Submitted:
26 February 2026
Posted:
27 February 2026
You are already at the latest version
Abstract
Keywords:
Introduction
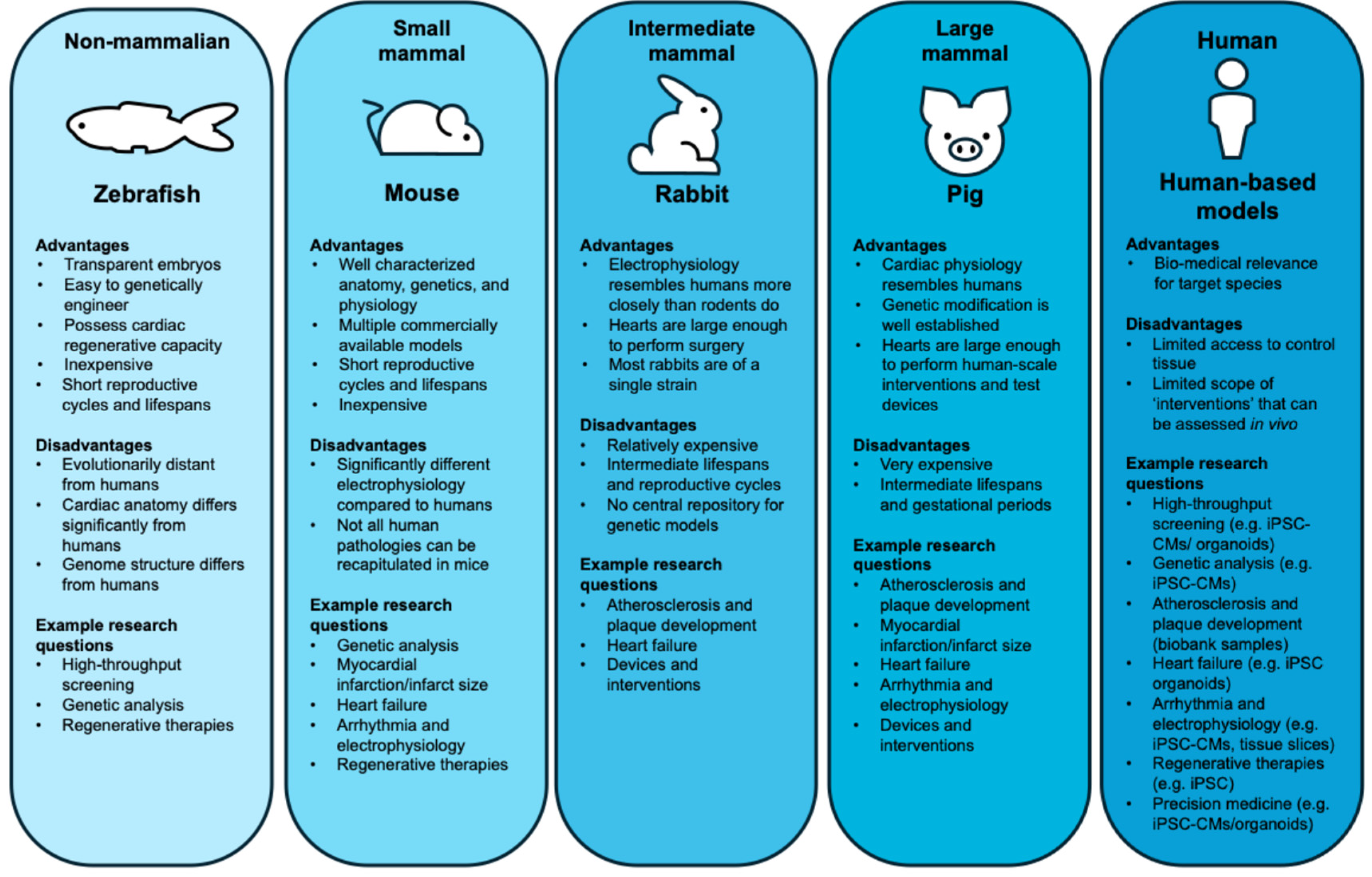
Non-Human Vertebrate Models of the Heart for Cardiac Research
Zebrafish: An Example of Non-Mammalian Model Systems
Mice: the Most Widely Used Small Mammalian Model
Rabbits: The Intermediate Size Mammalian Model
Pigs: A ‘Human-Sized’ Mammalian Model
Challenges of Using Non-Human Vertebrate Models of the Heart
Human Models for Cardiac Research
Human Induced Pluripotent Stem Cells and Human Engineered Heart Tissue
Human Isolated Primary Cardiomyocytes and Non-Cardiomyocytes
Myocardial Tissue Slices
In Silico Models
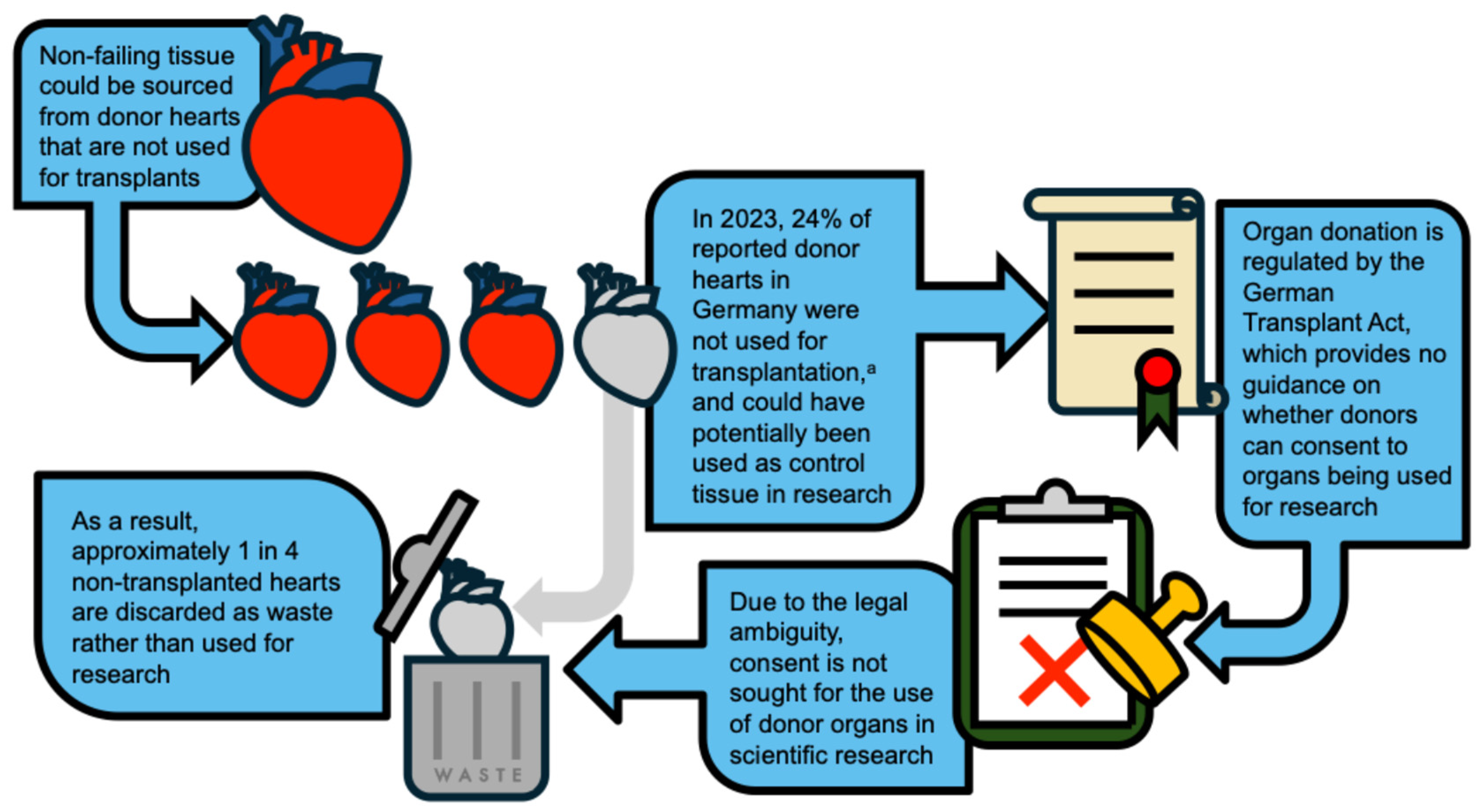
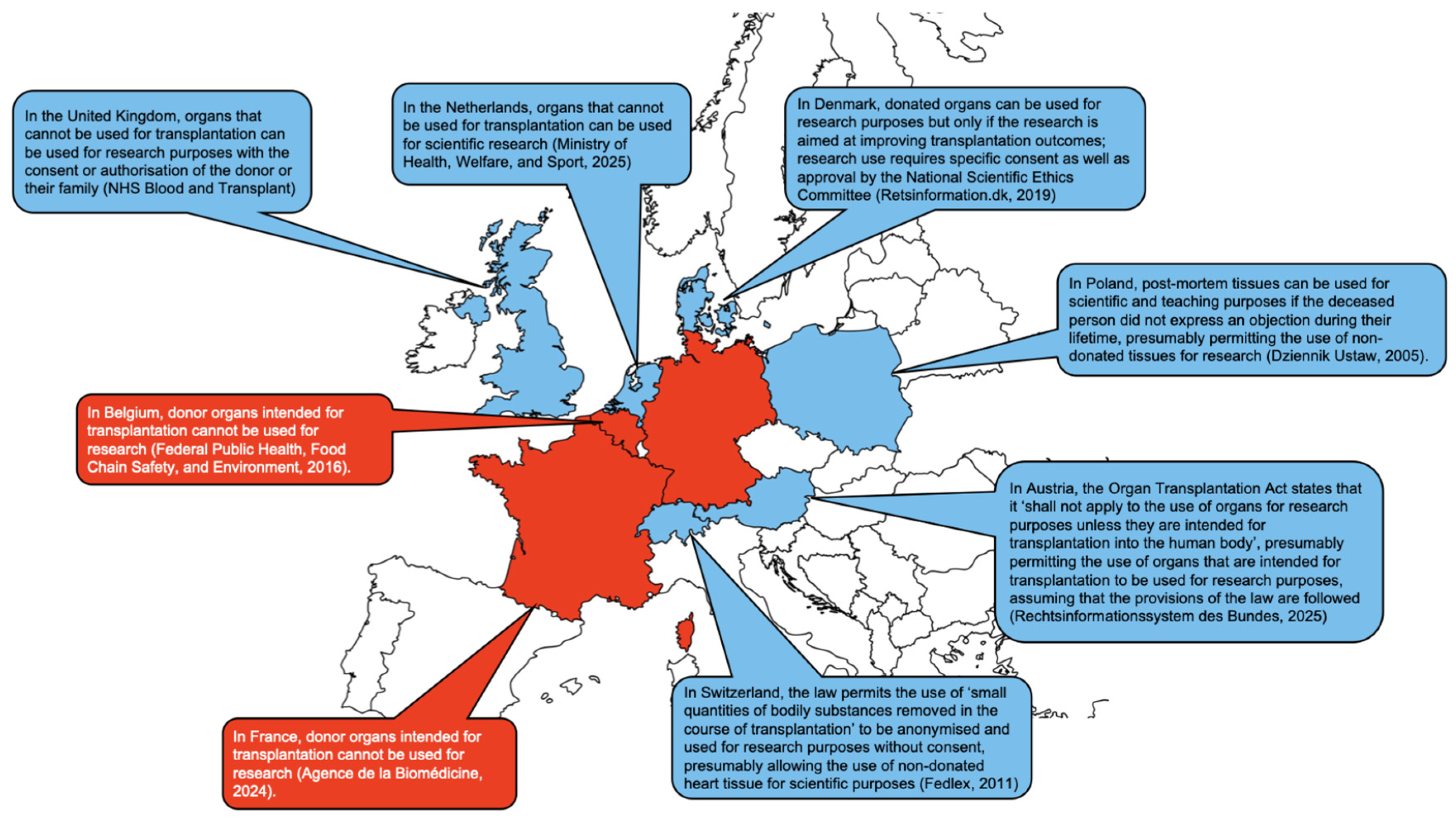
Availability of Human Tissue for Cardiac Research
Use of Non-Failing Human Heart Tissue for Scientific Research: Challenges and Opportunities
Conclusions
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest Statement
References
- Morehouse, LA; Sugarman, ED; Bourassa, PA; Sand, TM; Zimetti, F; Gao, F; et al. Inhibition of CETP activity by torcetrapib reduces susceptibility to diet-induced atherosclerosis in New Zealand White rabbits. J Lipid Res 2007, 48, 1263–72. [Google Scholar] [CrossRef]
- Barter, PJ; Caulfield, M; Eriksson, M; Grundy, SM; Kastelein, JJP; Komajda, M; et al. Effects of Torcetrapib in Patients at High Risk for Coronary Events. New England Journal of Medicine 2007, 357, 2109–22. [Google Scholar] [CrossRef]
- Harrison, RK. Phase II and phase III failures: 2013–2015. Nat Rev Drug Discov 2016, 15, 817–8. [Google Scholar] [CrossRef]
- Perry, CJ; Lawrence, AJ. Hurdles in Basic Science Translation. Front Pharmacol 2017, 8, 478. [Google Scholar] [CrossRef]
- Austin, CP. Opportunities and challenges in translational science. Clin Transl Sci 2021, 14, 1629–47. [Google Scholar] [CrossRef] [PubMed]
- Sun, D; Gao, W; Hu, H; Zhou, S. Why 90% of clinical drug development fails and how to improve it? Acta Pharm Sin B 2022, 12, 3049–62. [Google Scholar] [CrossRef] [PubMed]
- Ineichen, B V.; Furrer, E; Grüninger, SL; Zürrer, WE; Macleod, MR. Analysis of animal-to-human translation shows that only 5% of animal-tested therapeutic interventions obtain regulatory approval for human applications. PLoS Biol 2024, 22, e3002667. [Google Scholar] [CrossRef] [PubMed]
- Thomas, A; Desai, P; Takahashi, N. Translational research: A patient-centered approach to bridge the valley of death. Cancer Cell 2022, 40, 565–8. [Google Scholar] [CrossRef]
- Giacomotto, J; Ségalat, L. High-throughput screening and small animal models, where are we? Br J Pharmacol 2010, 160, 204–16. [Google Scholar] [CrossRef]
- Odening, KE; Gomez, A-M; Dobrev, D; Fabritz, L; Heinzel, FR; Mangoni, ME; et al. ESC working group on cardiac cellular electrophysiology position paper: relevance, opportunities, and limitations of experimental models for cardiac electrophysiology research. EP Europace 2021, 23, 1795–814. [Google Scholar] [CrossRef]
- Zaragoza, C; Gomez-Guerrero, C; Martin-Ventura, JL; Blanco-Colio, L; Lavin, B; Mallavia, B; et al. Animal models of cardiovascular diseases. J Biomed Biotechnol 2011, 2011, 497841. [Google Scholar] [CrossRef] [PubMed]
- Tsang, HG; Rashdan, NA; Whitelaw, CBA; Corcoran, BM; Summers, KM; MacRae, VE. Large animal models of cardiovascular disease. Cell Biochem Funct 2016, 34, 113–32. [Google Scholar] [CrossRef] [PubMed]
- van der Velden, J; Asselbergs, FW; Bakkers, J; Batkai, S; Bertrand, L; Bezzina, CR; et al. Animal models and animal-free innovations for cardiovascular research: current status and routes to be explored. Consensus document of the ESC Working Group on Myocardial Function and the ESC Working Group on Cellular Biology of the Heart. Cardiovasc Res 2022, 118, 3016–51. [Google Scholar] [CrossRef] [PubMed]
- Ray, AK; Priya, A; Malik, MdZ; Thanaraj, TA; Singh, AK; Mago, P; et al. A bioinformatics approach to elucidate conserved genes and pathways in C. elegans as an animal model for cardiovascular research. Sci Rep 2024, 14, 7471. [Google Scholar] [CrossRef]
- Benian, GM; Epstein, HF. Caenorhabditis elegans Muscle. Circ Res 2011, 109, 1082–95. [Google Scholar] [CrossRef]
- Paternostro, G; Vignola, C; Bartsch, D-U; Omens, JH; McCulloch, AD; Reed, JC. Age-Associated Cardiac Dysfunction in Drosophila melanogaster. Circ Res 2001, 88, 1053–8. [Google Scholar] [CrossRef]
- Feala, JD; Omens, JH; Paternostro, G; McCulloch, AD. Discovering Regulators of the Drosophila Cardiac Hypoxia Response Using Automated Phenotyping Technology. Ann N Y Acad Sci 2008, 1123, 169–77. [Google Scholar] [CrossRef]
- Wittig, JG; Münsterberg, A. The chicken as a model organism to study heart development. Cold Spring Harb Perspect Biol 2020, 12, 1–17. [Google Scholar] [CrossRef]
- Rees, J; Winkler, A; Huettemeister, J; Stengel, L; Spangler, P; Ramesh, G; et al. Rostafuroxin ameliorates cardiac glycoside-induced cardiomyocyte electrolyte imbalances and arrhythmia in ovo. Am J Physiol Heart Circ Physiol 2026. [Google Scholar] [CrossRef]
- MacRae, CA; Peterson, RT. Zebrafish as tools for drug discovery. Nat Rev Drug Discov 2015, 14, 721–31. [Google Scholar] [CrossRef]
- Asnani, A; Peterson, RT. The zebrafish as a tool to identify novel therapies for human cardiovascular disease. Dis Model Mech 2014, 7, 763–7. [Google Scholar] [CrossRef] [PubMed]
- Cesarovic, N; Lipski, M; Falk, V; Emmert, MY. Animals in cardiovascular research. Eur Heart J 2020, 41, 200–3. [Google Scholar] [CrossRef] [PubMed]
- Yang, D; Jian, Z; Tang, C; Chen, Z; Zhou, Z; Zheng, L; et al. Zebrafish Congenital Heart Disease Models: Opportunities and Challenges. Int J Mol Sci 2024, 25, 5943. [Google Scholar] [CrossRef] [PubMed]
- Vornanen, M; Hassinen, M. Zebrafish heart as a model for human cardiac electrophysiology. Channels 2016, 10, 101–10. [Google Scholar] [CrossRef]
- Hickman, DL; Johnson, J; Vemulapalli, TH; Crisler, JR; Shepherd, R. Commonly Used Animal Models. In Principles of Animal Research for Graduate and Undergraduate Students; Elsevier Inc., 2017; pp. 117–75. [Google Scholar] [CrossRef]
- Gurumurthy, CB; Kent Lloyd, KC. Generating mouse models for biomedical research: Technological advances. Dis Model Mech 2019, 12, dmm029462. [Google Scholar] [CrossRef]
- Gordon, JW; Scangos, GA; Plotkin, DJ; Barbosa, JA; Ruddle, FH. Genetic transformation of mouse embryos by microinjection of purified DNA. Proceedings of the National Academy of Sciences 1980, 77, 7380–4. [Google Scholar] [CrossRef]
- Hanahan, D; Wagner, EF; Palmiter, RD. The origins of oncomice: a history of the first transgenic mice genetically engineered to develop cancer. Genes Dev 2007, 21, 2258–70. [Google Scholar] [CrossRef]
- Justice, MJ; Dhillon, P. Using the mouse to model human disease: increasing validity and reproducibility. Dis Model Mech 2016, 9, 101–3. [Google Scholar] [CrossRef]
- Chowdhury, R; Ashraf, H; Melanson, M; Tanada, Y; Nguyen, M; Silberbach, M; et al. Mouse Model of Human Congenital Heart Disease: Progressive Atrioventricular Block Induced by a Heterozygous Nkx2-5 Homeodomain Missense Mutation. Circ Arrhythm Electrophysiol 2015, 8, 1255–64. [Google Scholar] [CrossRef]
- Lindsey, ML; Brunt, KR; Kirk, JA; Kleinbongard, P; Calvert, JW; de Castro Brás, LE; et al. Guidelines for in vivo mouse models of myocardial infarction. American Journal of Physiology-Heart and Circulatory Physiology 2021, 321, H1056–73. [Google Scholar] [CrossRef]
- Rao, KS; Kameswaran, V; Bruneau, BG. Modeling congenital heart disease: lessons from mice, hPSC-based models, and organoids. Genes Dev 2022, 36, 652–63. [Google Scholar] [CrossRef]
- Salerno, N; Scalise, M; Marino, F; Filardo, A; Chiefalo, A; Panuccio, G; et al. A Mouse Model of Dilated Cardiomyopathy Produced by Isoproterenol Acute Exposure Followed by 5-Fluorouracil Administration. J Cardiovasc Dev Dis 2023, 10, 225. [Google Scholar] [CrossRef] [PubMed]
- Fu, F; Pietropaolo, M; Cui, L; Pandit, S; Li, W; Tarnavski, O; et al. Lack of authentic atrial fibrillation in commonly used murine atrial fibrillation models. PLoS One 2022, 17, e0256512. [Google Scholar] [CrossRef] [PubMed]
- Emini Veseli, B; Perrotta, P; De Meyer, GRA; Roth, L; Van der Donckt, C; Martinet, W; et al. Animal models of atherosclerosis. Eur J Pharmacol 2017, 816, 3–13. [Google Scholar] [CrossRef] [PubMed]
- Nickel, AG; von Hardenberg, A; Hohl, M; Löffler, JR; Kohlhaas, M; Becker, J; et al. Reversal of Mitochondrial Transhydrogenase Causes Oxidative Stress in Heart Failure. Cell Metab 2015, 22, 472–84. [Google Scholar] [CrossRef]
- Schiattarella, GG; Altamirano, F; Tong, D; French, KM; Villalobos, E; Kim, SY; et al. Nitrosative stress drives heart failure with preserved ejection fraction. Nature 2019, 568, 351–6. [Google Scholar] [CrossRef]
- Müller, M; Bischof, C; Kapries, T; Wollnitza, S; Liechty, C; Geißen, S; et al. Right Heart Failure in Mice Upon Pressure Overload Is Promoted by Mitochondrial Oxidative Stress. JACC Basic Transl Sci 2022, 7, 658–77. [Google Scholar] [CrossRef]
- Kohlhaas, M; Sequeira, V; Parikh, S; Dietl, A; Richter, O; Bay, J; et al. Mitochondrial reactive oxygen species cause arrhythmias in hypertrophic cardiomyopathy. BioRxiv 2024, 2024, 10.02.616214. [Google Scholar] [CrossRef]
- Pepin, ME; Konrad, PJM; Nazir, S; Bazgir, F; Maack, C; Nickel, A; et al. Mitochondrial NNT Promotes Diastolic Dysfunction in Cardiometabolic HFpEF. Circ Res 2025, 136, 1564–78. [Google Scholar] [CrossRef]
- Rosshart, SP; Herz, J; Vassallo, BG; Hunter, A; Wall, MK; Badger, JH; et al. Laboratory mice born to wild mice have natural microbiota and model human immune responses. Science (1979) 2019, 365. [Google Scholar] [CrossRef]
- Eppig, JT; Motenko, H; Richardson, JE; Richards-Smith, B; Smith, CL. The International Mouse Strain Resource (IMSR): cataloging worldwide mouse and ES cell line resources. Mammalian Genome 2015, 26, 448–55. [Google Scholar] [CrossRef] [PubMed]
- Donahue, LR; Hrabe de Angelis, M; Hagn, M; Franklin, C; Lloyd, KCK; Magnuson, T; et al. Centralized mouse repositories. Mammalian Genome 2012, 23, 559–71. [Google Scholar] [CrossRef] [PubMed]
- Odening, KE; Kohl, P. Follow the white rabbit: Experimental and computational models of the rabbit heart provide insights into cardiac (patho-) physiology. Prog Biophys Mol Biol 2016, 121, 75–6. [Google Scholar] [CrossRef] [PubMed]
- Pogwizd, SM; Bers, DM. Rabbit models of heart disease. Drug Discov Today Dis Models 2008, 5, 185–93. [Google Scholar] [CrossRef] [PubMed]
- Panfilov, A V. Is heart size a factor in ventricular fibrillation? Or how close are rabbit and human hearts? Heart Rhythm 2006, 3, 862–4. [Google Scholar] [CrossRef]
- Odening, KE; Baczko, I; Brunner, M. Animals in cardiovascular research: Important role of rabbit models in cardiac electrophysiology. Eur Heart J 2020, 41, 2036. [Google Scholar] [CrossRef]
- Endoh, M. Force–frequency relationship in intact mammalian ventricular myocardium: physiological and pathophysiological relevance. Eur J Pharmacol 2004, 500, 73–86. [Google Scholar] [CrossRef]
- Milani-Nejad, N; Janssen, PML. Small and large animal models in cardiac contraction research: Advantages and disadvantages. Pharmacol Ther 2014, 141, 235–49. [Google Scholar] [CrossRef]
- Alpert, NR; Hasenfuss, G; Mulieri, LA; Blanchard, EM; Leavitt, BJ; Ittleman, F. The reorganization of the human and rabbit heart in response to haemodynamic overload. Eur Heart J 1992, 13, 9–16. [Google Scholar] [CrossRef]
- Cooper, LL; Odening, KE; Hwang, M-S; Chaves, L; Schofield, L; Taylor, CA; et al. Electromechanical and structural alterations in the aging rabbit heart and aorta. American Journal of Physiology-Heart and Circulatory Physiology 2012, 302, H1625–35. [Google Scholar] [CrossRef]
- Hornyik, T; Rieder, M; Castiglione, A; Major, P; Baczko, I; Brunner, M; et al. Transgenic rabbit models for cardiac disease research. Br J Pharmacol 2022, 179, 938–57. [Google Scholar] [CrossRef]
- Liu, Z; Chen, M; Chen, S; Deng, J; Song, Y; Lai, L; et al. Highly efficient RNA-guided base editing in rabbit. Nat Commun 2018, 9, 2717. [Google Scholar] [CrossRef] [PubMed]
- Yang, D; Xu, J; Chen, YE. Generation of Rabbit Models by Gene Editing Nucleases. Methods in Molecular Biology 2019, 1874, 327–45. [Google Scholar] [CrossRef]
- Davis, BT; Wang, XJ; Rohret, JA; Struzynski, JT; Merricks, EP; Bellinger, DA; et al. Targeted disruption of LDLR causes hypercholesterolemia and atherosclerosis in Yucatan miniature pigs. PLoS One 2014, 9, e93457. [Google Scholar] [CrossRef] [PubMed]
- Huang, L; Hua, Z; Xiao, H; Cheng, Y; Xu, K; Gao, Q; et al. CRISPR/Cas9-mediated ApoE-/-and LDLR-/-double gene knockout in pigs elevates serum LDL-C and TC levels. Oncotarget 2017, 8, 37751–60. [Google Scholar] [CrossRef] [PubMed]
- Montag, J; Petersen, B; Flögel, AK; Becker, E; Lucas-Hahn, A; Cost, GJ; et al. Successful knock-in of Hypertrophic Cardiomyopathy-mutation R723G into the MYH7 gene mimics HCM pathology in pigs. Sci Rep 2018, 8, 4786. [Google Scholar] [CrossRef]
- Sridharan, D; Pracha, N; Rana, SJ; Ahmed, S; Dewani, AJ; Alvi, SB; et al. Preclinical Large Animal Porcine Models for Cardiac Regeneration and Its Clinical Translation: Role of hiPSC-Derived Cardiomyocytes. Cells 2023, 12, 1090. [Google Scholar] [CrossRef]
- Liu, S; Li, K; Florencio, LW; Tang, L; Heallen, TR; Leach, JP; et al. Gene therapy knockdown of Hippo signaling induces cardiomyocyte renewal in pigs after myocardial infarction. Sci Transl Med 2021, 13, eabd6892. [Google Scholar] [CrossRef]
- Fischer, K; Schnieke, A. How genome editing changed the world of large animal research. Front Genome Ed 2023, 5, 1272687. [Google Scholar] [CrossRef]
- Barlabé, P; Aranguren, XL; Coppiello, G. Blastocyst complementation: current progress and future directions in xenogeneic organogenesis. Stem Cell Res Ther 2025, 16, 321. [Google Scholar] [CrossRef]
- Moretti, A; Fonteyne, L; Giesert, F; Hoppmann, P; Meier, AB; Bozoglu, T; et al. Somatic gene editing ameliorates skeletal and cardiac muscle failure in pig and human models of Duchenne muscular dystrophy. Nat Med 2020, 26, 207–14. [Google Scholar] [CrossRef] [PubMed]
- Rieblinger, B; Sid, H; Duda, D; Bozoglu, T; Klinger, R; Schlickenrieder, A; et al. Cas9-expressing chickens and pigs as resources for genome editing in livestock. Proceedings of the National Academy of Sciences 2021, 118, e2022562118. [Google Scholar] [CrossRef] [PubMed]
- Prather, RS; Lorson, M; Ross, JW; Whyte, JJ; Walters, E. Genetically engineered pig models for human diseases. Annu Rev Anim Biosci 2013, 1, 203–19. [Google Scholar] [CrossRef] [PubMed]
- Gao, X; Nowak-Imialek, M; Chen, X; Chen, D; Herrmann, D; Ruan, D; et al. Establishment of porcine and human expanded potential stem cells. Nat Cell Biol 2019, 21, 687–99. [Google Scholar] [CrossRef]
- Rawat, H; Kornherr, J; Zawada, D; Bakhshiyeva, S; Kupatt, C; Laugwitz, KL; et al. Recapitulating porcine cardiac development in vitro: from expanded potential stem cell to embryo culture models. Front Cell Dev Biol 2023, 11, 1111684. [Google Scholar] [CrossRef]
- Miller, JR; Henn, MC; Lancaster, TS; Lawrance, CP; Schuessler, RB; Shepard, M; et al. Pulmonary Valve Replacement With Small Intestine Submucosa-Extracellular Matrix in a Porcine Model. World J Pediatr Congenit Heart Surg 2016, 7, 475–83. [Google Scholar] [CrossRef]
- Cooper, DKC; Cozzi, E. Clinical Pig Heart Xenotransplantation—Where Do We Go From Here? Transplant International 2024, 37, 12592. [Google Scholar] [CrossRef]
- Garry, DJ; Garry, MG; Nakauchi, H; Masaki, H; Sachs, DH; Weiner, JI; et al. Allogeneic, Xenogeneic, and Exogenic Hearts for Transplantation. Methodist Debakey Cardiovasc J 2025, 21, 92–9. [Google Scholar] [CrossRef]
- Alibrandi, L; Lionetti, V. Interspecies differences in mitochondria: Implications for cardiac and vascular translational research. Vascul Pharmacol 2025, 159, 107476. [Google Scholar] [CrossRef]
- Horváth, B; Hézső, T; Szentandrássy, N; Kistamás, K; Árpádffy-Lovas, T; Varga, R; et al. Late sodium current in human, canine and guinea pig ventricular myocardium. J Mol Cell Cardiol 2020, 139, 14–23. [Google Scholar] [CrossRef]
- Linz, KW; Meyer, R. Profile and kinetics of L-type calcium current during the cardiac ventricular action potential compared in guinea-pigs, rats and rabbits. Pflügers Archiv 2000, 439, 588–99. [Google Scholar] [CrossRef] [PubMed]
- Pai, AA; Bell, JT; Marioni, JC; Pritchard, JK; Gilad, Y. A Genome-Wide Study of DNA Methylation Patterns and Gene Expression Levels in Multiple Human and Chimpanzee Tissues. PLoS Genet 2011, 7, e1001316. [Google Scholar] [CrossRef] [PubMed]
- Greiner, J; Sonak, F; Jones, WD; Madl, J; Ryeng, KA; Efimov, IR; et al. Architecture of the cardiac transverse-axial tubular system across different mammalian species. BioRxiv 2026, 2026.01, 10.698376. [Google Scholar] [CrossRef]
- Mulbjerg, H; Ringgaard, S; Agger, P. Diffusion tensor imaging reveals myocardial architectural differences between porcine and primate hearts with potential implications for cardiac xenotransplantation. Sci Rep 2025, 15, 28696. [Google Scholar] [CrossRef]
- Lee, KH; Lee, DW; Kang, BC. The ‘R’ principles in laboratory animal experiments. Lab Anim Res 2020, 36, 45. [Google Scholar] [CrossRef]
- Tannenbaum, J; Bennett, BT. Russell and Burch’s 3Rs then and now: the need for clarity in definition and purpose. J Am Assoc Lab Anim Sci 2015, 54, 120–32. [Google Scholar]
- Hubrecht, RC; Carter, E. The 3Rs and humane experimental technique: Implementing change. Animals 2019, 9, 754. [Google Scholar] [CrossRef]
- Viceconti, M; Emili, L; Afshari, P; Courcelles, E; Curreli, C; Famaey, N; et al. Possible Contexts of Use for In Silico Trials Methodologies: A Consensus-Based Review. IEEE J Biomed Health Inform 2021, 25, 3977–82. [Google Scholar] [CrossRef]
- Morotti, S; Liu, C; Hegyi, B; Ni, H; Fogli Iseppe, A; Wang, L; et al. Quantitative cross-species translators of cardiac myocyte electrophysiology: Model training, experimental validation, and applications. Sci Adv 2021, 7, eabg0927. [Google Scholar] [CrossRef]
- Cyganek, L; Tiburcy, M; Sekeres, K; Gerstenberg, K; Bohnenberger, H; Lenz, C; et al. Deep phenotyping of human induced pluripotent stem cell-derived atrial and ventricular cardiomyocytes. JCI Insight 2018, 3, e99941. [Google Scholar] [CrossRef]
- Protze, SI; Liu, J; Nussinovitch, U; Ohana, L; Backx, PH; Gepstein, L; et al. Sinoatrial node cardiomyocytes derived from human pluripotent cells function as a biological pacemaker. Nat Biotechnol 2017, 35, 56–68. [Google Scholar] [CrossRef] [PubMed]
- Wang, G; McCain, ML; Yang, L; He, A; Pasqualini, FS; Agarwal, A; et al. Modeling the mitochondrial cardiomyopathy of Barth syndrome with induced pluripotent stem cell and heart-on-chip technologies. Nat Med 2014, 20, 616–23. [Google Scholar] [CrossRef] [PubMed]
- Kime, C; Mandegar, MA; Srivastava, D; Yamanaka, S; Conklin, BR; Rand, TA. Efficient CRISPR/Cas9-Based Genome Engineering in Human Pluripotent Stem Cells. Curr Protoc Hum Genet 2016, 88, 21.4.1–21.4.23. [Google Scholar] [CrossRef] [PubMed]
- Bengel, P; Dybkova, N; Tirilomis, P; Ahmad, S; Hartmann, N; A. Mohamed, B; et al. Detrimental proarrhythmogenic interaction of Ca2+/calmodulin-dependent protein kinase II and NaV1.8 in heart failure. Nat Commun 2021, 12, 6586. [Google Scholar] [CrossRef]
- Stüdemann, T; Rössinger, J; Manthey, C; Geertz, B; Srikantharajah, R; von Bibra, C; et al. Contractile Force of Transplanted Cardiomyocytes Actively Supports Heart Function After Injury. Circulation 2022, 146, 1159–69. [Google Scholar] [CrossRef]
- Haertter, D; Hauke, L; Driehorst, T; Nishi, K; Zimmermann, W-H; Schmidt, CF. Sarcomere dynamic instability and stochastic heterogeneity drive robust cardiomyocyte contraction. BioRxiv 2024, 2024, 05.28.596183. [Google Scholar] [CrossRef]
- Moretti, A; Bellin, M; Welling, A; Jung, CB; Lam, JT; Bott-Flügel, L; et al. Patient-Specific Induced Pluripotent Stem-Cell Models for Long-QT Syndrome. New England Journal of Medicine 2010, 363, 1397–409. [Google Scholar] [CrossRef]
- Itzhaki, I; Maizels, L; Huber, I; Zwi-Dantsis, L; Caspi, O; Winterstern, A; et al. Modelling the long QT syndrome with induced pluripotent stem cells. Nature 2011, 471, 225–9. [Google Scholar] [CrossRef]
- Streckfuss-Bömeke, K; Tiburcy, M; Fomin, A; Luo, X; Li, W; Fischer, C; et al. Severe DCM phenotype of patient harboring RBM20 mutation S635A can be modeled by patient-specific induced pluripotent stem cell-derived cardiomyocytes. J Mol Cell Cardiol 2017, 113, 9–21. [Google Scholar] [CrossRef]
- Cuello, F; Knaust, AE; Saleem, U; Loos, M; Raabe, J; Mosqueira, D; et al. Impairment of the ER/mitochondria compartment in human cardiomyocytes with PLN p.Arg14del mutation. EMBO Mol Med 2021, 13. [Google Scholar] [CrossRef]
- Wu, X; Swanson, K; Yildirim, Z; Liu, W; Liao, R; Wu, JC. Clinical trials in-a-dish for cardiovascular medicine. Eur Heart J 2024, 45, 4275–90. [Google Scholar] [CrossRef] [PubMed]
- Prondzynski, M; Lemoine, MD; Zech, AT; Horváth, A; Di Mauro, V; Koivumäki, JT; et al. Disease modeling of a mutation in α-actinin 2 guides clinical therapy in hypertrophic cardiomyopathy. EMBO Mol Med 2019, 11, e11115. [Google Scholar] [CrossRef]
- Ma, N; Zhang, JZ; Itzhaki, I; Zhang, SL; Chen, H; Haddad, F; et al. Determining the Pathogenicity of a Genomic Variant of Uncertain Significance Using CRISPR/Cas9 and Human-Induced Pluripotent Stem Cells. Circulation 2018, 138, 2666–81. [Google Scholar] [CrossRef] [PubMed]
- Hinson, JT; Chopra, A; Nafissi, N; Polacheck, WJ; Benson, CC; Swist, S; et al. Titin mutations in iPS cells define sarcomere insufficiency as a cause of dilated cardiomyopathy. Science (1979) 2015, 349, 982–6. [Google Scholar] [CrossRef] [PubMed]
- Mannhardt, I; Breckwoldt, K; Letuffe-Brenière, D; Schaaf, S; Schulz, H; Neuber, C; et al. Human Engineered Heart Tissue: Analysis of Contractile Force. Stem Cell Reports 2016, 7, 29–42. [Google Scholar] [CrossRef]
- Kyrychenko, V; Kyrychenko, S; Tiburcy, M; Shelton, JM; Long, C; Schneider, JW; et al. Functional correction of dystrophin actin binding domain mutations by genome editing. JCI Insight 2017, 2, e95918. [Google Scholar] [CrossRef]
- Borchert, T; Hübscher, D; Guessoum, CI; Lam, T-DD; Ghadri, JR; Schellinger, IN; et al. Catecholamine-Dependent β-Adrenergic Signaling in a Pluripotent Stem Cell Model of Takotsubo Cardiomyopathy. J Am Coll Cardiol 2017, 70, 975–91. [Google Scholar] [CrossRef]
- Saleem, U; van Meer, BJ; Katili, PA; Mohd Yusof, NAN; Mannhardt, I; Garcia, AK; et al. Blinded, Multicenter Evaluation of Drug-induced Changes in Contractility Using Human-induced Pluripotent Stem Cell-derived Cardiomyocytes. Toxicological Sciences 2020, 176, 103–23. [Google Scholar] [CrossRef]
- Fomin, A; Gärtner, A; Cyganek, L; Tiburcy, M; Tuleta, I; Wellers, L; et al. Truncated titin proteins and titin haploinsufficiency are targets for functional recovery in human cardiomyopathy due to TTN mutations. Sci Transl Med 2021, 13, eabd3079. [Google Scholar] [CrossRef]
- Kyriakopoulou, E; Versteeg, D; de Ruiter, H; Perini, I; Seibertz, F; Döring, Y; et al. Therapeutic efficacy of AAV-mediated restoration of PKP2 in arrhythmogenic cardiomyopathy. Nature Cardiovascular Research 2023, 2, 1262–76. [Google Scholar] [CrossRef]
- Greer-Short, A; Greenwood, A; Leon, EC; Qureshi, TN; von Kraut, K; Wong, J; et al. AAV9-mediated MYBPC3 gene therapy with optimized expression cassette enhances cardiac function and survival in MYBPC3 cardiomyopathy models. Nat Commun 2025, 16, 2196. [Google Scholar] [CrossRef] [PubMed]
- Jebran, AF; Seidler, T; Tiburcy, M; Daskalaki, M; Kutschka, I; Fujita, B; et al. Engineered heart muscle allografts for heart repair in primates and humans. Nature 2025, 639, 503–11. [Google Scholar] [CrossRef] [PubMed]
- Tiburcy, M; Hudson, JE; Balfanz, P; Schlick, S; Meyer, T; Liao, MLC; et al. Defined engineered human myocardium with advanced maturation for applications in heart failure modeling and repair. Circulation 2017, 135, 1832–47. [Google Scholar] [CrossRef] [PubMed]
- Landau, S; Zhao, Y; Hamidzada, H; Kent, GM; Okhovatian, S; Lu, RXZ; et al. Primitive macrophages enable long-term vascularization of human heart-on-a-chip platforms. Cell Stem Cell 2024, 31, 1222–1238.e10. [Google Scholar] [CrossRef]
- Long, C; Li, H; Tiburcy, M; Rodriguez-Caycedo, C; Kyrychenko, V; Zhou, H; et al. Correction of diverse muscular dystrophy mutations in human engineered heart muscle by single-site genome editing. Sci Adv 2018, 4, eaap9004. [Google Scholar] [CrossRef]
- Mallapaty, S. Mini hearts, lungs and livers made in lab now grow their own blood vessels. Nature 2025, 643, 892–892. [Google Scholar] [CrossRef]
- Li, J; Wiesinger, A; Fokkert, L; Bakker, P; de Vries, DK; Tijsen, AJ; et al. Modeling the atrioventricular conduction axis using human pluripotent stem cell-derived cardiac assembloids. Cell Stem Cell 2024, 31, 1667–1684.e6. [Google Scholar] [CrossRef]
- Schneider, LV; Guobin, B; Methi, A; Jensen, O; Schmoll, KA; Setya, MG; et al. Bioengineering of a human innervated cardiac muscle model. BioRxiv 2023, 08.18.552653. [Google Scholar] [CrossRef]
- Emanuelli, G; Zoccarato, A; Reumiller, CM; Papadopoulos, A; Chong, M; Rebs, S; et al. A roadmap for the characterization of energy metabolism in human cardiomyocytes derived from induced pluripotent stem cells. J Mol Cell Cardiol 2022, 164, 136–47. [Google Scholar] [CrossRef]
- Feyen, DAM; McKeithan, WL; Bruyneel, AAN; Spiering, S; Hörmann, L; Ulmer, B; et al. Metabolic Maturation Media Improve Physiological Function of Human iPSC-Derived Cardiomyocytes. Cell Rep 2020, 32, 107925. [Google Scholar] [CrossRef]
- Li, W; Luo, X; Strano, A; Arun, S; Gamm, O; Poetsch, MS; et al. Comprehensive promotion of iPSC-CM maturation by integrating metabolic medium with nanopatterning and electrostimulation. Nat Commun 2025, 16, 2785. [Google Scholar] [CrossRef] [PubMed]
- Wickramasinghe, NM; Sachs, D; Shewale, B; Gonzalez, DM; Dhanan-Krishnan, P; Torre, D; et al. PPARdelta activation induces metabolic and contractile maturation of human pluripotent stem cell-derived cardiomyocytes. Cell Stem Cell 2022, 29, 559–576.e7. [Google Scholar] [CrossRef] [PubMed]
- Rebs, S; Eberl, H; Wagensohner, N; Dybkova, N; Unsöld, JK; Dudek, J; et al. Enhanced iPSC Cardiomyocyte Maturation via Combined 3D-Culture and Metabolic Cues. BioRxiv 2025, 2025, 10.29.684362. [Google Scholar] [CrossRef]
- Hirt, MN; Boeddinghaus, J; Mitchell, A; Schaaf, S; Börnchen, C; Müller, C; et al. Functional improvement and maturation of rat and human engineered heart tissue by chronic electrical stimulation. J Mol Cell Cardiol 2014, 74, 151–61. [Google Scholar] [CrossRef]
- Tan, SH; Ye, L. Maturation of Pluripotent Stem Cell-Derived Cardiomyocytes: a Critical Step for Drug Development and Cell Therapy. J Cardiovasc Transl Res 2018, 11, 375–92. [Google Scholar] [CrossRef]
- Yang, H; Yang, Y; Kiskin, FN; Shen, M; Zhang, JZ. Recent advances in regulating the proliferation or maturation of human-induced pluripotent stem cell-derived cardiomyocytes. Stem Cell Res Ther 2023, 14, 228. [Google Scholar] [CrossRef]
- Thomas, D; Cunningham, NJ; Shenoy, S; Wu, JC. Human-induced pluripotent stem cells in cardiovascular research: current approaches in cardiac differentiation, maturation strategies, and scalable production. Cardiovasc Res 2022, 118, 20–36. [Google Scholar] [CrossRef]
- Selfa Aspiroz, L; Mennecozzi, M; Batlle, L; Corneo, B; Healy, L; Kotter, M; et al. Promoting the adoption of best practices and standards to enhance quality and reproducibility of stem cell research. Stem Cell Reports 2025, 20, 102531. [Google Scholar] [CrossRef]
- Zhou, B; Shi, X; Tang, X; Zhao, Q; Wang, L; Yao, F; et al. Functional isolation, culture and cryopreservation of adult human primary cardiomyocytes. Signal Transduct Target Ther 2022, 7, 254. [Google Scholar] [CrossRef]
- Oh, JG; Kho, C; Hajjar, RJ; Ishikawa, K. Experimental models of cardiac physiology and pathology. Heart Fail Rev 2019, 24, 601–15. [Google Scholar] [CrossRef]
- Pitoulis, FG; Watson, SA; Perbellini, F; Terracciano, CM. Myocardial slices come to age: an intermediate complexity in vitro cardiac model for translational research. Cardiovasc Res 2020, 116, 1275–87. [Google Scholar] [CrossRef]
- Molina, CE; Abu-Taha, IH; Wang, Q; Roselló-Díez, E; Kamler, M; Nattel, S; et al. Profibrotic, electrical, and calcium-handling remodeling of the atria in heart failure patients with and without atrial fibrillation. Front Physiol 2018, 9, 1383. [Google Scholar] [CrossRef]
- Ahmad, S; Tirilomis, P; Pabel, S; Dybkova, N; Hartmann, N; Molina, CE; et al. The functional consequences of sodium channel NaV1.8 in human left ventricular hypertrophy. ESC Heart Fail 2019, 6, 154–63. [Google Scholar] [CrossRef] [PubMed]
- Grammatika Pavlidou, N; Dobrev, S; Beneke, K; Reinhardt, F; Pecha, S; Jacquet, E; et al. Phosphodiesterase 8 governs cAMP/PKA-dependent reduction of L-type calcium current in human atrial fibrillation: a novel arrhythmogenic mechanism. Eur Heart J 2023, 44, 2483–94. [Google Scholar] [CrossRef] [PubMed]
- Berisha, F; Götz, KR; Wegener, JW; Brandenburg, S; Subramanian, H; Molina, CE; et al. CAMP Imaging at Ryanodine Receptors Reveals β2-Adrenoceptor Driven Arrhythmias. Circ Res 2021, 129, 81–94. [Google Scholar] [CrossRef] [PubMed]
- Beneke, K; Molina, CE. Live Cell Imaging of Cyclic Nucleotides in Human Cardiomyocytes. Methods Mol Biol 2022, 2483, 195–204. [Google Scholar] [CrossRef]
- Pabel, S; Knierim, M; Stehle, T; Alebrand, F; Paulus, M; Sieme, M; et al. Effects of Atrial Fibrillation on the Human Ventricle. Circ Res 2022, 130, 994–1010. [Google Scholar] [CrossRef]
- Aceituno, C; Revuelta, D; Jiménez-Sábado, V; Ginel, A; Molina, CE; Hove-Madsen, L. Impact of Overnight Storage of Human Atrial Myocytes on Intracellular Calcium Homeostasis and Electrophysiological Utility. Biomolecules 2024, 14, 1415. [Google Scholar] [CrossRef]
- Mitcheson, JS; Hancox, JC; Levi, AJ. Action potentials, ion channel currents and transverse tubule density in adult rabbit ventricular myocytes maintained for 6 days in cell culture. Pflugers Arch 1996, 431, 814–27. [Google Scholar] [CrossRef]
- Banyasz, T; Lozinskiy, I; Payne, CE; Edelmann, S; Norton, B; Chen, B; et al. Transformation of adult rat cardiac myocytes in primary culture. Exp Physiol 2008, 93, 370–82. [Google Scholar] [CrossRef]
- Guo, GR; Chen, L; Rao, M; Chen, K; Song, JP; Hu, SS. A modified method for isolation of human cardiomyocytes to model cardiac diseases. J Transl Med 2018, 16, 288. [Google Scholar] [CrossRef] [PubMed]
- Seidel, T; Fiegle, DJ; Baur, TJ; Ritzer, A; Nay, S; Heim, C; et al. Glucocorticoids preserve the t-tubular system in ventricular cardiomyocytes by upregulation of autophagic flux. Basic Res Cardiol 2019, 114. [Google Scholar] [CrossRef] [PubMed]
- Greiner, J; Schiatti, T; Kaltenbacher, W; Dente, M; Semenjakin, A; Kok, T; et al. Consecutive-Day Ventricular and Atrial Cardiomyocyte Isolations from the Same Heart: Shifting the Cost–Benefit Balance of Cardiac Primary Cell Research. Cells 2022, 11. [Google Scholar] [CrossRef] [PubMed]
- Pfeilschifter, B; Martinez-Vilchez, A; Iqbal, Z; Potue, P; Fiegle, DJ; Morhenn, K; et al. Cold storage of mouse hearts prior to cardiomyocyte isolation preserves electromechanical function, microstructure, and gene expression for 24 h. Basic Res Cardiol 2025, 120, 1055–74. [Google Scholar] [CrossRef]
- Brandenburger, M; Wenzel, J; Bogdan, R; Richardt, D; Nguemo, F; Reppel, M; et al. Organotypic slice culture from human adult ventricular myocardium. Cardiovasc Res 2012, 93, 50–9. [Google Scholar] [CrossRef]
- Wang, Z; Tong, C; Xu, M; Feng, S; Dong, M; Rao, R; et al. Optimized methods for the functional cryopreservation of adult human primary cardiomyocytes. Cryobiology 2026, 122, 105583. [Google Scholar] [CrossRef]
- Fischer, C; Milting, H; Fein, E; Reiser, E; Lu, K; Seidel, T; et al. Long-term functional and structural preservation of precision-cut human myocardium under continuous electromechanical stimulation in vitro. Nat Commun 2019, 10, 117. [Google Scholar] [CrossRef]
- Watson, SA; Duff, J; Bardi, I; Zabielska, M; Atanur, SS; Jabbour, RJ; et al. Biomimetic electromechanical stimulation to maintain adult myocardial slices in vitro. Nat Commun 2019, 10, 2168. [Google Scholar] [CrossRef]
- Wang, K; Terrar, D; Gavaghan, DJ; Mu-u-min, R; Kohl, P; Bollensdorff, C. Living cardiac tissue slices: An organotypic pseudo two-dimensional model for cardiac biophysics research. Prog Biophys Mol Biol 2014, 115, 314–27. [Google Scholar] [CrossRef]
- Klumm, MJ; Heim, C; Fiegle, DJ; Weyand, M; Volk, T; Seidel, T. Long-Term Cultivation of Human Atrial Myocardium. Front Physiol 2022, 13, 839139. [Google Scholar] [CrossRef]
- van der Geest, JSA; de Boer, TP; Terracciano, CM; Thum, T; Dendorfer, A; Doevendans, PA; et al. Living myocardial slices: walking the path towards standardization. Cardiovasc Res 2025, 121, 1011–23. [Google Scholar] [CrossRef]
- Abu-Khousa, M; Fiegle, DJ; Sommer, ST; Minabari, G; Milting, H; Heim, C; et al. The Degree of t-System Remodeling Predicts Negative Force-Frequency Relationship and Prolonged Relaxation Time in Failing Human Myocardium. Front Physiol 2020, 11, 182. [Google Scholar] [CrossRef]
- Amesz, JH; Langmuur, SJJ; Zhang, L; Manintveld, OC; Schinkel, AFL; de Jong, PL; et al. Biomechanical response of ultrathin slices of hypertrophic cardiomyopathy tissue to myosin modulator mavacamten. Biomedicine and Pharmacotherapy 2024, 170, 116036. [Google Scholar] [CrossRef] [PubMed]
- Krammer, T; Baier, MJ; Hegner, P; Zschiedrich, T; Lukas, D; Wolf, M; et al. Cardioprotective effects of semaglutide on isolated human ventricular myocardium. Eur J Heart Fail 2025, 27, 1315–25. [Google Scholar] [CrossRef] [PubMed]
- Bierhuizen, MFA; Amesz, JH; Langmuur, SJJ; Lam, B; Knops, P; Veen, KM; et al. Acute Biomechanical Effects of Cardiac Contractility Modulation in Living Myocardial Slices from End-Stage Heart Failure Patients. Bioengineering 2025, 12, 174. [Google Scholar] [CrossRef] [PubMed]
- Shi, R; Reichardt, M; Fiegle, DJ; Küpfer, LK; Czajka, T; Sun, Z; et al. Contractility measurements for cardiotoxicity screening with ventricular myocardial slices of pigs. Cardiovasc Res 2023, 119, 2469–81. [Google Scholar] [CrossRef]
- Habeler, W; Peschanski, M; Monville, C. Organotypic heart slices for cell transplantation and physiological studies. Organogenesis 2009, 5, 62–6. [Google Scholar] [CrossRef]
- Baron, V; Sommer, ST; Fiegle, DJ; Pfeuffer, A-KM; Peyronnet, R; Volk, T; et al. Effects of electro-mechanical uncouplers, hormonal stimulation and pacing rate on the stability and function of cultured rabbit myocardial slices. Front Bioeng Biotechnol 2024, 12, 1363538. [Google Scholar] [CrossRef]
- Quinn, TA; Granite, S; Allessie, MA; Antzelevitch, C; Bollensdorff, C; Bub, G; et al. Minimum Information about a Cardiac Electrophysiology Experiment (MICEE): Standardised reporting for model reproducibility, interoperability, and data sharing. Prog Biophys Mol Biol 2011, 107, 4–10. [Google Scholar] [CrossRef]
- Niederer, SA; Lumens, J; Trayanova, NA. Computational models in cardiology. Nat Rev Cardiol 2019, 16, 100–11. [Google Scholar] [CrossRef]
- Trayanova, NA; Lyon, A; Shade, J; Heijman, J. Computational modeling of cardiac electrophysiology and arrhythmogenesis: toward clinical translation. Physiol Rev 2024, 104, 1265–333. [Google Scholar] [CrossRef]
- Lai, C; Yin, M; Kholmovski, EG; Sani, MM; Gilotra, NA; Chrispin, J; et al. Predicting Sudden Cardiac Death in Patients With Sarcoidosis Using a Multimodal Artificial Intelligence Model. JACC Clin Electrophysiol 2025, S2405-500X(25)00829-1. [Google Scholar] [CrossRef] [PubMed]
- Yamamoto, C; Sakata, K; Ali, SY; Loeffler, S; Prakosa, A; Tice, B; et al. Arrhythmogenic substrates in atrial fibrillation and the role of ablation lesions: A longitudinal biatrial digital twin study. Cardiovasc Res 2026. [Google Scholar] [CrossRef] [PubMed]
- Zhang, K; Magtibay, K; Trayanova, N; Vigmond, E. A model of β-adrenergic stimulation in human ventricular cells for tissue-scale simulations of sympathetically modulated tachycardias. J Physiol 2026. [Google Scholar] [CrossRef] [PubMed]
- Loewe, A; Hunter, PJ; Kohl, P. Computational modelling of biological systems now and then: revisiting tools and visions from the beginning of the century. Philosophical Transactions of the Royal Society A: Mathematical, Physical and Engineering Sciences 2025, 383, 20230384. [Google Scholar] [CrossRef]
- Quinn, TA; Kohl, P. Combining wet and dry research: experience with model development for cardiac mechano-electric structure-function studies. Cardiovasc Res 2013, 97, 601–11. [Google Scholar] [CrossRef]
- Roney, CH; Sim, I; Yu, J; Beach, M; Mehta, A; Alonso Solis-Lemus, J; et al. Predicting Atrial Fibrillation Recurrence by Combining Population Data and Virtual Cohorts of Patient-Specific Left Atrial Models. Circ Arrhythm Electrophysiol 2022, 15, e010253. [Google Scholar] [CrossRef]
- Corral-Acero, J; Margara, F; Marciniak, M; Rodero, C; Loncaric, F; Feng, Y; et al. The ‘Digital Twin’ to enable the vision of precision cardiology. Eur Heart J 2020, 41, 4556–64. [Google Scholar] [CrossRef]
- Shade, JK; Prakosa, A; Popescu, DM; Yu, R; Okada, DR; Chrispin, J; et al. Predicting risk of sudden cardiac death in patients with cardiac sarcoidosis using multimodality imaging and personalized heart modeling in a multivariable classifier. Sci Adv 2021, 7, eabi8020. [Google Scholar] [CrossRef]
- Sakata, K; Bradley, RP; Prakosa, A; Yamamoto, CAP; Ali, SY; Loeffler, S; et al. Assessing the arrhythmogenic propensity of fibrotic substrate using digital twins to inform a mechanisms-based atrial fibrillation ablation strategy. Nature Cardiovascular Research 2024, 3, 857–68. [Google Scholar] [CrossRef]
- Sakata, K; Yamamoto, CAP; Ali, SY; Loeffler, S; Prakosa, A; Tice, BM; et al. Assessment of persistent atrial fibrillation arrhythmogenesis in the right atrium using digital twins. Heart Rhythm 2026, 23, e123–32. [Google Scholar] [CrossRef]
- Waight, MC; Prakosa, A; Li, AC; Truong, A; Bunce, N; Marciniak, A; et al. Heart Digital Twins Predict Features of Invasive Reentrant Circuits and Ablation Lesions in Scar-Dependent Ventricular Tachycardia. Circ Arrhythm Electrophysiol 2025, 18, e013660. [Google Scholar] [CrossRef]
- Vicente, J; Zusterzeel, R; Johannesen, L; Mason, J; Sager, P; Patel, V; et al. Mechanistic Model-Informed Proarrhythmic Risk Assessment of Drugs: Review of the “CiPA” Initiative and Design of a Prospective Clinical Validation Study. Clin Pharmacol Ther 2018, 103, 54–66. [Google Scholar] [CrossRef] [PubMed]
- Li, Z; Mirams, GR; Yoshinaga, T; Ridder, BJ; Han, X; Chen, JE; et al. General Principles for the Validation of Proarrhythmia Risk Prediction Models: An Extension of the CiPA In Silico Strategy. Clin Pharmacol Ther 2020, 107, 102–11. [Google Scholar] [CrossRef] [PubMed]
- Food and Drug Administration. FDA Announces Plan to Phase Out Animal Testing Requirement for Monoclonal Antibodies and Other Drugs 2025. Available online: https://www.fda.gov/news-events/press-announcements/fda-announces-plan-phase-out-animal-testing-requirement-monoclonal-antibodies-and-other-drugs (accessed on 10 February 2026).
- National Institutes of Health. NIH to prioritize human-based research technologies 2025. Available online: https://www.nih.gov/news-events/news-releases/nih-prioritize-human-based-research-technologies (accessed on 10 February 2026).
- Ridder, BJ; Leishman, DJ; Bridgland-Taylor, M; Samieegohar, M; Han, X; Wu, WW; et al. A systematic strategy for estimating hERG block potency and its implications in a new cardiac safety paradigm. Toxicol Appl Pharmacol 2020, 394, 114961. [Google Scholar] [CrossRef] [PubMed]
- Assessing the credibility of computational modeling through verification and validation: application to medical devices; American Society of Mechanical Engineers, 2018.
- Toward Good Simulation Practice; Viceconti, M, Emili, L, Eds.; Springer Nature Switzerland: Cham, 2024. [Google Scholar] [CrossRef]
- Iaizzo, PA. The Visible Heart® project and free-access website “Atlas of Human Cardiac Anatomy”. Europace 2016, 18, iv163–72. [Google Scholar] [CrossRef]
- Bunnik, EM. Ethics of allocation of donor organs. Curr Opin Organ Transplant 2023, 28, 192–6. [Google Scholar] [CrossRef]
- Pick, F; Krug, N. Was Deutschland von anderen Ländern lernen kann. Spiegel. 2024. Available online: https://www.spiegel.de/gesundheit/organspende-register-was-deutschland-von-anderen-laendern-lernen-kann-a-d1eecb0f-27d0-4bc8-9d58-2c22bacf469a (accessed on 22 October 2024).
- Eurotransplant. Deutschland 2024. Available online: https://www.eurotransplant.org/region/deutschland/ (accessed on 9 October 2024).
- Eurotransplant Statistics Report Library. Heart donation, deceased donors in 2023, by country, by allocation phase. n.d. [Google Scholar]
- IRODaT. 2024. Available online: https://www.irodat.org/?p=database&c=DE#data (accessed on 9 October 2024).
- Statista. Number of postmortem organ donors in Germany from 1998 to 2023 2024. Available online: https://www.statista.com/statistics/1384771/postmortem-organ-donors-germany/ (accessed on 9 October 2024).
- Mumford, L. NHS BLOOD & TRANSPLANT RESEARCH, INNOVATION AND NOVEL TECHNOLOGIES ADVISORY GROUP RESEARCH CONSENT/AUTHORISATION RATES. 2022.
- Bledsoe, MJ; Grizzle, WE. Use of human specimens in research: the evolving United States regulatory, policy, and scientific landscape. Diagn Histopathol 2013, 19, 322–30. [Google Scholar] [CrossRef]
- Grizzle, WE. Issues in the Use of Human Tissues to Support Precision Medicine. J Health Care Poor Underserved 2019, 30, 66–78. [Google Scholar] [CrossRef]
- Lutomski, JE; Manders, P. From opt-out to opt-in consent for secondary use of medical data and residual biomaterial: An evaluation using the RE-AIM framework. PLoS One 2024, 19, e0299430. [Google Scholar] [CrossRef]
- Fitzpatrick, PE; McKenzie, KD; Beasley, A; Sheehan, JD. Patients attending tertiary referral urology clinics: Willingness to participate in tissue banking. BJU Int 2009, 104, 209–13. [Google Scholar] [CrossRef]
- Jack, AL; Womack, C. Why surgical patients do not donate tissue for commercial research: review of records. BMJ 2003, 327, 262. [Google Scholar] [CrossRef]
- Rehsmann, J. A revealing scandal: The German transplant scandal between structural failures, moralizing rules, and ambivalent manipulations. Journal of Liver Transplantation 2023, 11, 100168. [Google Scholar] [CrossRef]
- Deutscher Bundestag. Organspenden: Mehrheit für die Entscheidungslösung 2020. Available online: https://www.bundestag.de/dokumente/textarchiv/2020/kw03-de-transplantationsgesetz-674682 (accessed on 9 October 2024).
- Deutsche Welle. German lawmakers reject “opt-out” organ donor bill 2020. Available online: https://www.dw.com/en/german-parliament-explicit-consent-still-necessary-from-organ-donors/a-52022245 (accessed on 9 October 2024).
- Kappler, HE; Kohl, P. Wissenschaftliche Nutzung von nicht transplantiertem Humangewebe. CardioNews 2023. Available online: https://app.cardionews.de/Politik-Gesellschaft/Wissenschaftliche-Nutzung-von-nicht-transplantiertem-Humangewebe-442919.html (accessed on 23 February 2026).
- Deutscher Bundestag 20. Wahlperiode. Antwort der Bundesregierung auf die Kleine Anfrage der Fraktion der CDU/CSU – Drucksache 20/7480 – Nutzung postmortaler Spender-Organe zu Forschungszwecken. Drucksache 20/7731 2023. Available online: https://dserver.bundestag.de/btd/20/077/2007731.pdf (accessed on 23 February 2026).
- Deutscher Bundestag 20. Wahlperiode. Kleine Anfrage der Fraktion der CDU/CSU Nutzung postmortaler Spender-Organe zu Forschungszwecken. Drucksache 20/7480 2023. Available online: https://dserver.bundestag.de/btd/20/074/2007480.pdf (accessed on 23 February 2026).
- Fedlex. Federal Act of 30 September 2011 on Research involving Human Beings, SR 810. 30 2011. Available online: https://www.fedlex.admin.ch/eli/cc/2013/617/en (accessed on 6 January 2026).
- Federal Public Health Food Chain Safety and Environment. Organ donation and donation of body to science. Federal Public Health Food Chain Safety and Environment 2016. Available online: https://www.health.belgium.be/en/organ-donation-and-donation-body-science (accessed on 20 December 2025).
- Ministry of Health Welfare and Sport Donor Register. Scientific Research 2025. Available online: https://english.donorregister.nl/recording-your-choise/what-choices-do-you-have/scientific-research (accessed on 9 January 2026).
- Ustaw, Dziennik. Ustawa z dnia 1 lipca 2005 r. o pobieraniu, przechowywaniu i przeszczepianiu komórek, tkanek i narządów [Act of 1 July 2005 on the Collection, Storage, and Transplantation of Cells, Tissues, and Organs.] Dz. U. 2005 No. 169 item 141 n.d. Available online: https://isap.sejm.gov.pl/isap.nsf/DocDetails.xsp?id=WDU20051691411 (accessed on 9 January 2026).
- Retsinformation.dk. Vejledning om samtykke til transplantation fra afdøde personer og til transplantationsrelateret forskning [Guidance on consent for transplantation from deceased persons and for transplantation-related research], VEJ nr 10009 of 05/12/2019. RetsinformationDk n.d. Retsinformation.dk. Available online: https://www.retsinformation.dk/eli/retsinfo/2019/10099 (accessed on 9 January 2026).
- Agence de la biomédicine. Baromètre d’opinion 2024 2024. Available online: https://back.agence-biomedecine.fr/uploads/Dossier_de_presse_complet_en_PDF_6598766daa.pdf (accessed on 8 January 2026).
- Rechtsinformationssystem des Bundes. Organtransplantationsgesetz [Federal Act on the Transplantation of Human Organs], §2, Abs. 2. n.d. Available online: https://www.ris.bka.gv.at/GeltendeFassung.wxe?Abfrage=Bundesnormen&Gesetzesnummer=20008119 (accessed on 9 January 2026).
- NHS Blood and Transplant. Understanding consent for organ donation. n.d. Available online: https://www.organdonation.nhs.uk/about-organ-donation/consent/ (accessed on 23 February 2026).
| Parameter | Non-mammalian, e.g., zebrafish |
Small mammal, e.g., mouse |
Intermediate mammal, e.g., rabbit |
Large mammal, e.g., pig |
Human |
| Body weight | 0.5 g | 20–45 g | 1-5 kg | 40–90 kg | 50–100 kg |
| Heart weight | 0.5 mg | 0.15–0.2 g | 5–10 g | 145–450 g | 150–480 g |
| Resting heart rate | 110–170 bpm | 450–800 bpm | 140-280 bpm | 70–120 bpm | 60–100 bpm |
| Cardiac output | 0.09–0.172 mL/min | 13–15 mL/min | 260-420 mL/min | 2,720–3,620 mL/min | 3,340–9,790 mL/min |
| Ejection fraction | 40–50% | 65–90% | 55–65% | 50–55% | 55–60% |
| Ventricular action potential duration | 150–250 ms | 80–180 ms | 150–280 ms | 180–300 ms | 200–400 ms |
| Conduction velocity | 4–10 mm/s | 350–600 mm/s | 300-500 mm/s | 500–800 mm/s | 500–900 mm/s |
| Reentry space factora | 0.42 | 0.34 | 0.69 | 1.43 | 1.5 |
| Approximate cost per animal | €2.00–2.50/tank | €4–7 | €250–600 | €80–400+ | - |
| Husbandry notes | Swarm fish, density 4–5 animals per litre | Individually ventilated or standard cages | Larger pens | Large facilities or catheterisation laboratories, low volume but translational | - |
| Anaesthesia, and effects on cardiovascular endpoints | Tricaine: bradycardia, decreased heart rate and contractility, effects dose-dependent | Isoflurane and opioids: decreased heart rate and blood pressure | Volatile or injectable anaesthetics: altered haemodynamics, effects on heart rhythm | Isoflurane and similar: more frequent arrhythmias, infarct variability, effects on blood pressure effects | - |
| Typical n per group in cardiovascular studies | 30–100+ embryos | 8–15 | 6–10 | 6–12 | - |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




