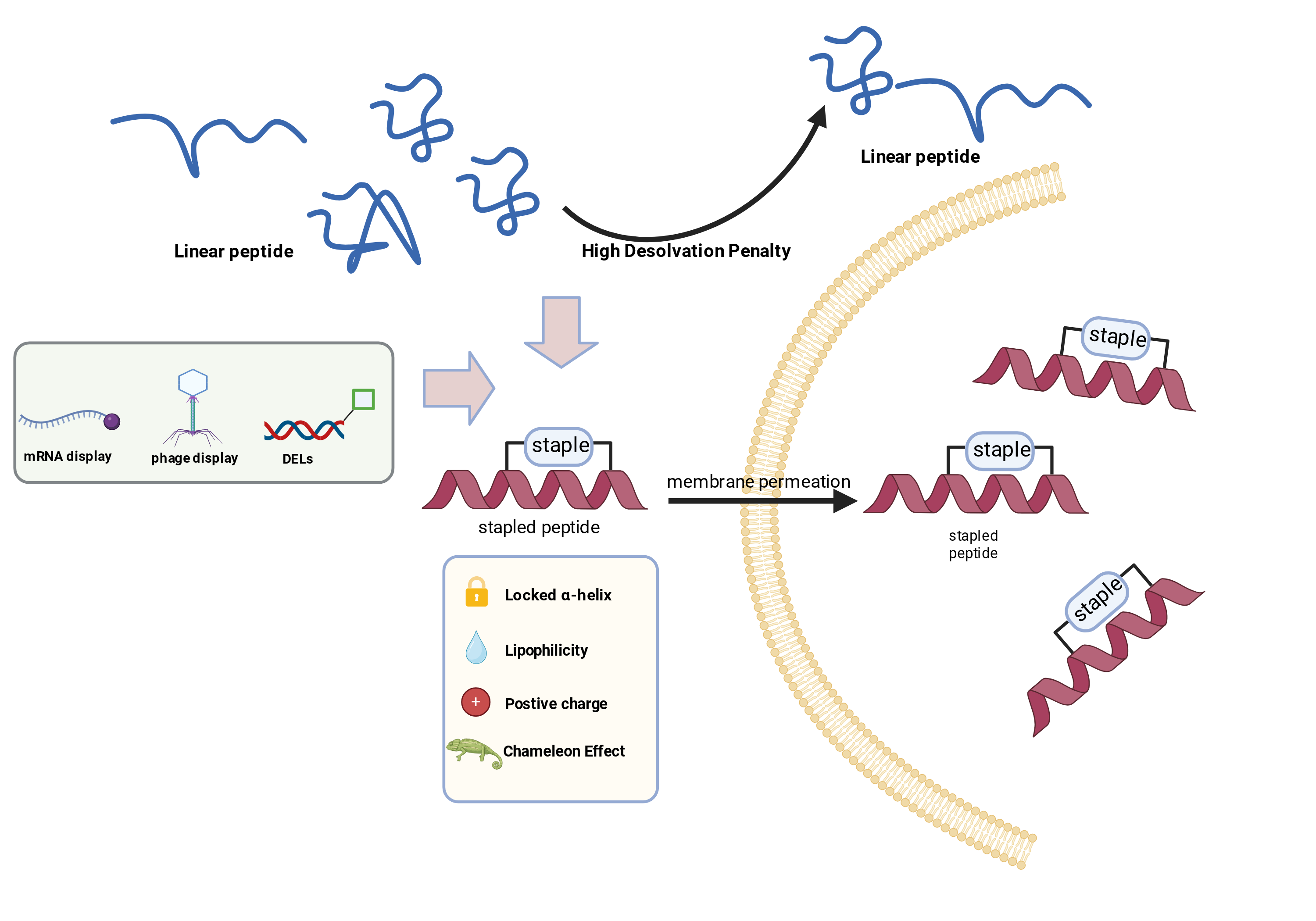
Keywords:
1. Introduction
2. Linkers Derived from In Vitro High-Throughput Screening
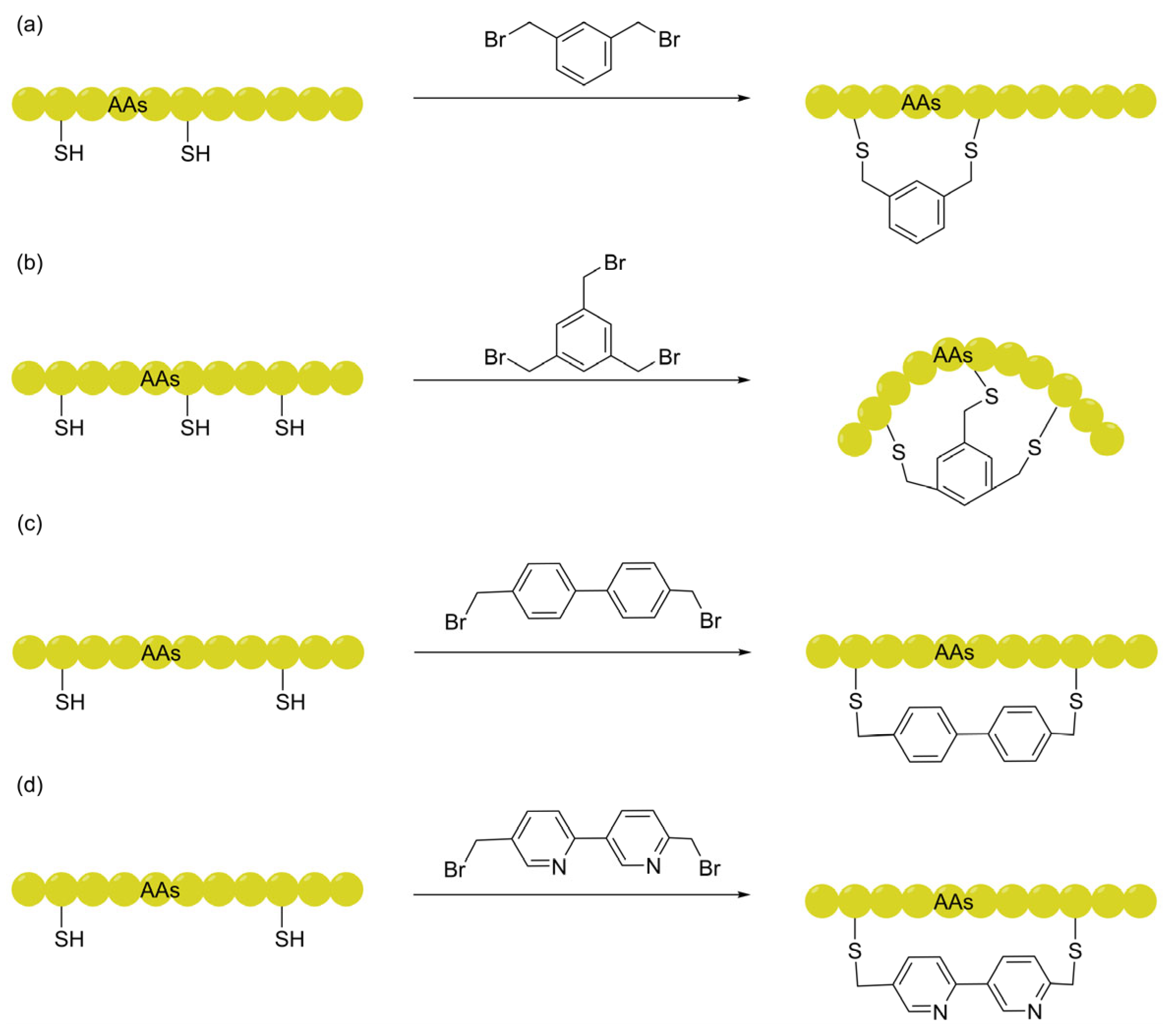
2.1. Arylene-Based Linkers via Thiol-Mediated Nucleophilic Substitution
2.1.1. Aryl Linkers: Conformational Rigidification
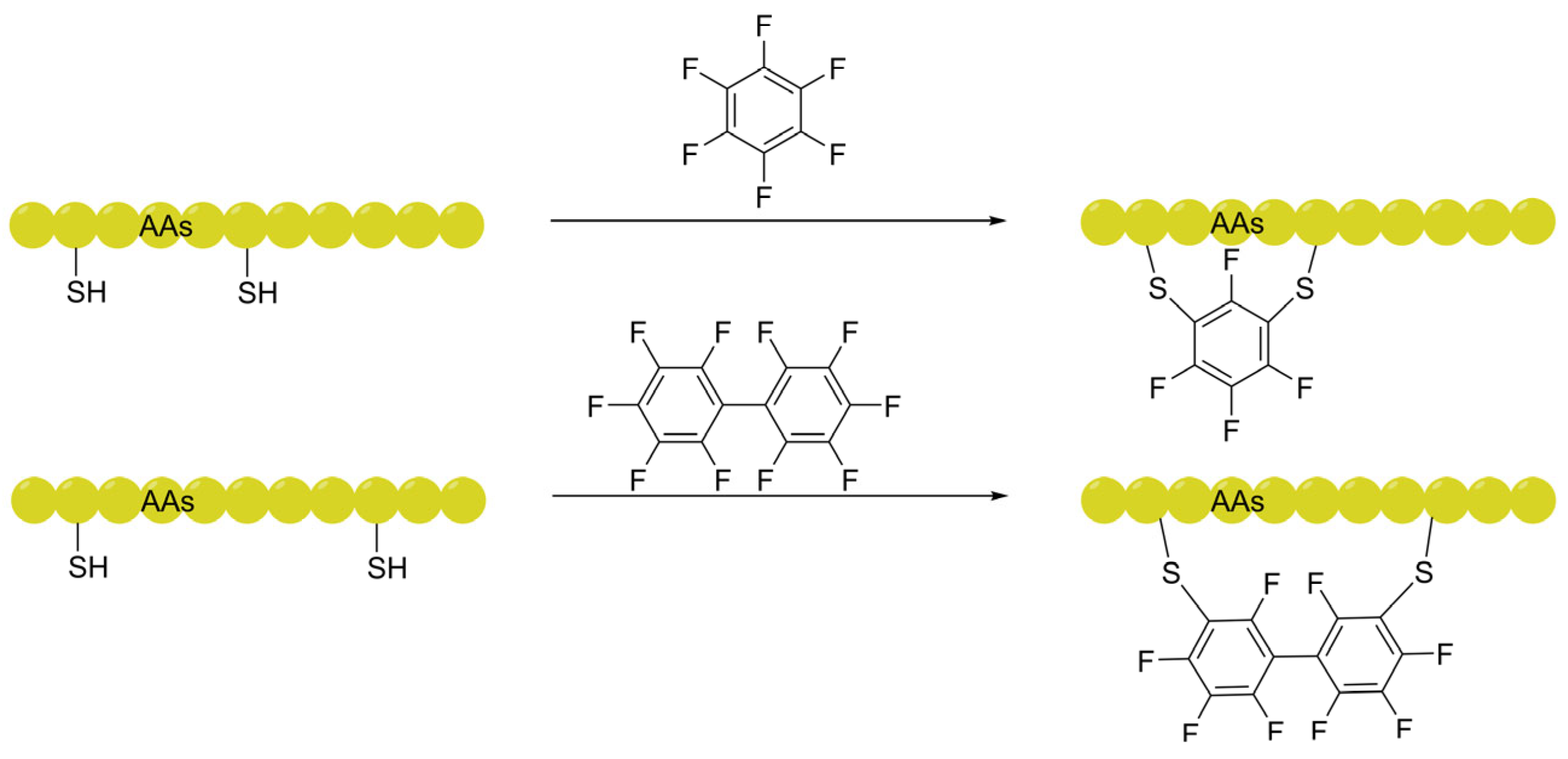
2.1.2. Perfluoroaromatic Linkers: Leveraging the "Fluorine Effect" for Translocation
2.1.3. Bipyridine Linkers: IMHB Shielding
2.2. Heteroaromatic Linkers Constructed from Unnatural Amino Acids (UAAs)
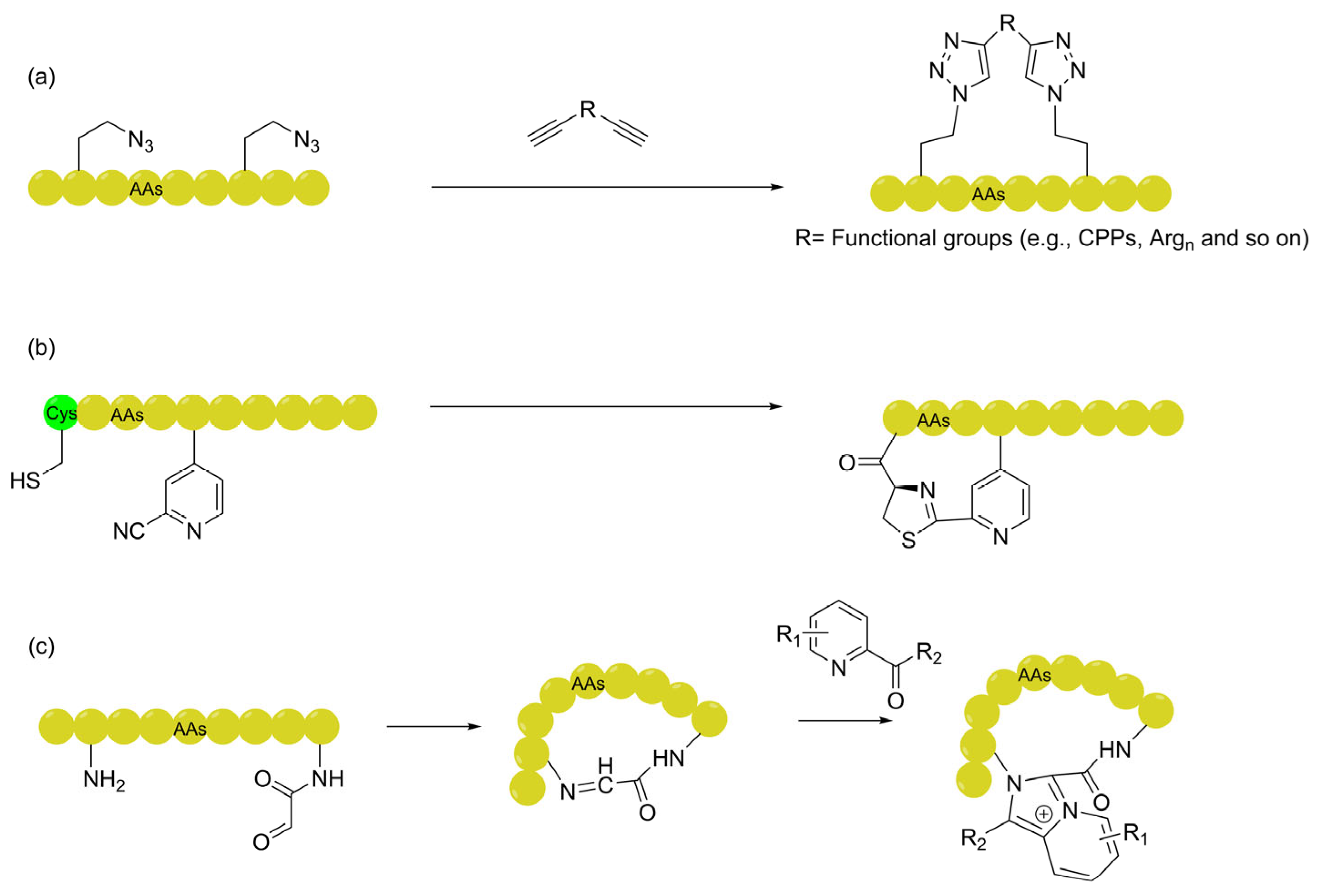
2.2.1. Triazole Linkers: From Structural Stabilization to Decoupled Functionalization
2.2.2. Thiazoline Linkers: Spontaneous Formation and Intramolecular Hydrogen Bond Shielding
2.2.3. Imidazopyridinium (IP+): Challenging Passive Diffusion Dogmas with Cationic Chameleons
3. Post-Screening Structural Refinement and Optimization Strategies
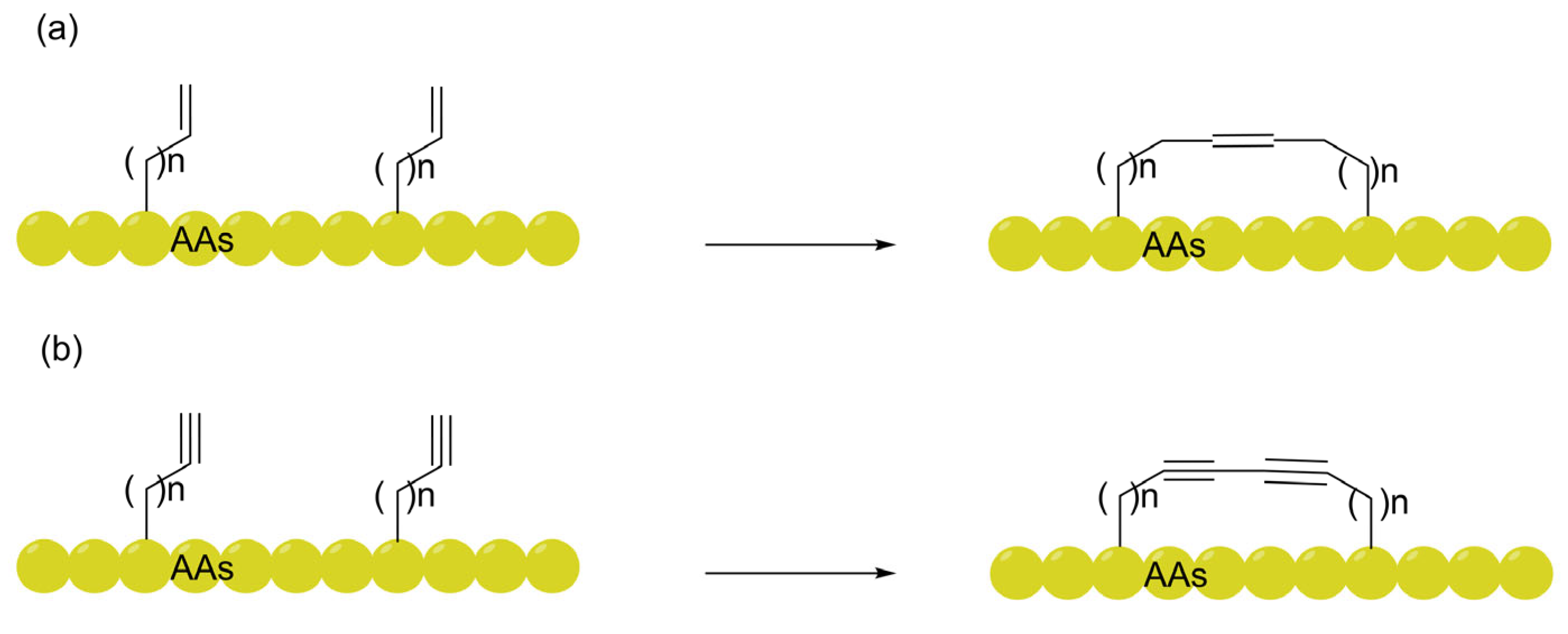
3.1. All-Hydrocarbon Staples: Refining the "Gold Standard" from Helical Stabilization to Metabolic Trapping
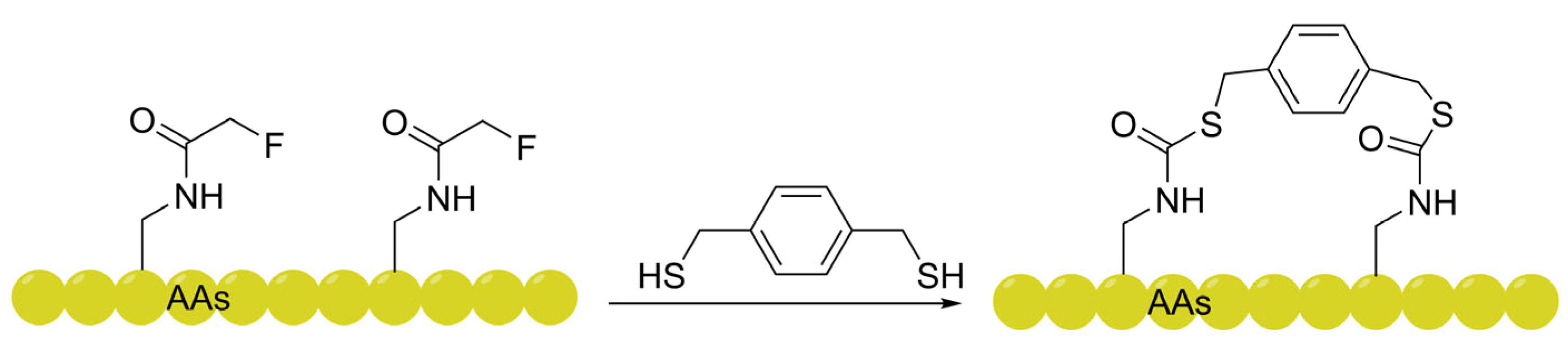
3.2. Fluorine-Thiol Displacement Reaction (FTDR): Leveraging Chameleonic Adaptability for Dynamic Shielding
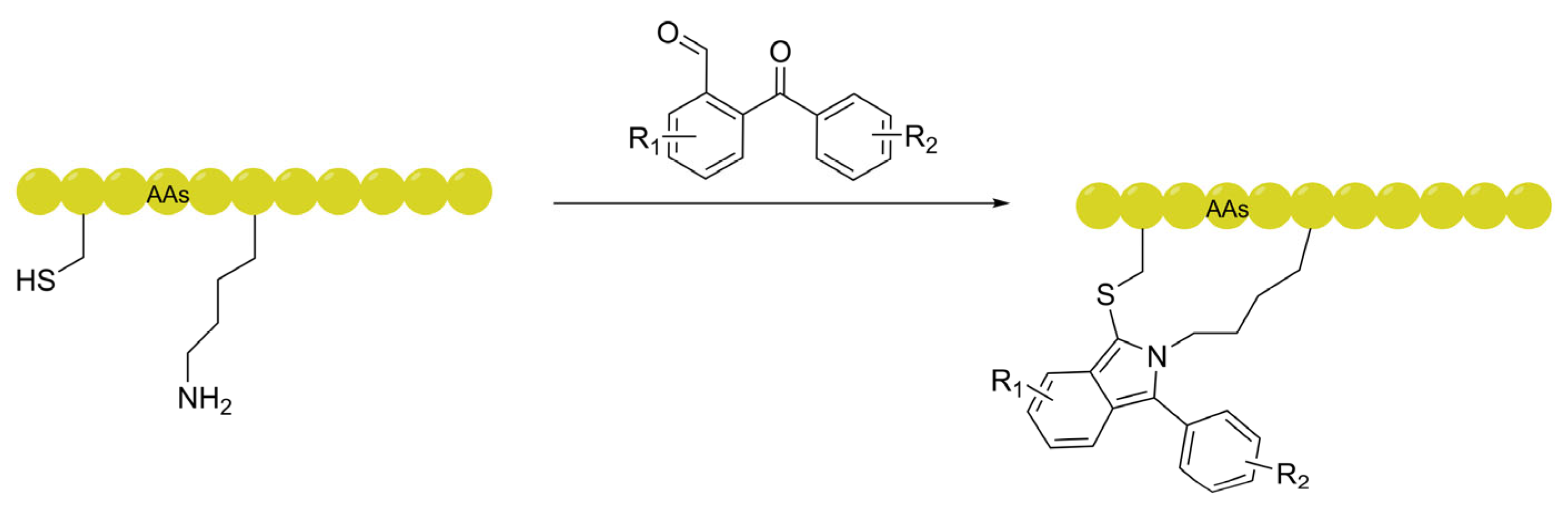
3.3. Self-Tracing Linkers: Eliminating Experimental Artifacts via Intrinsic Fluorescence
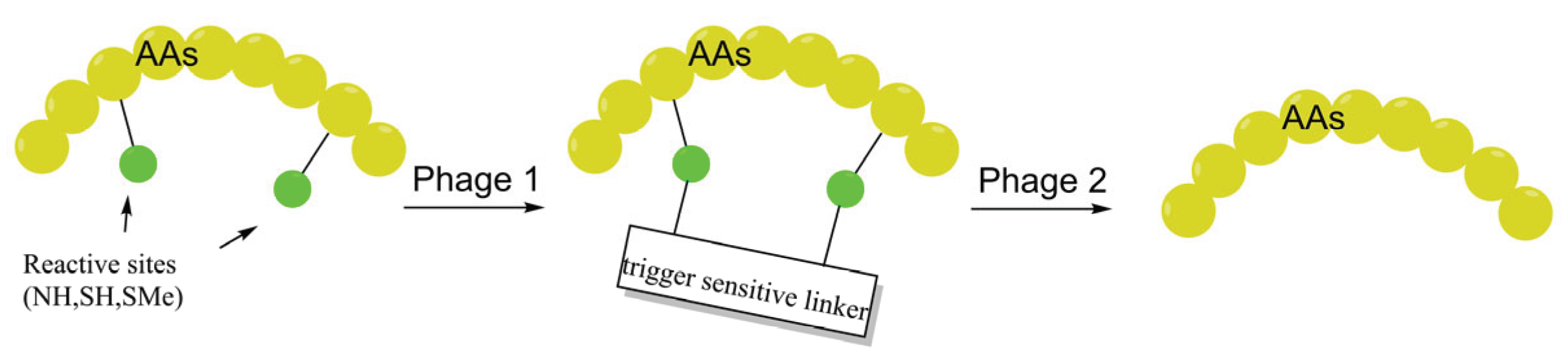
3.4. Stimuli-Responsive Reversible Staples: Harmonizing Permeability with Pharmacophore Regeneration
4. Discussion
5. Conclusions
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| PPI | Protein–protein interaction |
| HTS | High-throughput screening |
| HBDs | hydrogen-bond donors |
| DEL | DNA-encoded library |
| ArF | Perfluoroaryl |
| AlkF | perfluoroalkyl |
| IMHB | Intramolecular hydrogen bond |
| PSA | Polar surface area |
| RCM | Ring-closing metathesis |
| CPPs | cell-penetrating peptides |
| CLIPS | Chemical Linkage of Peptides onto Scaffolds |
| TBMB | 1,3,5-Tris(bromomethyl)benzene |
| BBB | Blood–brain barrier |
| BPy | Bipyridine |
| UAA | Unnatural amino acid |
| HSPG | Heparan Sulfate Proteoglycans |
| IP+ | Imidazopyridinium |
| PAMPA | Parallel Artificial Membrane Permeability Assay |
| CAPA | Chloroalkane Penetration Assay |
| FTDR | Fluorine–Thiol Displacement Reaction |
| OPA | ortho-phthalaldehyde |
| FlICk | Fluorescent Isoindole Crosslinking |
| AI | Artificial Intelligence |
References
- Greenblatt, J.F.; Alberts, B.M.; Krogan, N.J. Discovery and significance of protein-protein interactions in health and disease. Cell 2024, 187, 6501–6517. [Google Scholar] [CrossRef] [PubMed]
- Singh, K.; Ejaz, W.; Dutta, K.; Thayumanavan, S. Antibody Delivery for Intracellular Targets: Emergent Therapeutic Potential. Bioconjugate Chem 2019, 30, 1028–1041. [Google Scholar] [CrossRef] [PubMed]
- Dang, C.V.; Reddy, E.P.; Shokat, K.M.; Soucek, L. Drugging the 'undruggable' cancer targets. Nature reviews. Cancer 2017, 17, 502–508. [Google Scholar] [CrossRef]
- Hong, S.H.; Nguyen, T.; Ongkingco, J.F.; Nazzaro, A.; Arora, P.S. From Concepts to Inhibitors: A Blueprint for Targeting Protein-Protein Interactions. Chem Rev 2025, 125, 6819–6869. [Google Scholar] [CrossRef]
- Chan, A.; Tsourkas, A. Intracellular Protein Delivery: Approaches, Challenges, and Clinical Applications. BME frontiers 2024, 5, 35. [Google Scholar] [CrossRef]
- Buyanova, M.; Pei, D. Targeting intracellular protein-protein interactions with macrocyclic peptides. Trends Pharmacol Sci 2022, 43, 234–248. [Google Scholar] [CrossRef]
- Wang, S.; Faucher, F.F.; Bertolini, M.; Kim, H.; Yu, B.; Cao, L.; Roeltgen, K.; Lovell, S.; Shanker, V.; Boyd, S.D.; et al. Identification of Covalent Cyclic Peptide Inhibitors Targeting Protein-Protein Interactions Using Phage Display. bioRxiv: the preprint server for biology 2024, 2011–2024. [Google Scholar] [CrossRef] [PubMed]
- Otvos, L. The latest trends in peptide drug discovery and future challenges. Expert Opin Drug Dis 2024, 19, 869–872. [Google Scholar] [CrossRef]
- Anand, U.; Bandyopadhyay, A.; Jha, N.K.; Pérez De La Lastra, J.M.; Dey, A. Translational aspect in peptide drug discovery and development: An emerging therapeutic candidate. BioFactors (Oxford, England) 2023, 49, 251–269. [Google Scholar] [CrossRef]
- Muttenthaler, M.; King, G.F.; Adams, D.J.; Alewood, P.F. Trends in peptide drug discovery. Nature reviews. Drug discovery 2021, 20, 309–325. [Google Scholar] [CrossRef]
- Pei, D. Designing Cell-Permeable Peptide Therapeutics That Enter the Cell by Endocytosis. ACS symposium series. American Chemical Society 2022, 1417, 179–197. [Google Scholar] [CrossRef]
- Kremsmayr, T.; Aljnabi, A.; Blanco-Canosa, J.B.; Tran, H.N.T.; Emidio, N.B.; Muttenthaler, M. On the Utility of Chemical Strategies to Improve Peptide Gut Stability. J Med Chem 2022, 65, 6191–6206. [Google Scholar] [CrossRef]
- Rezai, T.; Yu, B.; Millhauser, G.L.; Jacobson, M.P.; Lokey, R.S. Testing the conformational hypothesis of passive membrane permeability using synthetic cyclic peptide diastereomers. J Am Chem Soc 2006, 128, 2510–2511. [Google Scholar] [CrossRef]
- Mathiowetz, A.M. Design Principles for Intestinal Permeability of Cyclic Peptides. Methods in molecular biology (Clifton, N.J.) 2019, 2001, 1–15. [Google Scholar] [CrossRef]
- Jiang, F.; Geng, H. Computational Methods for Studying Conformational Behaviors of Cyclic Peptides. Methods in molecular biology (Clifton, N.J.) 2019, 2001, 61–71. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.; Li, W.; Xu, Z. Improvement on Permeability of Cyclic Peptide/Peptidomimetic: Backbone N-Methylation as A Useful Tool. Mar Drugs 2021, 19, 311. [Google Scholar] [CrossRef] [PubMed]
- Chang, C.F.; Zehfus, M.H. The effect of N-methylation on helical peptides. Biopolymers 1996, 40, 609–616. [Google Scholar] [CrossRef]
- Chatterjee, J.; Rechenmacher, F.; Kessler, H. N-methylation of peptides and proteins: an important element for modulating biological functions. Angewandte Chemie (International ed. in English) 2013, 52, 254–269. [Google Scholar] [CrossRef]
- Ghosh, P.; Raj, N.; Verma, H.; Patel, M.; Chakraborti, S.; Khatri, B.; Doreswamy, C.M.; Anandakumar, S.R.; Seekallu, S.; Dinesh, M.B.; et al. An amide to thioamide substitution improves the permeability and bioavailability of macrocyclic peptides. Nat Commun 2023, 14, 6050. [Google Scholar] [CrossRef] [PubMed]
- Hosono, Y.; Uchida, S.; Shinkai, M.; Townsend, C.E.; Kelly, C.N.; Naylor, M.R.; Lee, H.; Kanamitsu, K.; Ishii, M.; Ueki, R.; et al. Amide-to-ester substitution as a stable alternative to N-methylation for increasing membrane permeability in cyclic peptides. Nat Commun 2023, 14, 1416. [Google Scholar] [CrossRef]
- Wang, C.K.; Swedberg, J.E.; Harvey, P.J.; Kaas, Q.; Craik, D.J. Conformational Flexibility Is a Determinant of Permeability for Cyclosporin. The journal of physical chemistry. B 2018, 122, 2261–2276. [Google Scholar] [CrossRef]
- Corbett, K.M.; Ford, L.; Warren, D.B.; Pouton, C.W.; Chalmers, D.K. Cyclosporin Structure and Permeability: From A to Z and Beyond. J Med Chem 2021, 64, 13131–13151. [Google Scholar] [CrossRef] [PubMed]
- Mathiowetz, A.M. Design Principles for Intestinal Permeability of Cyclic Peptides. Methods in molecular biology (Clifton, N.J.) 2019, 2001, 1–15. [Google Scholar] [CrossRef]
- Tang, X.; Kokot, J.; Waibl, F.; Fernández-Quintero, M.L.; Kamenik, A.S.; Liedl, K.R. Addressing Challenges of Macrocyclic Conformational Sampling in Polar and Apolar Solvents: Lessons for Chameleonicity. J Chem Inf Model 2023, 63, 7107–7123. [Google Scholar] [CrossRef] [PubMed]
- Fang, P.; Pang, W.; Xuan, S.; Chan, W.; Leung, K.C. Recent advances in peptide macrocyclization strategies. Chem Soc Rev 2024, 53, 11725–11771. [Google Scholar] [CrossRef] [PubMed]
- Bird, G.H.; Mazzola, E.; Opoku-Nsiah, K.; Lammert, M.A.; Godes, M.; Neuberg, D.S.; Walensky, L.D. Biophysical determinants for cellular uptake of hydrocarbon-stapled peptide helices. Nat Chem Biol 2016, 12, 845–852. [Google Scholar] [CrossRef]
- Li, L.; Li, R.; Jiang, Y.; Chao, J.; Chen, S.; Liao, H.; Li, X. Advances in Hydrocarbon Stapled Peptides via Ring-Closing Metathesis: Synthetic Strategies, Structural Diversity, and Therapeutic Applications. Chembiochem: a European journal of chemical biology 2025, 26, e202500527. [Google Scholar] [CrossRef]
- Chandramohan, A.; Josien, H.; Yuen, T.Y.; Duggal, R.; Spiegelberg, D.; Yan, L.; Juang, Y.A.; Ge, L.; Aronica, P.G.; Kaan, H.Y.K.; et al. Design-rules for stapled peptides with in vivo activity and their application to Mdm2/X antagonists. Nat Commun 2024, 15, 489. [Google Scholar] [CrossRef]
- Colas, K.; Bindl, D.; Suga, H. Selection of Nucleotide-Encoded Mass Libraries of Macrocyclic Peptides for Inaccessible Drug Targets. Chem Rev 2024, 124, 12213–12241. [Google Scholar] [CrossRef]
- Wang, X.S.; Chen, P.C.; Hampton, J.T.; Tharp, J.M.; Reed, C.A.; Das, S.K.; Wang, D.; Hayatshahi, H.S.; Shen, Y.; Liu, J.; et al. A Genetically Encoded, Phage-Displayed Cyclic-Peptide Library. Angewandte Chemie (International ed. in English) 2019, 58, 15904–15909. [Google Scholar] [CrossRef]
- Yao, X.; Aphicho, K.; Pani, S.; Rupanya, A.; Lan, T.; Dickinson, B.C. Discovery of Macrocyclic Peptide Binders, Covalent Modifiers, and Degraders of a Structured RNA by mRNA Display. J Am Chem Soc 2025, 147, 34256–34270. [Google Scholar] [CrossRef]
- Petrov, D.; Plais, L.; Schira, K.; Cai, J.; Keller, M.; Lessing, A.; Bassi, G.; Cazzamalli, S.; Neri, D.; Gloger, A.; et al. Flexibility-tuning of dual-display DNA-encoded chemical libraries facilitates cyclic peptide ligand discovery. Nat Commun 2025, 16, 3273. [Google Scholar] [CrossRef]
- Bai, L.; Dan, T.; Cheng, P.; Yang, X.; Xiang, H.; Zhai, W.; Chen, Y.; Huang, R.; Wang, Q.; Li, K.; et al. Isothiocyanate-mediated cyclization of phage-displayed peptides enables discovery of macrocyclic binders. Sci Adv 2026, 12, b7086. [Google Scholar] [CrossRef]
- Colas, K.; Bindl, D.; Suga, H. Selection of Nucleotide-Encoded Mass Libraries of Macrocyclic Peptides for Inaccessible Drug Targets. Chem Rev 2024, 124, 12213–12241. [Google Scholar] [CrossRef]
- Zhao, H.; Li, X.; Yao, X.; Zhang, S.; Bao, Y.; Lu, W.; Xing, M.; Wang, X.; Wang, X.; Zhao, Y.; et al. Influence of Macrocyclization Strategies on DNA-Encoded Cyclic Peptide Libraries. JACS Au 2025, 5, 3399–3407. [Google Scholar] [CrossRef] [PubMed]
- Chavali, S.S.; Mali, S.M.; Bonn, R.; Saseendran Anitha, A.; Bennett, R.P.; Smith, H.C.; Fasan, R.; Wedekind, J.E. Cyclic peptides with a distinct arginine-fork motif recognize the HIV trans-activation response RNA in?vitro and in cells. The Journal of biological chemistry 2021, 297, 101390. [Google Scholar] [CrossRef] [PubMed]
- Anananuchatkul, T.; Chang, I.V.; Miki, T.; Tsutsumi, H.; Mihara, H. Construction of a Stapled α-Helix Peptide Library Displayed on Phage for the Screening of Galectin-3-Binding Peptide Ligands. ACS omega 2020, 5, 5666–5674. [Google Scholar] [CrossRef]
- Walensky, L.D.; Kung, A.L.; Escher, I.; Malia, T.J.; Barbuto, S.; Wright, R.D.; Wagner, G.; Verdine, G.L.; Korsmeyer, S.J. Activation of apoptosis in vivo by a hydrocarbon-stapled BH3 helix. Science (New York, N.Y.) 2004, 305, 1466–1470. [Google Scholar] [CrossRef]
- Colas, K.; Bindl, D.; Suga, H. Selection of Nucleotide-Encoded Mass Libraries of Macrocyclic Peptides for Inaccessible Drug Targets. Chem Rev 2024, 124, 12213–12241. [Google Scholar] [CrossRef]
- Chen, F.; Pinnette, N.; Gao, J. Strategies for the Construction of Multicyclic Phage Display Libraries. Chembiochem: a European journal of chemical biology 2024, 25, e202400072. [Google Scholar] [CrossRef]
- Timmerman, P.; Beld, J.; Puijk, W.C.; Meloen, R.H. Rapid and quantitative cyclization of multiple peptide loops onto synthetic scaffolds for structural mimicry of protein surfaces. Chembiochem: a European journal of chemical biology 2005, 6, 821–824. [Google Scholar] [CrossRef]
- Li, Y.; Wu, M.; Fu, Y.; Xue, J.; Yuan, F.; Qu, T.; Rissanou, A.N.; Wang, Y.; Li, X.; Hu, H. Therapeutic stapled peptides: Efficacy and molecular targets. Pharmacol Res 2024, 203, 107137. [Google Scholar] [CrossRef] [PubMed]
- Muppidi, A.; Zhang, H.; Curreli, F.; Li, N.; Debnath, A.K.; Lin, Q. Design of antiviral stapled peptides containing a biphenyl cross-linker. Bioorg Med Chem Lett 2014, 24, 1748–1751. [Google Scholar] [CrossRef]
- Heinis, C.; Rutherford, T.; Freund, S.; Winter, G. Phage-encoded combinatorial chemical libraries based on bicyclic peptides. Nat Chem Biol 2009, 5, 502–507. [Google Scholar] [CrossRef]
- Villequey, C.; Zurmühl, S.S.; Cramer, C.N.; Bhusan, B.; Andersen, B.; Ren, Q.; Liu, H.; Qu, X.; Yang, Y.; Pan, J.; et al. An efficient mRNA display protocol yields potent bicyclic peptide inhibitors for FGFR3c: outperforming linear and monocyclic formats in affinity and stability. Chem Sci 2024, 15, 6122–6129. [Google Scholar] [CrossRef] [PubMed]
- Al Musaimi, O.; Lombardi, L.; Williams, D.R.; Albericio, F. Strategies for Improving Peptide Stability and Delivery. Pharmaceuticals (Basel, Switzerland) 2022, 15, 1283. [Google Scholar] [CrossRef] [PubMed]
- Luo, Y.; Schofield, J.A.; Na, Z.; Hann, T.; Simon, M.D.; Slavoff, S.A. Discovery of cellular substrates of human RNA-decapping enzyme DCP2 using a stapled bicyclic peptide inhibitor. Cell Chem Biol 2021, 28, 463–474. [Google Scholar] [CrossRef]
- Krishna Sudhakar, H.; Yau, J.T.K.; Alcock, L.J.; Lau, Y.H. Accessing diverse bicyclic peptide conformations using 1,2,3-TBMB as a linker. Org Biomol Chem 2024, 22, 6095–6102. [Google Scholar] [CrossRef] [PubMed]
- Kadota, K.; Kohata, A.; Sando, S.; Morimoto, J.; Aikawa, K.; Okazoe, T. Comparison of the effects of perfluoroalkyl and alkyl groups on cellular uptake in short peptides. Rsc Adv 2025, 15, 8189–8194. [Google Scholar] [CrossRef]
- Ni, C.; Hu, J. The unique fluorine effects in organic reactions: recent facts and insights into fluoroalkylations. Chem Soc Rev 2016, 45, 5441–5454. [Google Scholar] [CrossRef]
- Dognini, P.; Chaudhry, T.; Scagnetti, G.; Assante, M.; Hanson, G.S.M.; Ross, K.; Giuntini, F.; Coxon, C. R. 5,10,15,20-Tetrakis(pentafluorophenyl)porphyrin as a Functional Platform for Peptide Stapling and Multicyclisation. Chemistry (Weinheim an der Bergstrasse, Germany) 2023, 29, e202301410. [Google Scholar] [CrossRef]
- Miles, S.A.; Nillama, J.A.; Hunter, L. Tinker, Tailor, Soldier, Spy: The Diverse Roles That Fluorine Can Play within Amino Acid Side Chains. Molecules (Basel, Switzerland) 2023, 28, 6192. [Google Scholar] [CrossRef]
- Spokoyny, A.M.; Zou, Y.; Ling, J.J.; Yu, H.; Lin, Y.; Pentelute, B.L. A perfluoroaryl-cysteine S(N)Ar chemistry approach to unprotected peptide stapling. J Am Chem Soc 2013, 135, 5946–5949. [Google Scholar] [CrossRef]
- de Araujo, A.D.; Lim, J.; Wu, K.; Hoang, H.N.; Nguyen, H.T.; Fairlie, D.P. Landscaping macrocyclic peptides: stapling hDM2-binding peptides for helicity, protein affinity, proteolytic stability and cell uptake. RSC chemical biology 2022, 3, 895–904. [Google Scholar] [CrossRef]
- Kalhor-Monfared, S.; Jafari, M.R.; Patterson, J.T.; Kitov, P.I.; Dwyer, J.J.; Nuss, J.M.; Derda, R. Rapid biocompatible macrocyclization of peptides with decafluoro-diphenylsulfone. Chem Sci 2016, 7, 3785–3790. [Google Scholar] [CrossRef] [PubMed]
- Fadzen, C.M.; Wolfe, J.M.; Cho, C.; Chiocca, E.A.; Lawler, S.E.; Pentelute, B.L. Perfluoroarene-Based Peptide Macrocycles to Enhance Penetration Across the Blood-Brain Barrier. J Am Chem Soc 2017, 139, 15628–15631. [Google Scholar] [CrossRef] [PubMed]
- Peng, L.; Li, Q.; Li, Z.; Tian, Y.; Zhou, S. A Chemically Engineered Macrocyclic Peptide Shuttle Overcomes Blood-Brain Barrier in Aging Brain. Angewandte Chemie (International ed. in English) 2026, 65, e20279. [Google Scholar] [CrossRef] [PubMed]
- Chen, H.; Katoh, T.; Suga, H. Macrocyclic Peptides Closed by a Thioether-Bipyridyl Unit That Grants Cell Membrane Permeability. ACS bio & med chem Au 2023, 3, 429–437. [Google Scholar] [CrossRef]
- Peraro, L.; Deprey, K.L.; Moser, M.K.; Zou, Z.; Ball, H.L.; Levine, B.; Kritzer, J.A. Cell Penetration Profiling Using the Chloroalkane Penetration Assay. J Am Chem Soc 2018, 140, 11360–11369. [Google Scholar] [CrossRef]
- Zhan, W.; Duan, H.; Li, C. Recent Advances in Metal-Free Peptide Stapling Strategies. Chem & bio engineering 2024, 1, 593–605. [Google Scholar] [CrossRef]
- He, J.; Ghosh, P.; Nitsche, C. Biocompatible strategies for peptide macrocyclisation. Chem Sci 2024, 15, 2300–2322. [Google Scholar] [CrossRef]
- Hacker, D.E.; Abrigo, N.A.; Hoinka, J.; Richardson, S.L.; Przytycka, T.M.; Hartman, M.C.T. Direct, Competitive Comparison of Linear, Monocyclic, and Bicyclic Libraries Using mRNA Display. Acs Comb Sci 2020, 22, 306–310. [Google Scholar] [CrossRef]
- Kawamoto, S.A.; Coleska, A.; Ran, X.; Yi, H.; Yang, C.; Wang, S. Design of triazole-stapled BCL9 α-helical peptides to target the β-catenin/B-cell CLL/lymphoma 9 (BCL9) protein-protein interaction. J Med Chem 2012, 55, 1137–1146. [Google Scholar] [CrossRef] [PubMed]
- Gaikwad, V.; Choudhury, A.R.; Chakrabarti, R. Decoding the dynamics of BCL9 triazole stapled peptide. Biophys Chem 2024, 307, 107197. [Google Scholar] [CrossRef]
- Xu, W.; Lau, Y.H.; Fischer, G.; Tan, Y.S.; Chattopadhyay, A.; de la Roche, M.; Hyv Nen, M.; Verma, C.; Spring, D.R.; Itzhaki, L.S. Macrocyclized Extended Peptides: Inhibiting the Substrate-Recognition Domain of Tankyrase. J Am Chem Soc 2017, 139, 2245–2256. [Google Scholar] [CrossRef]
- Wu, Y.; Kaur, A.; Fowler, E.; Wiedmann, M.M.; Young, R.; Galloway, W.R.J.D.; Olsen, L.; Sore, H.F.; Chattopadhyay, A.; Kwan, T.T.; et al. Toolbox of Diverse Linkers for Navigating the Cellular Efficacy Landscape of Stapled Peptides. Acs Chem Biol 2019, 14, 526–533. [Google Scholar] [CrossRef]
- Lau, Y.H.; De Andrade, P.; Quah, S.T.; Rossmann, M.; Laraia, L.; Sköld, N.; Sum, T.J. Functionalised staple linkages for modulating the cellular activity of stapled peptides. Chem Sci 2014, 5, 1804–1809. [Google Scholar] [CrossRef]
- Liu, M.; Morewood, R.; Yoshisada, R.; Pascha, M.N.; Hopstaken, A.J.P.; Tarcoveanu, E.; Poole, D.A.R.; de Haan, C.A.M.; Nitsche, C.; Jongkees, S.A.K. Selective thiazoline peptide cyclisation compatible with mRNA display and efficient synthesis. Chem Sci 2023, 14, 10561–10569. [Google Scholar] [CrossRef] [PubMed]
- Takashi Tamura, M.I.Y.Y.; Kyosuke Tsumura, K.S.T.W. Selected Paper Chemical Synthesis and Cell-Free Expression of Thiazoline Ring-Bridged Cyclic Peptides and Their Properties on Biomembrane Permeability. B Chem Soc Jpn 2021, 95, 359–366. [Google Scholar] [CrossRef]
- Li, B.; Parker, J.; Tong, J.; Kodadek, T. Synthesis of Membrane-Permeable Macrocyclic Peptides via Imidazopyridinium Grafting. J Am Chem Soc 2024, 146, 14633–14644. [Google Scholar] [CrossRef]
- Nie, Q.; Xu, T.; Fang, X.; Dan, Y.; Zhang, G.; Li, Y.; Li, J.; Li, Y. The Furan-Thiol-Amine Reaction Facilitates DNA-Compatible Thiopyrrole-Grafted Macrocyclization and Late-Stage Amine Transformation. Org Lett 2025, 27, 498–503. [Google Scholar] [CrossRef]
- Bao, Y.; Xing, M.; Matthew, N.; Chen, X.; Wang, X.; Lu, X. Macrocyclizing DNA-Linked Peptides via Three-Component Cyclization and Photoinduced Chemistry. Org Lett 2024, 26, 2763–2767. [Google Scholar] [CrossRef] [PubMed]
- Chai, J.; Arico-Muendel, C.C.; Ding, Y.; Pollastri, M.P.; Scott, S.; Mantell, M.A.; Yao, G. Synthesis of a DNA-Encoded Macrocyclic Library Utilizing Intramolecular Benzimidazole Formation. Bioconjugate Chem 2023, 34, 988–993. [Google Scholar] [CrossRef]
- Brown, S.P.; Smith, A.B.R. Peptide/protein stapling and unstapling: introduction of s-tetrazine, photochemical release, and regeneration of the peptide/protein. J Am Chem Soc 2015, 137, 4034–4037. [Google Scholar] [CrossRef]
- Morewood, R.; Nitsche, C. Bioinspired peptide stapling generates stable enzyme inhibitors. Chemical communications (Cambridge, England) 2022, 58, 10817–10820. [Google Scholar] [CrossRef]
- Rivas, M.; Gevorgyan, V. Advances in Selected Heterocyclization Methods. Synlett: accounts and rapid communications in synthetic organic chemistry 2023, 34, 1554–1562. [Google Scholar] [CrossRef]
- Merz, M.L.; Habeshian, S.; Li, B.; David, J.G.L.; Nielsen, A.L.; Ji, X.; Il Khwildy, K.; Duany Benitez, M.M.; Phothirath, P.; Heinis, C. De novo development of small cyclic peptides that are orally bioavailable. Nat Chem Biol 2024, 20, 624–633. [Google Scholar] [CrossRef]
- Li, L.; Li, R.; Jiang, Y.; Chao, J.; Chen, S.; Liao, H.; Li, X. Advances in Hydrocarbon Stapled Peptides via Ring-Closing Metathesis: Synthetic Strategies, Structural Diversity, and Therapeutic Applications. Chembiochem: a European journal of chemical biology 2025, 26, e202500527. [Google Scholar] [CrossRef]
- Blackwell, H.E.; Grubbs, R.H. Highly Efficient Synthesis of Covalently Cross-Linked Peptide Helices by Ring-Closing Metathesis. Angewandte Chemie (International ed. in English) 1998, 37, 3281–3284. [Google Scholar] [CrossRef]
- Ma, B.; Liu, D.; Zheng, M.; Wang, Z.; Zhang, D.; Jian, Y.; Ma, J.; Fan, Y.; Chen, Y.; Gao, Y.; et al. Development of a Double-Stapled Peptide Stabilizing Both α-Helix and β-Sheet Structures for Degrading Transcription Factor AR-V7. JACS Au 2024, 4, 816–827. [Google Scholar] [CrossRef]
- Morgan, D.C.; McDougall, L.; Knuhtsen, A.; Jamieson, A.G. Development of Bifunctional, Raman Active Diyne-Girder Stapled α-Helical Peptides. Chemistry (Weinheim an der Bergstrasse, Germany) 2023, 29, e202300855. [Google Scholar] [CrossRef] [PubMed]
- Morgan, D.C.; McDougall, L.; Knuhtsen, A.; Buetow, L.; Steven, C.F.; Shepperson, O.A.; Huang, D.T.; Hulme, A.N.; Jamieson, A.G. Raman active diyne-girder conformationally constrained p53 stapled peptides bind to MDM2 for visualisation without fluorophores. RSC chemical biology 2025, 6, 394–403. [Google Scholar] [CrossRef] [PubMed]
- Islam, M.S.; Junod, S.L.; Zhang, S.; Buuh, Z.Y.; Guan, Y.; Zhao, M.; Kaneria, K.H.; Kafley, P.; Cohen, C.; Maloney, R.; et al. Unprotected peptide macrocyclization and stapling via a fluorine-thiol displacement reaction. Nat Commun 2022, 13, 350. [Google Scholar] [CrossRef]
- Maloney, R.; Junod, S.L.; Hagen, K.M.; Lewis, T.; Cheng, C.; Shajan, F.J.; Zhao, M.; Moore, T.W.; Truong, T.H.; Yang, W.; et al. Flexible fluorine-thiol displacement stapled peptides with enhanced membrane penetration for the estrogen receptor/coactivator interaction. The Journal of biological chemistry 2024, 300, 107991. [Google Scholar] [CrossRef]
- Dayanara, N.L.; Froelich, J.; Roome, P.; Perrin, D.M. Chemoselective, regioselective, and positionally selective fluorogenic stapling of unprotected peptides for cellular uptake and direct cell imaging. Chem Sci 2024, 16, 584–595. [Google Scholar] [CrossRef]
- Shi, X.; Zhao, R.; Jiang, Y.; Zhao, H.; Tian, Y.; Jiang, Y.; Li, J.; Qin, W.; Yin, F.; Li, Z. Reversible stapling of unprotected peptides via chemoselective methionine bis-alkylation/dealkylation. Chem Sci 2018, 9, 3227–3232. [Google Scholar] [CrossRef]
- Cai, X.; Zheng, W.; Shi, X.; Chen, L.; Liu, Z.; Li, Z. HBx-Derived Constrained Peptides Inhibit the Secretion of Hepatitis B Virus Antigens. Mol Pharmaceut 2018, 15, 5646–5652. [Google Scholar] [CrossRef]
- Zeng, Z.; Zhu, J.; Deng, X.; Chen, H.; Jin, Y.; Miclet, E.; Alezra, V.; Wan, Y. Customized Reversible Stapling for Selective Delivery of Bioactive Peptides. J Am Chem Soc 2022, 144, 23614–23621. [Google Scholar] [CrossRef]
- Qian, Z.; Rhodes, C.A.; McCroskey, L.C.; Wen, J.; Appiah-Kubi, G.; Wang, D.J.; Guttridge, D.C.; Pei, D. Enhancing the Cell Permeability and Metabolic Stability of Peptidyl Drugs by Reversible Bicyclization. Angewandte Chemie (International ed. in English) 2017, 56, 1525–1529. [Google Scholar] [CrossRef] [PubMed]
- Xiong, W.; Chen, P.; Liu, J.; Liao, Y.; Tang, M.; Zhao, J. Synthesis of Bis-urea-Bridged Cyclopeptides via Stapling of Unprotected Peptides. Org Lett 2025, 27, 14230–14235. [Google Scholar] [CrossRef] [PubMed]
- Neuville, M.; Bornez, M.M.; Bourgeais, M.; Samueli, H.; Mauran, L.; Goudreau, S.R.; Khatib, A.; Guichard, G.; Pasco, M. A General Synthesis Approach to Double-Guanidinium Stapled Peptides and Foldamers. Chemistry (Weinheim an der Bergstrasse, Germany) 2025, 31, e2273. [Google Scholar] [CrossRef]
- Neuville, M.; Bourgeais, M.; Buratto, J.; Saragaglia, C.; Li, B.; Galeano-Otero, I.; Mauran, L.; Varajao, L.; Goudreau, S.R.; Kauffmann, B.; et al. Optimal Stapling of a Helical Peptide-Foldamer Hybrid Using a C-Terminal 4-Mercaptoproline Enhances Protein Surface Recognition and Cellular Activity. Chemistry (Weinheim an der Bergstrasse, Germany) 2025, 31, e202403330. [Google Scholar] [CrossRef] [PubMed]
- Bakhshinejad, B.; Ghiasvand, S. A Beautiful Bind: Phage Display and the Search for Cell-Selective Peptides. Viruses 2025, 17, 975. [Google Scholar] [CrossRef] [PubMed]
- Monty, O.B.C.; Nyshadham, P.; Bohren, K.M.; Palaniappan, M.; Matzuk, M.M.; Young, D.W.; Simmons, N. Homogeneous and Functional Group Tolerant Ring-Closing Metathesis for DNA-Encoded Chemical Libraries. Acs Comb Sci 2020, 22, 80–88. [Google Scholar] [CrossRef] [PubMed]
- Blanco, C.O.; Cormier, S.K.; Koller, A.J.; Boros, E.; Fogg, D.E. Olefin Metathesis in Water: Speciation of a Leading Water-Soluble Catalyst Pinpoints Challenges and Opportunities for Chemical Biology. J Am Chem Soc 2025, 147, 9441–9448. [Google Scholar] [CrossRef]
- Zou, Z.; Kalvet, I.; Lozhkin, B.; Morris, E.; Zhang, K.; Chen, D.; Ernst, M.L.; Zhang, X.; Baker, D.; Ward, T.R. De novo design and evolution of an artificial metathase for cytoplasmic olefin metathesis. Nature catalysis 2025, 8, 1208–1219. [Google Scholar] [CrossRef] [PubMed]
| Linker Strategy & Examples | Primary Permeation Mechanism | Key Evaluation Assays | Ref. |
| Arylene-based (mDBMB, TBMB, BPh) | Enhances permeability via conformational rigidification; "topological condensation" reduces the molecular radius of gyration and entropic penalty. | FITC imaging, CETSA, Live-cell imaging. | [43] [47] |
| Perfluoro-aromatic (ArF, AlkF) | Leverages the "Fluorine Effect" to increase lipophilicity; drives lipid raft accumulation and triggers CMT. | 3D BBB sphere model, Flow cytometry, Confocal microscopy. | [53,54] [56,57] |
| Bipyridine (BPy) (BPy unit) | Pyridine nitrogens act as H-bond acceptors to form IMHBs with backbone amides, providing effective shielding and increasing lipophilicity. | CAPA, PAMPA. | [58] |
| Triazole-based (Dialkynyl linkers) | Employs a "decoupling" strategy to independently optimize permeability (via cationic motifs or CPPs) and binding affinity. | Confocal microscopy, Reporter gene assays. | [54,63,64] |
| Thiazoline (Cys-Nitrile condensation) | Converts an HBD into an HBA; induces a stable IMHB network in hydrophobic environments to compress the PSA and lower desolvation energy. | PAMPA, Temp-variable NMR, Computational simulations. | [69] |
| Imidazo-pyridinium (IP+ scaffold) | Acts as a "cationic chameleon" where the delocalized cation facilitates membrane enrichment while the fused-ring system enables "pseudo-hydrophobic sliding". | PAMPA, CAPA. | [70] |
| All-hydrocarbon (RCM-based staples) | Stabilizes α-helical structures to shield polar amides; utilizes metabolic-driven trapping where enzymatic cleavage of a Poly-Ala tail leads to cytosolic accumulation. | Nano-Click, LDH release, Cell Ratio (EC50/KD). | [28] [80,81,82] |
| FTDR-based (Aromatic dithiols) | Exhibits "chameleonic adaptability" with dynamic helicity increases (from 12% to 89%) upon membrane contact; bypasses HSPG-dependence via multiple endocytic pathways. | Flow cytometry, Trypan Blue quenching, Inhibitor panels. | [83,84] |
| Self-tracing (Isoindole, Diyne) | Eliminates artifacts from external dyes using intrinsic fluorescence (FlICk) or Raman signals in the "cell-silent" region. | Raman microscopy, Live-cell fluorescence imaging. | [85] |
| Stimuli-responsive (Sulfonium, Carbamate) | Enhances enrichment via cationic linkers followed by intracellular "unstapling" (GSH or H2O2 triggered) to regenerate the pharmacophore. | MFI, Co-localization, Flow cytometry. | [86,87,88,89] |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




