Submitted:
20 February 2026
Posted:
27 February 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
- Our Contributions.
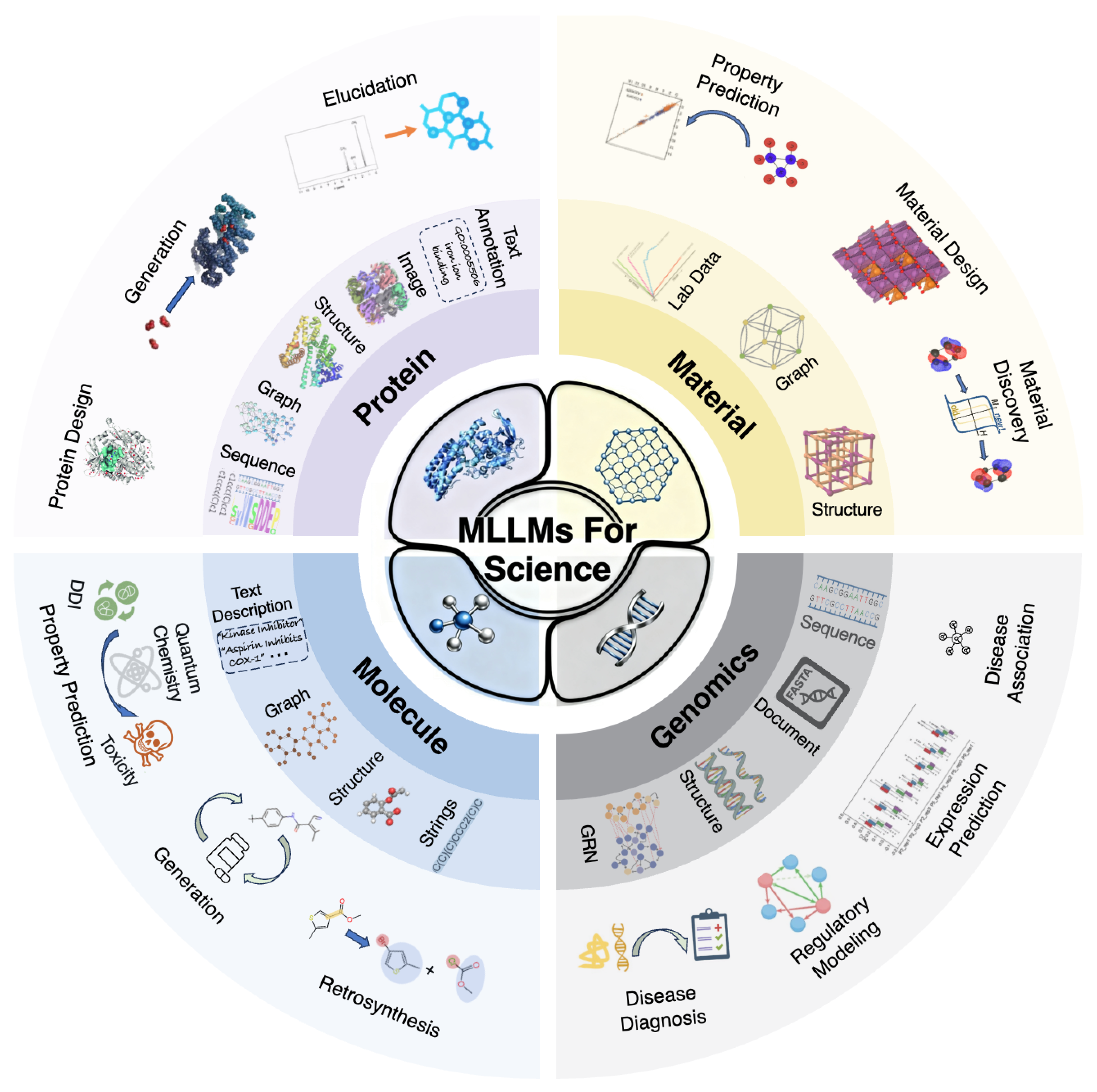
2. General Overview for LLMs and MLLMs
- Core Components of LLMs.
| Survey | Protein | Drug & Samll Molecule | Gene | Material | Biomedicine | Target Multimodal | Benchmarking |
|---|---|---|---|---|---|---|---|
| Our Survey | ✓ | ✓ | ✓ | ✓ | ✓ | ✓ | ✓ |
| LLMs/MLLMs for Science | |||||||
| [218] | ✓ | ✓ | ✓ | ✓ | |||
| [216] | ✓ | ✓ | ✓ | ✓ | |||
| [69] | ✓ | ✓ | ✓ | ✓ | ✓ | ||
| [20] | ✓ | ✓ | ✓ | ||||
| LLMs/MLLMs for Biomedicine | |||||||
| [186] | ✓ | ||||||
| [200] | ✓ | ||||||
| [164] | ✓ | ✓ | |||||
| [228] | ✓ | ||||||
| [16] | ✓ | ✓ | ✓ | ||||
| [226] | ✓ | ||||||
| [106] | ✓ | ||||||
| [60] | ✓ | ||||||
| [185] | ✓ | ||||||
| [167] | ✓ | ||||||
| [165] | ✓ | ||||||
| [157] | ✓ | ||||||
- Training Objectives and Techniques.
- Multimodal Large Language Models (MLLMs).
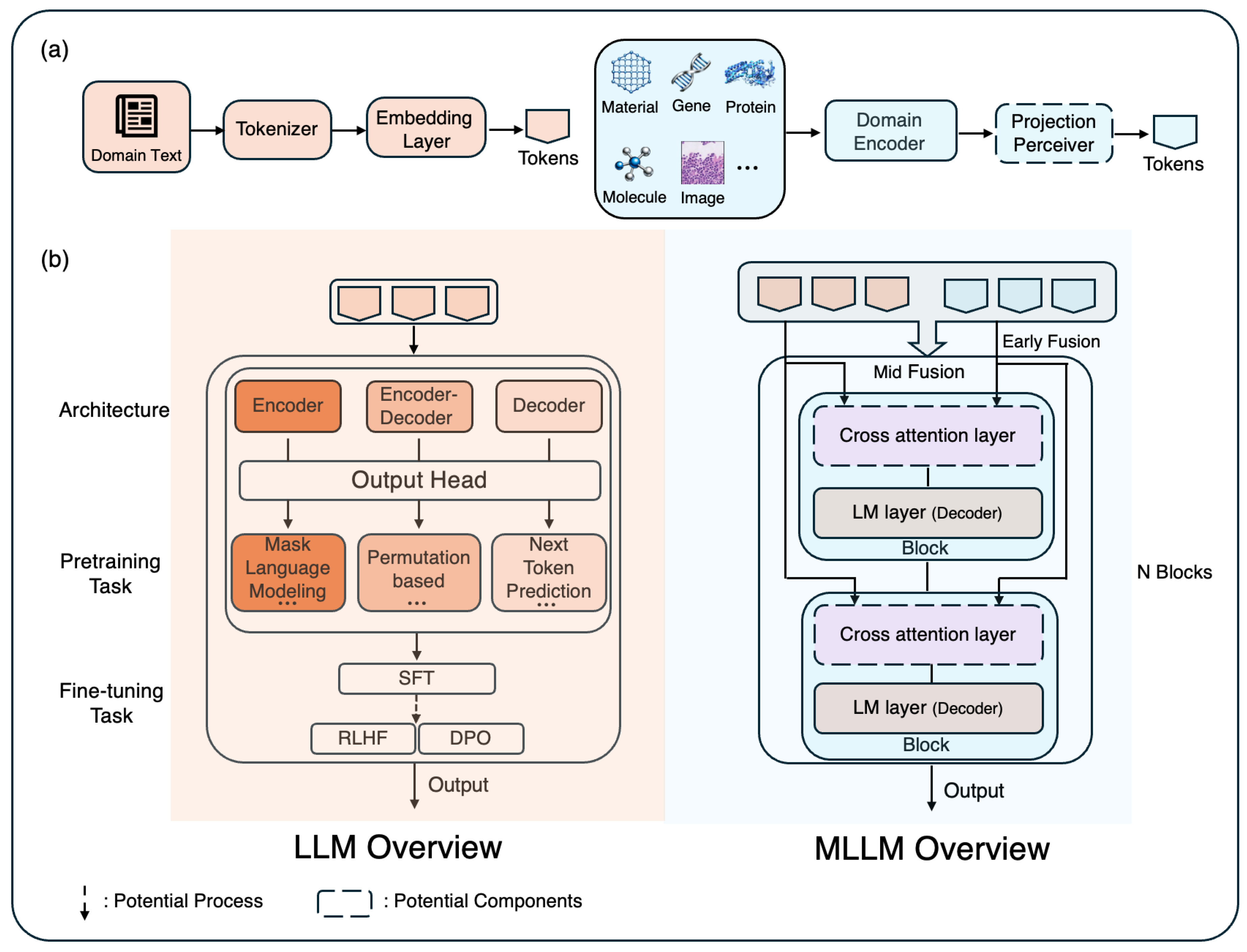
- Pretraining Datasets and Modalities.
- Common Use Cases Across Domains.
3. MLLMs for Molecule Science and Drug Design
3.1. LLMs for Molecule Representation and Design
| Survey | Protein | Drug & Samll Molecule | Gene | Material | Biomedicine | Target Multimodal | Benchmarking |
|---|---|---|---|---|---|---|---|
| Our Survey | ✓ | ✓ | ✓ | ✓ | ✓ | ✓ | ✓ |
| LLMs/MLLMs for Science | |||||||
| [218] | ✓ | ✓ | ✓ | ✓ | |||
| [216] | ✓ | ✓ | ✓ | ||||
| [69] | ✓ | ✓ | ✓ | ✓ | ✓ | ||
| [20] | ✓ | ✓ | |||||
| LLMs/MLLMs for Biomedicine | |||||||
| [186] | ✓ | ||||||
| [200] | ✓ | ✓ | |||||
| [164] | ✓ | ✓ | |||||
| [228] | ✓ | ||||||
| [16] | ✓ | ✓ | ✓ | ||||
| [226] | ✓ | ||||||
| [106] | ✓ | ||||||
| [60] | ✓ | ||||||
| [185] | ✓ | ||||||
| [167] | ✓ | ||||||
| [165] | ✓ | ||||||
| [157] | ✓ | ||||||
3.2. MLLMs for 1D and 2D Molecular Tasks
3.3. MLLMs with 3D Geometry Integration for Molecular Tasks
3.4. MLLMs for Chemistry-Focused Agents and Special Applications
4. MLLMs for Protein Science
4.1. LLMs for Protein Science
4.2. MLLMs for Protein Sequence–Language Integration
4.3. MLLMs for Protein Structure–Sequence–Language Integration
4.4. MLLMs for Protein Interactions and Specialized Applications
5. MLLMs for Genomics and Gene
5.1. LLMs for Genomics
5.2. MLLMs for Genomics and Gene Function Prediction
6. MLLMs for Material Science
6.1. LLMs for Material Discovery
6.2. MLLMs for Material Discovery
7. MLLMs Bridging Molecular Science and Biomedicine
7.1. LLMs for Biomedicine
7.2. MLLMs for Cross Modal Tasks
7.3. Outlook
8. Emerging Hot Topics and Future Directions
8.1. Emerging Hot Topics
8.1.1. Diffusion Large Language Models
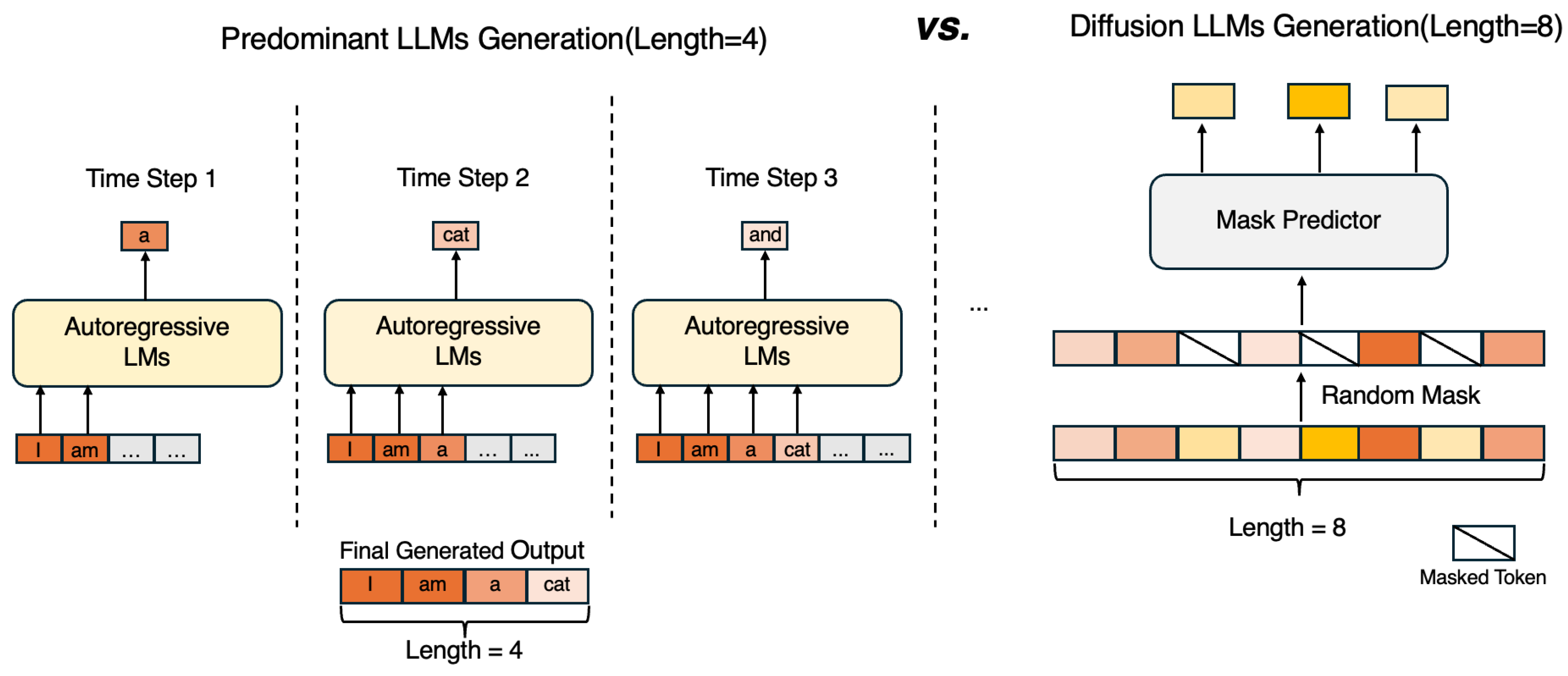
8.1.2. Diffusion Multi-Modal Large Language Models
9. Selected Benchmarking Evaluation
9.1. Molecular Property Prediction
9.2. Protein Property Prediction
10. Conclusion
Appendix A. Summary Model Tables
| Model | Year | Modality | Architecture | Size | Category | Main Task |
|---|---|---|---|---|---|---|
| MolPROP [143] | 2024/05/22 | SMILES, Graph | Encoder-Only | 46M | Property Prediction | Molecular property prediction |
| LLM-MPP [74] | 2025/05/20 | SMILES, Graph, Text | Decoder-Only | 8B | Property Prediction | Property prediction |
| interpretability | ||||||
| ModuLM [25] | 2025/06/01 | 1D, 2D, 3D, Text | Modular/Encoder | 14B | Property Prediction | Flexible property prediction |
| GIT-Mol [107] | 2023/08/14 | Graph, Image, Text | Encoder-Decoder | 700M | Property Prediction | Property prediction |
| generation | ||||||
| PolyLLMem [217] | 2025/03/29 | Polymer, Structure, Text | Encoder-Only | 8B | Polymer Informatics | Polymer property prediction |
| Molbind [188] | 2024/03/13 | Structure, Protein, Text | Encoder-Only | 150M | Property Prediction | Binding affinity prediction |
| BioMedGPT [120] | 2023/08/18 | Protein, Text | Encoder-Decoder | 10B | General-purpose | Biomedical QA |
| multi-modal tasks | ||||||
| InstructMol [18] | 2023/11/27 | Graph, Text | Encoder-Decoder | 2.2B | General-purpose | Instruction following |
| generation | ||||||
| UniMoT [211] | 2024/08/01 | Graph, Text | Encoder-Decoder | 7B | General-purpose | Generation |
| multi-task | ||||||
| Mol-LLM [85] | 2025/01/01 | SMILES, Graph, Text | Encoder-Decoder | 7B | General-purpose | Generation |
| multi-task | ||||||
| ChemVLM [90] | 2024/08/14 | Graph, Image, Text | Encoder-Decoder | 20B | General-purpose | Vision-language tasks |
| Token-Mol [166] | 2024/07/10 | SMILES, 2D/3D | Decoder-Only | N/A | General-purpose | Generative modeling |
| M3LLM [66] | 2025/08/03 | Graph, Text | Encoder-Decoder | 1.28B | General-purpose | Generation |
| granularity study | ||||||
| ChemCrow [12] | 2023/04/11 | Text, Tools | Agent (LLM+Tools) | 100B-1T | Agents & Special Tasks | Chemistry agent |
| ChatMolData [207] | 2024/11/19 | Text, Molecular Data | Agent (LLM+Modules) | 100B-1T | Agents & Special Tasks | Data analysis |
| retrieval | ||||||
| ChemToolAgent [204] | 2024/11/11 | Text, Tools | Agent (LLM+Tools) | 100B-1T | Agents & Special Tasks | Tool-use agent |
| ChemAgent [154] | 2025/01/11 | Text, Memory | Agent (LLM+Memory) | 100B-1T | Agents & Special Tasks | Agent with memory |
| ChemThinker [76] | 2024/09/28 | Text, Tools, Agents | Multi-Agent | 70B | Agents & Special Tasks | Multi-agent reasoning |
| MolPuzzle [57] | 2024/01/01 | Multimodal | Special Task | N/A | Puzzle Task | Structure elucidation |
| reasoning | ||||||
| MM-RCR [219] | 2024/07/21 | Text, Graph, SMILES | Encoder-Decoder | 7B | Reaction Condition | Reaction condition recommendation |
| Chem3DLLM [73] | 2025/08/14 | Text, 3D structure | Encoder-Decoder | ∼ 7B | Drug discovery | Generation |
| Model | Date | Modality | Architecture | Size | Category | Main Task |
|---|---|---|---|---|---|---|
| ProteinDT [108] | 2023/02/09 | Sequence, Text | Encoder-Decoder | 220M | Sequence-Text | Protein Design |
| ProtT3 [116] | 2024/05/21 | Sequence, Text | Encoder-Decoder | ∼1.3B | Sequence-Text | QA tasks, |
| Protein captioning | ||||||
| ProtCLIP [227] | 2024/12/28 | Sequence, Text | Encoder-Only | 770M | Sequence-Text | Function prediction |
| OntoProtein [212] | 2022/01/23 | Sequence, Graph | Encoder-Only | 220M | Sequence-Text | Multi prediction tasks |
| BioMedGPT [119] | 2023/05/26 | Sequence, Text, Graph | Encoder-Decoder | 10B | Sequence-Text | Different QA tasks |
| ProtLLM [236] | 2024/02/28 | Sequence, Text | Encoder-Decoder | 7B | Sequence-Text | Protein understanding, |
| Generation tasks | ||||||
| ProLLaMA [122] | 2024/02/26 | Sequence, Text | Encoder-Decoder | 7B | Sequence-Text | Protein understanding, |
| Generation tasks | ||||||
| InstructProtein [171] | 2023/10/05 | Sequence, Text, Graph | Decoder-Only | 1.3B / 7B | Sequence-Text | Protein design, |
| Prediction tasks | ||||||
| ESM-AA [224] | 2024/03/05 | Sequence, SMILES | Encoder-Only | 35M | Sequence-Text | Classification, |
| Property prediction tasks | ||||||
| BioT5 [135] | 2023/10/11 | Sequence, SMILES, Text | Encoder-Decoder | 252M | Sequence-Text | Diversity prediction, |
| Generation tasks | ||||||
| BioT5+ [134] | 2024/02/27 | Sequence, SMILES, Text | Encoder-Decoder | 252M | Sequence-Text | Diversity prediction, |
| Generation tasks | ||||||
| Galactica [155] | 2022/11/16 | Sequence, Text | Decoder-Only | 120B | Sequence-Text | Prediction, |
| QA tasks | ||||||
| ProteinChat [71] | 2024/08/19 | Sequence, Text | Encoder-Decoder | 14B | Sequence-Text | Function prediction, |
| categories | ||||||
| ESM3 [58] | 2025/01/16 | Sequence, Text, Structure | Encoder-Decoder | 1.4/7/98B | Geometric-Sequence-Text | Design, |
| Generation tasks | ||||||
| proseLM-XL [144] | 2024/08/03 | Sequence, Structure | Encoder-Decoder | 6.5B | Geometric-Sequence-Text | Protein Design |
| SaProt [153] | 2023/10/01 | Sequence, Structure | Encoder-Only | 650M | Geometric-Sequence-Text | Prediction tasks |
| FoldToken [47] | 2024/02/04 | Sequence, Structure | Encoder-Decoder | 280M | Geometric-Sequence-Text | Reconstruction, |
| Antibody Design | ||||||
| Evolla [231] | 2025/01/05 | Sequence, Text, Structure | Encoder-Decoder | 80B | Geometric-Sequence-Text | Diverse QA tasks |
| DPLM-2 [168] | 2024/10/17 | Sequence, Structure | Encoder-Decoder | 150/650M | Geometric-Sequence-Text | Protein generation, |
| Folding | ||||||
| ProTokens [99] | 2023/11/27 | Sequence, Structure | Encoder-Decoder | 7B | Geometric-Sequence-Text | Protein Design |
| ProSST [92] | 2024/04/15 | Sequence, Structure | Encoder-Decoder | 110M | Geometric-Sequence-Text | Prediction tasks |
| ProteinGPT [190] | 2024/08/21 | Sequence, Text, Structure | Encoder-Decoder | 10B | Geometric-Sequence-Text | Protein QA |
| Protein understanding | ||||||
| ProtChatGPT [163] | 2024/02/15 | Sequence, Text, Structure | Encoder-Decoder | 13B | Geometric-Sequence-Text | Protein QA, |
| Protein understanding | ||||||
| STELLA [187] | 2025/06/04 | Sequence, Text, Structure | Encoder-Decoder | ∼9B | Geometric-Sequence-Text | Structure understanding, |
| QA tasks | ||||||
| InstructBioMol [235] | 2024/10/10 | Sequence, Text, SMILES, Structure | Encoder-Decoder | ∼7B | Geometric-Sequence-Text | Protein Design, |
| QA tasks | ||||||
| BioBRIDGE [174] | 2023/10/05 | Sequence, Graph, Text | Encoder-Only | ∼3B | Special-case | PPI Prediction |
| LLaPA [230] | 2024/09/26 | Sequence, Graph, Text | Encoder-Decoder | ∼10B | Special-case | PPI Prediction |
| MolBind [189] | 2024/03/13 | Text, SMILES, Graph, Structure | Encoder-Only | N/A | Special-case | Retrieval tasks |
| BioTranslator [192] | 2023/02/10 | Text, Gene, Sequence, Graph | Encoder-Only | 230M | Special-case | Modal Translator |
| Model | Date | Modality | Architecture | Size | Category | Main Task |
|---|---|---|---|---|---|---|
| GeneChat [35] | 2025/06/05 | DNA, Text | DNABERT-2 + Adaptor | ∼13B | Function Prediction | Free-text gene function generation |
| + Vicuna-13B | ||||||
| ChatNT [141] | 2024/04/30 | DNA, RNA, | Nucleotide Transformer + | ∼7B | Multi-task Genomics | Multimodal sequence |
| Protein, Text | Perceiver + Vicuna-7B | Language Q&A | ||||
| Gene classification | ||||||
| Structure prediction | ||||||
| LLaMA-Gene [96] | 2024/11/30 | DNA, Protein, | LLaMA3-7B | ∼7B | Multi-task Genomics | MSA |
| Text | Function prediction | |||||
| Regression | ||||||
| OmniCellTOSG [210] | 2025/04/02 | RNA, Text | DeBERTa+DNAGPT+ | ∼16B | Multi-task Genomics | Predict cellular states |
| ProtGPT2+GAT | Predict cell types | |||||
| Geneverse [113] | 2024/07/21 | DNA, Protein, | Multi-model | ∼7/8/13B | Multi-task Genomics | Multi-modal gene/protein tasks |
| Text, Figure | LLM/MLLM collection | |||||
| GenoMAS [103] | 2025/07/08 | DNA, RNA, | LLM Agents | N/A | Gene Expression Analysis | (Un)conditional GTA |
| Text | Report Generation | |||||
| cGSA [173] | 2025/06/04 | DNA, Text | LLaMA 3.1-70B | ∼70B | Gene Expression Analysis | Gene pathway finding |
| GTA [63] | 2024/10/02 | DNA, Text | Sei Encoder + Token Alignment | ∼8B | Gene Expression Analysis | Long-range gene expression modeling |
| + Llama3-8B | ||||||
| LLM4GRN [2] | 2024/10/21 | RNA, Text | LLaMA3.1-70B | ∼70B | Regulatory Genomics | Gene regulatory network discovery |
| GeneBERT [126] | 2021/10/11 | DNA (1D), | BERT+ | ∼110M | Regulatory Genomics | Multi-modal self-supervised pre-training |
| TF-Region (2D) | Swin Transformer | |||||
| GeneCompass [196] | 2023/09/28 | RNA, Text | Transformer | N/A | Regulatory Genomics | GRN inference |
| Model | Date | Modality | Architecture | Size | Category | Main Task |
|---|---|---|---|---|---|---|
| CrystaLLM [7] | 2023/07/10 | Text | Decoder-Only | 25/200M | Crystal Structure | Generate crystal structures |
| LLMatDesign [72] | 2024/06/19 | Text | LLM Agent | N/A | Autonomous Discovery | Autonomous materials discovery |
| FlowLLM [151] | 2024/10/30 | Text | LLM+RFM | N/A | Material Design | Generate stable novel materials |
| GenMS [195] | 2024/09/10 | Text, Graph | LLM+Diffusion | N/A | Crystal Generation | Low-energy crystal structure generation |
| Mat2Seq [194] | 2024/12/01 | Text, Graph | Encoder-Decoder | 25/200M | Property Prediction | Crystal sequence representation |
| CrystaltextLLM [56] | 2024/02/06 | Text | Encoder-Decoder | ∼70B | Stability Prediction | Generate stable materials |
| ChatGPTMaterial [32] | 2024/02/12 | Text | Decoder-Only | 11B | Material Design | Suggest material compositions |
| ICGPT [104] | 2024/04/22 | Text | Transformer | N/A | Property Prediction | Accurate material property prediction |
| ELLM [54] | 2024/04/23 | Text | Encoder-Decoder | N/A | Material Selection | Expert recommendations for materials |
| ElaTBot [111] | 2024/11/19 | Text, Quantitative Data | Llama2-7B | ∼7B | Material Discovery | (Details TBD) |
| CrossMatAgent [158] | 2025/03/25 | Text,Image | Agent | N/A | Material Discovery | Multi-agent material design framework |
| AutoMEX [44] | 2025/03/– | Text,3D Document | Agent | N/A | Material Selection | Autonomous material extrusion workflow |
| Structure Data | ||||||
| LLM-Fusion [11] | 2024/12/19 | Text, SMILES, Fingerprints | Encoder-Decoder | N/A | Property Prediction | Multimodal property prediction |
| Cephalo [15] | 2024/05/29 | Image, Text | VLM | ∼600M | Bio-Inspired Design | Analyze bio-inspired materials |
| MaCBench [4] | 2024/10/08 | Text, Image | VLM | N/A | Material Discovery | Evaluate multimodal models’ performance |
| FMMD [136] | 2024 | Text, Image | Fusion Model | N/A | Material Prediction | Scalable property prediction |
| MatterGPT [24] | 2024/08/14 | Text | Transformer | 80M | Property Prediction | Generate solid-state materials |
| Model | Date | Modality | Architecture | Size | Main Tasks | |
|---|---|---|---|---|---|---|
| GenoMAS [103] | 2025/07/08 | DNA, RNA, Text | LLM agents | N/A | Gene expression analysis | |
| cGSA [173] | 2025/06/04 | DNA, Text | LlaMA 3.1-70B | ∼70B | Gene pathway findiing | |
| LLM4GRN [2] | 2024/10/21 | RNA, Text | LLaMA3.1-70B | ∼70B | Gene regulatory networks discovery | |
| GeneCompass [196] | 2023/09/28 | RNA, Text | Transformer | N/A | Gene Regulatory Network inference | |
| Geneverse [113] | 2024/07/21 | DNA, Protein | Multi-model LLM/MLLM collection | ∼7/8/13B | Multi-modal gene/protein tasks | |
| Text, Figure | ||||||
| Natural Language | BioMedGPT-LM+ | Protein Question Answering | ||||
| BioMedGPT [119] | 2024/11/25 | Molecular Graphs | Multimodal encoder | ∼10B | Molecule Question Answering | |
| Protein Sequences | ||||||
| Gene classification | ||||||
| LLaMA-Gene [96] | 2024/11/30 | DNA, Protein, Text | LLaMA3-7B | ∼7B | Gene structure prediction | |
| Multiple sequence analysis | ||||||
| Function prediction | ||||||
| OmniCellTOSG [210] | 2025/04/02 | RNA, Text | DeBERTa+DNAGPT | ∼16B | Cellular States Prediction | |
| +ProtGPT2+GAT | Cell Type Prediction | |||||
| Survival prediction | ||||||
| mSTAR [193] | 2024/07/22 | pathological images, | CLIP | Varies | Diagnosis | |
| RNA-seq, Text | Molecule prediction | |||||
| Report generation | ||||||
| ST-ALign [100] | 2024/11/25 | pathological images, gene | Image encoder + Gene encoder | N/A | Spatial clustering identification | |
| Spot Gene Expression Prediction | ||||||
| Pathological images | Spatial domain identification | |||||
| spEMO [112] | 2025/01/13 | spatial multi omics | PFM+LLM | N/A | Disease Prediction | |
| Report Generation | ||||||
| SpaLLM [91] | 2025/07/03 | Single-cell transcriptome data, | LLM+omics encoder+GNN | N/A | Region Identification | |
| Multi-omics data |
Appendix B. Summary Dataset Tables of MLLMs for Science
| Datasets | Year | Modality | Tasks | Source | Application | Stage |
|---|---|---|---|---|---|---|
| PubChem (77M SMILES) | – | SMILES, Text | MLM, MTR, caption/retrieval | Source | [143] [107] [84] [18] [211] [117] [25] [74] |
Pretraining |
| ChEBI-20 | 2021 | SMILES, Text | Captioning, generation | Source | [107] [211] [85] [18] |
Pretraining |
| ZINC | – | SMILES | Language modeling, generation | Source | [117] | Pretraining |
| USPTO (full/50k) | 2012/2017 | Reaction SMILES, Text | FS/RS/RP reaction modeling |
Source (full) Source (full) Source (50k) |
[85] [211] |
Pretraining/Instr. |
| Mol-Instructions | 2023 | Text, SMILES, Graph | FS, RS, RP, caption-guided gen | Source | [85] [211] |
Instruction |
| SMolInstruct | 2024 | Text, SMILES, Graph | FS, RS, RP, generation | Source | [85] | Instruction |
| PCdes | – | Molecule, Text | Retrieval (M2T/T2M) | Source | [211] | Instruction |
| MoMu | 2022 | Molecule, Text | Cross-modal retrieval | Source | [211] | Instruction |
| Molecule3D | 2021 | 3D | Conformations Graph–3D alignment |
Source Source |
[188] | Pretraining |
| GEOM | 2020 | 3D | Conformations Graph–3D alignment | Source | [188] | Pretraining |
| PDBBind | 2016 | Protein pockets, 3D | Conf.–Protein alignment | Source | [188] | Pretraining |
| CrossDock | 2019 | Protein pockets, 3D | Conf.–Protein alignment | Source | [188] | Pretraining |
| DrugBank | – | SMILES, Text (properties) | Molecular relational learning | Source | [25] | Pretraining |
| L+M-24 | 2024 | Image, Text | Captioning (Mol2Lang) | Source | [160] | Pretraining |
| Chem Exam | 2024–2025 | Image, Text | OCR, VQA, Chem QA | Source | [90] | Pretraining |
| Chem OCR | 2024–2025 | Image, Text | OCR, VQA, Chem QA | Source | [90] | Pretraining |
| Web-Chem | 2024–2025 | Image, Text | OCR, VQA, Chem QA | Source | [90] | Pretraining |
| PubMed abstracts | – | Text (biomedical) | Domain LM pretraining | Source | [118] | Pretraining |
| Datasets | Year | Modality | Tasks | Source | Application | Stage |
|---|---|---|---|---|---|---|
| ESOL (LogS) | 2012 | SMILES, Graph | Regression (solubility) | source | [143] [74] [85] [84] |
Downstream |
| FreeSolv | 2014 | SMILES, Graph | Regression (hydration free energy) | source | [143] [74] [25] |
Downstream |
| Lipophilicity (Lipo) | 2016 | SMILES, Graph | Regression (logD/logP) | source | [143] [74] [85] |
Downstream |
| QM7 | 2011 | SMILES, Graph | Regression (atomization energy) | source | [143] [74] |
Downstream |
| QM9 | 2014 | SMILES, Graph | Regression (HOMO/LUMO etc.) | source | [18] [85] |
Downstream |
| BBBP | 2018 | SMILES, Graph | Classification (BBB) | source | [143] [74] [85] [84] |
Downstream |
| BACE | 2016 | SMILES, Graph | Classification (binding) | source | [143] [74] [85] [84] |
Downstream |
| ClinTox | 2018 | SMILES, Graph | Classification (toxicity) | source | [143] [74] [85] [84] |
Downstream |
| Tox21 | 2014 | SMILES, Graph | Multi-task toxicity | source | [107] [211] [84] |
Downstream |
| ToxCast | 2013 | SMILES, Graph | Multi-task toxicity | source | [107] [211] |
Downstream |
| HIV | 2014 | SMILES, Graph | Classification (anti-HIV) | source | [85] [84] |
Downstream |
| SIDER | 2015 | SMILES, Graph | Multi-label side effects | source | [107] [85] [84] |
Downstream |
| MUV | 2013 | SMILES, Graph | Virtual screening | source | [84] | Downstream |
| ChEBI-20 | 2021 | SMILES, Text | Captioning, generation | source | [107] [85] [211] [84] |
Downstream |
| L+M-24 | 2024 | Image, Text | Captioning | source | [160] | Downstream |
| PubChem Captions | – | Image, SMILES, Text | Captioning, Image→SMILES | source | [107] | Downstream |
| USPTO-50k | 2017 | Reaction SMILES, Text | FS, RS, RP | source | [85] [18] |
Downstream |
| RetroBench | 2024 | Reaction network | Multi-step retrosynthesis | source | [78] | Downstream |
| ORDERly | 2024 | Reactions | OOD reaction evaluation | source | [85] | Downstream |
| AqSolDB | 2019 | SMILES | OOD solubility evaluation | source | [85] | Downstream |
| ChEMBL-02 | 2020 | Pairwise molecules | Molecule optimization | source | [84] | Downstream |
| PCdes | – | Molecule, Text | Retrieval (M2T/T2M) | source | [211] | Downstream |
| MoMu | 2022 | Molecule, Text | Cross-modal retrieval | source | [211] | Downstream |
| ZhangDDI | 2017 | SMILES, Graph | Drug–drug interaction | source | [25] | Downstream |
| ChChMiner | 2018 | SMILES, Graph | Drug–drug interaction | source | [25] | Downstream |
| DeepDDI | 2018 | SMILES, Graph | Drug–drug interaction | source | [25] | Downstream |
| TWOSIDES | 2012 | SMILES, Graph | Drug–drug interaction | source | [25] | Downstream |
| MNSol | 2020 | SMILES, Graph | Solute–solvent interaction | source | [25] | Downstream |
| CompSol | 2017 | SMILES, Graph | Solute–solvent interaction | source | [25] | Downstream |
| Abraham | 2010 | SMILES, Graph | Solute–solvent interaction | source | [25] | Downstream |
| CombiSolv | 2021 | SMILES, Graph | Solute–solvent interaction | source | [25] | Downstream |
| CombiSolv-QM | 2021 | SMILES, Graph (QM) | Solute–solvent interaction | source | [25] | Downstream |
| Chromophore | 2020 | SMILES, Graph | Chromophore–solvent interaction | source | [25] | Downstream |
| Datasets | Year | Modality | Tasks | Source | Application | Stage |
|---|---|---|---|---|---|---|
| SwissProt | 2000 | Sequence, Text | Sequence–text alignment, Captioning | Source |
[109] [116] [227] [71] [231] |
Pretraining |
| TrEMBL | 2000 | Sequence, Text | Sequence–text alignment | Source |
[227] [231] |
Pretraining |
| ProtAnno-S | 2024 | Sequence, Text | Contrastive alignment (sparse, curated) | Source | [227] | Pretraining |
| ProtAnno-D | 2024 | Sequence, Text | Contrastive alignment (dense, auto) | Source | [227] | Pretraining |
| ProteinKG25 | 2022 | Sequence, Graph, Text | KG-enhanced pretraining | Source |
[214] [116] |
Pretraining |
| PrimeKG | 2023 | Graph, Text | Biomedical KG bridging | Source | [174] | Pretraining |
| UniRef50 | 2007 | Sequence | Language modeling corpus | Source | [122] | Pretraining |
| UniRef90 | 2007 | Sequence | Language modeling corpus | Source | [168] | Pretraining |
| AlphaFold DB | 2022 | Structure (3D) | Structure-aware pretraining | Source |
[153] [224] [58] |
Pretraining |
| PDB | 2000 | Structure (3D) | Structure and token pretraining | Source |
[168] [99] |
Pretraining |
| PDBbind (v2019) | 2019 | Structure, Binding | Binding-aware pretraining | Source | [224] | Pretraining |
| S2ORC | 2020 | Text (scholarly) | Biomedical text pretraining | Source | [119] | Pretraining |
| PubMed abstracts | 1996 | Text (biomedical) | Biomedical text pretraining | Source |
[119] [236] [134] |
Pretraining |
| bioRxiv | 2013 | Text (preprints) | Biomedical text pretraining | Source | [134] | Pretraining |
| PubChem | 2004 | SMILES, Text | Chem–structure pretraining | Source |
[135] [134] |
Pretraining |
| ChEMBL | 2012 | SMILES, Bioactivity | Chem–structure pretraining | Source |
[224] [135] |
Pretraining |
| ZINC (ZINC15) | 2015 | SMILES | Generative pretraining | Source |
[135] [134] |
Pretraining |
| InterPT (instruction set) | 2024 | Sequence, Text | Protein–text instruction pretraining | Source | [236] | Instruction |
| ProteinChat Corpus | 2024 | Sequence, Text | Instruction/QA pretraining | Source | [71] | Instruction |
| SwissProtCLAP | 2023 | Sequence, Text | Sequence–text alignment | Source | [109] | Pretraining |
| Datasets | Year | Modality | Tasks | Source | Application | Stage |
|---|---|---|---|---|---|---|
| TAPE | 2019 | Sequence, Structure | SS, Contact, Homology, Fluorescence, Stability | Source |
[109] [214] [236] [224] [171] [144] [153] |
Downstream |
| DeepLoc | 2017 | Sequence, Text | Subcellular localization | Source |
[227] [171] |
Downstream |
| Solubility (DeepSol) | 2017 | Sequence | Solubility prediction | Source | [135] | Downstream |
| Localization | 2017 | Sequence | Membrane/soluble classification | Source | [135] | Downstream |
| SwissProt | 2000 | Sequence, Text | Function description classification | Source |
[171] [71] |
Downstream |
| CASP15 | 2022 | Structure | Protein folding | Source | [58] | Downstream |
| CB513 | 1999 | Sequence | Secondary structure prediction | Source |
[153] [92] |
Downstream |
| SCOPe | 2014 | Structure | Fold/superfamily classification | Source |
[122] [144] [92] |
Downstream |
| TAPE Stability | 2019 | Sequence | Stability prediction | Source | [144] | Downstream |
| TAPE Contact | 2019 | Structure | Contact map prediction | Source |
[153] [171] |
Downstream |
| STRING | 2021 | Graph (PPI) | PPI classification | Source |
[214] [236] [171] [174] [230] |
Downstream |
| SHS27k | 2019 | Sequence, Graph | PPI classification | Source |
[214] [236] [171] [174] |
Downstream |
| SHS148k | 2019 | Sequence, Graph | PPI classification | Source |
[214] [236] [171] [174] |
Downstream |
| BioGRID | 2003 | Graph | PPI classification | Source | [230] | Downstream |
| PPI (Yeast, Human) | 2019 | Sequence, Graph | PPI classification | Source | [135] | Downstream |
| BioSNAP | 2018 | Sequence, Graph | DTI, PPI prediction | Source | [135] | Downstream |
| DMS (-lac, AAV, Thermo, Flu, Sta) | 2018 | Sequence | Mutational effect prediction | Source | [227] | Downstream |
| ProteinGym | 2023 | Sequence | Mutational effect prediction | Source |
[58] [153] [92] |
Downstream |
| PubMedQA | 2019 | Text | Biomedical QA | Source |
[119] [155] [192] |
Downstream |
| MedMCQA | 2022 | Text | Biomedical QA | Source |
[119] [155] |
Downstream |
| USMLE | 2020 | Text | Medical exam QA | Source |
[119] [155] |
Downstream |
| UniProtQA | 2023 | Sequence, Text | Protein QA | Source |
[119] [155] [192] |
Downstream |
| ProteinQA benchmark | 2024 | Sequence, Text | Protein QA | Source |
[71] [190] [163] [187] |
Downstream |
| PDB-QA | 2024 | Structure, Text | Protein QA | Source | [116] | Downstream |
| MMLU-bio | 2021 | Text | Multitask biomedical QA | Source | [155] | Downstream |
| ChEBI-20 | 2019 | Molecule, Text | Molecule QA, Captioning | Source |
[119] [135] |
Downstream |
| ChemProt | 2019 | Text | Relation extraction | Source | [135] | Downstream |
| BindingDB | 2007 | Sequence, SMILES | Binding prediction | Source |
[224] [135] [189] |
Downstream |
| MoleculeNet | 2018 | Molecule | Property prediction | Source |
[224] [155] |
Downstream |
| USPTO | 2019 | SMILES, Text | Reaction prediction | Source | [155] | Downstream |
| PubChem BioAssay | 2014 | SMILES, Text | Retrieval | Source | [189] | Downstream |
| SAbDab | 2014 | Structure | Antibody design | Source | [47] | Downstream |
| Inverse folding sets | 2019 | Sequence, Structure | Inverse folding | Source | [99] | Downstream |
| Protein design benchmarks | 2024 | Sequence, Structure | Protein generation, Design | Source |
[58] [231] [235] |
Downstream |
| Datasets | Year | Modality | Tasks | Source | Application | Stage |
|---|---|---|---|---|---|---|
| NCBI Gene | 2005 | DNA, Text | Function modeling | source | [35] | Pretraining |
| NT | 2023 | DNA | Sequence classification | source | [141] | Pretraining |
| BEND | 2022 | DNA | Regulatory element classification | source | [141] | Pretraining |
| AgroNT | 2023 | DNA | Plant genomics tasks | source | [141] | Pretraining |
| ChromTransfer | 2022 | DNA | Regulatory element transfer | source | [141] | Pretraining |
| ATAC-seq fetal atlas | 2020 | DNA, TF-region | Chromatin accessibility | source | [126] | Pretraining |
| Sei | 2022 | DNA, Chromatin | Epigenomic feature extraction | source | [63] | Pretraining |
| SwissProt | 1986 | Protein | Protein sequence modeling | source | [96] | Pretraining |
| TrEMBL | 1996 | Protein | Protein sequence modeling | source | [96] | Pretraining |
| S2ORC | 2020 | Text | Scientific text modeling | source | [96] | Pretraining |
| scCompass-126M | 2024 | RNA | Cross-species modeling | source | [196] | Pretraining |
| Ensembl GRCh38 | 2013 | DNA | Genomic sequences | source | [113] | Pretraining |
| GTEx v8 | 2015 | RNA | Expression profiles | source | [113] | Pretraining |
| UniProt | 2023 | Protein | Protein sequences | source | [113] | Pretraining |
| PubMed abstracts | 1996 | Text | Biomedical language modeling | source | [113] | Pretraining |
| Datasets | Year | Modality | Tasks | Source | Application | Stage |
|---|---|---|---|---|---|---|
| NCBI Gene | 2005 | DNA, Text | Function prediction | source | [35] | Downstream |
| NT | 2023 | DNA | Sequence classification | source | [141] | Downstream |
| BEND | 2022 | DNA | Regulatory element classification | source | [141] | Downstream |
| AgroNT | 2023 | DNA | Plant genomics tasks | source | [141] | Downstream |
| ChromTransfer | 2022 | DNA | Regulatory element transfer | source | [141] | Downstream |
| DeepSTARR | 2019 | DNA | Enhancer activity prediction | source | [141] | Downstream |
| APARENT2 | 2022 | RNA | Polyadenylation prediction | source | [141] | Downstream |
| Saluki | 2022 | RNA | RNA degradation prediction | source | [141] | Downstream |
| GM12878 | 2012 | RNA | Expression prediction | source | [63] | Downstream |
| Geuvadis | 2013 | RNA | Expression prediction | source | [63] | Downstream |
| GenoTEX | 2025 | DNA, RNA | Gene–trait association | source | [103] | Downstream |
| GEO | 2002 | RNA | Expression prediction | source | [103] | Downstream |
| TCGA | 2008 | RNA, DNA | Expression prediction | source | [103] | Downstream |
| Curated gene sets (102) | 2025 | Gene sets | Pathway enrichment | source | [173] | Downstream |
| Case studies (melanoma, breast cancer) | 2025 | RNA, Text | Disease-specific analysis | source | [173] | Downstream |
| UniProt | 2023 | Protein | Function prediction | source | [96] | Downstream |
| Pfam | 1997 | Protein | Domain classification | source | [96] | Downstream |
| InterPro | 2000 | Protein | Domain classification | source | [96] | Downstream |
| PBMC-ALL | 2017 | RNA | GRN inference | source | [2] | Downstream |
| PBMC-CTL | 2017 | RNA | GRN inference | source | [2] | Downstream |
| BoneMarrow | 2019 | RNA | GRN inference | source | [2] | Downstream |
| OmniCellTOSG | 2025 | scRNA-seq, Text | Cellular state prediction | source | [210] | Downstream |
| HCA | 2017 | scRNA-seq | Cross-species GRN inference | source | [196] | Downstream |
| MCA | 2018 | scRNA-seq | Cross-species GRN inference | source | [196] | Downstream |
| Tabula Sapiens | 2022 | scRNA-seq | Cross-species GRN inference | source | [196] | Downstream |
| GO annotation | 2000 | DNA, Text | Function prediction | source | [113] | Downstream |
| UniProt | 2002 | Protein | Protein classification | source | [113] | Downstream |
| GTEx v8 | 2015 | RNA | Expression prediction | source | [113] | Downstream |
References
- Paul D Adams, Pavel V Afonine, Gábor Bunkóczi, Vincent B Chen, Nathaniel Echols, Jeffrey J Headd, Li-Wei Hung, Swati Jain, Gary J Kapral, Ralf W Grosse Kunstleve, et al. The phenix software for automated determination of macromolecular structures. Methods, 55(1):94–106, 2011.
- Tejumade Afonja, Ivaxi Sheth, Ruta Binkyte, Waqar Hanif, Thomas Ulas, Matthias Becker, and Mario Fritz. Llm4grn: Discovering causal gene regulatory networks with llms–evaluation through synthetic data generation. arXiv preprint arXiv:2410.15828, 2024.
- Genereux Akotenou and Achraf El Allali. Genomic language models (glms) decode bacterial genomes for improved gene prediction and translation initiation site identification. Briefings in Bioinformatics, 26(4):bbaf311, 2025.
- Nawaf Alampara, Mara Schilling-Wilhelmi, Martiño Ríos-García, Indrajeet Mandal, Pranav Khetarpal, Hargun Singh Grover, NM Krishnan, and Kevin Maik Jablonka. Probing the limitations of multimodal language models for chemistry and materials research. arXiv preprint arXiv:2411.16955, 2024.
- Jean-Baptiste Alayrac, Jeff Donahue, Pauline Luc, Antoine Miech, Iain Barr, Yana Hasson, Karel Lenc, Arthur Mensch, Katherine Millican, Malcolm Reynolds, et al. Flamingo: a visual language model for few-shot learning. Advances in neural information processing systems, 35:23716–23736, 2022.
- Ethan C Alley, Grigory Khimulya, Surojit Biswas, Mohammed AlQuraishi, and George M Church. Unified rational protein engineering with sequence-based deep representation learning. Nature methods, 16(12):1315–1322, 2019.
- Luis M Antunes, Keith T Butler, and Ricardo Grau-Crespo. Crystal structure generation with autoregressive large language modeling. Nature Communications, 15(1):10570, 2024.
- Marianne Arriola, Aaron Gokaslan, Justin T Chiu, Zhihan Yang, Zhixuan Qi, Jiaqi Han, Subham Sekhar Sahoo, and Volodymyr Kuleshov. Block diffusion: Interpolating between autoregressive and diffusion language models. arXiv preprint arXiv:2503.09573, 2025.
- Vivek Bagal, Rohit Aggarwal, Yash Deshmukh, and Alexander Noskov. MolGPT: Molecular generation using a transformer-decoder model. Journal of Chemical Information and Modeling, 61(11):5071–5080, 2021.
- Manojit Bhattacharya, Soumen Pal, Srijan Chatterjee, Sang-Soo Lee, and Chiranjib Chakraborty. Large language model to multimodal large language model: A journey to shape the biological macromolecules to biological sciences and medicine. Molecular Therapy Nucleic Acids, 35(3), 2024.
- Onur Boyar, Indra Priyadarsini, Seiji Takeda, and Lisa Hamada. Llm-fusion: A novel multimodal fusion model for accelerated material discovery. arXiv preprint arXiv:2503.01022, 2025.
- Andres M Bran, Sam Cox, Oliver Schilter, Carlo Baldassari, Andrew D White, and Philippe Schwaller. Chemcrow: Augmenting large-language models with chemistry tools. arXiv preprint arXiv:2304.05376, 2023.
- Naomi Brandes, Dan Ofer, Yuval Peleg, Nadav Rappoport, and Michal Linial. Proteinbert: A universal deep-learning model of protein sequence and function. Bioinformatics, 38(8):2102–2110, 2022.
- Tom Brown, Benjamin Mann, Nick Ryder, Melanie Subbiah, Jared D Kaplan, Prafulla Dhariwal, Arvind Neelakantan, Pranav Shyam, Girish Sastry, Amanda Askell, et al. Language models are few-shot learners. Advances in neural information processing systems, 33:1877–1901, 2020.
- Markus J Buehler. Cephalo: Multi-modal vision-language models for bio-inspired materials analysis and design. Advanced Functional Materials, 34(49):2409531, 2024.
- Lukas Buess, Matthias Keicher, Nassir Navab, Andreas Maier, and Soroosh Tayebi Arasteh. From large language models to multimodal ai: A scoping review on the potential of generative ai in medicine. Biomedical Engineering Letters, pages 1–19, 2025.
- Gábor Bunkóczi, Nathaniel Echols, Airlie J McCoy, Robert D Oeffner, Paul D Adams, and Randy J Read. Phaser. mrage: automated molecular replacement. Biological Crystallography, 69(11):2276–2286, 2013.
- He Cao, Zijing Liu, Xingyu Lu, Yuan Yao, and Yu Li. Instructmol: Multi-modal integration for building a versatile and reliable molecular assistant in drug discovery. arXiv preprint arXiv:2311.16208, 2023.
- Siwar Chaabene, Amal Boudaya, Bassem Bouaziz, and Lotfi Chaari. An overview of methods and techniques in multimodal data fusion with application to healthcare. International Journal of Data Science and Analytics, pages 1–25, 2025.
- Chiranjib Chakraborty, Manojit Bhattacharya, Soumen Pal, Srijan Chatterjee, Arpita Das, and Sang-Soo Lee. Ai-enabled language models (lms) to large language models (llms) and multimodal large language models (mllms) in drug discovery and development. Journal of Advanced Research, 2025.
- Jiayu Chang, Shiyu Wang, Chen Ling, Zhaohui Qin, and Liang Zhao. Gene-associated disease discovery powered by large language models. arXiv preprint arXiv:2401.09490, 2024.
- Bo Chen, Xingyi Cheng, Pan Li, Yangli-ao Geng, Jing Gong, Shen Li, Zhilei Bei, Xu Tan, Boyan Wang, Xin Zeng, et al. xtrimopglm: unified 100b-scale pre-trained transformer for deciphering the language of protein. arXiv preprint arXiv:2401.06199, 2024.
- Tianqi Chen, Shujian Zhang, and Mingyuan Zhou. Dlm-one: Diffusion language models for one-step sequence generation. arXiv preprint arXiv:2506.00290, 2025.
- Yan Chen, Xueru Wang, Xiaobin Deng, Yilun Liu, Xi Chen, Yunwei Zhang, Lei Wang, and Hang Xiao. Mattergpt: A generative transformer for multi-property inverse design of solid-state materials. arXiv preprint arXiv:2408.07608, 2024.
- Zhuo Chen, Yizhen Zheng, Huan Yee Koh, Hongxin Xiang, Linjiang Chen, Wenjie Du, and Yang Wang. Modulm: Enabling modular and multimodal molecular relational learning with large language models. arXiv preprint arXiv:2506.00880, 2025.
- Jiabei Cheng, Xiaoyong Pan, Yi Fang, Kaiyuan Yang, Yiming Xue, Qingran Yan, and Ye Yuan. Gexmolgen: cross-modal generation of hit-like molecules via large language model encoding of gene expression signatures. Briefings in Bioinformatics, 25(6):bbae525, 2024.
- Le Cheng and Shuangyin Li. Diffuspoll: Conditional text diffusion model for poll generation. In Findings of the Association for Computational Linguistics ACL 2024, pages 925–935, 2024.
- Vasudev Chenthamarakshan, Payel Das, Samuel C. Hoffman, Hendrik Strobelt, Kumar Padmanabhan, Patrick Riley, and Bonggun Kim. CogMol: Target-specific and selective drug design for covid-19 using deep generative models. arXiv preprint arXiv:2004.01215, 2020.
- Wei-Lin Chiang, Zhuohan Li, Zi Lin, Ying Sheng, Zhanghao Wu, Hao Zhang, Lianmin Zheng, Siyuan Zhuang, Yonghao Zhuang, Joseph E. Gonzalez, Ion Stoica, and Eric P. Xing. Vicuna: An open-source chatbot impressing gpt-4 with 90%* chatgpt quality, March 2023.
- Ananya Chithrananda, Gabriel J. Grand, and Bharath Ramsundar. ChemBERTa: Large-scale self-supervised learning for molecular property prediction. arXiv preprint arXiv:2010.09885, 2020.
- Hugo Dalla-Torre, Liam Gonzalez, Javier Mendoza-Revilla, Nicolas Lopez Carranza, Adam Henryk Grzywaczewski, Francesco Oteri, Christian Dallago, Evan Trop, Bernardo P de Almeida, Hassan Sirelkhatim, et al. Nucleotide transformer: building and evaluating robust foundation models for human genomics. Nature Methods, 22(2):287–297, 2025.
- Jyotirmoy Deb, Lakshi Saikia, Kripa Dristi Dihingia, and G Narahari Sastry. Chatgpt in the material design: Selected case studies to assess the potential of chatgpt. Journal of Chemical Information and Modeling, 64(3):799–811, 2024.
- Yifan Deng, Spencer S Ericksen, and Anthony Gitter. Chemical language model linker: blending text and molecules with modular adapters. arXiv preprint arXiv:2410.20182, 2024.
- Jacob Devlin, Ming-Wei Chang, Kenton Lee, and Kristina Toutanova. BERT: Pre-training of deep bidirectional transformers for language understanding. In Proceedings of NAACL-HLT, pages 4171–4186, 2019.
- Shashi Dhanasekar, Akash Saranathan, and Pengtao Xie. Genechat: A multi-modal large language model for gene function prediction. bioRxiv, pages 2025–06, 2025.
- Gautham Dharuman, Kyle Hippe, Alexander Brace, Sam Foreman, Väinö Hatanpää, Varuni K Sastry, Huihuo Zheng, Logan Ward, Servesh Muralidharan, Archit Vasan, et al. Mprot-dpo: Breaking the exaflops barrier for multimodal protein design workflows with direct preference optimization. In SC24: International Conference for High Performance Computing, Networking, Storage and Analysis, pages 1–13. IEEE, 2024.
- Zhengxiao Du, Yujie Qian, Xiao Liu, Ming Ding, Jiezhong Qiu, Zhilin Yang, and Jie Tang. Glm: General language model pretraining with autoregressive blank infilling. arXiv preprint arXiv:2103.10360, 2021.
- Chenrui Duan, Zelin Zang, Yongjie Xu, Hang He, Siyuan Li, Zihan Liu, Zhen Lei, Ju-Sheng Zheng, and Stan Z Li. Fgenebert: function-driven pre-trained gene language model for metagenomics. Briefings in Bioinformatics, 26(2):bbaf149, 2025.
- Ran Duan, Lin Gao, Yong Gao, Yuxuan Hu, Han Xu, Mingfeng Huang, Kuo Song, Hongda Wang, Yongqiang Dong, Chaoqun Jiang, et al. Evaluation and comparison of multi-omics data integration methods for cancer subtyping. PLoS computational biology, 17(8):e1009224, 2021.
- David K Duvenaud, Dougal Maclaurin, Jorge Iparraguirre, Rafael Bombarell, Timothy Hirzel, Alán Aspuru-Guzik, and Ryan P Adams. Convolutional networks on graphs for learning molecular fingerprints. Advances in neural information processing systems, 28, 2015.
- Ahmed Elnaggar, Michael Heinzinger, Christian Dallago, Ghalia Rehawi, Yu Wang, Llion Jones, Tom Gibbs, Tamas Feher, Christoph Angerer, Martin Steinegger, et al. Prottrans: Toward understanding the language of life through self-supervised learning. IEEE transactions on pattern analysis and machine intelligence, 44(10):7112–7127, 2021.
- Devlin et al. BERT: Pre-training of deep bidirectional transformers for language understanding. https://arxiv.org/abs/1810.04805, 2018.
- Benjamin Fabian, Simon Edlich, Hadrien Gaspar, Marwin H.S. Segler, Mark Ahmed, Kathrin Rother, Jan A. Hiss, and Gisbert Schneider. Molecular representation learning with language models and domain-relevant auxiliary tasks. Journal of Chemical Information and Modeling, 60(11):4894–4905, 2020.
- Haolin Fan, Junlin Huang, Jilong Xu, Yifei Zhou, Jerry Ying Hsi Fuh, Wen Feng Lu, and Bingbing Li. Automex: Streamlining material extrusion with ai agents powered by large language models and knowledge graphs. Materials & Design, 251:113644, 2025.
- Noelia Ferruz, Steffen Schmidt, and Birte Höcker. Protgpt2 is a deep unsupervised language model for protein design. Nature Communications, 13(4348), 2022.
- Patrick C Fricker, Marcus Gastreich, and Matthias Rarey. Automated drawing of structural molecular formulas under constraints. Journal of chemical information and computer sciences, 44(3):1065–1078, 2004.
- Zhangyang Gao, Cheng Tan, Jue Wang, Yufei Huang, Lirong Wu, and Stan Z Li. Foldtoken: Learning protein language via vector quantization and beyond. In Proceedings of the AAAI Conference on Artificial Intelligence, volume 39, pages 219–227, 2025.
- Amelia Glaese, Nat McAleese, Maja Trębacz, John Aslanides, Vlad Firoiu, Timo Ewalds, Maribeth Rauh, Laura Weidinger, Martin Chadwick, Phoebe Thacker, et al. Improving alignment of dialogue agents via targeted human judgements. arXiv preprint arXiv:2209.14375, 2022.
- Vladimir Golkov, Marcin J Skwark, Atanas Mirchev, Georgi Dikov, Alexander R Geanes, Jeffrey Mendenhall, Jens Meiler, and Daniel Cremers. 3d deep learning for biological function prediction from physical fields. In 2020 International Conference on 3D Vision (3DV), pages 928–937. IEEE, 2020.
- Shansan Gong, Shivam Agarwal, Yizhe Zhang, Jiacheng Ye, Lin Zheng, Mukai Li, Chenxin An, Peilin Zhao, Wei Bi, Jiawei Han, et al. Scaling diffusion language models via adaptation from autoregressive models. arXiv preprint arXiv:2410.17891, 2024.
- Shansan Gong, Mukai Li, Jiangtao Feng, Zhiyong Wu, and LingPeng Kong. Diffuseq: Sequence to sequence text generation with diffusion models. arXiv preprint arXiv:2210.08933, 2022.
- Shansan Gong, Ruixiang Zhang, Huangjie Zheng, Jiatao Gu, Navdeep Jaitly, Lingpeng Kong, and Yizhe Zhang. Diffucoder: Understanding and improving masked diffusion models for code generation. arXiv preprint arXiv:2506.20639, 2025.
- Google DeepMind. Gemini diffusion: Our state-of-the-art, experimental text diffusion model. Web page, 2025. May 20, 2025; experimental text diffusion model; accessed 2025-09-20.
- Daniele Grandi, Yash Patawari Jain, Allin Groom, Brandon Cramer, and Christopher McComb. Evaluating large language models for material selection. Journal of Computing and Information Science in Engineering, 25(2):021004, 2025.
- Aaron Grattafiori, Abhimanyu Dubey, Abhinav Jauhri, Abhinav Pandey, Abhishek Kadian, Ahmad Al-Dahle, Aiesha Letman, Akhil Mathur, Alan Schelten, Alex Vaughan, et al. The llama 3 herd of models. arXiv preprint arXiv:2407.21783, 2024.
- Nate Gruver, Anuroop Sriram, Andrea Madotto, Andrew Gordon Wilson, C Lawrence Zitnick, and Zachary Ulissi. Fine-tuned language models generate stable inorganic materials as text. arXiv preprint arXiv:2402.04379, 2024.
- Kehan Guo, Bozhao Nan, Yujun Zhou, Taicheng Guo, Zhichun Guo, Mihir Surve, Zhenwen Liang, Nitesh Chawla, Olaf Wiest, and Xiangliang Zhang. Can llms solve molecule puzzles? a multimodal benchmark for molecular structure elucidation. Advances in Neural Information Processing Systems, 37:134721–134746, 2024.
- Thomas Hayes, Roshan Rao, Halil Akin, Nicholas J Sofroniew, Deniz Oktay, Zeming Lin, Robert Verkuil, Vincent Q Tran, Jonathan Deaton, Marius Wiggert, et al. Simulating 500 million years of evolution with a language model. Science, 387(6736):850–858, 2025.
- Haohuai He, Bing He, Lei Guan, Yu Zhao, Feng Jiang, Guanxing Chen, Qingge Zhu, Calvin Yu-Chian Chen, Ting Li, and Jianhua Yao. De novo generation of sars-cov-2 antibody cdrh3 with a pre-trained generative large language model. Nature Communications, 15(1):6867, 2024.
- Kai He, Rui Mao, Qika Lin, Yucheng Ruan, Xiang Lan, Mengling Feng, and Erik Cambria. A survey of large language models for healthcare: from data, technology, and applications to accountability and ethics. Information Fusion, page 102963, 2025.
- Megha Hegde, Jean-Christophe Nebel, and Farzana Rahman. Language modelling techniques for analysing the impact of human genetic variation. arXiv preprint arXiv:2503.10655, 2025.
- Shion Honda, Shoi Shi, and Hiroki R Ueda. Smiles transformer: Pre-trained molecular fingerprint for low data drug discovery. arXiv preprint arXiv:1911.04738, 2019.
- Edouardo Honig, Huixin Zhan, Ying Nian Wu, and Zijun Frank Zhang. Long-range gene expression prediction with token alignment of large language model. arXiv preprint arXiv:2410.01858, 2024.
- Wenpin Hou, Xinyi Shang, and Zhicheng Ji. Benchmarking large language models for genomic knowledge with geneturing. bioRxiv, pages 2023–03, 2025.
- C Hsu, R Verkuil, J Liu, Z Lin, B Hie, T Sercu, A Lerer, and A Rives. Learning inverse folding from millions of predicted structures. biorxiv (2022). URL https://api. semanticscholar. org/CorpusID, 248151599, 2022.
- Chengxin Hu, Hao Li, Yihe Yuan, Jing Li, and Ivor Tsang. Exploring hierarchical molecular graph representation in multimodal llms. arXiv preprint arXiv:2411.04708, 2024.
- Edward J Hu, Yelong Shen, Phillip Wallis, Zeyuan Allen-Zhu, Yuanzhi Li, Shean Wang, Lu Wang, and Weizhu Chen. Lora: Low-rank adaptation of large language models. In International Conference on Learning Representations, 2022.
- Mengzhou Hu, Sahar Alkhairy, Ingoo Lee, Rudolf T Pillich, Dylan Fong, Kevin Smith, Robin Bachelder, Trey Ideker, and Dexter Pratt. Evaluation of large language models for discovery of gene set function. Nature methods, 22(1):82–91, 2025.
- Ming Hu, Chenglong Ma, Wei Li, Wanghan Xu, Jiamin Wu, Jucheng Hu, Tianbin Li, Guohang Zhuang, Jiaqi Liu, Yingzhou Lu, Ying Chen, Chaoyang Zhang, Cheng Tan, Jie Ying, Guocheng Wu, Shujian Gao, Pengcheng Chen, Jiashi Lin, Haitao Wu, Lulu Chen, Fengxiang Wang, Yuanyuan Zhang, Xiangyu Zhao, Feilong Tang, Encheng Su, Junzhi Ning, Xinyao Liu, Ye Du, Changkai Ji, Cheng Tang, Huihui Xu, Ziyang Chen, Ziyan Huang, Jiyao Liu, Pengfei Jiang, Yizhou Wang, Chen Tang, Jianyu Wu, Yuchen Ren, Siyuan Yan, Zhonghua Wang, Zhongxing Xu, Shiyan Su, Shangquan Sun, Runkai Zhao, Zhisheng Zhang, Yu Liu, Fudi Wang, Yuanfeng Ji, Yanzhou Su, Hongming Shan, Chunmei Feng, Jiahao Xu, Jiangtao Yan, Wenhao Tang, Diping Song, Lihao Liu, Yanyan Huang, Lequan Yu, Bin Fu, Shujun Wang, Xiaomeng Li, Xiaowei Hu, Yun Gu, Ben Fei, Zhongying Deng, Benyou Wang, Yuewen Cao, Minjie Shen, Haodong Duan, Jie Xu, Yirong Chen, Fang Yan, Hongxia Hao, Jielan Li, Jiajun Du, Yanbo Wang, Imran Razzak, Chi Zhang, Lijun Wu, Conghui He, Zhaohui Lu, Jinhai Huang, Yihao Liu, Fenghua Ling, Yuqiang Li, Aoran Wang, Qihao Zheng, Nanqing Dong, Tianfan Fu, Dongzhan Zhou, Yan Lu, Wenlong Zhang, Jin Ye, Jianfei Cai, Wanli Ouyang, Yu Qiao, Zongyuan Ge, Shixiang Tang, Junjun He, Chunfeng Song, Lei Bai, and Bowen Zhou. A survey of scientific large language models: From data foundations to agent frontiers, 2025.
- Shaohan Huang, Li Dong, Wenhui Wang, Yaru Hao, Saksham Singhal, Shuming Ma, and et al. Language is not all you need: Aligning perception with language models. arXiv:2302.14045, 2023.
- Mingjia Huo, Han Guo, Xingyi Cheng, Digvijay Singh, Hamidreza Rahmani, Shen Li, Philipp Gerlof, Trey Ideker, Danielle A Grotjahn, Elizabeth Villa, et al. Multi-modal large language model enables protein function prediction. bioRxiv, pages 2024–08, 2024.
- Shuyi Jia, Chao Zhang, and Victor Fung. Llmatdesign: Autonomous materials discovery with large language models. arXiv preprint arXiv:2406.13163, 2024.
- Lei Jiang, Shuzhou Sun, Biqing Qi, Yuchen Fu, Xiaohua Xu, Yuqiang Li, Dongzhan Zhou, and Tianfan Fu. Chem3dllm: 3d multimodal large language models for chemistry, 2025.
- Chang Jin, Siyuan Guo, Shuigeng Zhou, and Jihong Guan. Effective and explainable molecular property prediction by chain-of-thought enabled large language models and multi-modal molecular information fusion. Journal of Chemical Information and Modeling, 2025.
- Qiao Jin, Yifan Yang, Qingyu Chen, and Zhiyong Lu. Genegpt: Augmenting large language models with domain tools for improved access to biomedical information. Bioinformatics, 40(2):btae075, 2024.
- Jiaxin Ju, YIZHEN ZHENG, Huan Yee Koh, Can Wang, and Shirui Pan. Chemthinker: Thinking like a chemist with multi-agent LLMs for deep molecular insights, 2024.
- John Jumper, Richard Evans, Alexander Pritzel, ..., and Demis Hassabis. Highly accurate protein structure prediction with alphafold. Nature, 596:583–589, 2021.
- Chenglong Kang, Xiaoyi Liu, and Fei Guo. Retrointext: A multimodal large language model enhanced framework for retrosynthetic planning via in-context representation learning. In The Thirteenth International Conference on Learning Representations, 2025.
- Taushif Khan, Mohammed Toufiq, Marina Yurieva, Nitaya Indrawattana, Akanitt Jittmittraphap, Nathamon Kosoltanapiwat, Pornpan Pumirat, Passanesh Sukphopetch, Muthita Vanaporn, Karolina Palucka, et al. Automating candidate gene prioritization with large language models: Development and benchmarking of an api-driven workflow leveraging gpt-4. bioRxiv, pages 2024–12, 2024.
- Junyoung Kim, Kai Wang, Chunhua Weng, and Cong Liu. Assessing the utility of large language models for phenotype-driven gene prioritization in the diagnosis of rare genetic disease. The American Journal of Human Genetics, 111(10):2190–2202, 2024.
- Takeshi Kojima, Shixiang Shane Gu, Machel Reid, Yutaka Matsuo, and Yusuke Iwasawa. Large language models are zero-shot reasoners. Advances in neural information processing systems, 35:22199–22213, 2022.
- Lingkai Kong, Yuanqi Du, Wenhao Mu, Kirill Neklyudov, Valentin De Bortoli, Dongxia Wu, Haorui Wang, Aaron Ferber, Yi-An Ma, Carla P Gomes, et al. Diffusion models as constrained samplers for optimization with unknown constraints. arXiv preprint arXiv:2402.18012, 2024.
- Mario Krenn, Florian Häse, Akshat Nigam, Pascal Friederich, and Alán Aspuru-Guzik. SELFIES: a robust representation of semantically constrained graphs. Machine Learning: Science and Technology, 1(4):045024, 2020.
- Khiem Le, Zhichun Guo, Kaiwen Dong, Xiaobao Huang, Bozhao Nan, Roshni Iyer, Xiangliang Zhang, Olaf Wiest, Wei Wang, and Nitesh V Chawla. Molx: Enhancing large language models for molecular learning with a multi-modal extension. arXiv preprint arXiv:2406.06777, 2024.
- Chanhui Lee, Yuheon Song, YongJun Jeong, Hanbum Ko, Rodrigo Hormazabal, Sehui Han, Kyunghoon Bae, Sungbin Lim, and Sungwoong Kim. Mol-llm: Generalist molecular llm with improved graph utilization. arXiv preprint arXiv:2502.02810, 2025.
- Chunyuan Li, Cliff Wong, Sheng Zhang, Naoto Usuyama, Haotian Liu, Jianwei Yang, Tristan Naumann, Hoifung Poon, and Jianfeng Gao. LLaVA-Med: Training a large language-and-vision assistant for biomedicine in one day. arXiv preprint arXiv:2306.00890, 2023.
- Chunyuan Li, Cliff Wong, Sheng Zhang, Naoto Usuyama, Haotian Liu, Jianwei Yang, Tristan Naumann, Hoifung Poon, and Jianfeng Gao. Llava-med: Training a large language-and-vision assistant for biomedicine in one day. Advances in Neural Information Processing Systems, 36:28541–28564, 2023.
- Hao Li, Yizheng Sun, Viktor Schlegel, Kailai Yang, Riza Batista-Navarro, and Goran Nenadic. Arg-llada: Argument summarization via large language diffusion models and sufficiency-aware refinement. arXiv preprint arXiv:2507.19081, 2025.
- Junnan Li, Dongxu Li, Silvio Savarese, and Steven Hoi. Blip-2: Bootstrapping language-image pre-training with frozen image encoders and large language models. arXiv preprint arXiv:2301.12597, 2023.
- Junxian Li, Di Zhang, Xunzhi Wang, Zeying Hao, Jingdi Lei, Qian Tan, Cai Zhou, Wei Liu, Yaotian Yang, Xinrui Xiong, et al. Chemvlm: Exploring the power of multimodal large language models in chemistry area. In Proceedings of the AAAI Conference on Artificial Intelligence, volume 39, pages 415–423, 2025.
- Longyi Li, Liyan Dong, Hao Zhang, Dong Xu, and Yongli Li. spallm: enhancing spatial domain analysis in multi-omics data through large language model integration. Briefings in Bioinformatics, 26(4):bbaf304, 07 2025.
- Mingchen Li, Yang Tan, Xinzhu Ma, Bozitao Zhong, Huiqun Yu, Ziyi Zhou, Wanli Ouyang, Bingxin Zhou, Pan Tan, and Liang Hong. Prosst: Protein language modeling with quantized structure and disentangled attention. Advances in Neural Information Processing Systems, 37:35700–35726, 2024.
- Peng-Hsuan Li, Yih-Yun Sun, Hsueh-Fen Juan, Chien-Yu Chen, Huai-Kuang Tsai, and Jia-Hsin Huang. A large language model framework for literature-based disease–gene association prediction. Briefings in Bioinformatics, 26(1):bbaf070, 2025.
- Yuesen Li, Chengyi Gao, Xin Song, Xiangyu Wang, Yungang Xu, and Suxia Han. Druggpt: A gpt-based strategy for designing potential ligands targeting specific proteins. bioRxiv, pages 2023–06, 2023.
- Lungang Liang, Yulan Chen, Taifu Wang, Dan Jiang, Jishuo Jin, Yanmeng Pang, Qin Na, Qiang Liu, Xiaosen Jiang, Wentao Dai, et al. Genetic transformer: An innovative large language model driven approach for rapid and accurate identification of causative variants in rare genetic diseases. medRxiv, pages 2024–07, 2024.
- Wang Liang. Llama-gene: A general-purpose gene task large language model based on instruction fine-tuning. arXiv preprint arXiv:2412.00471, 2024.
- Wang Liang. Llama-gene: A general-purpose gene task large language model based on instruction fine-tuning, 2024.
- Zijing Liang, Yanjie Xu, Yifan Hong, Penghui Shang, Qi Wang, Qiang Fu, and Ke Liu. A survey of multimodel large language models. In Proceedings of the 3rd International Conference on Computer, Artificial Intelligence and Control Engineering, pages 405–409, 2024.
- Xiaohan Lin, Zhenyu Chen, Yanheng Li, Xingyu Lu, Chuanliu Fan, Ziqiang Cao, Shihao Feng, Yi Qin Gao, and Jun Zhang. Protokens: A machine-learned language for compact and informative encoding of protein 3d structures. 2023.
- Yuxiang Lin, Ling Luo, Ying Chen, Xushi Zhang, Zihui Wang, Wenxian Yang, Mengsha Tong, and Rongshan Yu. St-align: A multimodal foundation model for image-gene alignment in spatial transcriptomics, 2024.
- Zeming Lin, Halil Akin, Roshan Rao, Brian Hie, Zhongkai Zhu, Wenting Lu, Nikita Smetanin, Robert Verkuil, Ori Kabeli, Yaniv Shmueli, et al. Evolutionary-scale prediction of atomic-level protein structure with a language model. Science, 379(6637):1123–1130, 2023.
- Bowen Liu, Bharath Ramsundar, Prasad Kawthekar, Jade Shi, Joseph Gomes, Quang Luu Nguyen, Stephen Ho, Jack Sloane, Paul Wender, and Vijay Pande. Retrosynthetic reaction prediction using neural sequence-to-sequence models. ACS central science, 3(10):1103–1113, 2017.
- Haoyang Liu, Yijiang Li, and Haohan Wang. Genomas: A multi-agent framework for scientific discovery via code-driven gene expression analysis. arXiv preprint arXiv:2507.21035, 2025.
- Hongxuan Liu, Haoyu Yin, Zhiyao Luo, and Xiaonan Wang. Integrating chemistry knowledge in large language models via prompt engineering. Synthetic and Systems Biotechnology, 10(1):23–38, 2025.
- Huaqing Liu, Shuxian Zhou, Peiyi Chen, Jiahui Liu, Ku-Geng Huo, and Lanqing Han. Exploring genomic large language models: Bridging the gap between natural language and gene sequences. bioRxiv, pages 2024–02, 2024.
- Lei Liu, Xiaoyan Yang, Junchi Lei, Xiaoyang Liu, Yue Shen, Zhiqiang Zhang, Peng Wei, Jinjie Gu, Zhixuan Chu, Zhan Qin, et al. A survey on medical large language models: Technology, application, trustworthiness, and future directions. arXiv preprint arXiv:2406.03712, 2024.
- Pengfei Liu, Yiming Ren, Jun Tao, and Zhixiang Ren. Git-mol: A multi-modal large language model for molecular science with graph, image, and text. Computers in biology and medicine, 171:108073, 2024.
- Shengchao Liu, Yanjing Li, Zhuoxinran Li, Anthony Gitter, Yutao Zhu, Jiarui Lu, Zhao Xu, Weili Nie, Arvind Ramanathan, Chaowei Xiao, et al. A text-guided protein design framework. Nature Machine Intelligence, pages 1–12, 2025.
- Shengchao Liu, Yanjing Li, Zhuoxinran Li, Anthony Gitter, Yutao Zhu, Jiarui Lu, Zhao Xu, Weili Nie, Arvind Ramanathan, Chaowei Xiao, Jian Tang, Hongyu Guo, and Anima Anandkumar. A text-guided protein design framework (proteindt). Nature Machine Intelligence, 2025. Advance online publication.
- Shengchao Liu, Weili Nie, Chengpeng Wang, Jiarui Lu, Zhuoran Qiao, Ling Liu, Jian Tang, Chaowei Xiao, and Animashree Anandkumar. Multi-modal molecule structure–text model for text-based retrieval and editing. Nature Machine Intelligence, 5(12):1447–1457, 2023.
- Siyu Liu, Tongqi Wen, Beilin Ye, Zhuoyuan Li, and David J. Srolovitz. Large language models for material property predictions: elastic constant tensor prediction and materials design, 2024.
- Tianyu Liu, Tinglin Huang, Rex Ying, and Hongyu Zhao. spemo: Exploring the capacity of foundation models for analyzing spatial multi-omic data. 2025.
- Tianyu Liu, Yijia Xiao, Xiao Luo, Hua Xu, W Jim Zheng, and Hongyu Zhao. Geneverse: A collection of open-source multimodal large language models for genomic and proteomic research. arXiv preprint arXiv:2406.15534, 2024.
- Xianggen Liu, Yan Guo, Haoran Li, Jin Liu, Shudong Huang, Bowen Ke, and Jiancheng Lv. Drugllm: Open large language model for few-shot molecule generation. arXiv preprint arXiv:2405.06690, 2024.
- Xiaoran Liu, Zhigeng Liu, Zengfeng Huang, Qipeng Guo, Ziwei He, and Xipeng Qiu. Longllada: Unlocking long context capabilities in diffusion llms. arXiv preprint arXiv:2506.14429, 2025.
- Zhiyuan Liu, An Zhang, Hao Fei, Enzhi Zhang, Xiang Wang, Kenji Kawaguchi, and Tat-Seng Chua. Prott3: Protein-to-text generation for text-based protein understanding. arXiv preprint arXiv:2405.12564, 2024.
- Micha Livne, Zulfat Miftahutdinov, Elena Tutubalina, Maksim Kuznetsov, Daniil Polykovskiy, Annika Brundyn, Aastha Jhunjhunwala, Anthony Costa, Alex Aliper, Alán Aspuru-Guzik, et al. nach0: multimodal natural and chemical languages foundation model. Chemical Science, 15(22):8380–8389, 2024.
- Renqian Luo, Liai Sun, Yingce Xia, Tao Qin, Sheng Zhang, Hoifung Poon, and Tie-Yan Liu. Biogpt: generative pre-trained transformer for biomedical text generation and mining. Briefings in bioinformatics, 23(6):bbac409, 2022.
- Yizhen Luo, Jiahuan Zhang, Siqi Fan, Kai Yang, Yushuai Wu, Mu Qiao, and Zaiqing Nie. Biomedgpt: Open multimodal generative pre-trained transformer for biomedicine. arXiv preprint arXiv:2308.09442, 2023.
- Yizhen Luo, Jiahuan Zhang, Siqi Fan, Kai Yang, Yushuai Wu, Mu Qiao, and Zaiqing Nie. Biomedgpt: Open multimodal generative pre-trained transformer for biomedicine. arXiv preprint arXiv:2308.09442, 2023.
- Omer Luxembourg, Haim Permuter, and Eliya Nachmani. Plan for speed–dilated scheduling for masked diffusion language models. arXiv preprint arXiv:2506.19037, 2025.
- Liuzhenghao Lv, Zongying Lin, Hao Li, Yuyang Liu, Jiaxi Cui, Calvin Yu-Chian Chen, Li Yuan, and Yonghong Tian. Prollama: A protein large language model for multi-task protein language processing. IEEE Transactions on Artificial Intelligence, 2025.
- Ali Madani, Ben Krause, Eric R. Greene, Subu Subramanian, Benjamin P. Mohr, James M. Holton, Jose L. Olmos, Caiming Xiong, Zachary Z. Sun, Richard Socher, James S. Fraser, and Nikhil Naik. Large language models generate functional protein sequences across diverse families. Nature Biotechnology, 41:1099–1106, 2023.
- Ali Madani, Ben Krause, Eric R Greene, Subu Subramanian, Benjamin P Mohr, James M Holton, Jose Luis Olmos Jr, Caiming Xiong, Zachary Z Sun, Richard Socher, et al. Large language models generate functional protein sequences across diverse families. Nature biotechnology, 41(8):1099–1106, 2023.
- Somshubra Majumdar, Vahid Noroozi, Mehrzad Samadi, Sean Narenthiran, Aleksander Ficek, Wasi Uddin Ahmad, Jocelyn Huang, Jagadeesh Balam, and Boris Ginsburg. Genetic instruct: Scaling up synthetic generation of coding instructions for large language models. arXiv preprint arXiv:2407.21077, 2024.
- Shentong Mo, Xi Fu, Chenyang Hong, Yizhen Chen, Yuxuan Zheng, Xiangru Tang, Zhiqiang Shen, Eric P Xing, and Yanyan Lan. Multi-modal self-supervised pre-training for regulatory genome across cell types. arXiv preprint arXiv:2110.05231, 2021.
- Su Mu, Meng Cui, and Xiaodi Huang. Multimodal data fusion in learning analytics: A systematic review. Sensors, 20(23):6856, 2020.
- Jorge Navaza and Pedro Saludjian. [33] amore: An automated molecular replacement program package. In Methods in enzymology, volume 276, pages 581–594. Elsevier, 1997.
- Shen Nie, Fengqi Zhu, Zebin You, Xiaolu Zhang, Jingyang Ou, Jun Hu, Jun Zhou, Yankai Lin, Ji-Rong Wen, and Chongxuan Li. Large language diffusion models. arXiv preprint arXiv:2502.09992, 2025.
- Erik Nijkamp, Jeffrey A Ruffolo, Eli N Weinstein, Nikhil Naik, and Ali Madani. Progen2: exploring the boundaries of protein language models. Cell systems, 14(11):968–978, 2023.
- Irene MA Nooren and Janet M Thornton. Diversity of protein–protein interactions. The EMBO journal, 2003.
- OpenAI. Gpt-4 technical report. arXiv:2303.08774, 2023.
- Long Ouyang, Jeffrey Wu, Xu Jiang, Diogo Almeida, Carroll Wainwright, Pamela Mishkin, Chong Zhang, Sandhini Agarwal, and et al. Training language models to follow instructions with human feedback. In Advances in Neural Information Processing Systems (NeurIPS), 2022.
- Qizhi Pei, Lijun Wu, Kaiyuan Gao, Xiaozhuan Liang, Yin Fang, Jinhua Zhu, Shufang Xie, Tao Qin, and Rui Yan. Biot5+: Towards generalized biological understanding with iupac integration and multi-task tuning. arXiv preprint arXiv:2402.17810, 2024.
- Qizhi Pei, Wei Zhang, Jinhua Zhu, Kehan Wu, Kaiyuan Gao, Lijun Wu, Yingce Xia, and Rui Yan. Biot5: Enriching cross-modal integration in biology with chemical knowledge and natural language associations. arXiv preprint arXiv:2310.07276, 2023.
- Edward O Pyzer-Knapp, Matteo Manica, Peter Staar, Lucas Morin, Patrick Ruch, Teodoro Laino, John R Smith, and Alessandro Curioni. Foundation models for materials discovery–current state and future directions. Npj Computational Materials, 11(1):61, 2025.
- Alec Radford, Jeffrey Wu, Rewon Child, David Luan, Dario Amodei, and Ilya Sutskever. Language models are unsupervised multitask learners. https://cdn.openai.com/better-language-models/language_models_are_unsupervised_multitask_learners.pdf, 2019. OpenAI Technical Report.
- Rafael Rafailov, Archit Sharma, Eric Mitchell, Christopher D Manning, Stefano Ermon, and Chelsea Finn. Direct preference optimization: Your language model is secretly a reward model. Advances in neural information processing systems, 36:53728–53741, 2023.
- Roshan Rao, Nicholas Bhattacharya, Neil Thomas, Yan Duan, Peter Chen, John Canny, Pieter Abbeel, and Yun Song. Evaluating protein transfer learning with tape. Advances in neural information processing systems, 32, 2019.
- Roshan M Rao, Jason Liu, Robert Verkuil, Joshua Meier, John Canny, Pieter Abbeel, Tom Sercu, and Alexander Rives. Msa transformer. In International conference on machine learning, pages 8844–8856. PMLR, 2021.
- Guillaume Richard, Bernardo P de Almeida, Hugo Dalla-Torre, Christopher Blum, Lorenz Hexemer, Priyanka Pandey, Stefan Laurent, Marie Lopez, Alexandre Laterre, Maren Lang, et al. Chatnt: A multimodal conversational agent for dna, rna and protein tasks. bioRxiv, pages 2024–04, 2024.
- Alexander Rives, Joshua Meier, Tom Sercu, Siddharth Goyal, Zeming Lin, Jason Liu, Demi Guo, Myle Ott, C. Lawrence Zitnick, Jerry Ma, and Rob Fergus. Biological structure and function emerge from scaling unsupervised learning to 250 million protein sequences. Proceedings of the National Academy of Sciences, 118(15):e2016239118, 2021.
- Zachary A Rollins, Alan C Cheng, and Essam Metwally. Molprop: Molecular property prediction with multimodal language and graph fusion. Journal of Cheminformatics, 16(1):56, 2024.
- Jeffrey A Ruffolo, Aadyot Bhatnagar, Joel Beazer, Stephen Nayfach, Jordan Russ, Emily Hill, Riffat Hussain, Joseph Gallagher, and Ali Madani. Adapting protein language models for structure-conditioned design. BioRxiv, pages 2024–08, 2024.
- Daan Schouten, Giulia Nicoletti, Bas Dille, Catherine Chia, Pierpaolo Vendittelli, Megan Schuurmans, Geert Litjens, and Nadieh Khalili. Navigating the landscape of multimodal ai in medicine: a scoping review on technical challenges and clinical applications. Medical Image Analysis, page 103621, 2025.
- Christoph Schuhmann, Romain Beaumont, Richard Vencu, Cade Gordon, Ross Wightman, Mehdi Cherti, Theo Coombes, Aarush Katta, Clayton Mullis, Mitchell Wortsman, et al. Laion-5b: An open large-scale dataset for training next generation image-text models. arXiv preprint arXiv:2210.08402, 2022.
- Zhang Shengyu, Dong Linfeng, Li Xiaoya, Zhang Sen, Sun Xiaofei, Wang Shuhe, Li Jiwei, Runyi Hu, Zhang Tianwei, Fei Wu, et al. Instruction tuning for large language models: A survey. arXiv preprint arXiv:2308.10792, 2023.
- Aleksei Shmelev, Artem Shadskiy, Yuri Kuratov, Mikhail Burtsev, Olga Kardymon, and Veniamin Fishman. Genatator: de novo gene annotation with dna language model. In ICLR 2025 Workshop on AI for Nucleic Acids.
- Richard W Shuai, Jeffrey A Ruffolo, and Jeffrey J Gray. Iglm: Infilling language modeling for antibody sequence design. Cell Systems, 14(11):979–989, 2023.
- Yuerong Song, Xiaoran Liu, Ruixiao Li, Zhigeng Liu, Zengfeng Huang, Qipeng Guo, Ziwei He, and Xipeng Qiu. Sparse-dllm: Accelerating diffusion llms with dynamic cache eviction. arXiv preprint arXiv:2508.02558, 2025.
- Anuroop Sriram, Benjamin Miller, Ricky TQ Chen, and Brandon Wood. Flowllm: Flow matching for material generation with large language models as base distributions. Advances in Neural Information Processing Systems, 37:46025–46046, 2024.
- Jin Su, Chenchen Han, Yuyang Zhou, Junjie Shan, Xibin Zhou, and Fajie Yuan. Saprot: Protein language modeling with structure-aware vocabulary. BioRxiv, pages 2023–10, 2023.
- Jin Su, Chenchen Han, Yuyang Zhou, Junjie Shan, Xibin Zhou, and Fajie Yuan. Saprot: Protein language modeling with structure-aware vocabulary. BioRxiv, pages 2023–10, 2023.
- Xiangru Tang, Tianyu Hu, Muyang Ye, Yanjun Shao, Xunjian Yin, Siru Ouyang, Wangchunshu Zhou, Pan Lu, Zhuosheng Zhang, Yilun Zhao, et al. Chemagent: Self-updating library in large language models improves chemical reasoning. arXiv preprint arXiv:2501.06590, 2025.
- Ross Taylor, Marcin Kardas, Guillem Cucurull, Thomas Scialom, Anthony Hartshorn, Elvis Saravia, Andrew Poulton, Viktor Kerkez, and Robert Stojnic. Galactica: A large language model for science. arXiv preprint arXiv:2211.09085, 2022.
- Igor V. Tetko, Pavel Karpov, Ruud Van Deursen, and Gaston Godin. State-of-the-art augmented NLP transformer models for direct and single-step retrosynthesis. Journal of Chemical Information and Modeling, 60(12):5744–5752, 2020.
- Arun James Thirunavukarasu, Darren Shu Jeng Ting, Kabilan Elangovan, Laura Gutierrez, Ting Fang Tan, and Daniel Shu Wei Ting. Large language models in medicine. Nature medicine, 29(8):1930–1940, 2023.
- Jie Tian, Martin Taylor Sobczak, Dhanush Patil, Jixin Hou, Lin Pang, Arunachalam Ramanathan, Libin Yang, Xianyan Chen, Yuval Golan, Xiaoming Zhai, Hongyue Sun, Kenan Song, and Xianqiao Wang. A multi-agent framework integrating large language models and generative ai for accelerated metamaterial design, 2025.
- Mohammed Toufiq, Darawan Rinchai, Eleonore Bettacchioli, Basirudeen Syed Ahamed Kabeer, Taushif Khan, Bishesh Subba, Olivia White, Marina Yurieva, Joshy George, Noemie Jourde-Chiche, et al. Harnessing large language models (llms) for candidate gene prioritization and selection. Journal of translational medicine, 21(1):728, 2023.
- Duong Tran, Nhat Truong Pham, Nguyen Nguyen, and Balachandran Manavalan. Mol2lang-vlm: Vision-and text-guided generative pre-trained language models for advancing molecule captioning through multimodal fusion. In Proceedings of the 1st Workshop on Language+ Molecules (L+ M 2024), pages 97–102, 2024.
- Michel van Kempen, Stephanie S Kim, Charlotte Tumescheit, Milot Mirdita, Cameron LM Gilchrist, Johannes Söding, and Martin Steinegger. Foldseek: fast and accurate protein structure search. Biorxiv, pages 2022–02, 2022.
- Ashish Vaswani, Noam Shazeer, Niki Parmar, Jakob Uszkoreit, Llion Jones, Aidan N. Gomez, Lukasz Kaiser, and Illia Polosukhin. Attention is all you need. In Advances in Neural Information Processing Systems (NeurIPS), volume 30, pages 5998–6008, 2017.
- Chao Wang, Hehe Fan, Ruijie Quan, Lina Yao, and Yi Yang. Protchatgpt: Towards understanding proteins with hybrid representation and large language models. In Proceedings of the 48th International ACM SIGIR Conference on Research and Development in Information Retrieval, pages 1076–1086, 2025.
- Chong Wang, Mengyao Li, Junjun He, Zhongruo Wang, Erfan Darzi, Zan Chen, Jin Ye, Tianbin Li, Yanzhou Su, Jing Ke, et al. A survey for large language models in biomedicine. arXiv preprint arXiv:2409.00133, 2024.
- Dandan Wang and Shiqing Zhang. Large language models in medical and healthcare fields: applications, advances, and challenges. Artificial Intelligence Review, 57(11):299, 2024.
- Jike Wang, Rui Qin, Mingyang Wang, Meijing Fang, Yangyang Zhang, Yuchen Zhu, Qun Su, Qiaolin Gou, Chao Shen, Odin Zhang, et al. Token-mol 1.0: tokenized drug design with large language models. Nature Communications, 16(1):1–19, 2025.
- Peng Wang, Wenpeng Lu, Chunlin Lu, Ruoxi Zhou, Min Li, and Libo Qin. Large language model for medical images: A survey of taxonomy, systematic review, and future trends. Big Data Mining and Analytics, 8(2):496–517, 2025.
- X Wang, Z Zheng, F Ye, D Xue, S Huang, and Q Gu. Dplm-2: a multimodal diffusion protein language model. arxiv. arXiv preprint arXiv:2410.13782, 2024.
- Xinyou Wang, Zaixiang Zheng, Fei Ye, Dongyu Xue, Shujian Huang, and Quanquan Gu. Diffusion language models are versatile protein learners. arXiv preprint arXiv:2402.18567, 2024.
- Yue Wang and Xueying Tian. Qwendy: Gene regulatory network inference enhanced by large language model and transformer. arXiv preprint arXiv:2503.09605, 2025.
- Zeyuan Wang, Qiang Zhang, Keyan Ding, Ming Qin, Xiang Zhuang, Xiaotong Li, and Huajun Chen. Instructprotein: Aligning human and protein language via knowledge instruction. arXiv preprint arXiv:2310.03269, 2023.
- Zhenzhong Wang, Haowei Hua, Wanyu Lin, Ming Yang, and Kay Chen Tan. Crystalline material discovery in the era of artificial intelligence. arXiv preprint arXiv:2408.08044, 2024.
- Zhizheng Wang, Chi-Ping Day, Chih-Hsuan Wei, Qiao Jin, Robert Leaman, Yifan Yang, Shubo Tian, Aodong Qiu, Yin Fang, Qingqing Zhu, et al. Knowledge-guided contextual gene set analysis using large language models. arXiv preprint arXiv:2506.04303, 2025.
- Zifeng Wang, Zichen Wang, Balasubramaniam Srinivasan, Vassilis N Ioannidis, Huzefa Rangwala, and Rishita Anubhai. Biobridge: Bridging biomedical foundation models via knowledge graphs. arXiv preprint arXiv:2310.03320, 2023.
- Jason Wei, Maarten Bosma, Vincent Y Zhao, Kelvin Guu, Adams Wei Yu, Brian Lester, Nan Du, Andrew M Dai, and Quoc V Le. Finetuned language models are zero-shot learners. arXiv preprint arXiv:2109.01652, 2021.
- Jason Wei, Yi Tay, Rishi Bommasani, Colin Raffel, Barret Zoph, Sebastian Borgeaud, Dani Yogatama, Maarten Bosma, Denny Zhou, Donald Metzler, et al. Emergent abilities of large language models. arXiv preprint arXiv:2206.07682, 2022.
- David Weininger. Smiles, a chemical language and information system. 1. introduction to methodology and encoding rules. Journal of chemical information and computer sciences, 28(1):31–36, 1988.
- Zichen Wen, Jiashu Qu, Dongrui Liu, Zhiyuan Liu, Ruixi Wu, Yicun Yang, Xiangqi Jin, Haoyun Xu, Xuyang Liu, Weijia Li, et al. The devil behind the mask: An emergent safety vulnerability of diffusion llms. arXiv preprint arXiv:2507.11097, 2025.
- Daniel S Wigh, Jonathan M Goodman, and Alexei A Lapkin. A review of molecular representation in the age of machine learning. Wiley Interdisciplinary Reviews: Computational Molecular Science, 12(5):e1603, 2022.
- Chengyue Wu, Hao Zhang, Shuchen Xue, Zhijian Liu, Shizhe Diao, Ligeng Zhu, Ping Luo, Song Han, and Enze Xie. Fast-dllm: Training-free acceleration of diffusion llm by enabling kv cache and parallel decoding. arXiv preprint arXiv:2505.22618, 2025.
- Jiayang Wu, Wensheng Gan, Zefeng Chen, Shicheng Wan, and Philip S Yu. Multimodal large language models: A survey. In 2023 IEEE International Conference on Big Data (BigData), pages 2247–2256. IEEE, 2023.
- Kevin E Wu, Kathryn Yost, Bence Daniel, Julia Belk, Yu Xia, Takeshi Egawa, Ansuman Satpathy, Howard Chang, and James Zou. Tcr-bert: learning the grammar of t-cell receptors for flexible antigen-binding analyses. In Machine Learning in Computational Biology, pages 194–229. PMLR, 2024.
- Zhenqin Wu, Bharath Ramsundar, Evan N Feinberg, Joseph Gomes, Caleb Geniesse, Aneesh S Pappu, Karl Leswing, and Vijay Pande. Moleculenet: a benchmark for molecular machine learning. Chemical science, 9(2):513–530, 2018.
- Zhenxing Wu, Odin Zhang, Xiaorui Wang, Li Fu, Huifeng Zhao, Jike Wang, Hongyan Du, Dejun Jiang, Yafeng Deng, Dongsheng Cao, et al. Leveraging language model for advanced multiproperty molecular optimization via prompt engineering. Nature Machine Intelligence, pages 1–11, 2024.
- Hanguang Xiao, Feizhong Zhou, Xingyue Liu, Tianqi Liu, Zhipeng Li, Xin Liu, and Xiaoxuan Huang. A comprehensive survey of large language models and multimodal large language models in medicine. Information Fusion, page 102888, 2024.
- Hanguang Xiao, Feizhong Zhou, Xingyue Liu, Tianqi Liu, Zhipeng Li, Xin Liu, and Xiaoxuan Huang. A comprehensive survey of large language models and multimodal large language models in medicine. Information Fusion, 117:102888, 2025.
- Hongwang Xiao, Wenjun Lin, Xi Chen, Hui Wang, Kai Chen, Jiashan Li, Yuancheng Sun, Sicheng Dai, Boya Wu, and Qiwei Ye. Stella: Towards protein function prediction with multimodal llms integrating sequence-structure representations. arXiv preprint arXiv:2506.03800, 2025.
- Teng Xiao, Chao Cui, Huaisheng Zhu, and Vasant G Honavar. Molbind: Multimodal alignment of language, molecules, and proteins. arXiv preprint arXiv:2403.08167, 2024.
- Teng Xiao, Chao Cui, Huaisheng Zhu, and Vasant G Honavar. Molbind: Multimodal alignment of language, molecules, and proteins. arXiv preprint arXiv:2403.08167, 2024.
- Yijia Xiao, Edward Sun, Yiqiao Jin, Qifan Wang, and Wei Wang. Proteingpt: Multimodal llm for protein property prediction and structure understanding. arXiv preprint arXiv:2408.11363, 2024.
- Zhen Xiong, Yujun Cai, Zhecheng Li, and Yiwei Wang. Unveiling the potential of diffusion large language model in controllable generation. arXiv preprint arXiv:2507.04504, 2025.
- Hanwen Xu, Addie Woicik, Hoifung Poon, Russ B Altman, and Sheng Wang. Multilingual translation for zero-shot biomedical classification using biotranslator. Nature Communications, 14(1):738, 2023.
- Yingxue Xu, Yihui Wang, Fengtao Zhou, Jiabo Ma, Cheng Jin, Shu Yang, Jinbang Li, Zhengyu Zhang, Chenglong Zhao, Huajun Zhou, Zhenhui Li, Huangjing Lin, Xin Wang, Jiguang Wang, Anjia Han, Ronald Cheong Kin Chan, Li Liang, Xiuming Zhang, and Hao Chen. A multimodal knowledge-enhanced whole-slide pathology foundation model, 2025.
- Keqiang Yan, Xiner Li, Hongyi Ling, Kenna Ashen, Carl Edwards, Raymundo Arróyave, Marinka Zitnik, Heng Ji, Xiaofeng Qian, Xiaoning Qian, et al. Invariant tokenization of crystalline materials for language model enabled generation. Advances in Neural Information Processing Systems, 37:125050–125072, 2024.
- Sherry Yang, Simon Batzner, Ruiqi Gao, Muratahan Aykol, Alexander Gaunt, Brendan C McMorrow, Danilo Jimenez Rezende, Dale Schuurmans, Igor Mordatch, and Ekin Dogus Cubuk. Generative hierarchical materials search. Advances in Neural Information Processing Systems, 37:38799–38819, 2024.
- Xiaodong Yang, Guole Liu, Guihai Feng, Dechao Bu, Pengfei Wang, Jie Jiang, Shubai Chen, Qinmeng Yang, Hefan Miao, Yiyang Zhang, et al. Genecompass: deciphering universal gene regulatory mechanisms with a knowledge-informed cross-species foundation model. Cell Research, 34(12):830–845, 2024.
- Zhilin Yang, Zihang Dai, Yiming Yang, Jaime Carbonell, Ruslan Salakhutdinov, and Quoc V Le. Xlnet: Generalized autoregressive pretraining for language understanding. arXiv preprint arXiv:1906.08237, 2019.
- Jiacheng Ye, Jiahui Gao, Shansan Gong, Lin Zheng, Xin Jiang, Zhenguo Li, and Lingpeng Kong. Beyond autoregression: Discrete diffusion for complex reasoning and planning. arXiv preprint arXiv:2410.14157, 2024.
- Jiacheng Ye, Shansan Gong, Liheng Chen, Lin Zheng, Jiahui Gao, Han Shi, Chuan Wu, Xin Jiang, Zhenguo Li, Wei Bi, et al. Diffusion of thought: Chain-of-thought reasoning in diffusion language models. Advances in Neural Information Processing Systems, 37:105345–105374, 2024.
- Jiarui Ye and Hao Tang. Multimodal large language models for medicine: A comprehensive survey. arXiv preprint arXiv:2504.21051, 2025.
- Shukang Yin, Chaoyou Fu, Sirui Zhao, Ke Li, Xing Sun, Tong Xu, and Enhong Chen. A survey on multimodal large language models. National Science Review, 11(12), November 2024.
- Hyunwoo Yoo. Can large language models predict antimicrobial resistance gene? arXiv preprint arXiv:2503.04413, 2025.
- Zebin You, Shen Nie, Xiaolu Zhang, Jun Hu, Jun Zhou, Zhiwu Lu, Ji-Rong Wen, and Chongxuan Li. Llada-v: Large language diffusion models with visual instruction tuning. arXiv preprint arXiv:2505.16933, 2025.
- Botao Yu, Frazier N Baker, Ziru Chen, Garrett Herb, Boyu Gou, Daniel Adu-Ampratwum, Xia Ning, and Huan Sun. Tooling or not tooling? the impact of tools on language agents for chemistry problem solving. arXiv preprint arXiv:2411.07228, 2024.
- Runpeng Yu, Qi Li, and Xinchao Wang. Discrete diffusion in large language and multimodal models: A survey. arXiv preprint arXiv:2506.13759, 2025.
- Runpeng Yu, Xinyin Ma, and Xinchao Wang. Dimple: Discrete diffusion multimodal large language model with parallel decoding. arXiv preprint arXiv:2505.16990, 2025.
- Yi Yu, Huien Wang, Libin Zong, Bo Chen, Yaqin Li, and Xiaohui Yu. Chatmoldata: A multimodal agent for automatic molecular data processing. Advanced Intelligent Systems, page 2401089, 2024.
- Haolong Zeng, Chaoyi Yin, Chunyang Chai, Yuezhu Wang, Qi Dai, and Huiyan Sun. Cancer gene identification through integrating causal prompting large language model with omics data–driven causal inference. Briefings in Bioinformatics, 26(2), 2025.
- Zheni Zeng, Bangchen Yin, Shipeng Wang, Jiarui Liu, Cheng Yang, Haishen Yao, Xingzhi Sun, Maosong Sun, Guotong Xie, and Zhiyuan Liu. Chatmol: interactive molecular discovery with natural language. Bioinformatics, 40(9):btae534, 2024.
- Heming Zhang, Tim Xu, Dekang Cao, Shunning Liang, Lars Schimmelpfennig, Levi Kaster, Di Huang, Carlos Cruchaga, Guangfu Li, Michael Province, et al. Omnicelltosg: The first cell text-omic signaling graphs dataset for joint llm and gnn modeling. arXiv preprint arXiv:2504.02148, 2025.
- Juzheng Zhang, Yatao Bian, Yongqiang Chen, and Quanming Yao. Unimot: Unified molecule-text language model with discrete token representation. arXiv preprint arXiv:2408.00863, 2024.
- Ningyu Zhang, Zhen Bi, Xiaozhuan Liang, Siyuan Cheng, Haosen Hong, Shumin Deng, Jiazhang Lian, Qiang Zhang, and Huajun Chen. Ontoprotein: Protein pretraining with gene ontology embedding. arXiv preprint arXiv:2201.11147, 2022.
- Ningyu Zhang, Zhen Bi, Xiaozhuan Liang, Siyuan Cheng, Haosen Hong, Shumin Deng, Jiazhang Lian, Qiang Zhang, and Huajun Chen. Ontoprotein: Protein pretraining with gene ontology embedding. arXiv preprint arXiv:2201.11147, 2022.
- Ningyu Zhang, Zhen Bi, Xiaozhuan Liang, Siyuan Cheng, Haosen Hong, Shumin Deng, Qiang Zhang, Jiazhang Lian, and Huajun Chen. Ontoprotein: Protein pretraining with gene ontology embedding. In International Conference on Learning Representations (ICLR), 2022.
- Qiang Zhang, Keyan Ding, Tianwen Lv, Xinda Wang, Qingyu Yin, Yiwen Zhang, Jing Yu, Yuhao Wang, Xiaotong Li, Zhuoyi Xiang, et al. Scientific large language models: A survey on biological & chemical domains. ACM Computing Surveys, 57(6):1–38, 2025.
- Qiang Zhang, Keyang Ding, Tianwen Lyv, Xinda Wang, Qingyu Yin, Yiwen Zhang, Jing Yu, Yuhao Wang, Xiaotong Li, Zhuoyi Xiang, Kehua Feng, Xiang Zhuang, Zeyuan Wang, Ming Qin, Mengyao Zhang, Jinlu Zhang, Jiyu Cui, Tao Huang, Pengju Yan, Renjun Xu, Hongyang Chen, Xiaolin Li, Xiaohui Fan, Huabin Xing, and Huajun Chen. Scientific large language models: A survey on biological & chemical domains, 2024.
- Tianren Zhang and Dai-Bei Yang. Multimodal machine learning with large language embedding model for polymer property prediction. arXiv preprint arXiv:2503.22962, 2025.
- Yu Zhang, Xiusi Chen, Bowen Jin, Sheng Wang, Shuiwang Ji, Wei Wang, and Jiawei Han. A comprehensive survey of scientific large language models and their applications in scientific discovery. arXiv preprint arXiv:2406.10833, 2024.
- Yu Zhang, Ruijie Yu, Kaipeng Zeng, Ding Li, Feng Zhu, Xiaokang Yang, Yaohui Jin, and Yanyan Xu. Text-augmented multimodal llms for chemical reaction condition recommendation. arXiv preprint arXiv:2407.15141, 2024.
- Yuanhe Zhang, Fangzhou Xie, Zhenhong Zhou, Zherui Li, Hao Chen, Kun Wang, and Yufei Guo. Jailbreaking large language diffusion models: Revealing hidden safety flaws in diffusion-based text generation. arXiv preprint arXiv:2507.19227, 2025.
- Zuobai Zhang, Chuanrui Wang, Minghao Xu, Vijil Chenthamarakshan, Aurélie Lozano, Payel Das, and Jian Tang. A systematic study of joint representation learning on protein sequences and structures. arXiv preprint arXiv:2303.06275, 2023.
- Siyan Zhao, Devaansh Gupta, Qinqing Zheng, and Aditya Grover. d1: Scaling reasoning in diffusion large language models via reinforcement learning. arXiv preprint arXiv:2504.12216, 2025.
- Wayne Xin Zhao, Kun Zhou, Junyi Li, Tianyi Tang, Xiaolei Wang, Yupeng Hou, Yingqian Min, Beichen Zhang, Junjie Zhang, Zican Dong, et al. A survey of large language models. arXiv preprint arXiv:2303.18223, 1(2), 2023.
- Kangjie Zheng, Siyu Long, Tianyu Lu, Junwei Yang, Xinyu Dai, Ming Zhang, Zaiqing Nie, Wei-Ying Ma, and Hao Zhou. Esm all-atom: multi-scale protein language model for unified molecular modeling. arXiv preprint arXiv:2403.12995, 2024.
- Lin Zheng, Jianbo Yuan, Lei Yu, and Lingpeng Kong. A reparameterized discrete diffusion model for text generation. arXiv preprint arXiv:2302.05737, 2023.
- Yanxin Zheng, Wensheng Gan, Zefeng Chen, Zhenlian Qi, Qian Liang, and Philip S Yu. Large language models for medicine: a survey. International Journal of Machine Learning and Cybernetics, 16(2):1015–1040, 2025.
- Hanjing Zhou, Mingze Yin, Wei Wu, Mingyang Li, Kun Fu, Jintai Chen, Jian Wu, and Zheng Wang. Protclip: Function-informed protein multi-modal learning. In Proceedings of the AAAI Conference on Artificial Intelligence, volume 39, pages 22937–22945, 2025.
- Hongjian Zhou, Fenglin Liu, Boyang Gu, Xinyu Zou, Jinfa Huang, Jinge Wu, Yiru Li, Sam S Chen, Peilin Zhou, Junling Liu, et al. A survey of large language models in medicine: Progress, application, and challenge. arXiv preprint arXiv:2311.05112, 2023.
- Jiaming Zhou, Hongjie Chen, Shiwan Zhao, Jian Kang, Jie Li, Enzhi Wang, Yujie Guo, Haoqin Sun, Hui Wang, Aobo Kong, et al. Diffa: Large language diffusion models can listen and understand. arXiv preprint arXiv:2507.18452, 2025.
- Peng Zhou, Pengsen Ma, Jianmin Wang, Xibao Cai, Haitao Huang, Wei Liu, Longyue Wang, Lai Hou Tim, and Xiangxiang Zeng. Large language and protein assistant for protein-protein interactions prediction. In Proceedings of the 63rd Annual Meeting of the Association for Computational Linguistics (Volume 1: Long Papers), pages 11312–11327, 2025.
- Xibin Zhou, Chenchen Han, Yingqi Zhang, Jin Su, Kai Zhuang, Shiyu Jiang, Zichen Yuan, Wei Zheng, Fengyuan Dai, Yuyang Zhou, et al. Decoding the molecular language of proteins with evolla. bioRxiv, pages 2025–01, 2025.
- Zhihan Zhou, Yanrong Ji, Weijian Li, Pratik Dutta, Ramana Davuluri, and Han Liu. Dnabert-2: Efficient foundation model and benchmark for multi-species genome. arXiv preprint arXiv:2306.15006, 2023.
- Deyao Zhu, Jun Chen, Xiaoqian Shen, Xiang Li, and Mohamed Elhoseiny. Minigpt-4: Enhancing vision-language understanding with advanced large language models. arXiv:2304.10592, 2023.
- Yi-Heng Zhu, Chengxin Zhang, Dong-Jun Yu, and Yang Zhang. Integrating unsupervised language model with triplet neural networks for protein gene ontology prediction. PLOS Computational Biology, 18(12):e1010793, 2022.
- Xiang Zhuang, Keyan Ding, Tianwen Lyu, Yinuo Jiang, Xiaotong Li, Zhuoyi Xiang, Zeyuan Wang, Ming Qin, Kehua Feng, Jike Wang, et al. Instructbiomol: Advancing biomolecule understanding and design following human instructions. arXiv preprint arXiv:2410.07919, 2024.
- Le Zhuo, Zewen Chi, Minghao Xu, Heyan Huang, Heqi Zheng, Conghui He, Xian-Ling Mao, and Wentao Zhang. Protllm: An interleaved protein-language llm with protein-as-word pre-training. arXiv preprint arXiv:2403.07920, 2024.
- Daniel M Ziegler, Nisan Stiennon, Jeffrey Wu, Tom B Brown, Alec Radford, Dario Amodei, Paul Christiano, and Geoffrey Irving. Fine-tuning language models from human preferences. arXiv preprint arXiv:1909.08593, 2019.
- Peter H Zwart, Pavel V Afonine, Ralf W Grosse-Kunstleve, Li-Wei Hung, Thomas R Ioerger, Airlie J McCoy, Erik McKee, Nigel W Moriarty, Randy J Read, James C Sacchettini, et al. Automated structure solution with the PHENIX suite. Springer, 2008.
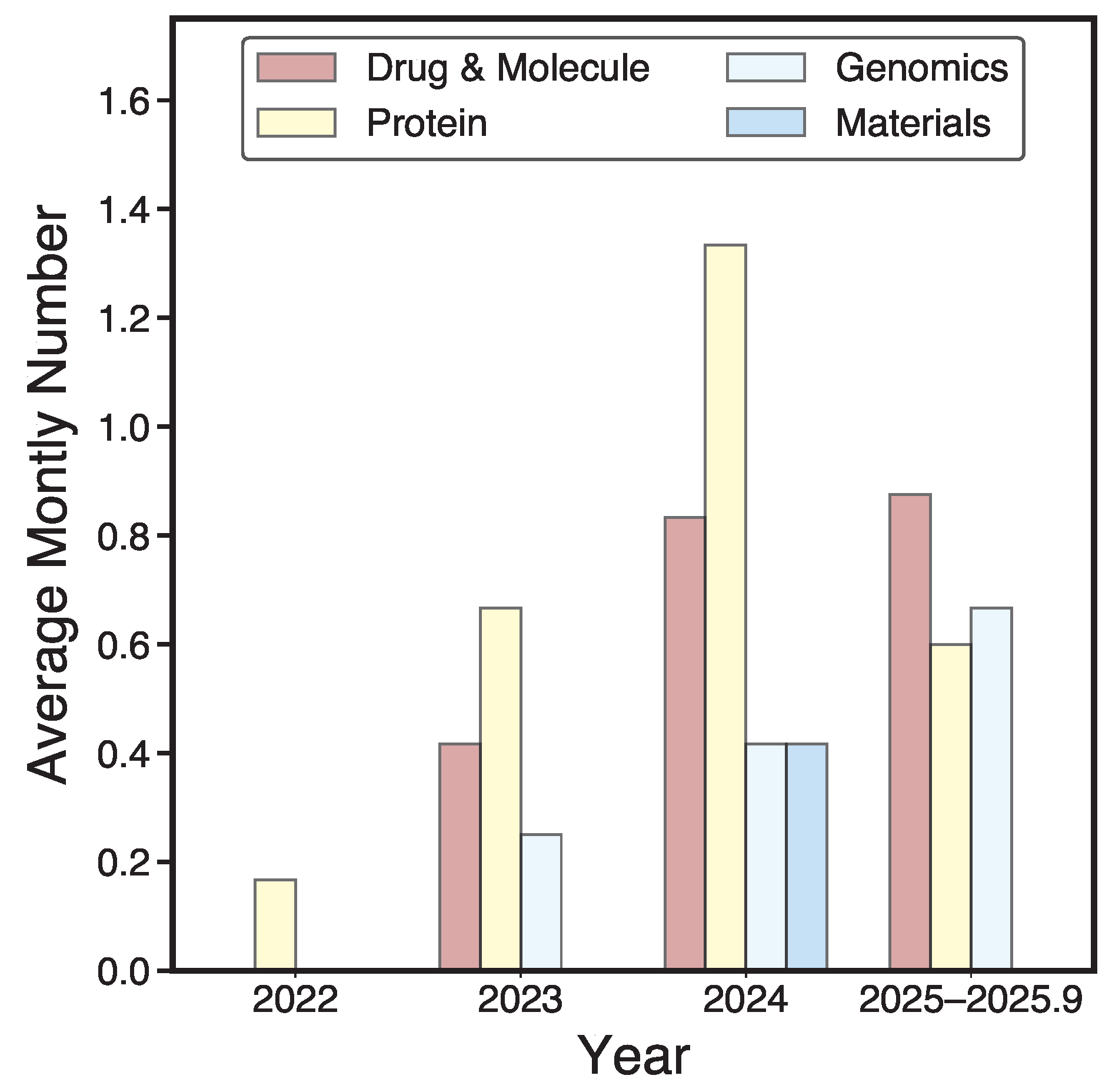
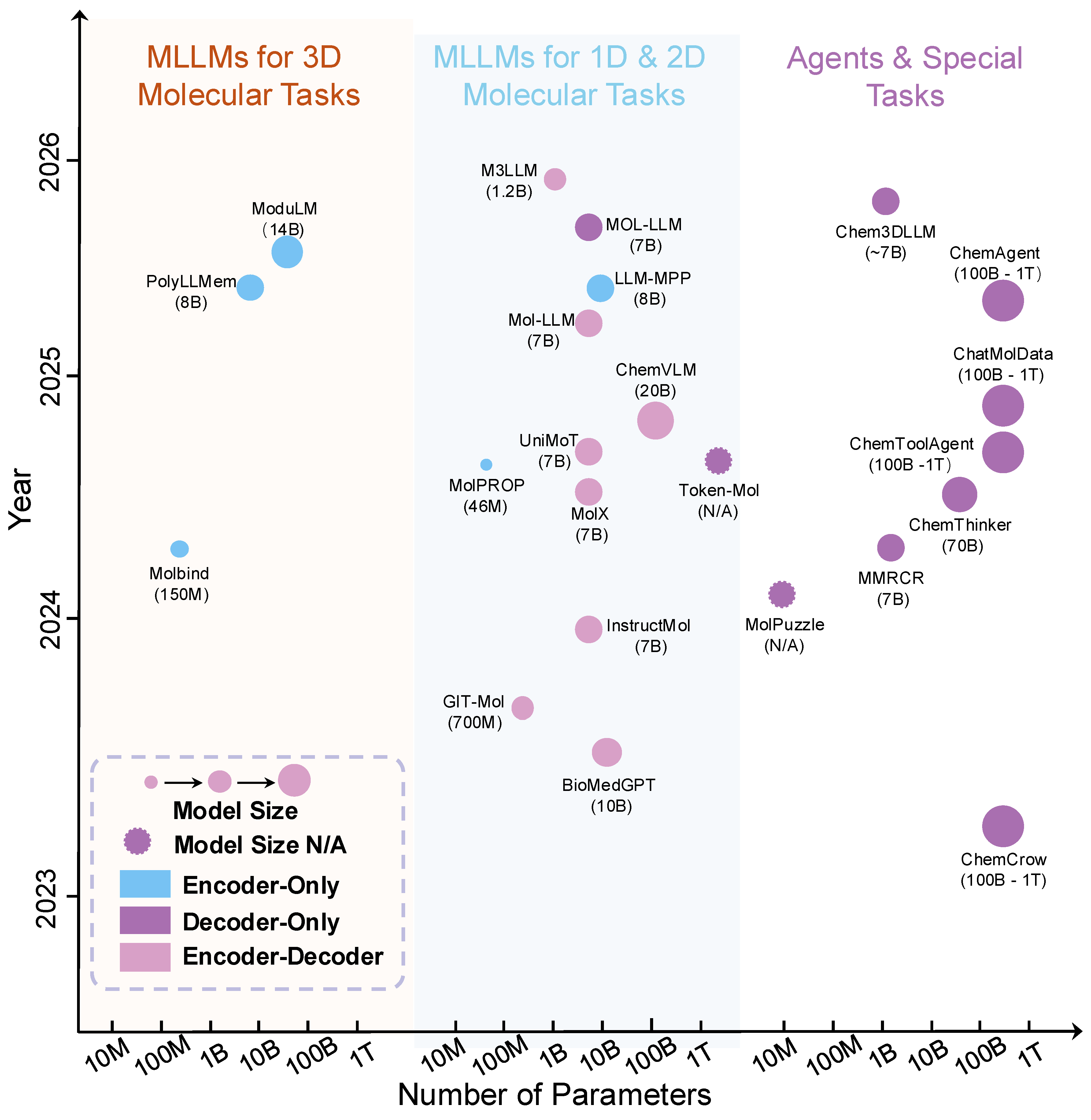
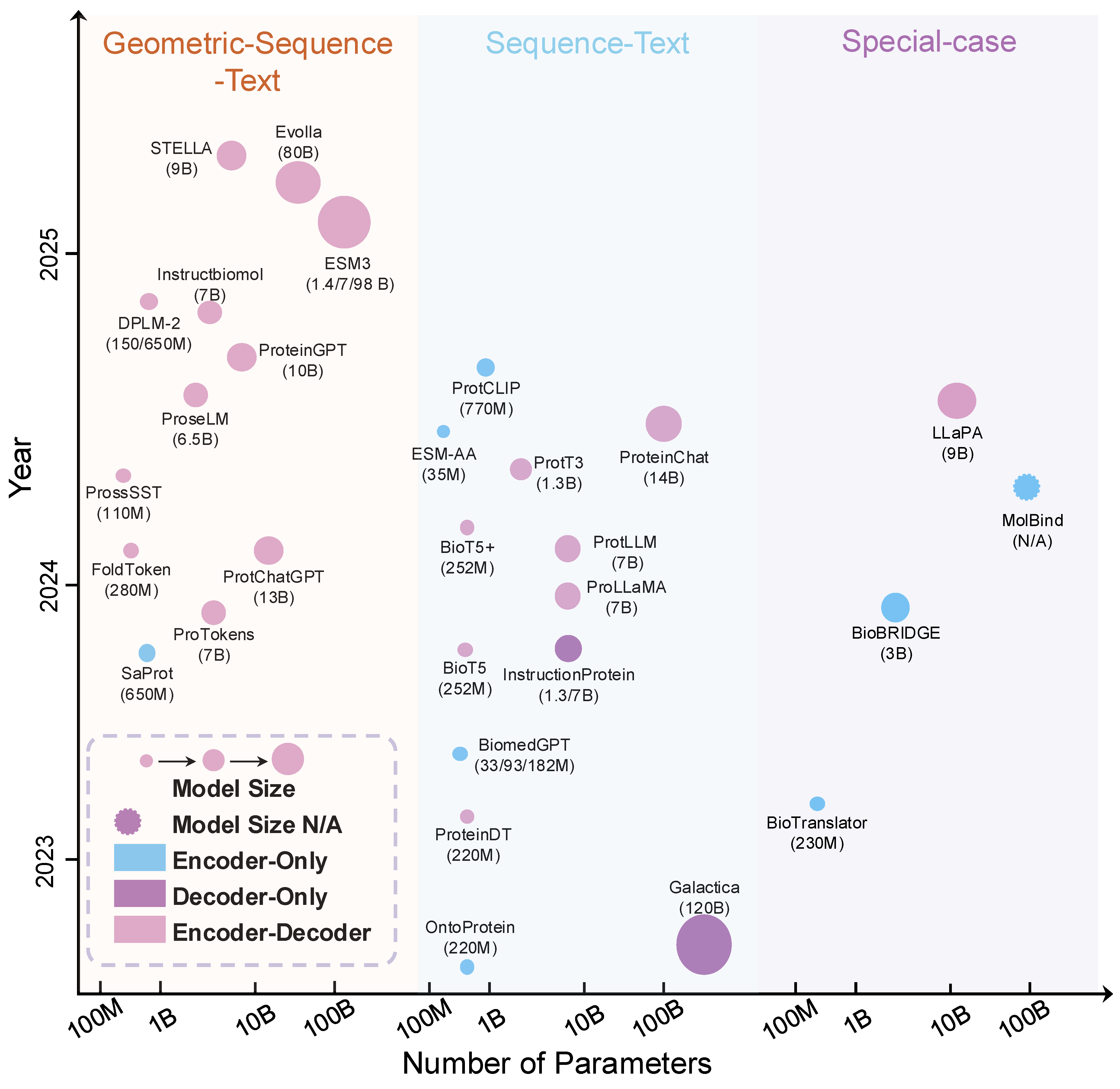
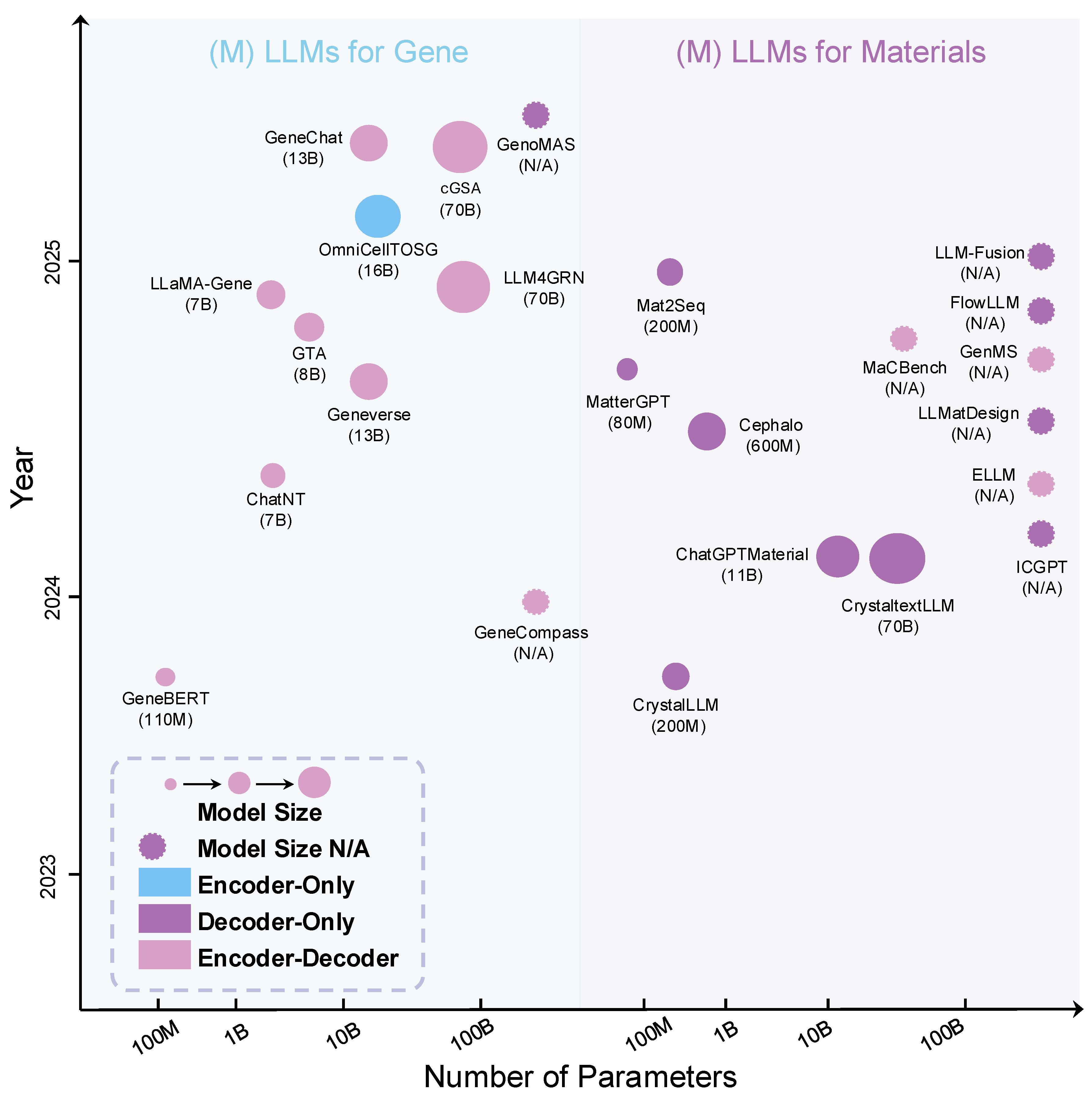
| Method | BACE ↑ 1513 |
BBBP ↑ 2039 |
HIV ↑ 41127 |
|---|---|---|---|
| Specialist Models | |||
| ChemBERTa v2 | 73.5 | 69.8 | 79.3 |
| DMP(TF+GNN) | 89.4 | 77.8 | 81.4 |
| KV-PLM | 78.5 | 70.5 | 71.8 |
| GraphCL | 75.3 | 69.7 | 78.5 |
| GraphMVP-C | 81.2 | 72.4 | 77.0 |
| MoMu | 76.7 | 70.5 | 75.9 |
| MolFM | 83.9 | 72.9 | 78.8 |
| Uni-Mol | 85.7 | 72.9 | 80.8 |
| LLM Based Generalist Models | |||
| Galactica-6.7B | 58.4 | 53.5 | 72.2 |
| Vicuna-v1.5-13b-16k (4-shot) | 49.2 | 52.7 | 50.5 |
| Vicuna-v1.3-7B* | 68.3 | 60.1 | 58.1 |
| LLaMA-2-7B-chat* | 74.8 | 65.6 | 62.3 |
| MolCA(1D) | 79.3 | 70.8 | – |
| MolCA(1D + 2D) | 79.8 | 70.0 | – |
| Instruct-G | 84.3 (±0.6) | 68.6 (±0.3) | 74.0 (±0.1) |
| Instruct-GS | 82.1 (±0.1) | 72.4 (±0.3) | 68.9 (±0.3) |
| MoleculeSTM (Graph) | 80.77 (±1.34) | 69.98 (±0.52) | 76.93 (±1.84) |
| MoleculeSTM (Smiles) | 81.99 (±0.41) | 70.75 (±1.90) | 76.23 (±0.80) |
| Token-Mol (averaged across five runs) | 89.52 (±1.32) | 91.67 (±0.98) | 82.40 (±0.17) |
| Method | Structure | Evolutionary | Engineering | |||
|---|---|---|---|---|---|---|
| SS-Q3↑ | SS-Q8↑ | Contact↑ | Homology↑ | Fluorescence↑ | Stability↑ | |
| LSTM | 0.75 | 0.59 | 0.26 | 0.26 | 0.67 | 0.69 |
| TAPE Transformer | 0.73 | 0.59 | 0.25 | 0.21 | 0.68 | 0.73 |
| ResNet | 0.75 | 0.58 | 0.25 | 0.17 | 0.21 | 0.73 |
| MSA Transformer | - | 0.73 | 0.49 | - | - | - |
| ProtBERT | 0.81 | 0.67 | 0.59 | 0.29 | 0.61 | 0.82 |
| OntoProtein | 0.82 | 0.68 | 0.56 | 0.24 | 0.66 | 0.75 |
| ProteinDT-ProteinCLAP-InfoNCE | 0.8354 | 0.6912 | 0.6011 | 0.3109 | 0.6047 | 0.8110 |
| ProteinDT-ProteinCLAP-EBM-NCE | 0.8310 | 0.6941 | 0.6023 | 0.2865 | 0.6127 | 0.7978 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).





