Submitted:
22 February 2026
Posted:
28 February 2026
You are already at the latest version
Abstract
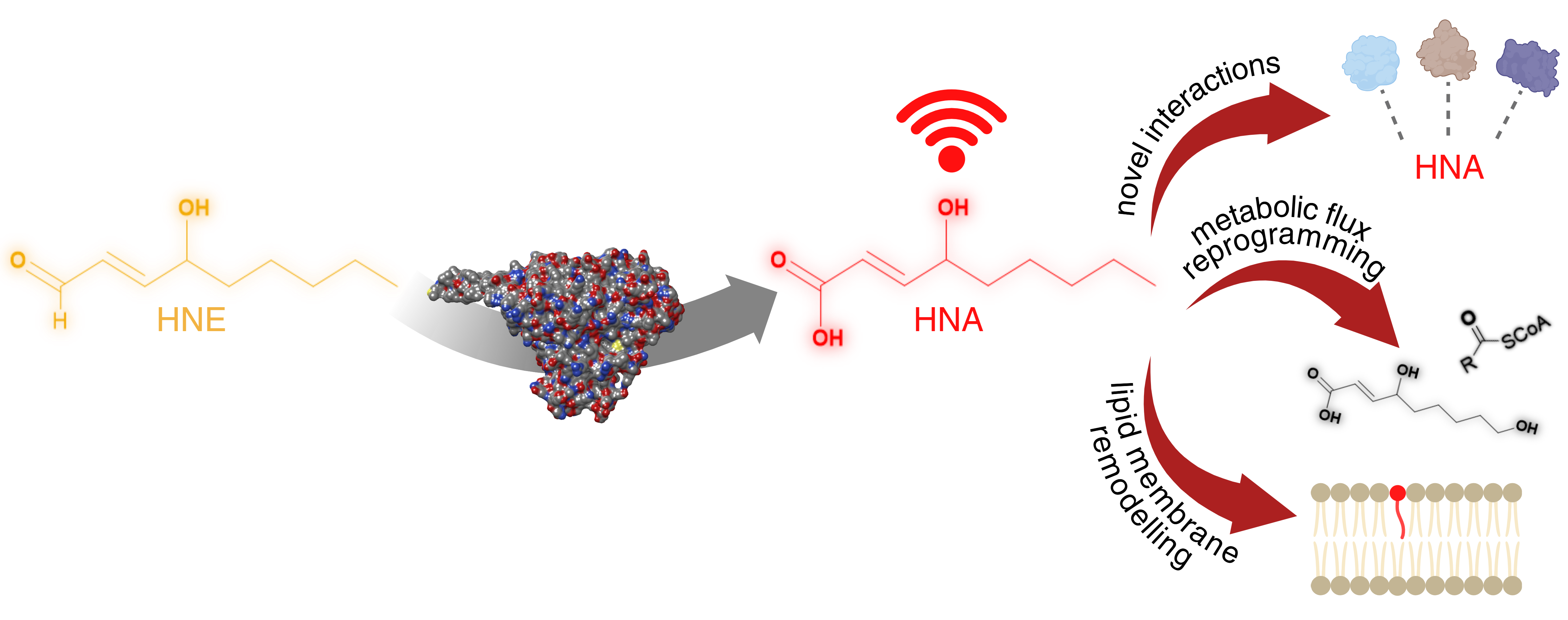
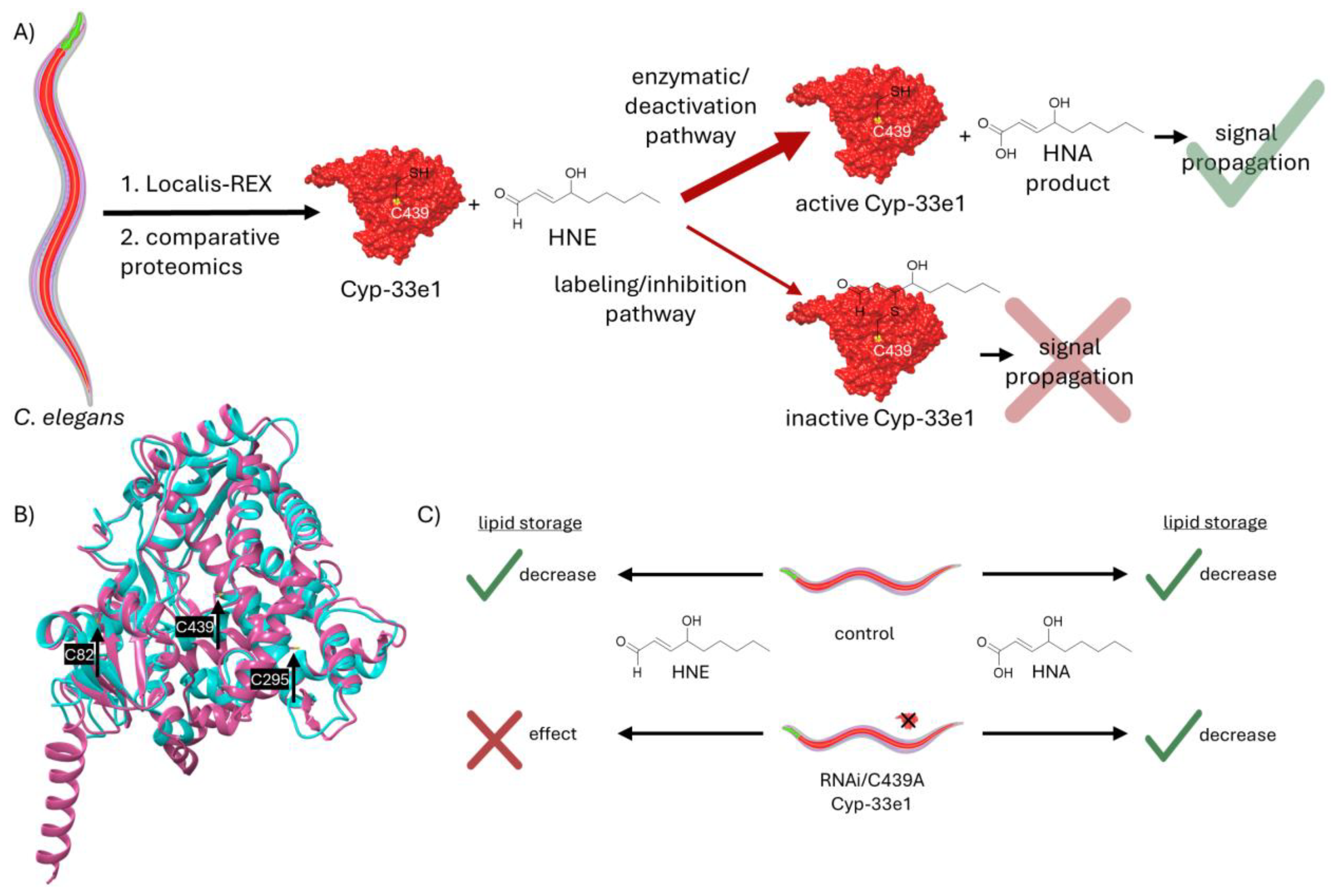
Discovered ~60years ago, the lipid metabolite 4-hydroxynonenal (HNE) is linked to a plethora of macromolecular targets and biological functions. For a molecule that weighs 156Da and possesses a single H-bond donor, this is quite a feat. Despite its chemical simplicity, HNE contains an α,β-unsaturated aldehyde system, endowing it with the capability to react covalently with numerous biological functional groups and bestowing on it pleiotropic properties. Regardless of the specific entity engaging with HNE, it is covalent bond formation that has dominated thought on HNE behavior. Indeed, cells possess a flurry of detoxifying enzymes that convert HNE to less reactive chemicals lacking the α,β-unsaturated aldehyde. For instance, the cell can either reduce or oxidize the aldehyde within HNE, deactivating HNE’s chemical reactivity. Here, we discuss one of our recent papers that discovered that HNE can modify the detoxification enzyme, Cyp-33e1, in C. elegans, using a customized tissue-specific screen for HNE-sensor proteins. Consistent with concepts of active site partitioning, HNE also emerged as a substrate of Cyp-33e1. We next discovered that HNE changed lipid storage in worms in a Cyp-33e1-dependent manner. We proposed that the product of Cyp-33e1 detoxifying HNE was responsible for this change in lipid storage, and were able to show that 4-hydroxynonenoic acid (HNA), the product of Cyp-33e1 oxidation of HNE, causes this phenotype. We have dubbed this new signaling mode, “deactivation signaling”. It sets an important precedent for how the bioactivity of HNE is considered, and we discuss the ramifications of this result in the paper.
Keywords:
1. It’s Difficult to Study “Natural” HNE Signaling
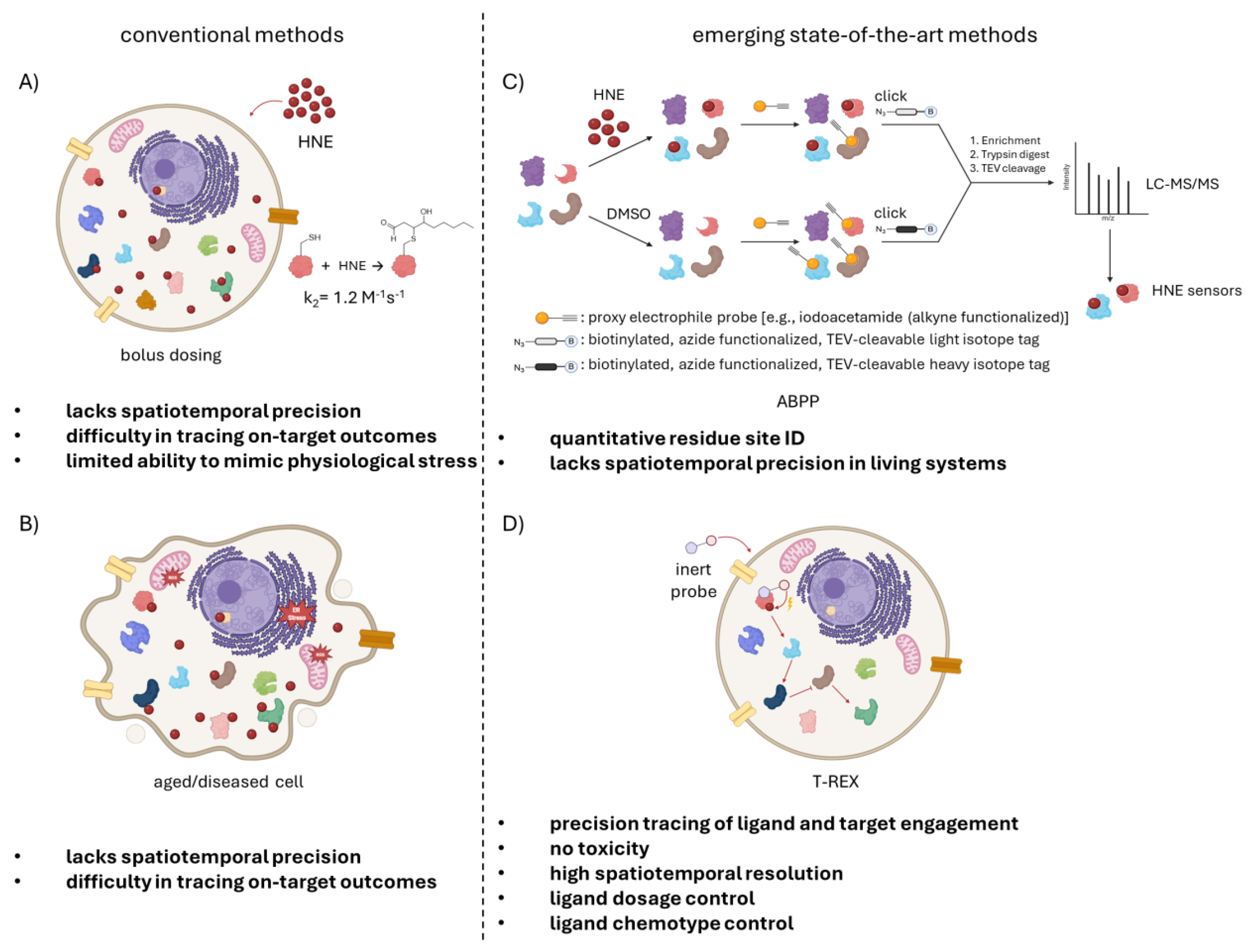
2. Chemical Biology Methods Increased Our Understanding of HNE Signaling
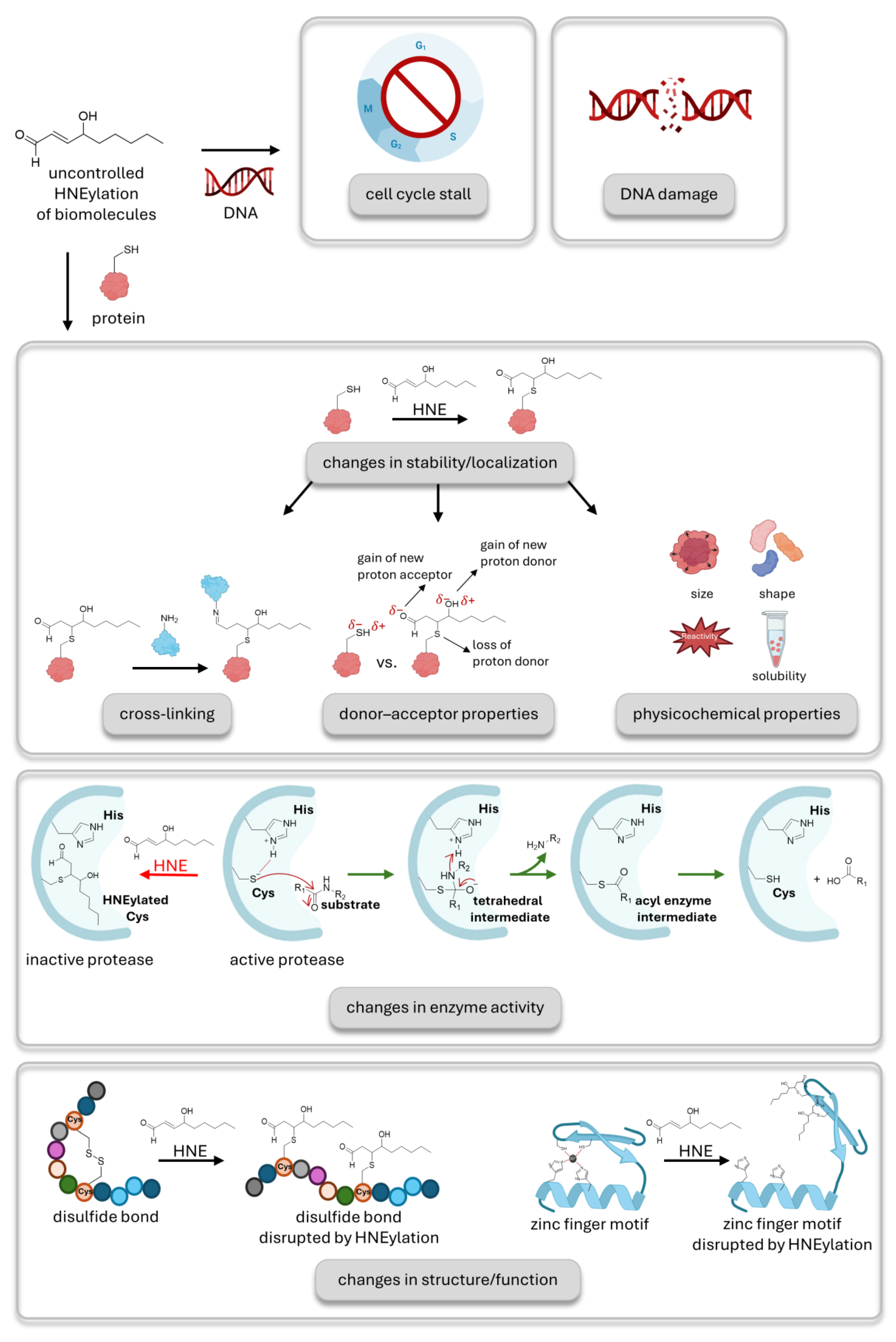
3. Breaking the Bonds of General Models for HNE-Mediated Signaling
4. HNEylation Profiling Can Inform on Non-Conjugative Signaling Modes
5. Biological Roles of Deactivation-Induced Signaling
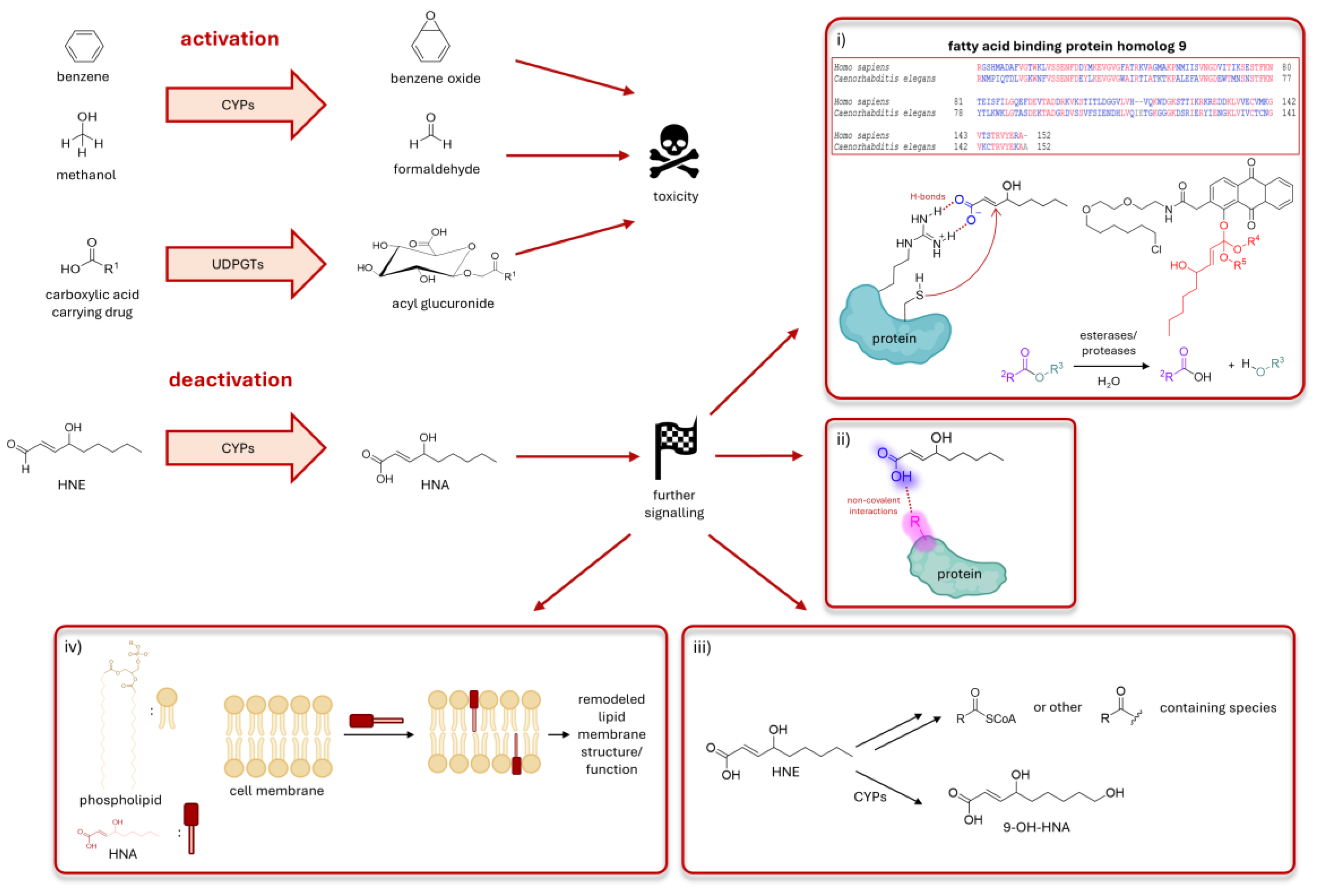
- (i)
- What deactivation?
- (ii)
- Less reactivity but new and strong non-covalent binding
- (iii)
- HNA is further metabolized
- (iv)
- HNA can affect lipid membrane biophysics or trafficking
6. Conclusions
Author Contributions
Acknowledgments
Conflicts of Interest
References
- Benedetti, A.; Comporti, M.; Esterbauer, H. Identification of 4-hydroxynonenal as a cytotoxic product originating from the peroxidation of liver microsomal lipids. Biochim. et Biophys. Acta (BBA) - Lipids Lipid Metab. 1980, 620, 281–296. [Google Scholar] [CrossRef]
- Uchida, K.; Stadtman, E.R. Modification of histidine residues in proteins by reaction with 4-hydroxynonenal. Proc. Natl. Acad. Sci. 1992, 89, 4544–4548. [Google Scholar] [CrossRef]
- Uchida, K.; Stadtman, E.R. Selective cleavage of thioether linkage in proteins modified with 4-hydroxynonenal. Proc. Natl. Acad. Sci. 1992, 89, 5611–5615. [Google Scholar] [CrossRef]
- Schopfer, F.J.; Cipollina, C.; Freeman, B.A. Formation and Signaling Actions of Electrophilic Lipids. Chem. Rev. 2011, 111, 5997–6021. [Google Scholar] [CrossRef] [PubMed]
- Parvez, S.; Long, M.J.C.; Poganik, J.R.; Aye, Y. Redox Signaling by Reactive Electrophiles and Oxidants. Chem. Rev. 2018, 118, 8798–8888. [Google Scholar] [CrossRef] [PubMed]
- Gensler, H.L.; Bernstein, H. DNA Damage as the Primary Cause of Aging. Q. Rev. Biol. 1981, 56, 279–303. [Google Scholar] [CrossRef] [PubMed]
- Benamira, M.; Marnett, L.J. The lipid peroxidation product 4-hydroxynonenal is a potent inducer of the SOS response. Mutat. Res. Repair 1992, 293, 1–10. [Google Scholar] [CrossRef]
- Uchida, K.; I Szweda, L.; Chae, H.Z.; Stadtman, E.R. Immunochemical detection of 4-hydroxynonenal protein adducts in oxidized hepatocytes. Proc. Natl. Acad. Sci. 1993, 90, 8742–8746. [Google Scholar] [CrossRef]
- Stadtman, E.R. Protein Oxidation and Aging. Science 1992, 257, 1220–1224. [Google Scholar] [CrossRef]
- Ho, T.-L. Hard soft acids bases (HSAB) principle and organic chemistry. Chem. Rev. 1975, 75, 1–20. [Google Scholar] [CrossRef]
- Crabb, J.W.; O'NEil, J.; Miyagi, M.; West, K.; Hoff, H.F. Hydroxynonenal inactivates cathepsin B by forming Michael adducts with active site residues. Protein Sci. 2002, 11, 831–840. [Google Scholar] [CrossRef] [PubMed]
- Salsman, S.J.; Hensley, K.; Floyd, R.A. Sensitivity of Protein Tyrosine Phosphatase Activity to the Redox Environment, Cytochrome c, and Microperoxidase. Antioxidants Redox Signal. 2005, 7, 1078–1088. [Google Scholar] [CrossRef] [PubMed]
- Grune, T.; Davies, K.J. The proteasomal system and HNE-modified proteins. Mol. Asp. Med. 2003, 24, 195–204. [Google Scholar] [CrossRef] [PubMed]
- Castro, J.P.; Jung, T.; Grune, T.; Siems, W. 4-Hydroxynonenal (HNE) modified proteins in metabolic diseases. Free Radic. Biol. Med. 2017, 111, 309–315. [Google Scholar] [CrossRef]
- Grune, T.; Reinheckel, T.; Joshi, M.; Davies, K.J. Proteolysis in Cultured Liver Epithelial Cells during Oxidative Stress. J. Biol. Chem. 1995, 270, 2344–2351. [Google Scholar] [CrossRef]
- Botzen, D.; Grune, T. Degradation of HNE-modified proteins – possible role of ubiquitin. Redox Rep. 2007, 12, 63–67. [Google Scholar] [CrossRef]
- Liu, X.; Long, M.J.C.; Aye, Y. Proteomics and Beyond: Cell Decision-Making Shaped by Reactive Electrophiles. Trends Biochem. Sci. 2019, 44, 75–89. [Google Scholar] [CrossRef]
- Wang, Y.; Lu, J.; Carisey, A.F.; Chadchan, S.B.; Lee, H.W.; Malireddi, R.S.; Sharma, B.R.; Pandian, N.; Tweedell, R.E.; Palacios, G.; et al. Innate immune and metabolic signals induce mitochondria-dependent membrane lysis via mitoxyperiosis. Cell 2025, 188, 7155–7174.e25. [Google Scholar] [CrossRef]
- Cubillos-Ruiz, J.R.; Silberman, P.C.; Rutkowski, M.R.; Chopra, S.; Perales-Puchalt, A.; Song, M.; Zhang, S.; Bettigole, S.E.; Gupta, D.; Holcomb, K.; et al. ER Stress Sensor XBP1 Controls Anti-tumor Immunity by Disrupting Dendritic Cell Homeostasis. Cell 2015, 161, 1527–1538. [Google Scholar] [CrossRef]
- Matsunaga, T.; Kawabata, S.; Yanagihara, Y.; Kezuka, C.; Kato, M.; Morikawa, Y.; Endo, S.; Chen, H.; Iguchi, K.; Ikari, A. Pathophysiological roles of autophagy and aldo-keto reductases in development of doxorubicin resistance in gastrointestinal cancer cells. Chem. Interactions 2019, 314, 108839. [Google Scholar] [CrossRef]
- Roy, B.; Pan, G.; Giri, S.; Thandavarayan, R.A.; Palaniyandi, S.S. Aldehyde dehydrogenase 2 augments adiponectin signaling in coronary angiogenesis in HFpEF associated with diabetes. FASEB J. 2022, 36, e22440. [Google Scholar] [CrossRef]
- Xiong, L.; Hu, H.; Zhu, F.; Shi, H.; Fan, X.; Pan, S.; Zhu, F.; Zhang, J.; Yu, Z.; Shi, Y. New insight for SS-31 in treating diabetic cardiomyopathy: Activation of mitoGPX4 and alleviation of mitochondria-dependent ferroptosis. Int. J. Mol. Med. 2024, 54, 1–13. [Google Scholar] [CrossRef]
- Monroe, T.B.; Hertzel, A.V.; Dickey, D.M.; Hagen, T.; Santibanez, S.V.; Berdaweel, I.A.; Halley, C.; Puchalska, P.; Anderson, E.J.; Camell, C.D.; et al. Lipid peroxidation products induce carbonyl stress, mitochondrial dysfunction, and cellular senescence in human and murine cells. Aging Cell 2024, 24, e14367. [Google Scholar] [CrossRef]
- Li, Y.; Zhao, T.; Li, J.; Xia, M.; Li, Y.; Wang, X.; Liu, C.; Zheng, T.; Chen, R.; Kan, D.; et al. Oxidative Stress and 4-hydroxy-2-nonenal (4-HNE): Implications in the Pathogenesis and Treatment of Aging-related Diseases. J. Immunol. Res. 2022, 2022, 1–12. [Google Scholar] [CrossRef] [PubMed]
- Delerue, T.; Khosrobakhsh, F.; Daloyau, M.; Emorine, L.J.; Dedieu, A.; Herbert, C.J.; Bonnefoy, N.; Arnauné-Pelloquin, L.; Belenguer, P. Loss of Msp1p in Schizosaccharomyces pombe induces a ROS-dependent nuclear mutator phenotype that affects mitochondrial fission genes. FEBS Lett. 2016, 590, 3544–3558. [Google Scholar] [CrossRef] [PubMed]
- Doorn, J.A.; Petersen, D.R. Covalent Modification of Amino Acid Nucleophiles by the Lipid Peroxidation Products 4-Hydroxy-2-nonenal and 4-Oxo-2-nonenal. Chem. Res. Toxicol. 2002, 15, 1445–1450. [Google Scholar] [CrossRef] [PubMed]
- Moss, R.A.; Ho, G.J. Kinetics of a 1,3-CH carbene insertion reaction: Tert-butylchlorocarbene. J. Am. Chem. Soc. 1990, 112, 5642–5644. [Google Scholar] [CrossRef]
- Moss, R.A.; Yan, S. Absolute kinetics of phenylchlorocarbene CH insertion reactions. Tetrahedron Lett. 1998, 39, 9381–9384. [Google Scholar] [CrossRef]
- Weerapana, E.; Wang, C.; Simon, G.M.; Richter, F.; Khare, S.; Dillon, M.B.D.; Bachovchin, D.A.; Mowen, K.; Baker, D.; Cravatt, B.F. Quantitative reactivity profiling predicts functional cysteines in proteomes. Nature 2010, 468, 790–795. [Google Scholar] [CrossRef]
- Jay, D.G. Selective destruction of protein function by chromophore-assisted laser inactivation. Proc. Natl. Acad. Sci. 1988, 85, 5454–5458. [Google Scholar] [CrossRef]
- Beck, S.; Sakurai, T.; Eustace, B.K.; Beste, G.; Schier, R.; Rudert, F.; Jay, D.G. Fluorophore-assisted light inactivation: A high-throughput tool for direct target validation of proteins. Proteomics 2002, 2, 247–255. [Google Scholar] [CrossRef]
- Parvez, S.; Long, M.J.C.; Lin, H.-Y.; Zhao, Y.; A Haegele, J.; Pham, V.N.; Lee, D.K.; Aye, Y. T-REX on-demand redox targeting in live cells. Nat. Protoc. 2016, 11, 2328–2356. [Google Scholar] [CrossRef] [PubMed]
- Parvez, S.; Fu, Y.; Li, J.; Long, M.J.C.; Lin, H.-Y.; Lee, D.K.; Hu, G.S.; Aye, Y. Substoichiometric Hydroxynonenylation of a Single Protein Recapitulates Whole-Cell-Stimulated Antioxidant Response. J. Am. Chem. Soc. 2014, 137, 10–13. [Google Scholar] [CrossRef] [PubMed]
- Lin, H.-Y.; Haegele, J.A.; Disare, M.T.; Lin, Q.; Aye, Y. A Generalizable Platform for Interrogating Target- and Signal-Specific Consequences of Electrophilic Modifications in Redox-Dependent Cell Signaling. J. Am. Chem. Soc. 2015, 137, 6232–6244. [Google Scholar] [CrossRef] [PubMed]
- Long, M.J.; Lin, H.-Y.; Parvez, S.; Zhao, Y.; Poganik, J.R.; Huang, P.; Aye, Y. β-TrCP1 Is a Vacillatory Regulator of Wnt Signaling. Cell Chem. Biol. 2017, 24, 944–957.e7. [Google Scholar] [CrossRef]
- Van Hall-Beauvais, A.; Poganik, J.R.; Huang, K.-T.; Parvez, S.; Zhao, Y.; Lin, H.-Y.; Liu, X.; Long, M.J.C.; Aye, Y. Z-REX uncovers a bifurcation in function of Keap1 paralogs. eLife 2022, 11. [Google Scholar] [CrossRef]
- Huang, K.-T.; Poganik, J.R.; Parvez, S.; Raja, S.; Miller, B.; Long, M.J.C.; Fetcho, J.R.; Aye, Y. Author Correction: Z-REX: Shepherding reactive electrophiles to specific proteins expressed tissue specifically or ubiquitously, and recording the resultant functional electrophile-induced redox responses in larval fish. Nat. Protoc. 2023, 18, 3155. [Google Scholar] [CrossRef]
- Long, M.J.C.; Parvez, S.; Zhao, Y.; Surya, S.L.; Wang, Y.; Zhang, S.; Aye, Y. Akt3 is a privileged first responder in isozyme-specific electrophile response. Nat. Chem. Biol. 2017, 13, 333–338. [Google Scholar] [CrossRef]
- Liu, X.; Long, M.J.C.; Hopkins, B.D.; Luo, C.; Wang, L.; Aye, Y. Precision Targeting of pten-Null Triple-Negative Breast Tumors Guided by Electrophilic Metabolite Sensing. ACS Central Sci. 2020, 6, 892–902. [Google Scholar] [CrossRef]
- Zhao, Y.; Herrera, P.A.M.; Chang, D.; Hamelin, R.; Long, M.J.C.; Aye, Y. Function-guided proximity mapping unveils electrophilic-metabolite sensing by proteins not present in their canonical locales. Proc. Natl. Acad. Sci. 2022, 119. [Google Scholar] [CrossRef]
- Liu, J.; Kulkarni, A.; Gao, Y.-Q.; Urul, D.A.; Hamelin, R.; Novotny, B.Á.; Long, M.J.; Aye, Y. Organ-specific electrophile responsivity mapping in live C. elegans. Cell 2024, 187, 7450–7469.e29. [Google Scholar] [CrossRef]
- Chang, D.; Assari, M.; Suwathep, C.; Sappakhaw, K.; Uttamapinant, C.; Long, M.J.C.; Aye, Y. NCBP1 stress signaling drives alternative S6K1 splicing inhibiting translation. Nat. Chem. Biol. 2026. [Google Scholar] [CrossRef] [PubMed]
- Honzatko, A.; Brichac, J.; Picklo, M.J. Quantification of trans-4-hydroxy-2-nonenal enantiomers and metabolites by LC–ESI-MS/MS. J. Chromatogr. B 2007, 857, 115–122. [Google Scholar] [CrossRef] [PubMed]
- Zhao, M.; Ma, J.; Li, M.; Zhang, Y.; Jiang, B.; Zhao, X.; Huai, C.; Shen, L.; Zhang, N.; He, L.; et al. Cytochrome P450 Enzymes and Drug Metabolism in Humans. Int. J. Mol. Sci. 2021, 22, 12808. [Google Scholar] [CrossRef] [PubMed]
- Tipparaju, S.M.; Barski, O.A.; Srivastava, S.; Bhatnagar, A. Catalytic Mechanism and Substrate Specificity of the β-Subunit of the Voltage-Gated Potassium Channel. Biochemistry 2008, 47, 8840–8854. [Google Scholar] [CrossRef]
- Wan, L.; Essuman, K.; Anderson, R.G.; Sasaki, Y.; Monteiro, F.; Chung, E.-H.; Nishimura, E.O.; DiAntonio, A.; Milbrandt, J.; Dangl, J.L.; et al. TIR domains of plant immune receptors are NAD + -cleaving enzymes that promote cell death. Science 2019, 365, 799–803. [Google Scholar] [CrossRef]
- Wang, Y.; Kavran, J.M.; Chen, Z.; Karukurichi, K.R.; Leahy, D.J.; Cole, P.A. Regulation of S-Adenosylhomocysteine Hydrolase by Lysine Acetylation. J. Biol. Chem. 2014, 289, 31361–31372. [Google Scholar] [CrossRef]
- Hellberg, K.; Grimsrud, P.A.; Kruse, A.C.; Banaszak, L.J.; Ohlendorf, D.H.; Bernlohr, D.A. X-ray crystallographic analysis of adipocyte fatty acid binding protein (aP2) modified with 4-hydroxy-2-nonenal. Protein Sci. 2010, 19, 1480–1489. [Google Scholar] [CrossRef]
- Xu, M.; Joo, H.-J.; Paik, Y.-K. Novel Functions of Lipid-binding Protein 5 in Caenorhabditis elegans Fat Metabolism. J. Biol. Chem. 2011, 286, 28111–28118. [Google Scholar] [CrossRef]
- Homan, R.A.; Lapek, J.D.; Woo, C.M.; Niessen, S.; Jones, L.H.; Parker, C.G. Photoaffinity labelling with small molecules. Nat. Rev. Methods Prim. 2024, 4, 1–23. [Google Scholar] [CrossRef]
- Zhao, Y.; Long, M.J.C.; Wang, Y.; Zhang, S.; Aye, Y. Ube2V2 Is a Rosetta Stone Bridging Redox and Ubiquitin Codes, Coordinating DNA Damage Responses. ACS Central Sci. 2018, 4, 246–259. [Google Scholar] [CrossRef]
- Sezgin, E.; Levental, I. Membranes in focus. Biophys. J. 2023, 122, E1–E4. [Google Scholar] [CrossRef]
- Renne, M.F.; Ernst, R. Membrane homeostasis beyond fluidity: Control of membrane compressibility. Trends Biochem. Sci. 2023, 48, 963–977. [Google Scholar] [CrossRef]
- Abouelkheir, M.; Roy, T.; Krzyscik, M.A.; Özdemir, E.; Hristova, K. Investigations of membrane protein interactions in cells using fluorescence microscopy. Curr. Opin. Struct. Biol. 2024, 86, 102816. [Google Scholar] [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.



