Submitted:
17 February 2026
Posted:
03 March 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
1.1. Issues and Context
1.2. Goal and Objectives
1.3. Suggested Method
1.4. Paper Structure
- Theoretical Framework: Formalization of the volumetric segmentation problem and presentation of the mathematical foundations of the preprocessing process, the custom architecture of the neural network and optimization functions.
- Implementation: A description of the software implementation of the proposed structure, including 3D data processing, tensor manipulation strategies, and a specific implementation of the proposed algorithms in Python/TensorFlow.
- Experimental Results: Presentation of quantitative performance indicators (Dice similarity coefficients) obtained by cross-validation with the exclusion of one subject, as well as a qualitative visual assessment of segmentation contours.
- Related Work: The proposed lightweight approach is compared with existing best practices.
- Conclusion: The results are summarized and the directions of further development are described.
2. Theoretical Aspect
2.1. Problem Formulation
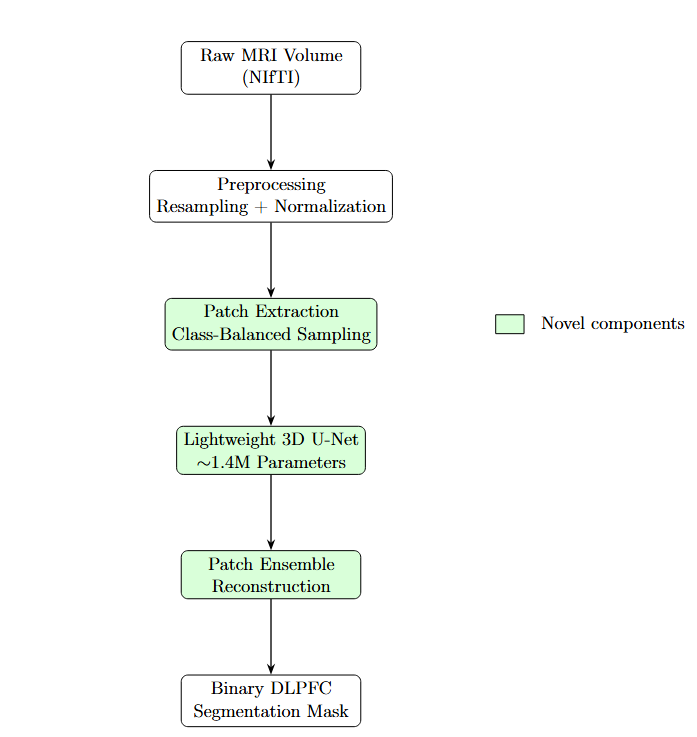
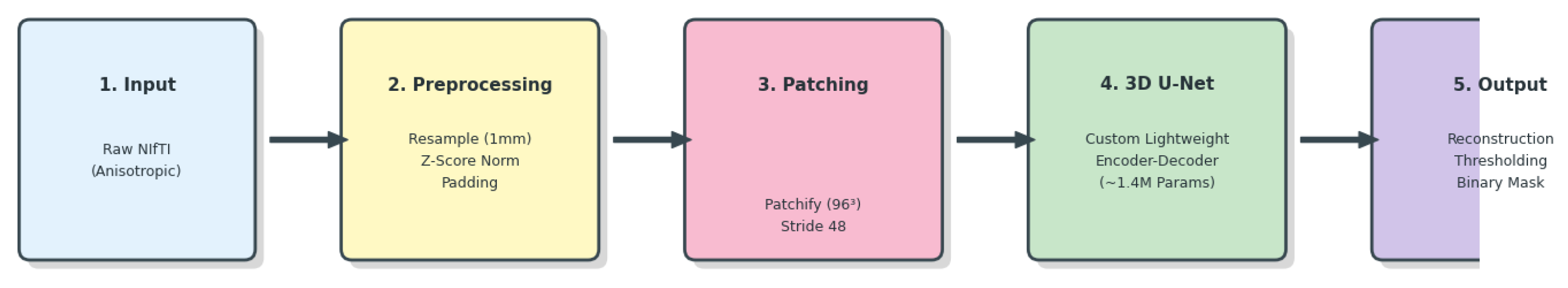
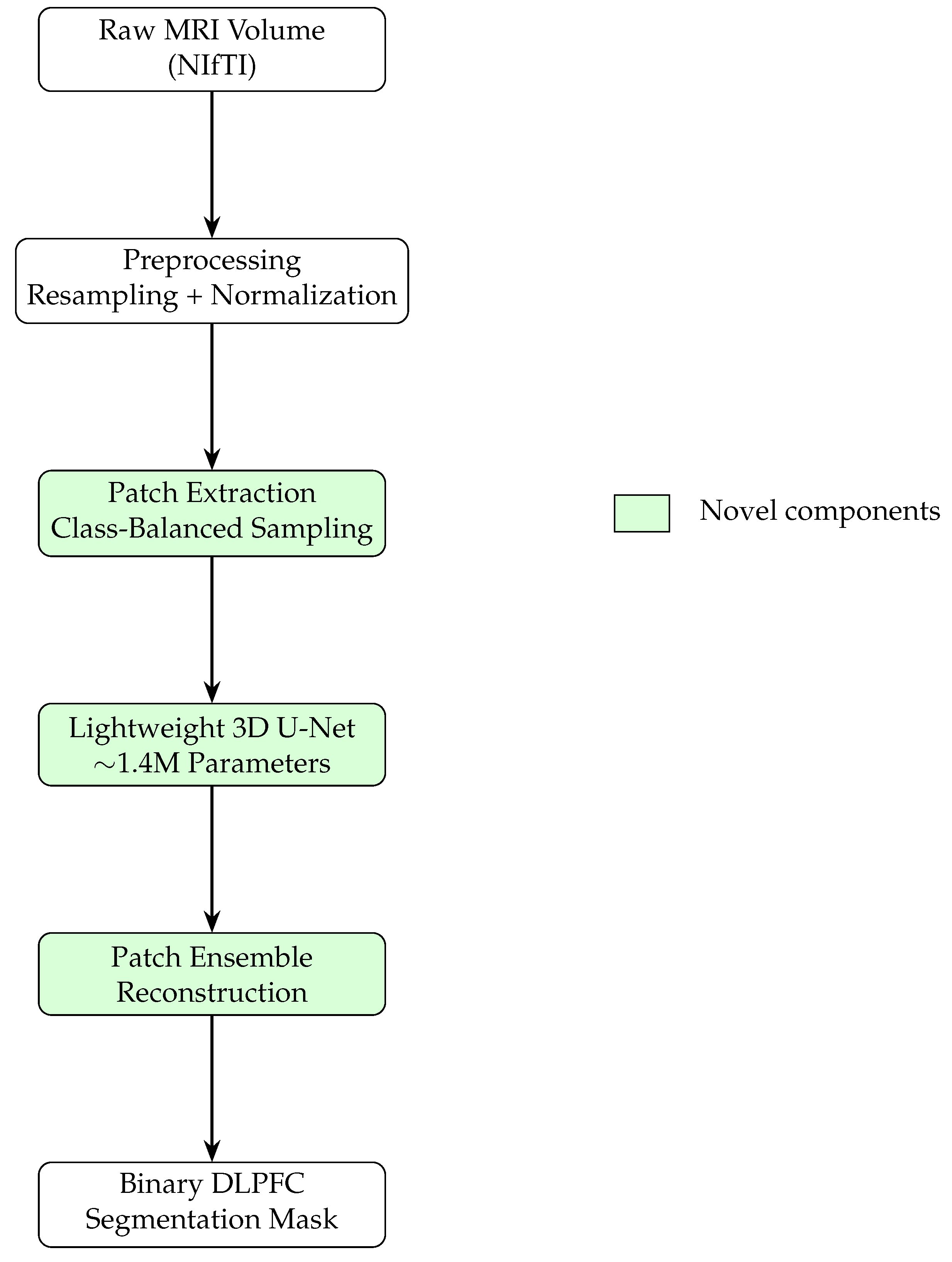
2.2. System Overview
2.3. Volumetric Preprocessing Models
Affine Registration and Resampling.
Z-Score Normalization.
2.4. Patch-Based Learning Strategy
2.5. Custom Lightweight 3D U-Net Architecture
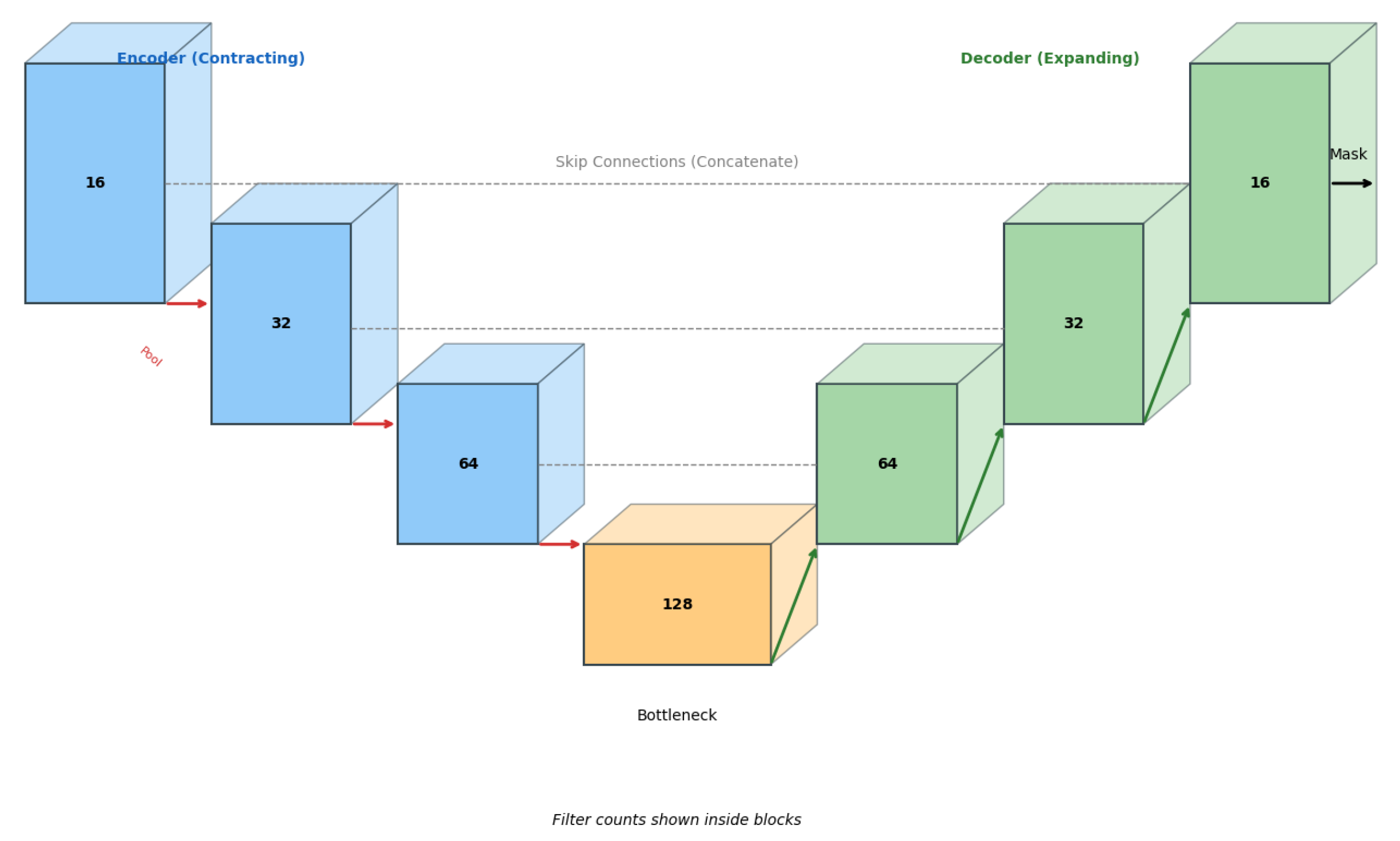
2.6. Optimization Functions
2.7. Mathematical Interpretation of Patch-Based Learning
2.8. Generalization in the Extreme Low-Data Regime
2.9. Novelty of the Proposed Method
- A lightweight 3D U-Net with ∼1.4M parameters,
- Aggressive class-balanced patch sampling,
- Training from scratch without pretraining,
- Patch ensemble reconstruction for robust inference.
3. Implementation
3.1. Development Environment and System Configuration
- TensorFlow/Keras (v2.x) for model definition, optimization, and automatic differentiation;
- SimpleITK for medical image input/output, resampling, and spatial transformations;
- Patchify for efficient extraction and reconstruction of volumetric patches;
- Nibabel for handling NIfTI data, affine transformations, and metadata preservation.
3.2. Tensor Manipulation and 3D Data Pipeline
3.2.1. Data Ingestion and Spatial Resampling
3.2.2. Dimensional Standardisation
3.2.3. Intensity Normalisation
3.3. Training Workflow Implementation
3.3.1. Patch Generation and Data Augmentation
3.3.2. Optimization and Training Control
3.4. Inference and Volumetric Reconstruction
3.5. Implementation Traceability
4. Experimental Results
4.1. Evaluation Metrics
4.2. Validation Protocol
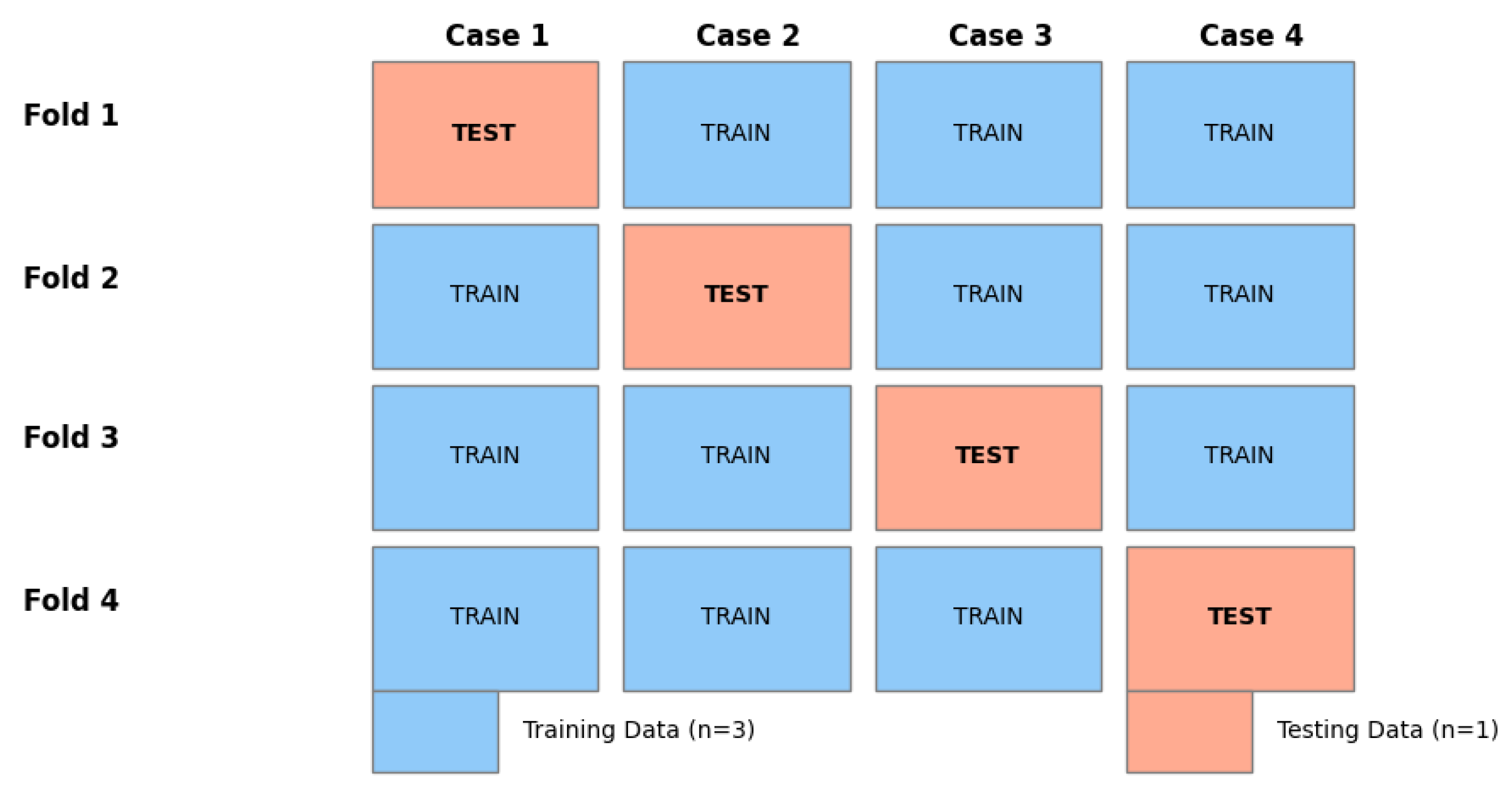
4.3. Quantitative Segmentation Performance
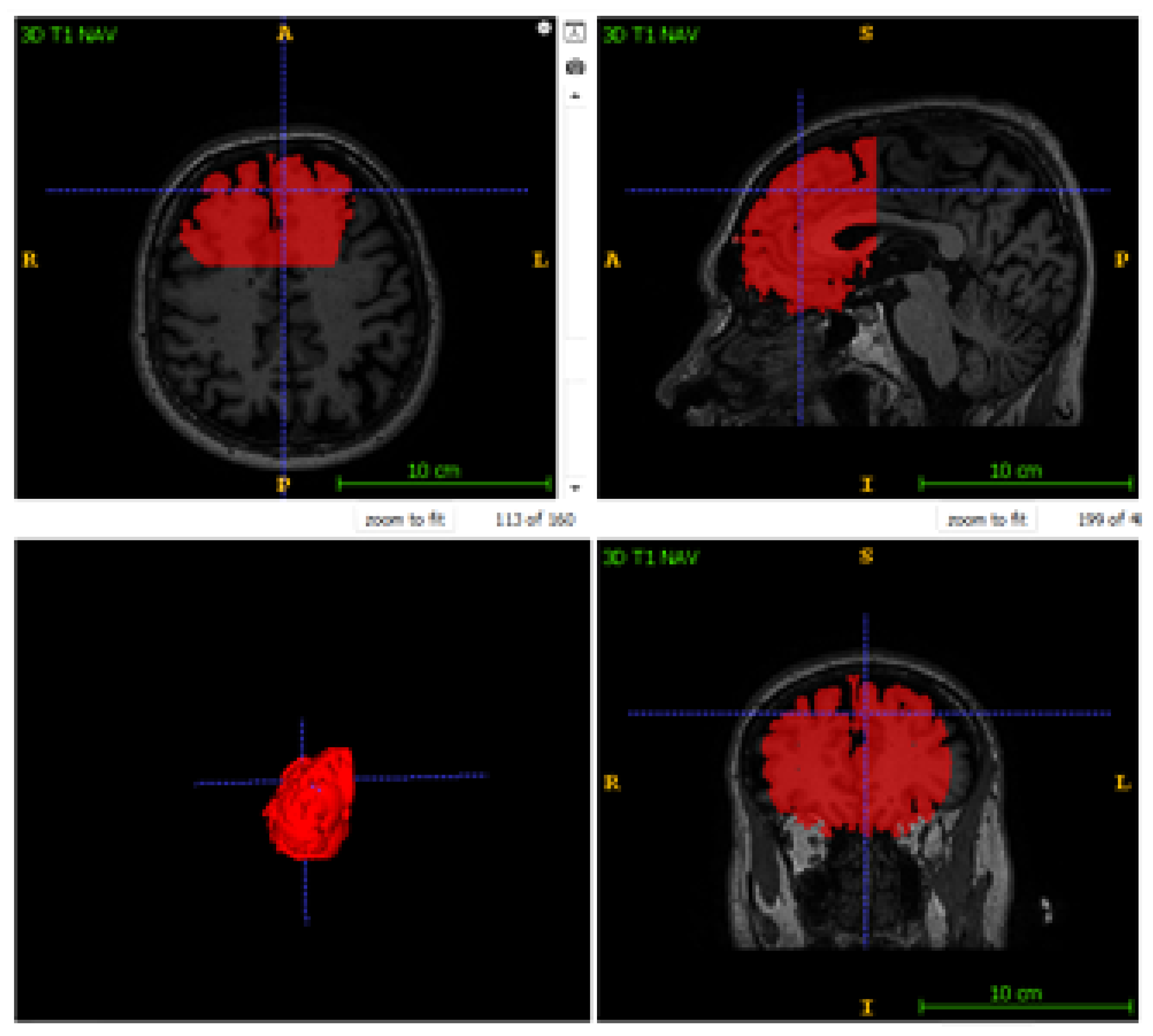
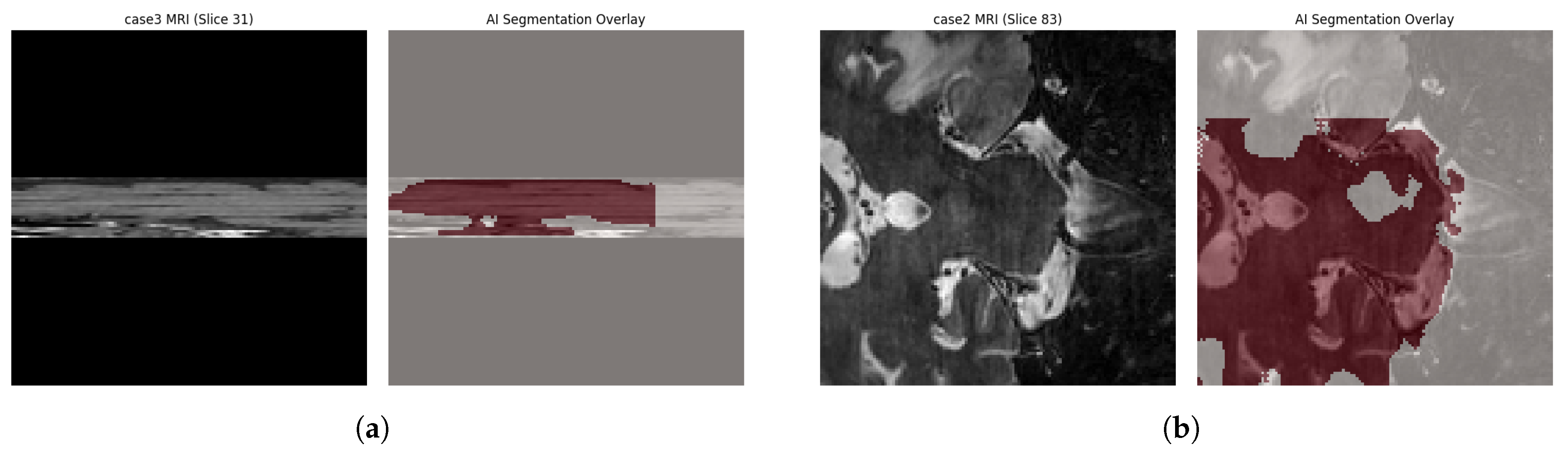
4.4. Qualitative Visual Assessment
4.5. Failure Mode and Error Pattern Analysis
4.6. Ablation Analysis of Design Choices
4.7. Efficiency of Computation and Operational Viability
5. Summary of Related Works and Discussion
5.1. Comparison with Other Methods
5.1.1. Comparison with Clinical Heuristics
5.1.2. Comparison with State-of-the-Art Deep Learning Models
- 1.
- Constrains Complexity: Uses only 1.4 million parameters (<10% of standard models).
- 2.
- Requires Minimal Data: Demonstrates convergence with only 4 subjects, whereas SOTA models fail or overfit without massive data augmentation.
- 3.
- Enables Accessibility: Deployable on consumer-grade hardware (standard clinical workstations) rather than specialized GPU servers.
5.2. Clinical Interpretation of Findings
5.2.1. Clinical Acceptability of Segmentation Precision
5.2.2. Workflow Integration
5.3. Ethical and Regulatory Considerations
6. Conclusion
Comparative Impact
7. Plan for Future Development
- 1.
- Dataset Scaling: To mitigate generalisation limitations observed in atypical anatomical cases, the dataset size will be expanded from to subjects. This will be achieved through multi-site data aggregation using publicly available repositories such as the Human Connectome Project (HCP) and OpenNeuro, thereby improving anatomical diversity and robustness.
- 2.
- Inference Optimisation: To ensure complete volumetric coverage during sliding-window inference, dynamic padding strategies will be incorporated. This enhancement will eliminate distal boundary artefacts observed in the current implementation and guarantee full brain-volume reconstruction.
- 3.
- Transition Toward Functional Targeting: While structural segmentation represents a necessary foundational step, the ultimate objective is functionally optimised rTMS targeting. Future extensions will integrate resting-state functional MRI (rs-fMRI) to identify subject-specific DLPFC subregions exhibiting maximal anti-correlation with the subgenual anterior cingulate cortex (sgACC).
References
- Fox, M. D.; Buckner, R. L.; White, M. P.; Greicius, M. D.; Pascual-Leone, A. Efficacy of TMS targets for depression is related to intrinsic functional connectivity with the subgenual cingulate. Biological Psychiatry 2012, vol. 72(no. 7), 595–603. [Google Scholar] [CrossRef] [PubMed]
- Herwig, U.; Satrapi, P.; Schönfeldt-Lecuona, C. Transcranial magnetic stimulation: Five-centimeter rule vs. neuronavigation. Psychiatry Research: Neuroimaging 2001, vol. 108(no. 1), 87–96. [Google Scholar]
- Ronneberger, O.; Fischer, P.; Brox, T. U-Net: Convolutional networks for biomedical image segmentation. International Conference on Medical Image Computing and Computer-Assisted Intervention (MICCAI), 2015; Springer; pp. 234–241. [Google Scholar]
- Çiçek, Ö.; Abdulkadir, A.; Lienkamp, S. S.; Brox, T.; Ronneberger, O. 3D U-Net: learning dense volumetric segmentation from sparse annotation. In MICCAI; Springer, 2016; pp. 424–432. [Google Scholar]
- Beam, W.; Borckardt, J. J.; Reeves, S. T.; George, M. S. An efficient method for positioning the coil for transcranial magnetic stimulation of the dorsolateral prefrontal cortex. Brain Stimulation 2009, vol. 2(no. 1), 50–54. [Google Scholar] [CrossRef] [PubMed]
- Milletari, F.; Navab, N.; Ahmadi, S. A. V-Net: Fully convolutional neural networks for volumetric medical image segmentation. 2016 Fourth International Conference on 3D Vision (3DV), 2016; IEEE; pp. 565–571. [Google Scholar]
- Mylius, V. Transcranial magnetic stimulation of the dorsolateral prefrontal cortex: variability in site selection and effects on pain perception. Brain Stimulation 2013, vol. 6(no. 5), 740–747. [Google Scholar]
- Yaniv, Z.; Lowekamp, B. C.; Johnson, H. J.; Beare, R. SimpleITK: a simplified layer over the Insight Segmentation and Registration Toolkit. Journal of Digital Imaging 2018, vol. 31(no. 2), 172–185. [Google Scholar]
- Tustison, N. J. N4ITK: Nick’s N3 ITK implementation for MRI bias field correction. IEEE Transactions on Medical Imaging 2010, vol. 29(no. 6), 1310–1320. [Google Scholar] [CrossRef] [PubMed]
- Dice, L. R. Measures of the amount of ecologic association between species. Ecology 1945, vol. 26(no. 3), 297–302. [Google Scholar] [CrossRef]

| Report Section | Functional Description | Script / Module | Key Functions / Code Components |
|---|---|---|---|
| 3.1 | Environment setup and reproducibility control | 3d_u_net.py | Random seed initialization; TensorFlow backend configuration |
| 3.2.1 | NIfTI ingestion and isotropic resampling | 3d_u_net.py | load_nifti(); sitk_resample_to_spacing() |
| 3.2.2 | Dimensional standardization | 3d_u_net.py | center_crop_or_pad() |
| 3.2.3 | Foreground-aware intensity normalization | 3d_u_net.py | zscore_normalize() |
| 3.3 | Streaming-based training pipeline | 3d_u_net.py | generator(); build_tf_dataset() |
| 3.3.1 | Patch extraction and data augmentation | 3d_u_net.py | sample_random_patch_from_volume(); rand_flip_rotate_3d() |
| 3.3.2 | Optimization and training control | 3d_u_net.py | Adam optimizer; bce_dice_loss(); Dice metric; Keras callbacks |
| 3.4 | Sliding-window inference and reconstruction | 3d_u_net.py | make_patches(); reconstruct_from_patches() |
| 3.4 | Thresholding and NIfTI export | 3d_u_net.py | Probability thresholding; Nifti1Image; save() |
| Validation Protocol | Leave-One-Subject-Out validation | 3d_u_net.py | Fold-wise training and evaluation loop |
| Validation Fold | Method | Dice Score | Interpretation |
|---|---|---|---|
| Fold 1 (Case 1) | Lightweight U-Net | 0.49 | Moderate Agreement |
| Fold 2 (Case 2) | Lightweight U-Net | 0.38 | Poor (Atypical Anatomy) |
| Fold 3 (Case 3) | Lightweight U-Net | 0.64 | Strong Agreement |
| Fold 4 (Case 4) | Lightweight U-Net | 0.62 | Strong Agreement |
| Mean | – | 0.53 ± 0.11 | – |
| Method | Processing Time (Mean) | Hardware |
|---|---|---|
| Manual Expert Tracing | min | Human Operator |
| AI Inference (Ours) | s | NVIDIA T4 GPU |
| Improvement Factor | ≈77× Faster | – |
| Method | MRI-based | Personalized | Time | Target Error |
|---|---|---|---|---|
| 5-cm Rule | No | No | <1 min | 20–30 mm |
| Beam F3 | No | No | <5 min | 15–25 mm |
| Manual Segmentation | Yes | Yes | 30–45 min | 2–3 mm |
| Proposed Method | Yes | Yes | 35 s | 3–6 mm |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




