Submitted:
16 February 2026
Posted:
28 February 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
Study Protocol
Clinical Evaluation
Ultrasound Evaluation
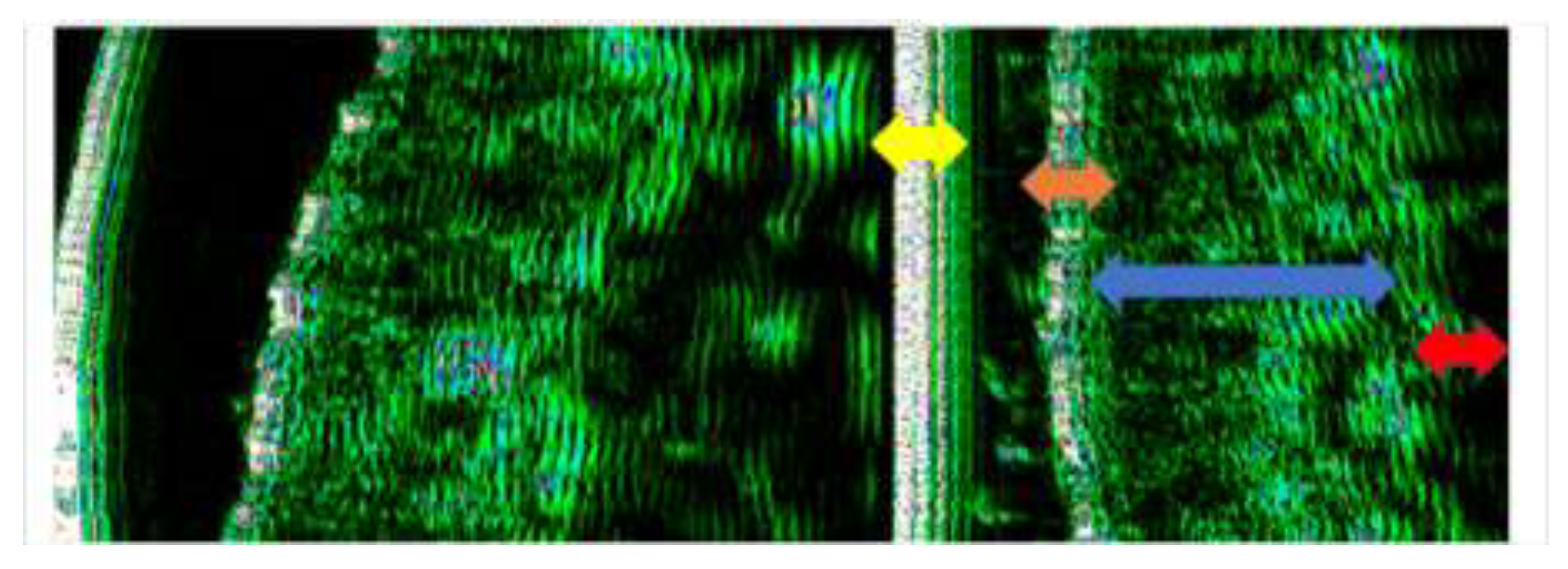

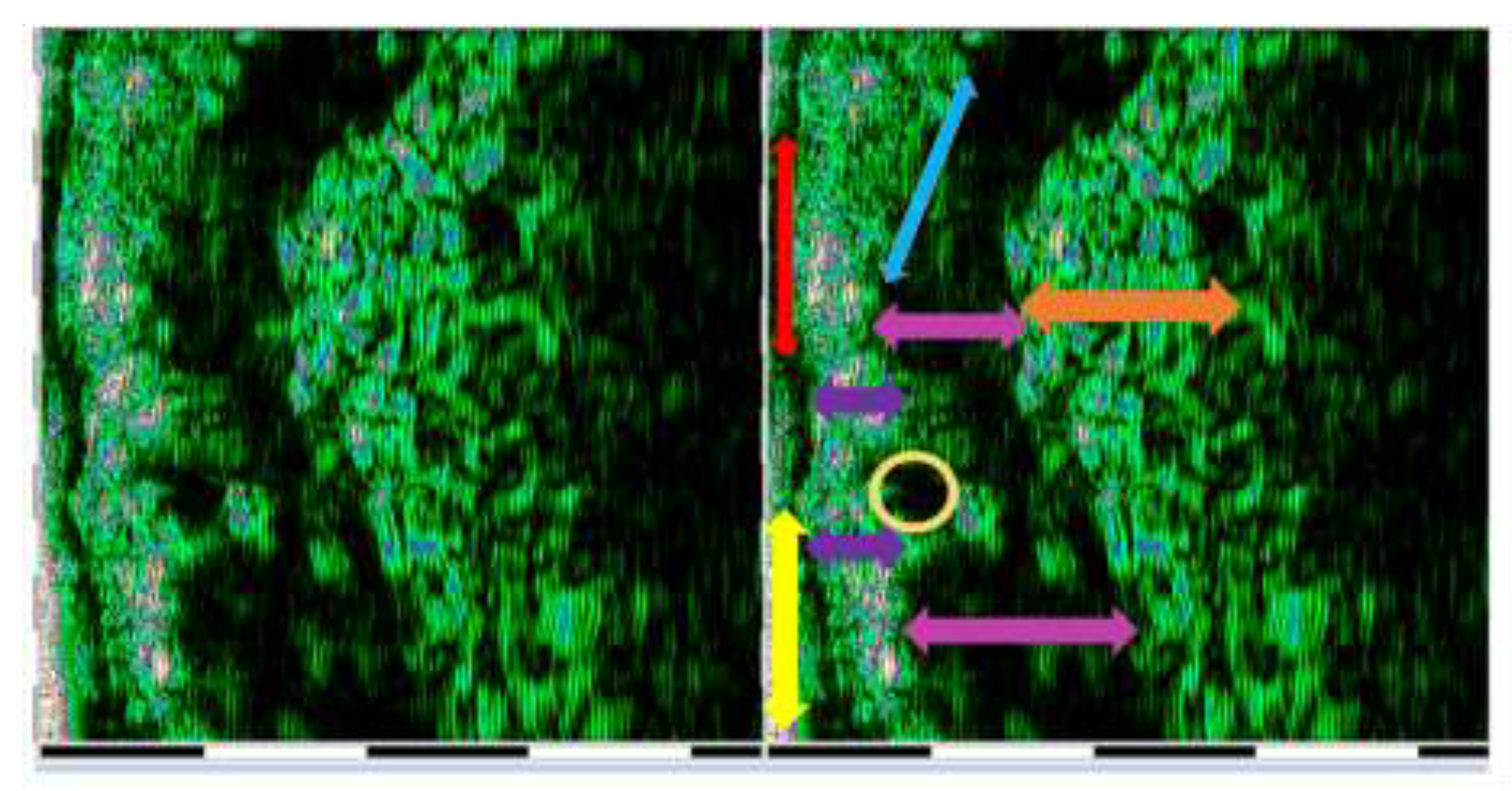
Statistical Analysis
3. Results
4. Discussion
5. Conclusions
Author Contributions
Funding
Informed Consent Statement
Conflicts of Interest
Abbreviations
References
- Constitution of the World Health Organization. Definition of Health. 2006. [Google Scholar] [CrossRef]
- Balkau, B.; Charles, M.A.; Drivsholm, T.; Borch-Johnsen, K.; Wareham, N.; Yudkin, J.S.; Morris, R.; Zavaroni., I.; van Dam, R.; Feskins, E.; et al. European Group For The Study Of Insulin Resistance (EGIR). Frequency of the WHO metabolic syndrome in European cohorts, and an alternative definition of an insulin resistance syndrome. Diabetes Metab 2002, 28, 364–376. [Google Scholar] [PubMed]
- Carrión-Barberà, I.; Triginer, L.; Tío, L.; Pérez-García, C.; Ribes, A.; Abad, V.; Pros, A.; Bermúdez-López, M.; Castro-Boqué, E.; Lecube, A.; et al. Role of Advanced Glycation End Products as New Biomarkers in Systemic Lupus Erythematosus. Int J Mol Sci 2024, 25, 3022. [Google Scholar] [CrossRef] [PubMed]
- Israelsen, N.M.; Michael, M.; Mogensen, M.; Bojesen, S.; Jensen, M.; Hædersdal, M.; Podoleanu, A.; Bang, O. The value of ultrahigh resolution OCT in dermatology - delineating the dermo-epidermal junction, capillaries in the dermal papillae and vellus hairs. Physics 2018. [Google Scholar] [CrossRef] [PubMed]
- Momma, H.; Niu, K.; Kobayashi, Y.; Guan, L.; Sato, M.; Guo, H.; Chujo, M.; Otomo, A.; Yufei, C.; Tadaura, H.; et al. Skin advanced glycation end-product accumulation is negatively associated with calcaneal osteo-sono assessment index among non-diabetic adult Japanese men. Osteoporos Int 2012, 23, 1673–1681. [Google Scholar] [CrossRef] [PubMed]
- Chen, J.; Arshi, B.; Waqas, K.; Lu, T.; Bos, D.; Ikram, M.A.; Uitterlinden, A.G.; Kavousi, M.; Zillikens, M.C. Advanced glycation end products measured by skin autofluorescence and subclinical cardiovascular disease: The Rotterdam Study. Cardiovasc Diabetol 2023, 22, 326. [Google Scholar] [CrossRef]
- Băbțan, A.M.; Vesa, Ș.C.; Boșca, B.A.; Crișan, M.; Mihu, C.M.; Băciuț, M.F.; Dinu, C.; Crișan, B.; Câmpian, R.S.; Feurdean, C.N.; et al. High-Frequency Ultrasound Assessment of Skin and Oral Mucosa in Metabolic Syndrome Patients-A Cross-Sectional Study. J Clin Med 2021, 10, 4461. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Alimova, S.; Sharobaro, V.; Yukhno, A.; Bondarenko, E. Possibilities of ultrasound examination in the assessment of age-related changes in the soft tissues of the face and neck: A review. Appl Sci 2023. [Google Scholar] [CrossRef]
- Machet, L.; et al. High resolution ultrasound imaging of melanocytic and other pigmented lesions of the skin. Ultrasound Imaging 2021. [Google Scholar] [CrossRef]
- Czajkowska, J.; Juszczyk, J.; Bugdol, M.N.; et al. High-frequency ultrasound in anti-aging skin therapy monitoring. Sci Rep 2023, 13, 17799. [Google Scholar] [CrossRef]
- Anderson, T. “Producte der trocknen Destillation thierischer Materien” [Products of the dry distillation of animal matter]. Annalen der Chemie und Pharmacie (in German). 1849, 70, 32–38. [Google Scholar] [CrossRef]
- Aeschbacher, H.U.; Wolleb, U.; Löliger, J.; Spadone, J.C.; Liardon, R. Contribution of coffee aroma constituents to the mutagenicity of coffee”. Food and Chemical Toxicology 1989, 27(4), 227–232. [Google Scholar] [CrossRef] [PubMed]
- Buttery, R.G.; Seifert, R.M.; Guadagni, D.G.; Ling, L.C. Characterization of Volatile Pyrazine and Pyridine Components of Potato Chips. In Journal of Agricultural and Food Chemistry; ACS: Washington, DC, 1971; Volume 19, pp. 969–971. [Google Scholar]
- Ho, C.T.; Lee, K.N.; Jin, Q.Z. Isolation and identification of volatile flavor compounds in fried bacon. Journal of Agricultural and Food Chemistry 1983, 31(2), 336. [Google Scholar] [CrossRef]
- Kostelc, J.G.; Preti, G.; Nelson, P.R.; Brauner, L.; Baehni, P. Oral Odors in Early Experimental Gingivitis”. Journal of Periodontal Research 1984, 19(3), 303–312. [Google Scholar] [CrossRef]
- Muraoka, M.Y.; Justino, A.B.; Caixeta, D.C.; Queiroz, J.S.; Sabino-Silva, R.; Espindola Salmen, F. Fructose and methylglyoxal-induced glycation alters structural and functional properties of salivary proteins, albumin and lysozyme. PLoS One 2022, 17, e0262369. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Morris, S. Arginine Metabolism Revisited. J Nutr 2016, 146, 2579S–2586S. [Google Scholar] [CrossRef]
- Kleinebenne, L.; Röhrig, W.; Ebach, F.; Reutter, H.; Pankraz, A.; Heuchel, K.M.; Müller, A.; Hilger, A.C. The components of arginine and methylarginine metabolism are indicative of altered kidney function in intrauterine growth-restricted neonates. J Hypertens 2024, 42, 1940–1947. [Google Scholar] [CrossRef]
- Emrich, I.E.; Zawada, A.M.; Martens-Lobenhoffer, J.; Fliser, D.; Wagenpfeil, S.; Heine, G.H.; Bode-Boger, S.M. Symmetric dimethylarginine (SDMA) outperforms asymmetric dimethylarginine (ADMA) and other methylarginines as predictor of renal and cardiovascular outcome in nondialysis chronic kidney disease. Clin Res Cardiol 2018, 107, 201–213. [Google Scholar] [CrossRef]
- Rabbani, N.; Thornalley, P.J. Advanced glycation end products in the pathogenesis of chronic kidney disease. Kidney Int 2018, 93, 803–813. [Google Scholar] [CrossRef]
- Kaleta, D.; Polańska, K.; Korytkowski, P.; Usidame, B.; Bąk-Romaniszyn, L. Patterns of nicotine dependence in four Eastern European countries. BMC Public Health 2015, 15, 1189. [Google Scholar] [CrossRef]
- Maciejczyk, M.; Nesterowicz, M.; Szulimowska, J.; Zalewska, A. Oxidation, Glycation, and Carbamylation of Salivary Biomolecules in Healthy Children, Adults, and the Elderly: Can Saliva Be Used in the Assessment of Aging? J Inflamm Res 2022, 15, 2051–2073. [Google Scholar] [CrossRef]
- Maciejczyk, M.; Bielas, M.; Zalewska, A.; Gerreth, K. Salivary Biomarkers of Oxidative Stress and Inflammation in Stroke Patients: From Basic Research to Clinical Practice. Oxid Med Cell Longev 2021, 2021, 5545330. [Google Scholar] [CrossRef]
- Christidis, G.; Küppers, F.; Karatayli, S.C.; et al. Skin advanced glycation end-products as indicators of the metabolic profile in diabetes mellitus: Correlations with glycemic control, liver phenotypes and metabolic biomarkers. BMC Endocr Disord 2024, 24, 31. [Google Scholar] [CrossRef]
- Monnier, V.M.; Sun, W.; Gao, X.; Sell, D.R.; Cleary, P.A.; Lachin, J.M.; et al. Skin collagen advanced glycation endproducts (AGEs) and the long-term progression of subclinical cardiovascular disease. Diabetes Care 2015, 38, 1170–1175. [Google Scholar]
- Gkogkolou, P.; Böhm, M. Advanced glycation end products: Key players in skin aging? Dermatoendocrinol 2012, 4, 259–270. [Google Scholar] [CrossRef]
- Du, T.; Brandl, B.; Hauner, H.; Skurk, T. Skin Autofluorescence Mirrors Surrogate Parameters of Vascular Aging: An Enable Study. Nutrients 2023, 15, 1597. [Google Scholar] [CrossRef] [PubMed]
- Meerwaldt, R.; Hartog, J.W.; Graaff, R.; Huisman, R.J.; Links, T.P.; den Hollander, N.C.; et al. Skin autofluorescence, a measure of cumulative metabolic stress and advanced glycation end products, predicts mortality in hemodialysis patients. J Am Soc Nephrol 2005, 6, 3687–3693. [Google Scholar] [CrossRef] [PubMed]
- Petrofsky, J.S.; Lee, S.; Hamdan, T.; et al. High-frequency ultrasound analysis of skin structure and its relation to glycation. J Clin Med 2021, 10, 4461. [Google Scholar]
- Saremi, A.; Howell, S.; Schwenke, D.C.; Bahn, G.; Beisswenger, P.J.; Reaven, P.D. for the VADT Investigators. Advanced Glycation End Products, Oxidation Products, and the Extent of Atherosclerosis During the VA Diabetes Trial and Follow-up Study. Diabetes Care 2017, 40, 591–598. [Google Scholar] [CrossRef]
- Zhao, X.W.; Yue, W.X.; Zhang, S.W.; Chen, Q. Correlation between the accumulation of skin glycosylation end products and the development of type 2 diabetic peripheral neuropathy. BMC Endocr Disord 2022, 22, 106. [Google Scholar] [CrossRef]
- Reurean-Pintilei, D.; Stoian, A.P.; Potcovaru, C.G.; et al. Skin autofluorescence as a potential adjunctive marker for cardiovascular risk assessment in type 2 diabetes: A systematic review. Int J Mol Sci 2024, 25, 3889. [Google Scholar] [CrossRef]
- Martinovic, D.; Tokic, D.; Usljebrka, M.; Lupi-Ferandin, S.; Cigic, L.; Vanjaka Rogosic, L.; Ercegovic, S.; Kontic, M.; Kumrić, M.; Rusic, D.; et al. The Association between the Level of Advanced Glycation End Products and Objective Skin Quality Parameters. Life 2023, 13, 256. [Google Scholar] [CrossRef]
| Characteristic | N (=113) | |
|---|---|---|
| Age | 52 (36; 60) | |
| Gender | Male | 37 (32.7%) |
| Female | 76 (67.3%) | |
| Epidermis depth | 281 (243; 325) | |
| Epidermis nr pixels | 5361.5 (4062; 4760) | |
| Epidermis density | 63.31 (47.61; 82.69) | |
| Epidermis density % | 26.1300 (19.8; 32.7) | |
| Uv dermis exposure damage depth uv aged dermis | 547 (394.5; 684) | |
| Uv dermis exposure damage nr pixels | 8806 (6312.5; 10979) | |
| Uv dermis exposure damage density | 9.8 (5.4; 17.5) | |
| Uv dermis exposure damage density % | 8.4 (6; 14.6) | |
| Dermis Depth | 1450 (1269.5; 1680) | |
| Dermis nr pixels | 23681 (20185; 27083.5) | |
| Dermis density % | 19.8 (15.2; 26.2) | |
| Sc tissue depth | 1594 (1094; 2164) | |
| Sc tissue nr pixels | 25320 (18000; 33201) | |
| Sc tissue density | 7.5 (5.2; 12.1) | |
| plasma FruLys [µg/ml] | 267 (146.7; 423.2) | |
| plasma Pyr [ng/ml] | 28.3 (24.9; 28.9) | |
| plasma MG-H1 [ng/ml] | 33.1 (31.3; 36.1) | |
| plasma CEL [ng/ml] | 13.2 (12.9; 13.4) | |
| plasma CML [ng/ml] | 49.1 (46.2; 51.8) | |
| plasma Arg [µg/ml] | 3.6 (2.6; 6.5) | |
| plasma Lys [µg/ml] | 6 (5.2; 7.2) | |
| serum FruLys [µg/ml] | 293.5 (205; 380.9) | |
| serum Pyr [ng/ml] | 28.4 (25.2; 29) | |
| serum MG-H1 [ng/ml] | 33.2 (31.3; 36.3) | |
| serum CEL [ng/ml] | 13.3 (12.8; 13.6) | |
| serum CML [ng/ml] | 46.9 (44.1; 49) | |
| serum Arg [µg/ml] | 5.4 (4.1; 7) | |
| serum Lys [µg/ml] | 6.4 (5.7; 7.6) | |
| urine FruLys [µg/mg Kreatinin] | 3.2 (2.46; 5.3) | |
| urine Pyr [µg/mg Kreatinin] | 0.11 (0.07; 0.18) | |
| urine MG-H1 [µg/mg Kreatinin] | 3.2 (2.1; 5.1) | |
| urine CEL [µg/mg Kreatinin] | 0.3 (0.22; 0.38) | |
| urine CML [µg/mg Kreatinin] | 1.4 (1.1; 2.1) | |
| urine Arg [µg/mg Kreatinin] | 2.2 (1.6; 2.9) | |
| urine Lys [µg/mg Kreatinin] | 8.1 (4.9; 14.5) | |
| saliva FruLys [ng/ml] | 21.9 (21; 24.3) | |
| saliva Pyr [ng/ml] | 1.7 (1.6; 1.7) | |
| saliva MG-H1 [ng/ml] | 3 (2.5; 4.2715) | |
| saliva CEL [ng/ml] | 1.1 (1; 1.3) | |
| saliva CML [ng/ml] | 3.9 (3.2; 4.8) | |
| saliva Arg [µg/ml] | 0.49 (0.24; 0.83) | |
| saliva Lys [µg/ml] | 0.59 (0.26; 0.89) | |
| Variable | EPIDERMIS DEPTH (N=113) | |
|---|---|---|
| R | p | |
| plasma FruLys [µg/ml] | 0.170 | 0.071 |
| plasma Pyr [ng/ml] | -0.167 | 0.077 |
| plasma MG-H1 [ng/ml] | 0.208* | 0.027 |
| plasma CEL [ng/ml] | 0.055 | 0.564 |
| plasma CML [ng/ml] | 0.181 | 0.055 |
| plasma Arg [µg/ml] | -0.189* | 0.045 |
| plasma Lys [µg/ml] | -0.170 | 0.071 |
| serum FruLys [µg/ml] | 0.011 | 0.904 |
| serum Pyr [ng/ml] | -0.128 | 0.177 |
| serum MG-H1 [ng/ml] | -0.047 | 0.625 |
| serum CEL [ng/ml] | -0.248** | 0.008 |
| serum CML [ng/ml] | -0.402** | 0.000 |
| serum Arg [µg/ml] | -0.089 | 0.349 |
| serum Lys [µg/ml] | -0.138 | 0.145 |
| saliva FruLys [ng/ml] | 0.045 | 0.637 |
| saliva Pyr [ng/ml] | 0.051 | 0.593 |
| saliva MG-H1 [ng/ml] | 0.204* | 0.030 |
| saliva CEL [ng/ml] | -0.011 | 0.907 |
| saliva Arg [µg/ml] | 0.223* | 0.018 |
| saliva Lys [µg/ml] | 0.115 | 0.224 |
| urine FruLys [µg/mg Kreatinin] | 0.048 | 0.612 |
| urine Pyr [µg/mg Kreatinin] | 0.034 | 0.725 |
| urine MG-H1 [µg/mg Kreatinin] | 0.109 | 0.250 |
| urine CEL [µg/mg Kreatinin] | 0.157 | 0.097 |
| urine CML [µg/mg Kreatinin] | 0.159 | 0.093 |
| urine Arg [µg/mg Kreatinin] | 0.212* | 0.024 |
| urine Lys [µg/mg Kreatinin] | -0.027 | 0.780 |
| Variable | EPIDERMIS DENSITY (N=113) | |
|---|---|---|
| r | p | |
| plasma FruLys [µg/ml] | 0.066 | 0.488 |
| plasma Pyr [ng/ml] | -0.010 | 0.913 |
| plasma MG-H1 [ng/ml] | -0.032 | 0.734 |
| plasma CEL [ng/ml] | -0.056 | 0.554 |
| plasma CML [ng/ml] | -0.045 | 0.639 |
| plasma Arg [µg/ml] | 0.006 | 0.949 |
| plasma Lys [µg/ml] | -0.019 | 0.838 |
| serum FruLys [µg/ml] | 0.111 | 0.241 |
| serum Pyr [ng/ml] | -0.082 | 0.386 |
| serum MG-H1 [ng/ml] | -0.003 | 0.972 |
| serum CEL [ng/ml] | 0.015 | 0.877 |
| serum CML [ng/ml] | 0.031 | 0.744 |
| serum Arg [µg/ml] | 0.000 | 0.999 |
| serum Lys [µg/ml] | -0.004 | 0.962 |
| saliva FruLys [ng/ml] | 0.045 | 0.637 |
| saliva Pyr [ng/ml] | 0.051 | 0.593 |
| saliva MG-H1 [ng/ml] | 0.204* | 0.030 |
| saliva CEL [ng/ml] | -0.011 | 0.907 |
| saliva Arg [µg/ml] | 0.223* | 0.018 |
| saliva Lys [µg/ml] | 0.115 | 0.224 |
| urine FruLys [µg/mg Kreatinin] | 0.018 | 0.852 |
| urine Pyr [µg/mg Kreatinin] | -0.003 | 0.975 |
| urine MG-H1 [µg/mg Kreatinin] | 0.012 | 0.896 |
| urine CEL [µg/mg Kreatinin] | 0.086 | 0.366 |
| urine CML [µg/mg Kreatinin] | 0.160 | 0.091 |
| urine Arg [µg/mg Kreatinin] | -0.007 | 0.937 |
| urine Lys [µg/mg Kreatinin] | 0.088 | 0.355 |
| Variable | UV DERMIS EXPOSURE DAMAGE DEPTH UV AGED DERMIS (N=113) | |
|---|---|---|
| r | p | |
| plasma FruLys [µg/ml] | -0.118 | 0.213 |
| plasma Pyr [ng/ml] | 0.125 | 0.188 |
| plasma MG-H1 [ng/ml] | -0.052 | 0.585 |
| plasma CEL [ng/ml] | 0.145 | 0.126 |
| plasma CML [ng/ml] | -0.065 | 0.492 |
| plasma Arg [µg/ml] | 0.091 | 0.337 |
| plasma Lys [µg/ml] | 0.151 | 0.110 |
| serum FruLys [µg/ml] | -0.070 | 0.464 |
| serum Pyr [ng/ml] | 0.227* | 0.016 |
| serum MG-H1 [ng/ml] | -0.126 | 0.183 |
| serum CEL [ng/ml] | 0.015 | 0.877 |
| serum CML [ng/ml] | -0.115 | 0.224 |
| serum Arg [µg/ml] | -0.010 | 0.912 |
| serum Lys [µg/ml] | 0.071 | 0.453 |
| saliva FruLys [ng/ml] | 0.184 | 0.051 |
| saliva Pyr [ng/ml] | 0.026 | 0.787 |
| saliva MG-H1 [ng/ml] | 0.285** | 0.002 |
| saliva CEL [ng/ml] | 0.039 | 0.685 |
| saliva CML [ng/ml] | 0.295** | 0.001 |
| saliva Arg [µg/ml] | 0.207* | 0.028 |
| saliva Lys [µg/ml] | 0.200* | 0.034 |
| urine FruLys [µg/mg Kreatinin] | -0.035 | 0.710 |
| urine Pyr [µg/mg Kreatinin] | -0.126 | 0.183 |
| urine MG-H1 [µg/mg Kreatinin] | -0.194* | 0.039 |
| urine CEL [µg/mg Kreatinin] | -0.020 | 0.829 |
| urine CML [µg/mg Kreatinin] | -0.252** | 0.007 |
| urine Arg [µg/mg Kreatinin] | -0.070 | 0.460 |
| urine Lys [µg/mg Kreatinin] | 0.064 | 0.498 |
| Variable | UV DERMIS EXPOSURE DAMAGE NR PIXELS (N=113) | |
|---|---|---|
| r | p | |
| plasma FruLys [µg/ml] | -0.029 | 0.763 |
| plasma Pyr [ng/ml] | 0.063 | 0.509 |
| plasma MG-H1 [ng/ml] | -0.075 | 0.432 |
| plasma CEL [ng/ml] | 0.112 | 0.238 |
| plasma CML [ng/ml] | -0.048 | 0.614 |
| plasma Arg [µg/ml] | 0.042 | 0.659 |
| plasma Lys [µg/ml] | 0.106 | 0.263 |
| serum FruLys [µg/ml] | -0.006 | 0.947 |
| serum Pyr [ng/ml] | 0.149 | 0.115 |
| serum MG-H1 [ng/ml] | -0.079 | 0.406 |
| serum CEL [ng/ml] | 0.047 | 0.619 |
| serum CML [ng/ml] | -0.032 | 0.738 |
| serum Arg [µg/ml] | -0.032 | 0.733 |
| serum Lys [µg/ml] | 0.065 | 0.494 |
| saliva FruLys [ng/ml] | 0.104 | 0.272 |
| saliva Pyr [ng/ml] | -0.007 | 0.942 |
| saliva MG-H1 [ng/ml] | 0.193* | 0.041 |
| saliva CEL [ng/ml] | 0.018 | 0.851 |
| saliva CML [ng/ml] | 0.227* | 0.016 |
| saliva Arg [µg/ml] | 0.115 | 0.226 |
| saliva Lys [µg/ml] | 0.123 | 0.193 |
| urine FruLys [µg/mg Kreatinin] | -0.037 | 0.700 |
| urine Pyr [µg/mg Kreatinin] | -0.114 | 0.229 |
| urine MG-H1[µg/mg Kreatinin] | -0.168 | 0.075 |
| urine CEL [µg/mg Kreatinin] | -0.030 | 0.753 |
| urine CML [µg/mg Kreatinin] | -0.240* | 0.011 |
| urine Arg [µg/mg Kreatinin] | -0.072 | 0.452 |
| urine Lys [µg/mg Kreatinin] | 0.058 | 0.542 |
| Variable | UV DERMIS EXPOSURE DAMAGE DENSITY (N=111) | |
|---|---|---|
| r | p | |
| plasma FruLys [µg/ml] | 0.363** | 0.000 |
| plasma Pyr [ng/ml] | -0.417** | 0.000 |
| plasma MG-H1 [ng/ml] | -0.036 | 0.709 |
| plasma CEL [ng/ml] | -0.054 | 0.576 |
| plasma CML [ng/ml] | 0.048 | 0.619 |
| plasma Arg [µg/ml] | -0.456** | 0.000 |
| plasma Lys [µg/ml] | -0.394** | 0.000 |
| serum FruLys [µg/ml] | 0.008 | 0.937 |
| serum Pyr [ng/ml] | -0.547** | 0.000 |
| serum MG-H1 [ng/ml] | -0.093 | 0.332 |
| serum CEL [ng/ml] | -0.241* | 0.011 |
| serum CML [ng/ml] | -0.140 | 0.143 |
| serum Arg [µg/ml] | -0.369** | 0.000 |
| serum Lys [µg/ml] | -0.270** | 0.004 |
| saliva FruLys [ng/ml] | 0.104 | 0.272 |
| saliva Pyr [ng/ml] | -0.007 | 0.942 |
| saliva MG-H1 [ng/ml] | 0.193* | 0.041 |
| saliva CEL [ng/ml] | 0.018 | 0.851 |
| saliva CML [ng/ml] | 0.227* | 0.016 |
| saliva Arg [µg/ml] | 0.115 | 0.226 |
| saliva Lys [µg/ml] | 0.123 | 0.193 |
| urine FruLys [µg/mg Kreatinin] | -0.037 | 0.700 |
| urine Pyr [µg/mg Kreatinin] | -0.114 | 0.229 |
| urine MG-H1 [µg/mg Kreatinin] | -0.168 | 0.075 |
| urine CEL [µg/mg Kreatinin] | -0.030 | 0.753 |
| urine CML [µg/mg Kreatinin] | -0.240* | 0.011 |
| urine Arg [µg/mg Kreatinin] | -0.072 | 0.452 |
| urine Lys [µg/mg Kreatinin] | 0.058 | 0.542 |
| Variable | UV DERMIS EXPOSURE DAMAGE DENSITY (N=111) | |
|---|---|---|
| r | p | |
| saliva FruLys [ng/ml] | -0.295** | 0.002 |
| saliva Pyr [ng/ml] | -0.006 | 0.951 |
| saliva MG-H1 [ng/ml] | -0.170* | 0.075 |
| saliva CEL [ng/ml] | -0.101 | 0.292 |
| saliva CML [ng/ml] | -0.195* | 0.041 |
| saliva Arg [µg/ml] | -0.180 | 0.059 |
| saliva Lys [µg/ml] | -0.233* | 0.014 |
| urine FruLys [µg/mg Kreatinin] | 0.012 | 0.904 |
| urine Pyr [µg/mg Kreatinin] | 0.137 | 0.152 |
| urine MG-H1 [µg/mg Kreatinin] | 0.196* | 0.039 |
| urine CEL [µg/mg Kreatinin] | 0.135 | 0.156 |
| urine CML [µg/mg Kreatinin] | 0.075 | 0.432 |
| urine Arg [µg/mg Kreatinin] | 0.432 | 0.744 |
| urine Lys [µg/mg Kreatinin] | -0.073 | 0.445 |
| Variable | DERMIS DEPTH N=113 | |
|---|---|---|
| r | p | |
| plasma FruLys [µg/ml] | 0.113 | 0.233 |
| plasma Pyr [ng/ml] | -0.075 | 0.429 |
| plasma MG-H1 [ng/ml] | -0.145 | 0.125 |
| plasma CEL [ng/ml] | -0.086 | 0.364 |
| Plasma CML | -0.203* | 0.031 |
| plasma Arg [µg/ml] | 0.042 | 0.662 |
| plasma Lys [µg/ml] | 0.090 | 0.342 |
| serum FruLys [µg/ml] | 0.073 | 0.443 |
| serum Pyr [ng/ml] | -0.017 | 0.861 |
| serum MG-H1 [ng/ml] | -0.184 | 0.052 |
| serum CEL [ng/ml] | -0.178 | 0.059 |
| serum CML [ng/ml] | -0.145 | 0.126 |
| serum Arg [µg/ml] | -0.071 | 0.458 |
| serum Lys [µg/ml] | -0.044 | 0.644 |
| saliva FruLys [ng/ml] | 0.158 | 0.095 |
| saliva Pyr [ng/ml] | 0.013 | 0.890 |
| saliva MG-H1 [ng/ml] | 0.158 | 0.096 |
| saliva CEL [ng/ml] | 0.050 | 0.596 |
| Saliva CML | 0.0170 | 0.072 |
| saliva Arg [µg/ml] | 0.154 | 0.104 |
| saliva Lys [µg/ml] | 0.228* | 0.015 |
| urine FruLys [µg/mg Kreatinin] | -0.216* | 0.022 |
| urine Pyr [µg/mg Kreatinin] | -0.111 | 0.243 |
| urine MG-H1 [µg/mg Kreatinin] | -0.224* | 0.017 |
| urine CEL [µg/mg Kreatinin] | -0.172 | 0.069 |
| urine CML [µg/mg Kreatinin] | -0.186* | 0.048 |
| urine Arg [µg/mg Kreatinin] | -0.159 | 0.094 |
| urine Lys [µg/mg Kreatinin] | 0.042 | 0.660 |
| Variable | DERMIS nr pixels N=113 | |
|---|---|---|
| r | p | |
| plasma FruLys [µg/ml] | 0.256** | 0.006 |
| plasma Pyr [ng/ml] | -0.181 | 0.055 |
| plasma MG-H1 [ng/ml] | -0.174 | 0.066 |
| plasma CEL [ng/ml] | -0.123 | 0.194 |
| Plasma CML | -0.177 | 0.061 |
| plasma Arg [µg/ml] | -0.001 | 0.989 |
| plasma Lys [µg/ml] | 0.036 | 0.703 |
| serum FruLys [µg/ml] | 0.174 | 0.066 |
| serum Pyr [ng/ml] | -0.159 | 0.092 |
| serum MG-H1 [ng/ml] | -0.093 | 0.329 |
| serum CEL [ng/ml] | -0.221* | 0.019 |
| serum CML [ng/ml] | 0.005 | 0.957 |
| serum Arg [µg/ml] | -0.077 | 0.420 |
| serum Lys [µg/ml] | -0.033 | 0.033 |
| saliva FruLys [ng/ml] | -0.004 | 0.968 |
| saliva Pyr [ng/ml] | -0.014 | 0.880 |
| saliva MG-H1 [ng/ml] | 0.002 | 0.985 |
| saliva CEL [ng/ml] | -0.012 | 0.896 |
| Saliva CML | 0.046 | 0.629 |
| saliva Arg [µg/ml] | -0.021 | 0.826 |
| saliva Lys [µg/ml] | 0.077 | 0.418 |
| urine FruLys [µg/mg Kreatinin] | -0.221* | 0.019 |
| urine Pyr [µg/mg Kreatinin] | -0.102 | 0.283 |
| urine MG-H1 [µg/mg Kreatinin] | -0.180 | 0.057 |
| urine CEL [µg/mg Kreatinin] | -0.179 | 0.058 |
| urine CML [µg/mg Kreatinin] | -0.177 | 0.061 |
| urine Arg [µg/mg Kreatinin] | -0.158 | 0.095 |
| urine Lys [µg/mg Kreatinin] | 0.038 | 0.690 |
| Variable | DERMIS density N=113 | |
|---|---|---|
| r | p | |
| plasma FruLys [µg/ml] | 0.098 | 0.304 |
| plasma Pyr [ng/ml] | -0.066 | 0.489 |
| plasma MG-H1 [ng/ml] | 0.066 | 0.485 |
| plasma CEL [ng/ml] | -0.089 | 0.350 |
| Plasma CML | 0.061 | 0.519 |
| plasma Arg [µg/ml] | -0.109 | 0.250 |
| plasma Lys [µg/ml] | -0.210* | 0.026 |
| serum FruLys [µg/ml] | 0.203* | 0.031 |
| serum Pyr [ng/ml] | -0.182 | 0.054 |
| serum MG-H1 [ng/ml] | 0.067 | 0.478 |
| serum CEL [ng/ml] | -0.057 | 0.551 |
| serum CML [ng/ml] | 0.002 | 0.979 |
| serum Arg [µg/ml] | -0.032 | 0.737 |
| serum Lys [µg/ml] | -0.052 | 0.586 |
| saliva FruLys [ng/ml] | -0.053 | 0.580 |
| saliva Pyr [ng/ml] | 0.112 | 0.236 |
| saliva MG-H1 [ng/ml] | 0.050 | 0.601 |
| saliva CEL [ng/ml] | -0.086 | 0.365 |
| Saliva CML [µg/ml] | 0.057 | 0.549 |
| saliva Arg [µg/ml] | -0.041 | 0.669 |
| saliva Lys [µg/ml] | 0.032 | 0.734 |
| urine FruLys [µg/mg Kreatinin] | 0.003 | 0.972 |
| urine Pyr [µg/mg Kreatinin] | 0.150 | 0.112 |
| urine MG-H1 [µg/mg Kreatinin] | 0.171 | 0.071 |
| urine CEL [µg/mg Kreatinin] | 0.220* | 0.019 |
| urine CML [µg/mg Kreatinin] | 0.133 | 0.159 |
| urine Arg [µg/mg Kreatinin] | 0.131 | 0.168 |
| urine Lys [µg/mg Kreatinin] | 0.026 | 0.785 |
| Variable | SC TISSUE DEPTH N=111 | |
|---|---|---|
| r | p | |
| plasma FruLys [µg/ml] | 0.046 | 0.635 |
| plasma Pyr [ng/ml] | 0.014 | 0.881 |
| plasma MG-H1 [ng/ml] | 0.208* | 0.029 |
| plasma CEL [ng/ml] | 0.008 | 0.936 |
| Plasma CML | 0.135 | 0.158 |
| plasma Arg [µg/ml] | 0.043 | 0.651 |
| plasma Lys [µg/ml] | 0.132 | 0.166 |
| serum FruLys [µg/ml] | -0.063 | 0.513 |
| serum Pyr [ng/ml] | 0.013 | 0.893 |
| serum MG-H1 [ng/ml] | -0.111 | 0.248 |
| serum CEL [ng/ml] | -0.062 | 0.520 |
| serum CML [ng/ml] | -0.290** | 0.002 |
| serum Arg [µg/ml] | -0.118 | 0.217 |
| serum Lys [µg/ml] | 0.083 | 0.388 |
| Variable | SC TISSUE NR PIXELS N=111 | |
|---|---|---|
| r | p | |
| plasma FruLys [µg/ml] | 0.134 | 0.162 |
| plasma Pyr [ng/ml] | -0.049 | 0.613 |
| plasma MG-H1 [ng/ml] | 0.201* | 0.035 |
| plasma CEL [ng/ml] | -0.035 | 0.712 |
| Plasma CML | 0.140 | 0.142 |
| plasma Arg [µg/ml] | 0.026 | 0.786 |
| plasma Lys [µg/ml] | 0.103 | 0.283 |
| serum FruLys [µg/ml] | -0.007 | 0.942 |
| serum Pyr [ng/ml] | -0.061 | 0.522 |
| serum MG-H1 [ng/ml] | -0.063 | 0.511 |
| serum CEL [ng/ml] | -0.124 | 0.195 |
| serum CML [ng/ml] | -0.213* | 0.025 |
| serum Arg [µg/ml] | -0.128 | 0.180 |
| serum Lys [µg/ml] | 0.086 | 0.370 |
| Variable | SC TISSUE DENSITY N=111 | |
|---|---|---|
| r | p | |
| plasma FruLys [µg/ml] | 0.034 | 0.719 |
| plasma Pyr [ng/ml] | -0.089 | 0.351 |
| plasma MG-H1 [ng/ml] | -0.117 | 0.220 |
| plasma CEL [ng/ml] | 0.009 | 0.925 |
| Plasma CML | -0.120 | 0.210 |
| plasma Arg [µg/ml] | -0.117 | 0.220 |
| plasma Lys [µg/ml] | -0.109 | 0.256 |
| serum FruLys [µg/ml] | -0.160 | 0.093 |
| serum Pyr [ng/ml] | -0.114 | 0.233 |
| serum MG-H1 [ng/ml] | 0.087 | 0.363 |
| serum CEL [ng/ml] | 0.746 | 0.746 |
| serum CML [ng/ml] | 0.049 | 0.606 |
| serum Arg [µg/ml] | -0.075 | 0.434 |
| serum Lys [µg/ml] | -0.191* | 0.045 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.




