Submitted:
21 February 2026
Posted:
27 February 2026
You are already at the latest version
Abstract
Keywords:
Introduction and Background
CAR-T vs TCR-T vs ICB
Literature Review
Biology of T-Cell Exhaustion and Transcriptional Control (TOX, NR4A, TCF1, T-BET/EOMES)
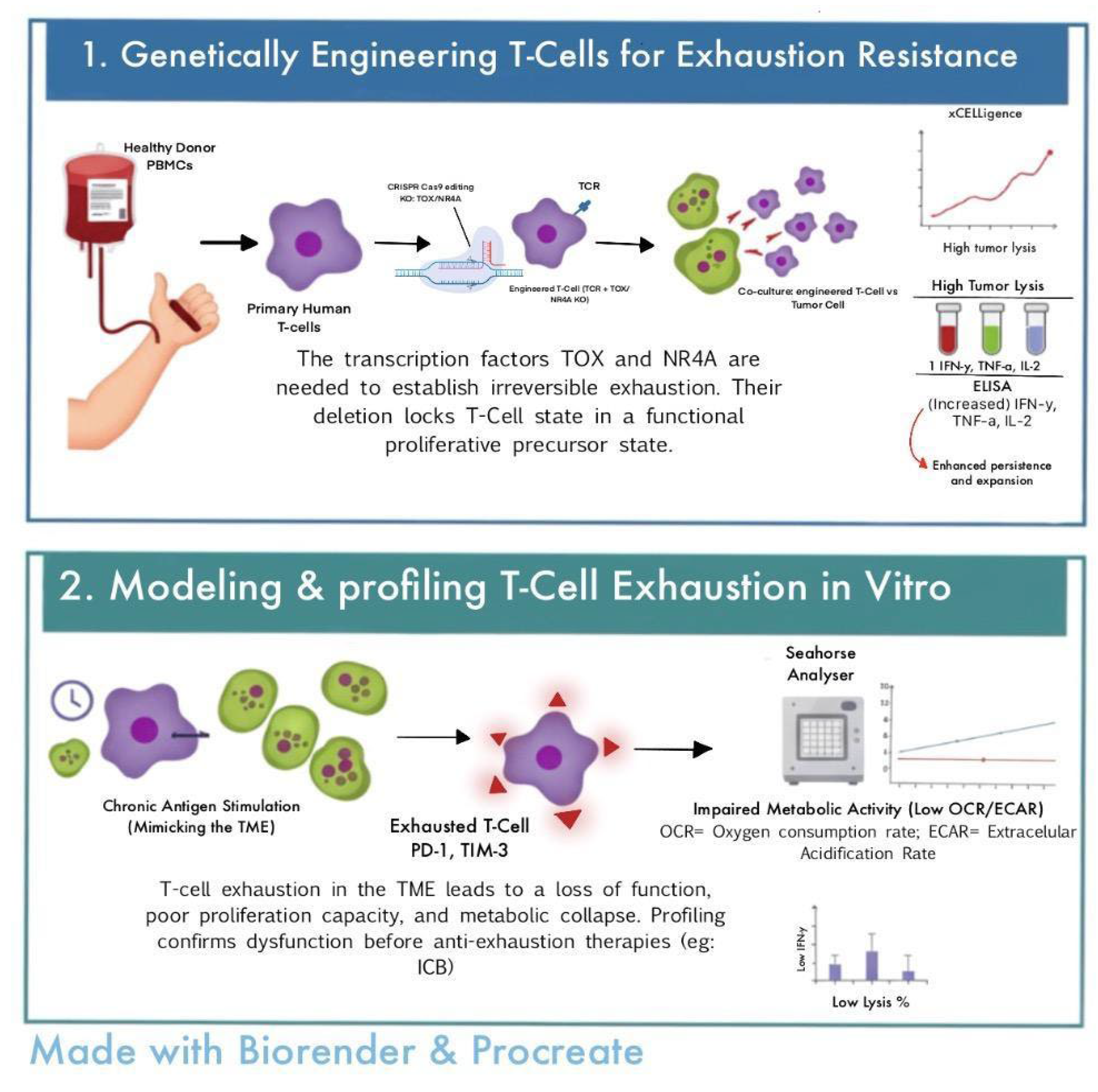
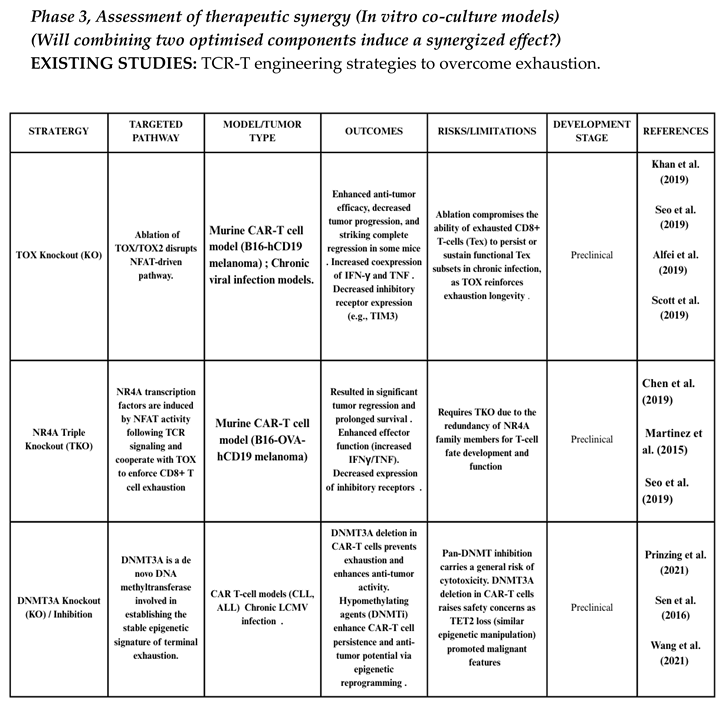
TCR-T Engineering Strategies to Overcome Exhaustion
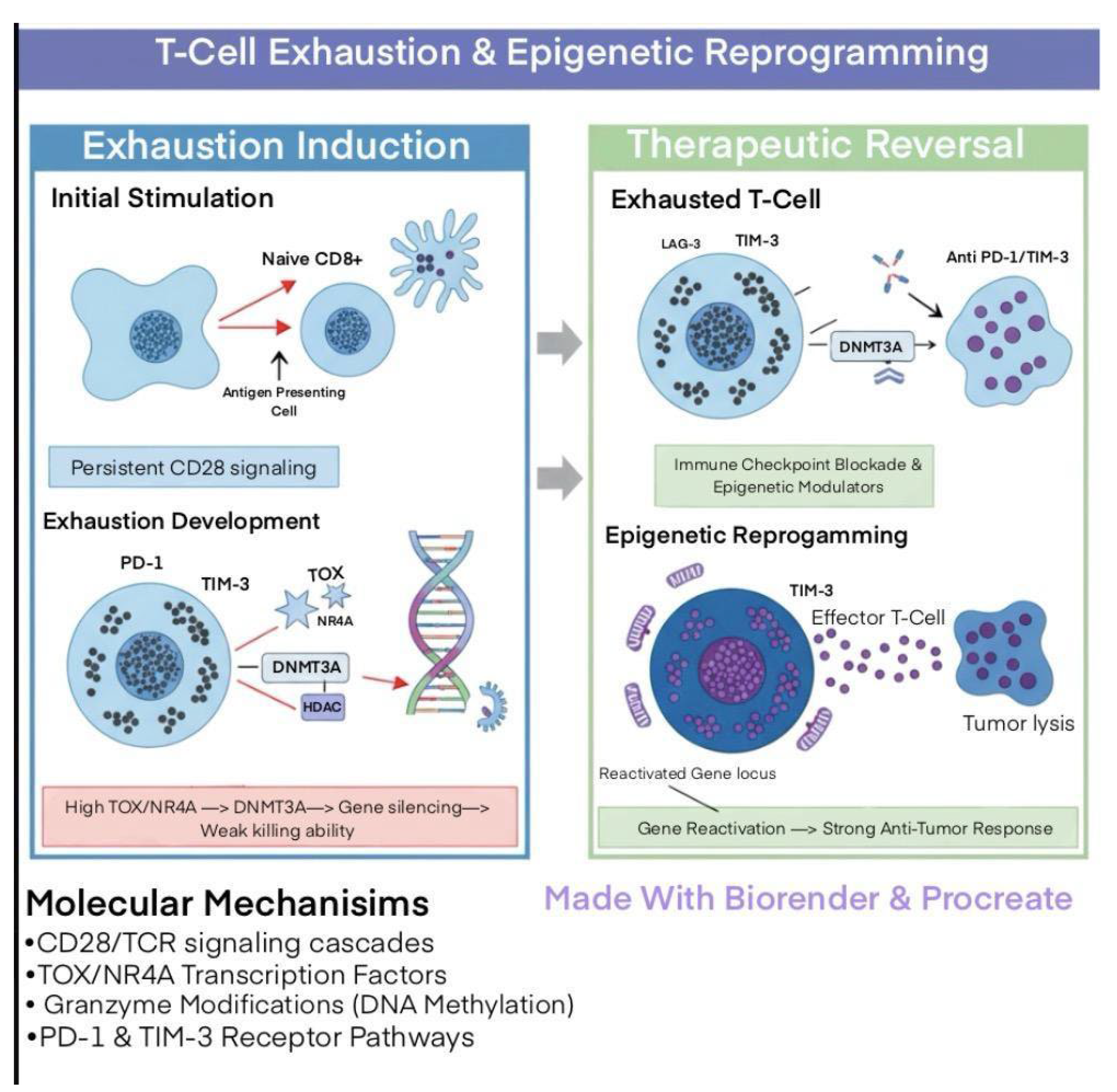
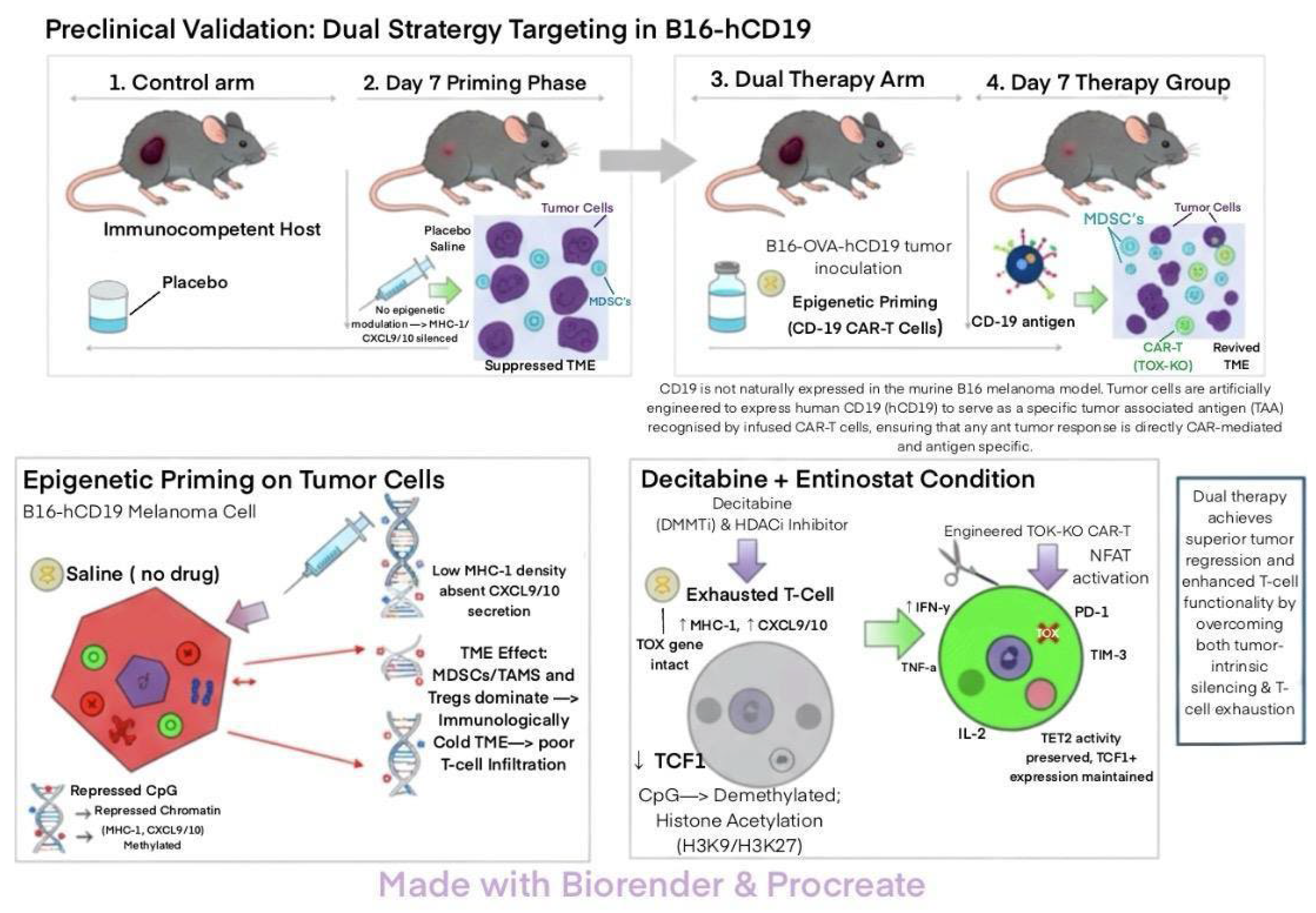
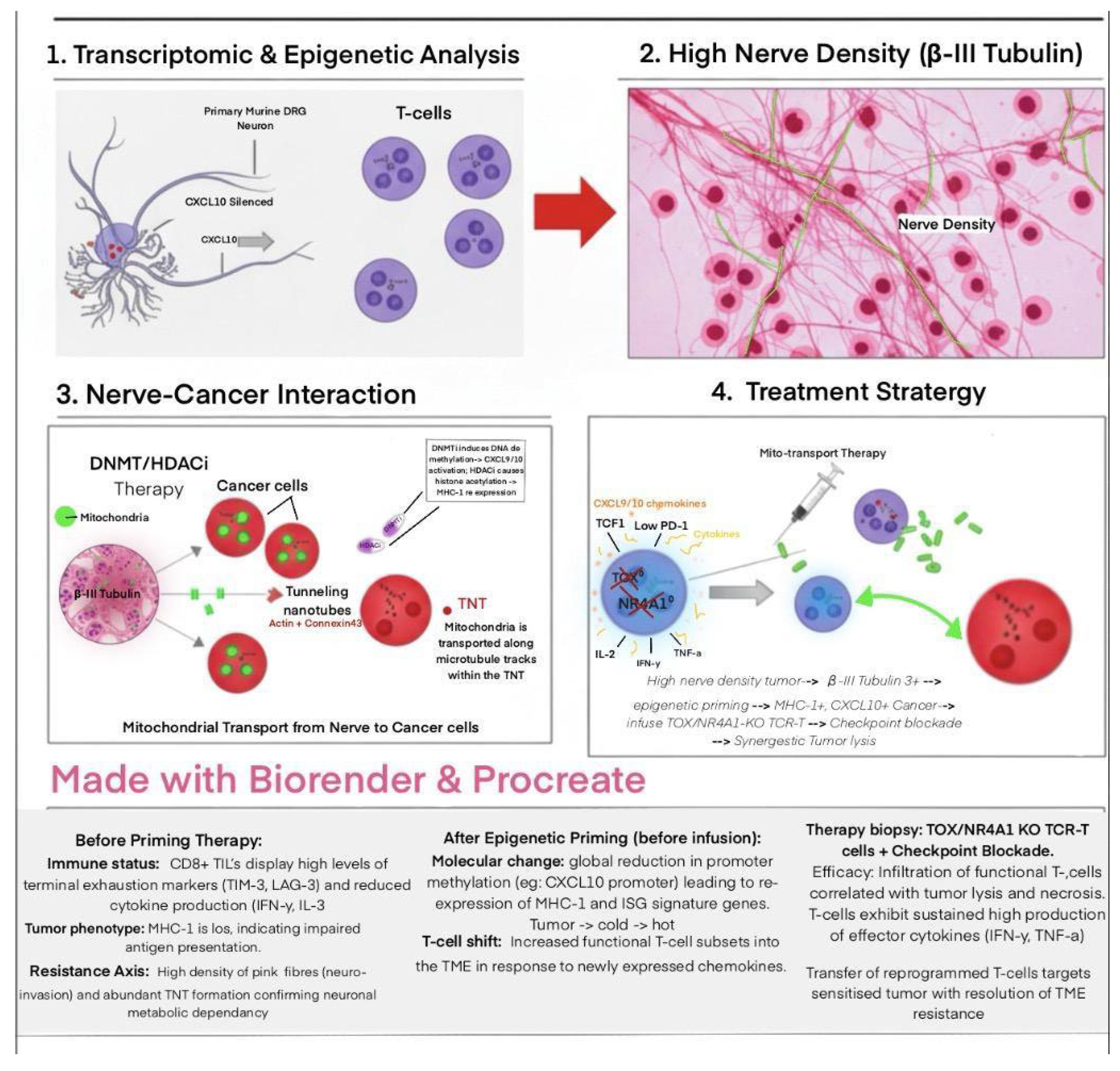
Epigenetic Reprogramming of Tumors to Restore Immunogenicity
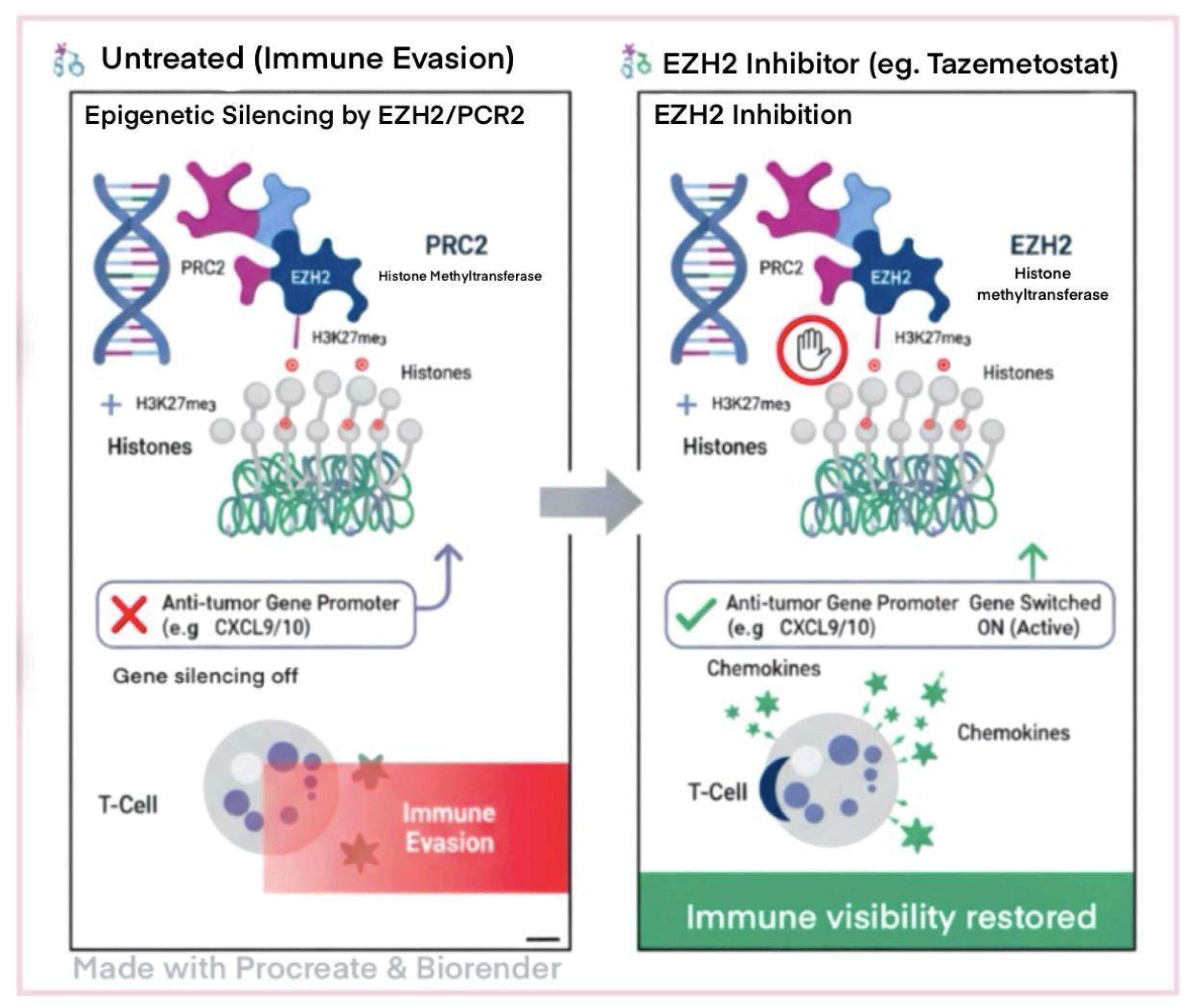
Evidence for Combination Strategies with T-Cell Therapies and ICB
Personalized Biomarker-Guided Selection and Monitoring
Limitations, Safety, Translational, and Regulatory Considerations
Challenges, Safety, and Regulatory Considerations
Future Directions and Open Questions
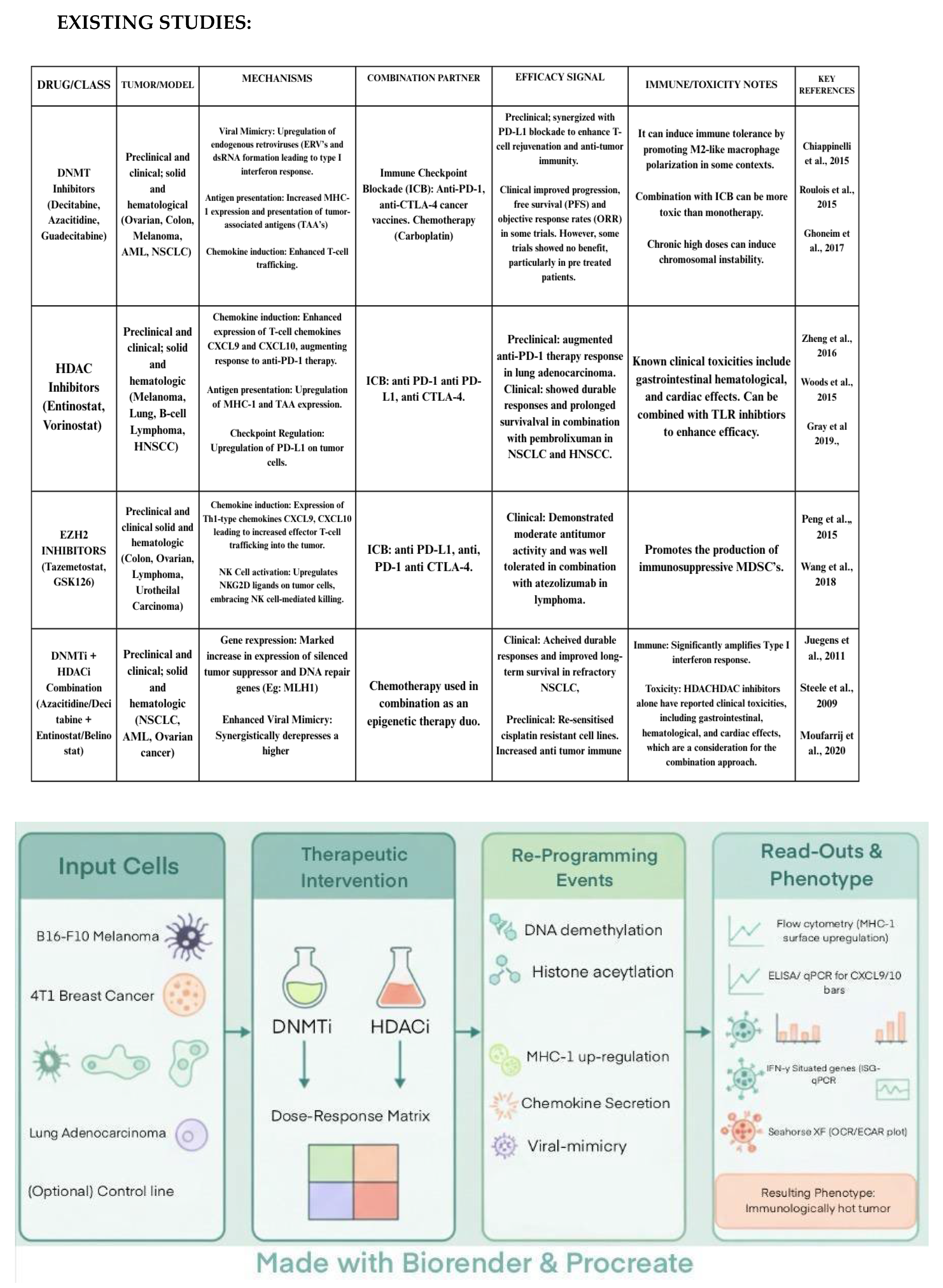
Conclusions
References
- Ai, L.; et al. BPQDs@Lipo-YSA Nanoplatform Triggers Mitophagy via PRKN/AKT1 to Drive Immunogenic Cell Death in Lung Adenocarcinoma. Journal of Nanobiotechnology 2025, 23(1). Available online: https://link.springer.com/article/10.1186/s12951-025-03496-7. [CrossRef]
- Chen, M.; Xie, S. Therapeutic targeting of cellular stress responses in cancer. Thoracic Cancer 2018, 9(12), 1575–1582. Available online: https://pmc.ncbi.nlm.nih.gov/articles/PMC6275842/. [CrossRef] [PubMed]
- Yang, H.; et al. The role of cellular reactive oxygen species in cancer chemotherapy. Journal of Experimental & Clinical Cancer Research 2018, 37(1). Available online: https://pmc.ncbi.nlm.nih.gov/articles/PMC6211502/. [CrossRef] [PubMed]
- Schöckel, L.; et al. Targeting mitochondrial complex I using BAY 87-2243 reduces melanoma tumor growth. Cancer & Metabolism 2015, 3(1). Available online: https://pubmed.ncbi.nlm.nih.gov/26500770/.
- DasGupta, R. K.; et al. A review of CD19-targeted immunotherapies for relapsed or refractory acute lymphoblastic leukemia. Journal of Oncology Pharmacy Practice 2017, 24(6), 453–467. Available online: https://pubmed.ncbi.nlm.nih.gov/28583018/. [CrossRef]
- Aldoss, I.; et al. Correlates of resistance and relapse during blinatumomab therapy for relapsed/refractory acute lymphoblastic leukemia. American Journal of Hematology 2017, 92(9), 858–865. Available online: https://onlinelibrary.wiley.com/doi/10.1002/ajh.24783. [CrossRef]
- Lemoine, J.; Ruella, M.; Houot, R. Born to survive: how cancer cells resist CAR T cell therapy. Journal of Hematology & Oncology 2021, 14(1). Available online: https://link.springer.com/article/10.1186/s13045-021-01209-9. [CrossRef] [PubMed]
- Maalej, K. M.; et al. CAR-cell therapy in the era of solid tumor treatment: current challenges and emerging therapeutic advances. In Molecular Cancer; 2023; 22, 1. Available online: https://pmc.ncbi.nlm.nih.gov/articles/PMC9885707/.
- Das, S.; Johnson, D. B. Immune-related adverse events and anti‑tumor efficacy of immune checkpoint inhibitors. Journal for ImmunoTherapy of Cancer 2019, 7, 306. Available online: https://pubmed.ncbi.nlm.nih.gov/31730012/. [CrossRef]
- Zhang, J.; Wang, L. The Emerging World of TCR-T Cell Trials Against Cancer: A Systematic Review. Technology in Cancer Research & Treatment 2019, 18. Available online: https://pmc.ncbi.nlm.nih.gov/articles/PMC6391541/.
- Li, Y.; Halladay, T.; Yang, L. Immune evasion in cell-based immunotherapy: unraveling challenges and novel strategies. Journal of Biomedical Science 2024, 31(1). Available online: https://link.springer.com/article/10.1186/s12929-024-00998-8. [CrossRef]
- Sekine, et al. TOX is expressed by exhausted and polyfunctional human effector memory CD8+ T cells. Science Immunology 2020, 5(49), eaba7918, https://www.science.org/doi/10.1126/sciimmunol.aba7918. [Google Scholar] [CrossRef]
- Wang, B.; et al. Overcoming acquired resistance to cancer immune checkpoint therapy: potential strategies based on molecular mechanisms. Cell & Bioscience 2023, 13(1). Available online: https://link.springer.com/article/10.1186/s13578-023-01073-9. [CrossRef]
- Kim, J.; et al. Targeted deletion of CD244 on monocytes promotes differentiation into anti-tumorigenic macrophages and potentiates PD-L1 blockade in melanoma. Molecular Cancer 2024, 23(1). Available online: https://link.springer.com/article/10.1186/s12943-024-01936-w. [CrossRef]
- Moseman, E. A.; et al. The TCF1-Bcl6 axis counteracts type I interferon to repress exhaustion and maintain T cell stemness. Science Immunology 2016, 1(6), eaai8593. Available online: https://pmc.ncbi.nlm.nih.gov/articles/PMC5179228/. [CrossRef] [PubMed]
- Knudson, K. M.; et al. NFκB–Pim-1–Eomesodermin axis is critical for maintaining CD8 T-cell memory quality. Proceedings of the National Academy of Sciences 2017, 114(9). Available online: https://pmc.ncbi.nlm.nih.gov/articles/PMC5338529/. [CrossRef] [PubMed]
- Li, F.; et al. Metabolic plasticity and regulation of T cell exhaustion. Immunology 2022, 167(4), 482–494. Available online: https://onlinelibrary.wiley.com/doi/10.1111/imm.13575. [CrossRef] [PubMed]
- Seo, H.; et al. TOX and TOX2 transcription factors cooperate with NR4A transcription factors to impose CD8 + T cell exhaustion. Proceedings of the National Academy of Sciences 2019, 116(25), 12410–12415. Available online: https://pmc.ncbi.nlm.nih.gov/articles/PMC6589758/. [CrossRef]
- Hu, Y.; Dong, Z.; Liu, K. Unraveling the complexity of STAT3 in cancer: molecular understanding and drug discovery. Journal of Experimental & Clinical Cancer Research 2024, 43(1). Available online: https://pubmed.ncbi.nlm.nih.gov/38245798/. [CrossRef]
- Casey, S. C.; et al. MYC regulates the antitumor immune response through CD47 and PD-L1. Science 2016, 352(6282), 227–231. Available online: https://pubmed.ncbi.nlm.nih.gov/26966191/. [CrossRef]
- Lodewijk, I.; et al. Tackling tumor microenvironment through epigenetic tools to improve cancer immunotherapy. Clinical Epigenetics 2021, 13(1). Available online: https://pubmed.ncbi.nlm.nih.gov/33761971/. [CrossRef]
- Jin, N.; et al. Advances in epigenetic therapeutics with a focus on solid tumors. Clinical Epigenetics 2021, 13(1). Available online: https://pmc.ncbi.nlm.nih.gov/articles/PMC8056722/.
- Sun, X.; et al. Orchestrated Cu2+-coordinated tetracycline-porphyrin self-assembly remodels tumor microenvironment for photo-enhanced immuno-chemodynamic therapy. Journal of Nanobiotechnology 2025, 23(1). Available online: https://pmc.ncbi.nlm.nih.gov/articles/PMC12139202/. [CrossRef]
- Lodewijk, I.; et al. Tackling tumor microenvironment through epigenetic tools to improve cancer immunotherapy; 2021. [Google Scholar]
- Liu, Q.; et al. Adoptive cellular immunotherapy for solid neoplasms beyond car-t. Molecular Cancer 2023, 22(1). [Google Scholar] [CrossRef]
- Emami Nejad, A.; et al. The role of hypoxia in the tumor microenvironment and development of cancer stem cell: A novel approach to developing treatment. Cancer Cell International 2021, 21(1). [Google Scholar] [CrossRef]
- Tie, Y.; et al. Immunosuppressive cells in cancer: Mechanisms and potential therapeutic targets. Journal of Hematology & Oncology 2022, 27 15(1). [Google Scholar] [CrossRef]
- Topalian, S. L.; Taube, J. M.; Pardoll, D. M. Neoadjuvant checkpoint blockade for cancer immunotherapy Neoadjuvant checkpoint blockade for cancer immunotherapy | Science. Science 2020, 367(6477). [Google Scholar]
- Zou, S.; et al. Targeting Stat3 in cancer immunotherapy. Molecular Cancer 2020, 29 19(1). [Google Scholar] [CrossRef]
- BioRender BioRender [Graphic design software]. 2024. Available online: https://www.biorender.com.
- Tillé, L. Activation of the transcription factor NFAT5 in the tumor microenvironment enforces CD8+ T cell exhaustion. Nat Immunol 2023, 24, 1645–1653. [Google Scholar] [CrossRef] [PubMed]
- Sekiya, T. Comparison Between Nr4a Transcription Factor Regulation and Function in Lymphoid and Tumor Treg Cells. Front. Immunol. 2022, 13, 866339. [Google Scholar] [CrossRef]
- Wu, Y.; Wu, Y.; Gao, Z.; et al. Revitalizing T cells: breakthroughs and challenges in overcoming T cell exhaustion. Sig Transduct Target Ther 2026, 11, 2. [Google Scholar] [CrossRef]
- Bradley, C.A. PRC2-mediated MHC-I silencing drives immune evasion. Nat Rev Cancer 2019, 19, 664. [Google Scholar] [CrossRef]
- Tu, Y.; Zhu, Qy.; Huang, Wj.; et al. DNMT inhibition epigenetically restores the cGAS-STING pathway and activates RIG-I/MDA5-MAVS to enhance antitumor immunity. Acta Pharmacol Sin47 2026, 197–208. [Google Scholar] [CrossRef]
- Matei; et al. 2012 Epigenetic Resensitization to Platinum in Ovarian Cancer. Cancer Res 1 May 2012, 72(9), 2197–2205. [Google Scholar] [CrossRef] [PubMed]
- Vukadin, S; Khaznadar, F; Kizivat, T; Vcev, A; Smolic, M. Molecular Mechanisms of Resistance to Immune Checkpoint Inhibitors in Melanoma Treatment: An Update. Biomedicines 2021, 9(7), 835. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Hoover, G.; Gilbert, S.; Curley, O.; et al. Nerve-to-cancer transfer of mitochondria during cancer metastasis. Nature 2025, 644, 252–262. [Google Scholar] [CrossRef]
- Baulu, E.; Gardet, C.; Chuvin, N.; Depil, S. TCR-engineered T cell therapy in solid tumors: State of the art and perspectives. 2023. [Google Scholar] [CrossRef]
- Daskalakis, M.; Brocks, D.; Sheng, Y.; Islam, M.; Ressnerova, A.; Assenov, Y.; Milde, T.; Oehme, I.; Witt, O.; Goyal, A.; Kühn, A.; Hartmann, M.; Weichenhan, D.; Jung, M.; Plass, C. Reactivation of endogenous retroviral elements via treatment with DNMT- and HDAC-inhibitors. 2018. [Google Scholar] [CrossRef] [PubMed]
- Duan, R.; Du, W.; Guo, W. EZH2: a novel target for cancer treatment. 2020. [Google Scholar] [CrossRef] [PubMed]
- Pauken, K.; Sammons, M.; Odorizzi, P.; Manne, S.; Godec, J.; Khan, O.; Drake, A.; Chen, Z.; Sen, D.; Kurachi, M.; Barnitz, R.; Bartman, C.; Bengsch, B.; Huang, A.; Schenkel, J.; Vahedi, G.; Haining, W.; Berger, S.; Wherry, E. Epigenetic stability of exhausted T cells limits durability of reinvigoration by PD-1 blockade. 2016. [Google Scholar] [CrossRef]
- Shao, W.; Yao, Y.; Yang, L.; Li, X.; Ge, T.; Zheng, Y.; Zhu, Q.; Ge, S.; Gu, X.; Jia, R.; Song, X.; Zhuang, A. Novel insights into TCR-T cell therapy in solid neoplasms: optimizing adoptive immunotherapy. 2024. [Google Scholar] [CrossRef]
- Tufail, M.; Jiang, C.; Li, N. Altered metabolism in cancer: insights into energy pathways and therapeutic targets. 2024. [Google Scholar] [CrossRef]
- Wang, L.; Zhu, S.; Ding, Z.; Hong, J.; Zheng, L.; Sun, J.; Tang, Y.; Chen, X.; Yong, X.; Hu, M.; Zhimin, J.; Liu, Y.; Wang, D.; Tang, H. Cell-type specific activation of the cGAS-STING pathway in tumor immunotherapy: mechanisms and therapeutic implications. 2026. [Google Scholar] [CrossRef]
- Yu, Y.; Yao, X.; Wang, Q.; Yang, M.; Li, R.; Qin, J.; Zhuang, J.; Sun, C. T Cell Exhaustion in Cancer Immunotherapy: Heterogeneity, Mechanisms, and Therapeutic Opportunities. 2026. [Google Scholar] [CrossRef] [PubMed]
- Xu, K.; et al. Glycolysis fuels phosphoinositide 3-kinase signaling to bolster T cell immunity. Science 2021, 371(6527), 405–410. [Google Scholar] [CrossRef] [PubMed]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.



