Submitted:
25 February 2026
Posted:
26 February 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
2.1. Study Design and Patient Cohorts
- CT examinations in which the L3 axial slice did not encompass complete abdominal coverage,
- DICOM studies that could not be exported due to PACS technical issues,
- Absence of abdominal CT (abdominal MRI only), and
- Ineligibility related to the source database (patients who did not undergo a TIPS procedure).
2.2. Image Preprocessing and Reference Standard Generation
2.3. Model Architecture and Training Protocol
2.4. Automated Inference Workflow and User Interface
2.5. Statistical Analysis
3. Results
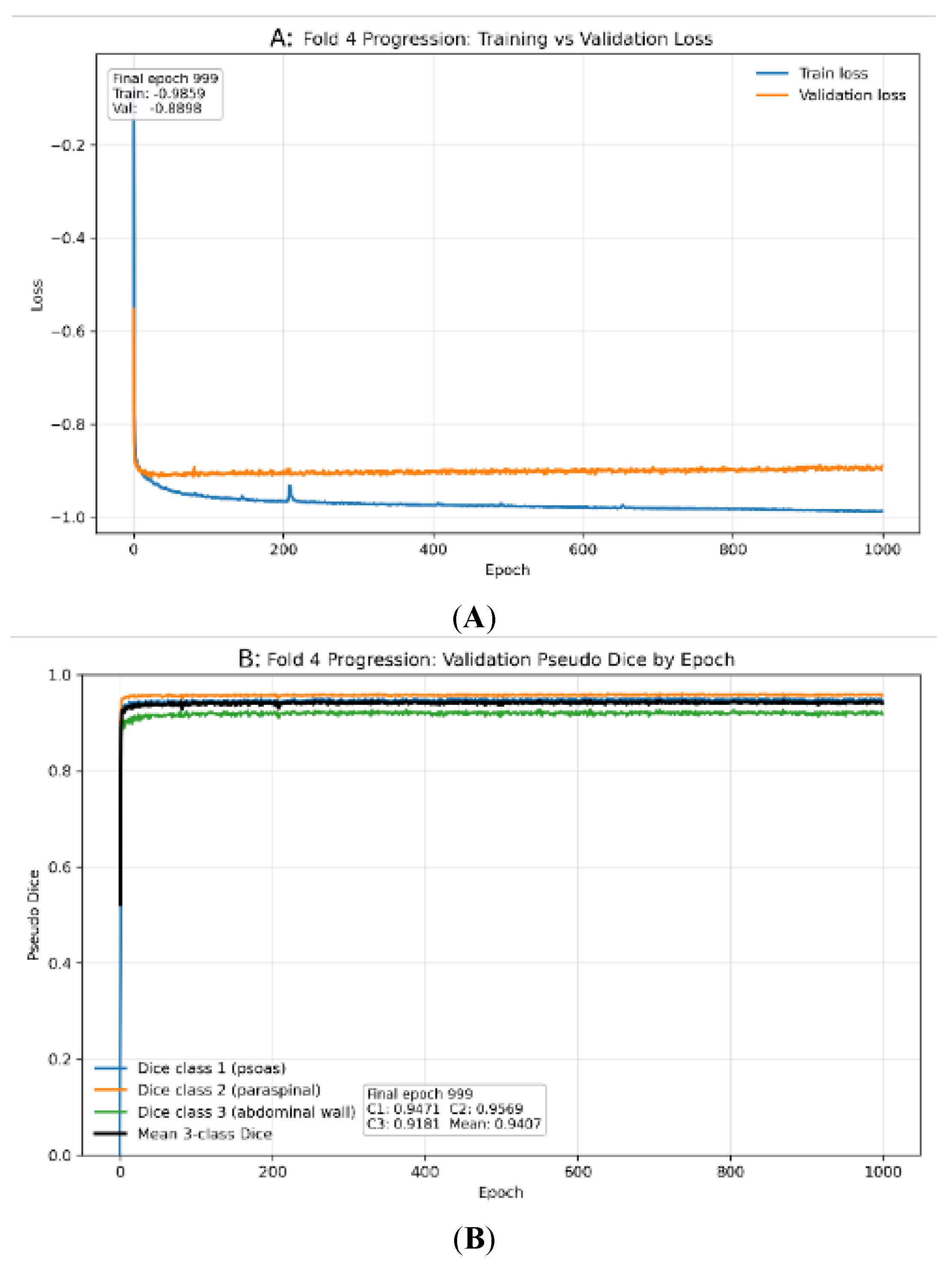
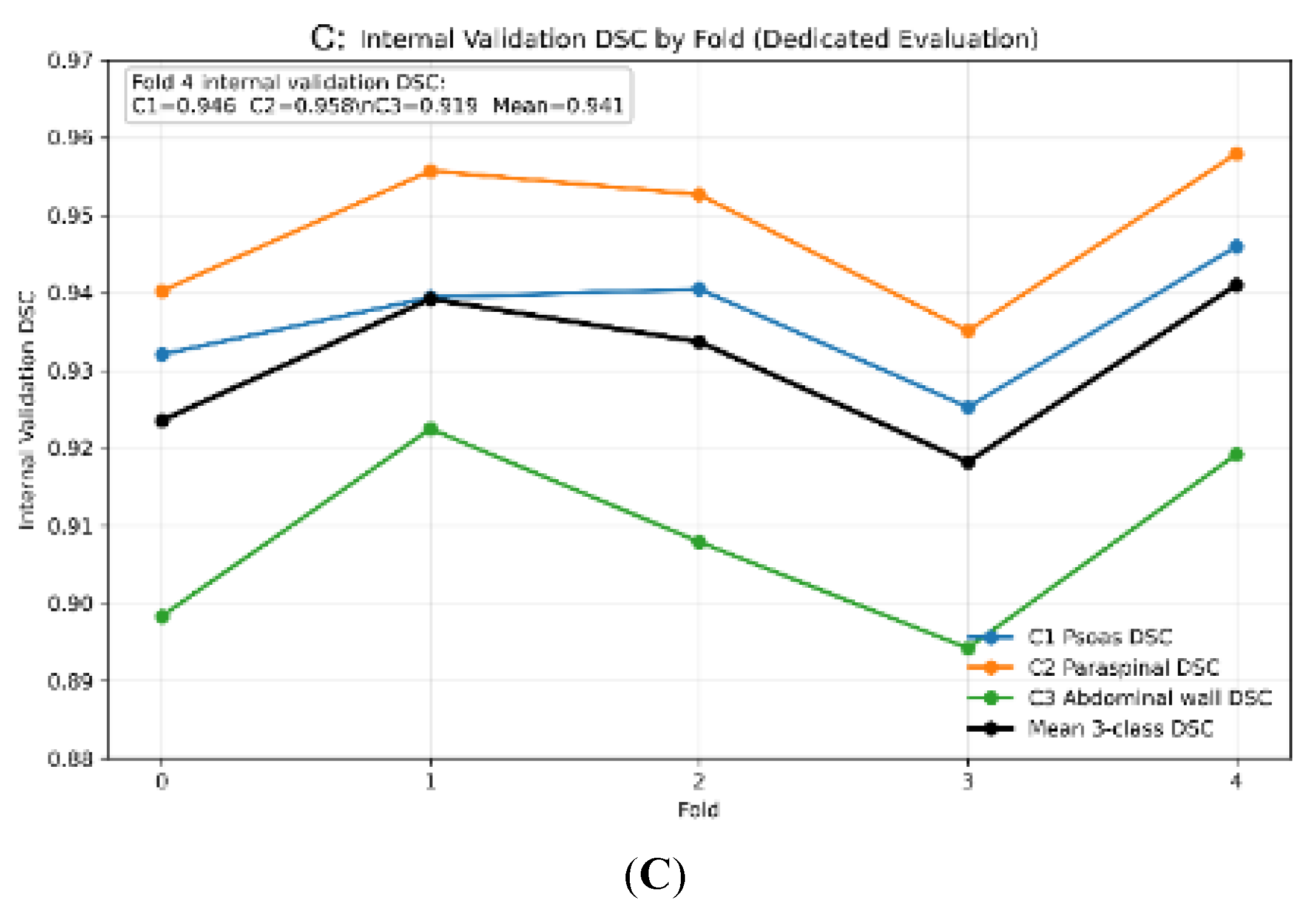
3.1. Internal Cross-Validation Performance
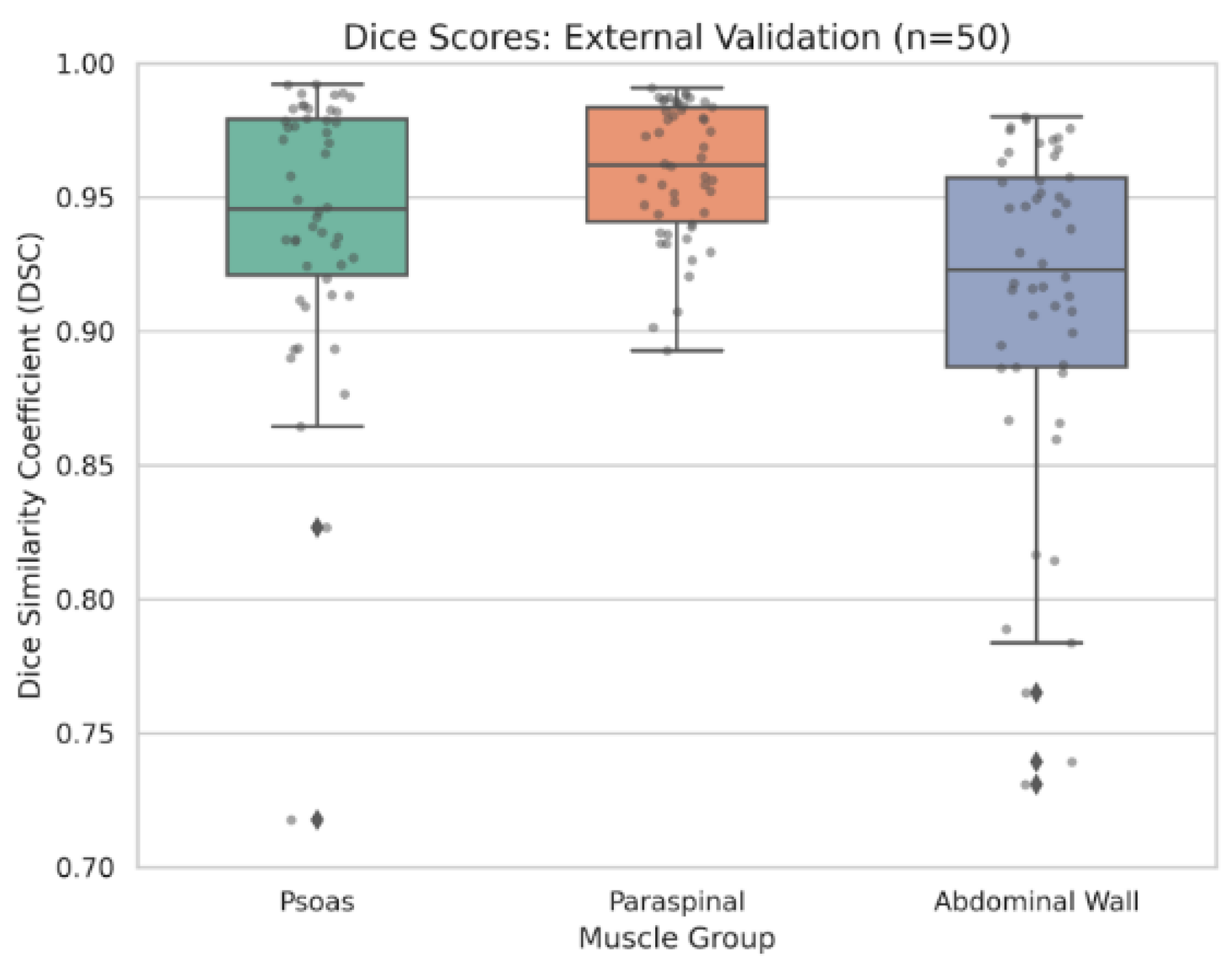
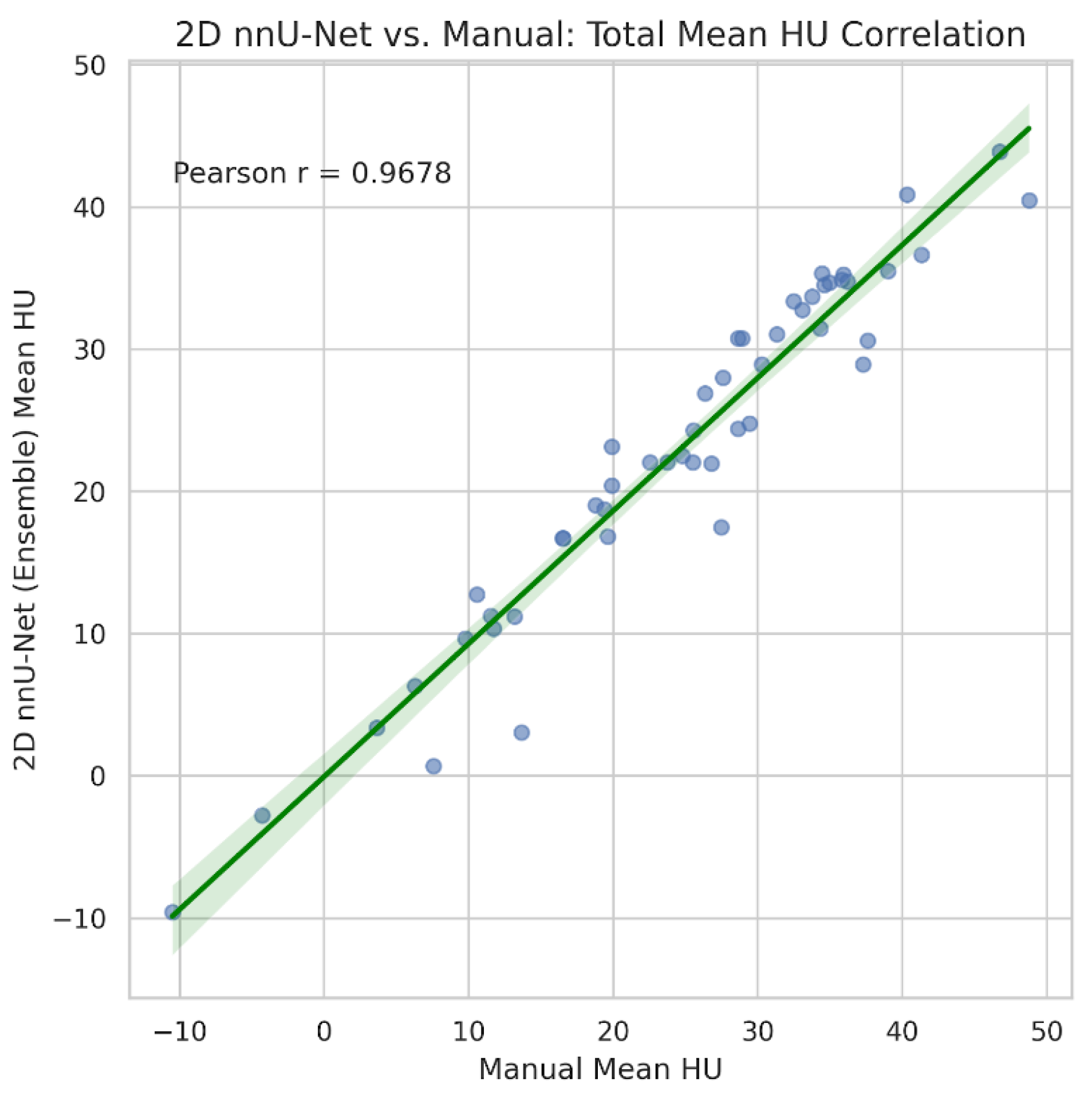
3.2. External Validation and Ensemble Efficacy
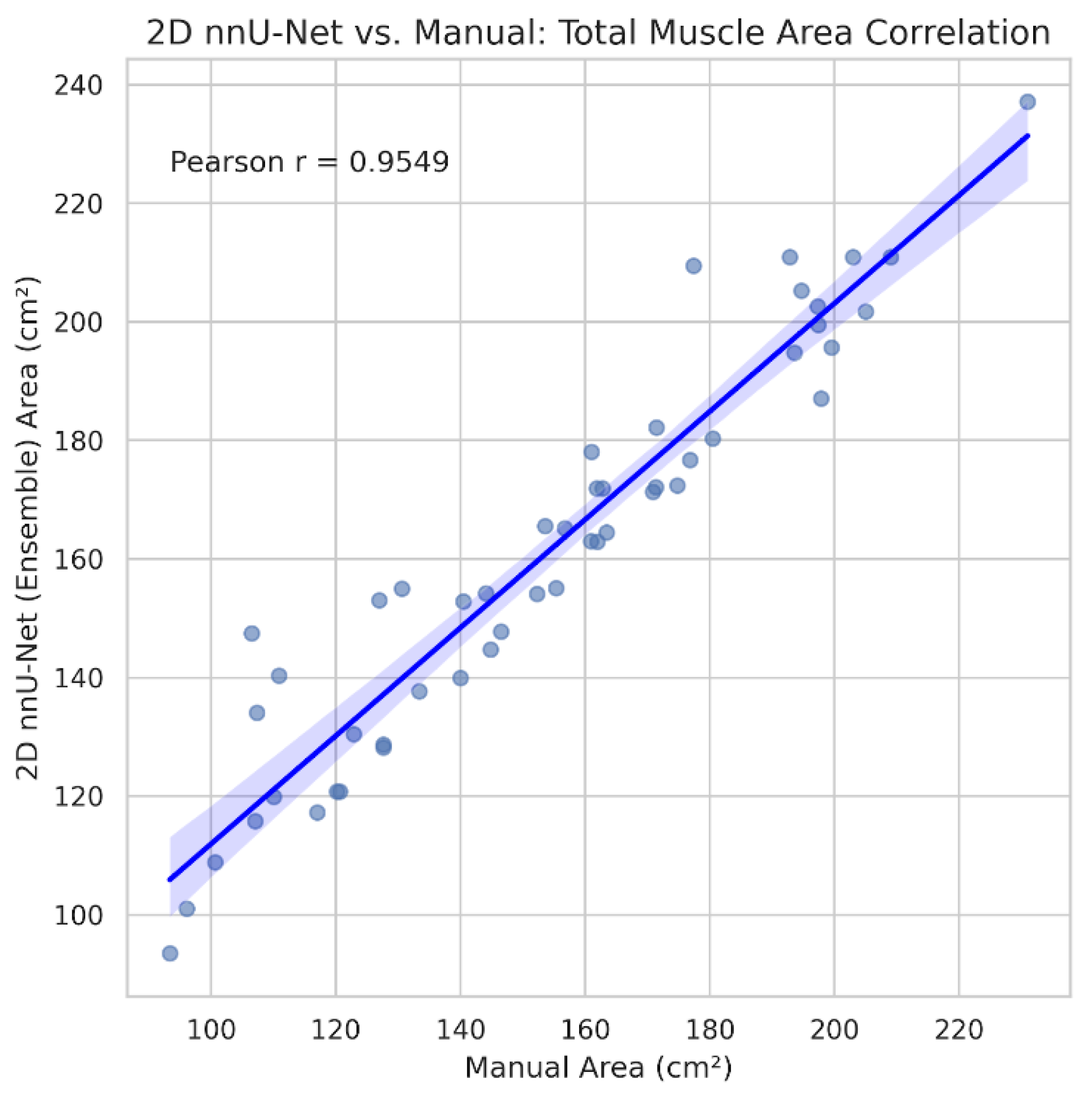
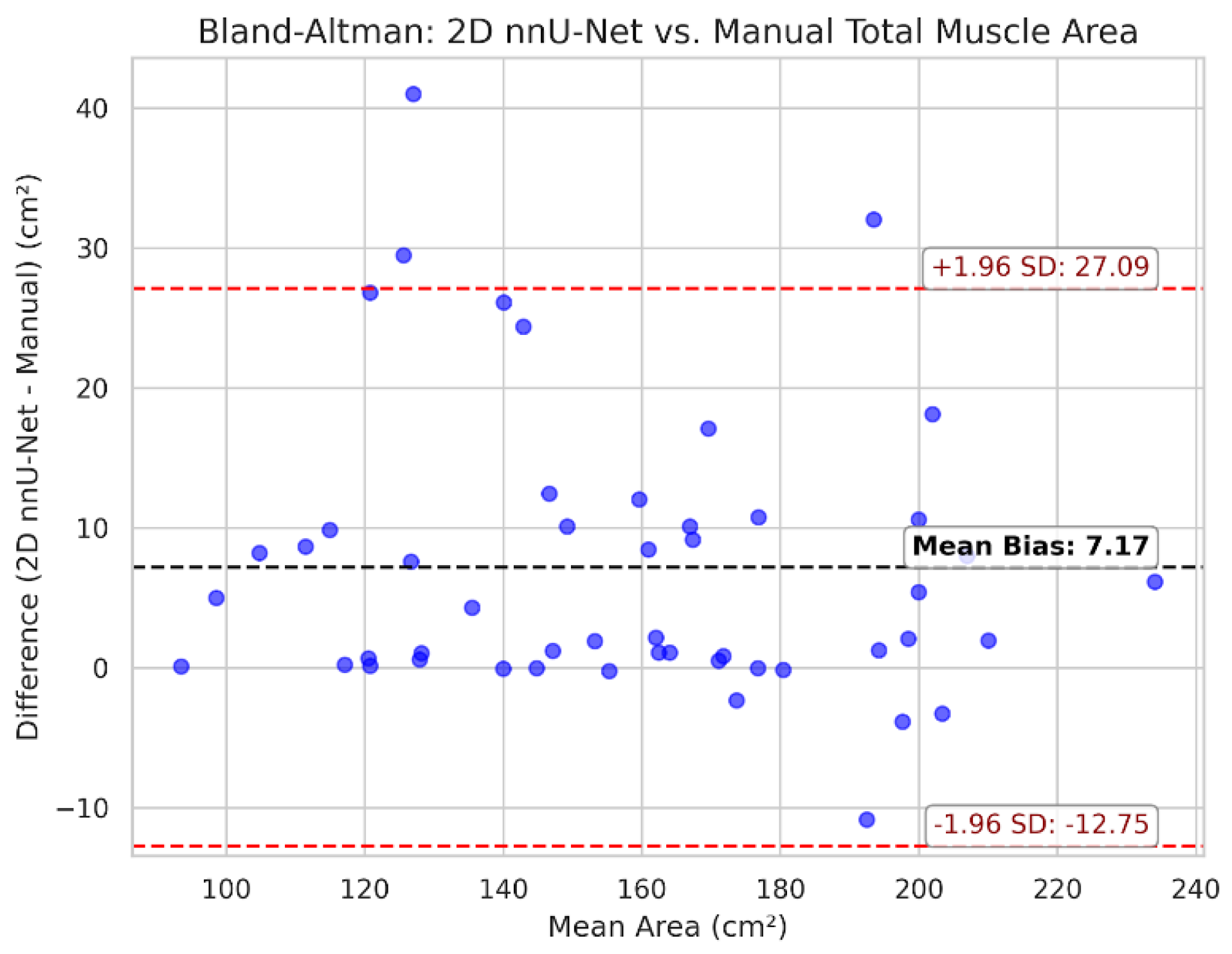
3.3. Clinical Agreement and Bias Analysis
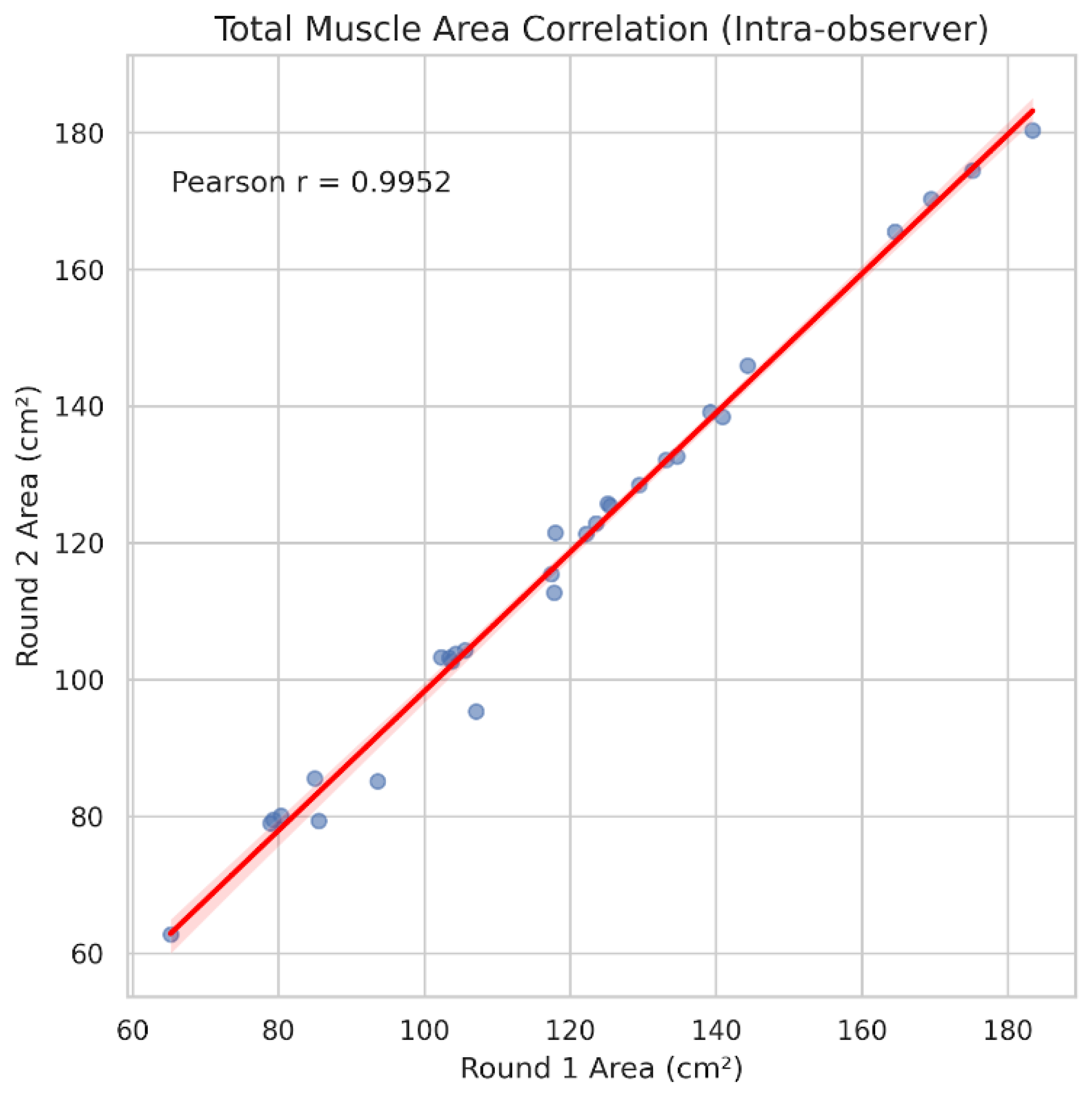
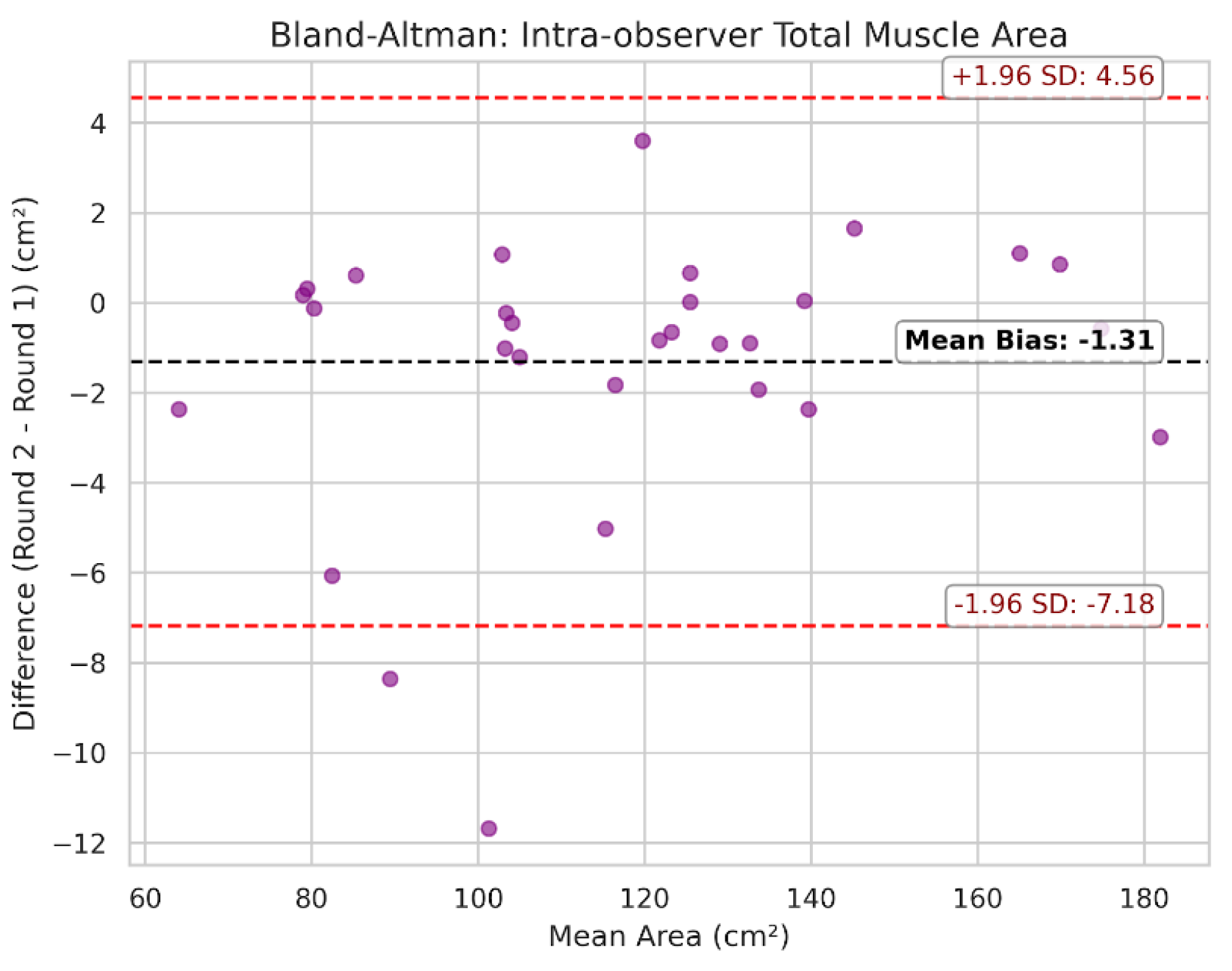
3.4. Intra-Observer Reliability and Time Efficiency
3.5. Qualitative Assessment and Error Analysis
4. Discussion
5. Conclusions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Rosenberg, I.H. Sarcopenia: origins and clinical relevance. J Nutr 1997, 127, 990S–1S. [Google Scholar] [CrossRef] [PubMed]
- Ribeiro, S.M.; Kehayias, J.J. Sarcopenia and the analysis of body composition. Adv Nutr 2014, 5, 260–267. [Google Scholar] [CrossRef] [PubMed]
- Derstine, B.A.; Holcombe, S.A.; Ross, B.E.; Wang, N.C.; Su, G.L.; Wang, S.C. Skeletal muscle cutoff values for sarcopenia diagnosis using T10 to L5 measurements in a healthy US population. Sci Rep 2018, 8, 11369. [Google Scholar] [CrossRef]
- Prado, C.M.; Lieffers, J.R.; McCargar, L.J.; Reiman, T.; Sawyer, M.B.; Martin, L.; Baracos, V.E. Prevalence and clinical implications of sarcopenic obesity in patients with solid tumours of the respiratory and gastrointestinal tracts: a population-based study. Lancet Oncol 2008, 9, 629–635. [Google Scholar] [CrossRef] [PubMed]
- Prado, C.M.; Baracos, V.E.; McCargar, L.J.; Reiman, T.; Mourtzakis, M.; Tonkin, K.; Mackey, J.R.; Koski, S.; Pituskin, E.; Sawyer, M.B. Sarcopenia as a determinant of chemotherapy toxicity and time to tumor progression in metastatic breast cancer patients receiving capecitabine treatment. Clin Cancer Res 2009, 15, 2920–2926. [Google Scholar] [CrossRef]
- Martin, L.; Birdsell, L.; Macdonald, N.; Reiman, T.; Clandinin, M.T.; McCargar, L.J.; Murphy, R.; Ghosh, S.; Sawyer, M.B.; Baracos, V.E. Cancer cachexia in the age of obesity: skeletal muscle depletion is a powerful prognostic factor, independent of body mass index. J Clin Oncol 2013, 31, 1539–1547. [Google Scholar] [CrossRef]
- Prado, C.M.; Heymsfield, S.B. Lean tissue imaging: a new era for nutritional assessment and intervention. JPEN J Parenter Enteral Nutr 2014, 38, 940–953. [Google Scholar] [CrossRef]
- Cruz-Jentoft, A.J.; Baeyens, J.P.; Bauer, J.M.; Boirie, Y.; Cederholm, T.; Landi, F.; Martin, F.C.; Michel, J.P.; Rolland, Y.; Schneider, S.M.; et al. Sarcopenia: European consensus on definition and diagnosis: Report of the European Working Group on Sarcopenia in Older People. Age Ageing 2010, 39, 412–423. [Google Scholar] [CrossRef]
- Wilson, M.M.; Morley, J.E. Invited review: Aging and energy balance. J Appl Physiol (1985) 2003, 95, 1728–1736. [Google Scholar] [CrossRef]
- Bauer, J.M.; Sieber, C.C. Sarcopenia and frailty: a clinician’s controversial point of view. Exp Gerontol 2008, 43, 674–678. [Google Scholar] [CrossRef]
- Baumgartner, R.N.; Koehler, K.M.; Gallagher, D.; Romero, L.; Heymsfield, S.B.; Ross, R.R.; Garry, P.J.; Lindeman, R.D. Epidemiology of sarcopenia among the elderly in New Mexico. Am J Epidemiol 1998, 147, 755–763. [Google Scholar] [CrossRef] [PubMed]
- Goodpaster, B.H.; Carlson, C.L.; Visser, M.; Kelley, D.E.; Scherzinger, A.; Harris, T.B.; Stamm, E.; Newman, A.B. Attenuation of skeletal muscle and strength in the elderly: The Health ABC Study. J Appl Physiol (1985) 2001, 90, 2157–2165. [Google Scholar] [CrossRef]
- Gewandter, J.S.; Dale, W.; Magnuson, A.; Pandya, C.; Heckler, C.E.; Lemelman, T.; Roussel, B.; Ifthikhar, R.; Dolan, J.; Noyes, K.; et al. Associations between a patient-reported outcome (PRO) measure of sarcopenia and falls, functional status, and physical performance in older patients with cancer. J Geriatr Oncol 2015, 6, 433–441. [Google Scholar] [CrossRef]
- Kim, T.N.; Choi, K.M. Sarcopenia: definition, epidemiology, and pathophysiology. J Bone Metab 2013, 20, 1–10. [Google Scholar] [CrossRef]
- Reisinger, K.W.; van Vugt, J.L.; Tegels, J.J.; Snijders, C.; Hulsewe, K.W.; Hoofwijk, A.G.; Stoot, J.H.; Von Meyenfeldt, M.F.; Beets, G.L.; Derikx, J.P.; et al. Functional compromise reflected by sarcopenia, frailty, and nutritional depletion predicts adverse postoperative outcome after colorectal cancer surgery. Ann Surg 2015, 261, 345–352. [Google Scholar] [CrossRef]
- Yip, C.; Dinkel, C.; Mahajan, A.; Siddique, M.; Cook, G.J.; Goh, V. Imaging body composition in cancer patients: visceral obesity, sarcopenia and sarcopenic obesity may impact on clinical outcome. Insights Imaging 2015, 6, 489–497. [Google Scholar] [CrossRef]
- Grotenhuis, B.A.; Shapiro, J.; van Adrichem, S.; de Vries, M.; Koek, M.; Wijnhoven, B.P.; van Lanschot, J.J. Sarcopenia/Muscle Mass is not a Prognostic Factor for Short- and Long-Term Outcome After Esophagectomy for Cancer. World J Surg 2016, 40, 2698–2704. [Google Scholar] [CrossRef]
- Tamandl, D.; Paireder, M.; Asari, R.; Baltzer, P.A.; Schoppmann, S.F.; Ba-Ssalamah, A. Markers of sarcopenia quantified by computed tomography predict adverse long-term outcome in patients with resected oesophageal or gastro-oesophageal junction cancer. Eur Radiol 2016, 26, 1359–1367. [Google Scholar] [CrossRef] [PubMed]
- Mukund, A.; Bhardwaj, V.; Jindal, A.; Patidar, Y.; Sarin, S.K. Transjugular Intrahepatic Portosystemic Shunt Related Hepatic Encephalopathy in Cirrhotics With Refractory Ascites: Incidence and Correlation With TIPS Stent Diameter and Pre-TIPS Sarcopenia. J Clin Exp Hepatol 2026, 16, 103125. [Google Scholar] [CrossRef]
- Zhou, B.; Song, Y.; Chen, C.; Chen, X.; Tao, T. Preoperative Prediction of Sarcopenia in Patients Scheduled for Gastric and Colorectal Cancer Surgery. J Gastrointest Cancer 2025, 56, 82. [Google Scholar] [CrossRef] [PubMed]
- Liao, Y. Sarcopenia with muscle wasting in hepatic cancer predicts therapeutic outcome after hepatic artery intervention. Int J Clin Pharmacol Ther 2025, 63, 70–76. [Google Scholar] [CrossRef]
- Kesby, N.; Chia, P.; Yang, J.; Chapuis, P.H.; Ng, K.S. Sarcopenia kinetics and colorectal cancer outcomes: Post-operative development of sarcopenia is a poor prognostic indicator of survival following colorectal cancer surgery. Colorectal Dis 2025, 27, e70258. [Google Scholar] [CrossRef]
- Kim, T.Y.; Kim, M.Y.; Sohn, J.H.; Kim, S.M.; Ryu, J.A.; Lim, S.; Kim, Y. Sarcopenia as a useful predictor for long-term mortality in cirrhotic patients with ascites. J Korean Med Sci 2014, 29, 1253–1259. [Google Scholar] [CrossRef]
- Taha, A.A.; Hanbury, A. Metrics for evaluating 3D medical image segmentation: analysis, selection, and tool. BMC Med Imaging 2015, 15, 29. [Google Scholar] [CrossRef]
- Hamaguchi, Y.; Kaido, T.; Okumura, S.; Kobayashi, A.; Hammad, A.; Tamai, Y.; Inagaki, N.; Uemoto, S. Proposal for new diagnostic criteria for low skeletal muscle mass based on computed tomography imaging in Asian adults. Nutrition 2016, 32, 1200–1205. [Google Scholar] [CrossRef]
- van der Werf, A.; Langius, J.A.E.; de van der Schueren, M.A.E.; Nurmohamed, S.A.; van der Pant, K.; Blauwhoff-Buskermolen, S.; Wierdsma, NJ. Percentiles for skeletal muscle index, area and radiation attenuation based on computed tomography imaging in a healthy Caucasian population. Eur J Clin Nutr 2018, 72, 288–296. [Google Scholar] [CrossRef] [PubMed]
- Mitsiopoulos, N.; Baumgartner, R.N.; Heymsfield, S.B.; Lyons, W.; Gallagher, D.; Ross, R. Cadaver validation of skeletal muscle measurement by magnetic resonance imaging and computerized tomography. J Appl Physiol (1985) 1998, 85, 115–122. [Google Scholar] [CrossRef]
- Mourtzakis, M.; Prado, C.M.; Lieffers, J.R.; Reiman, T.; McCargar, L.J.; Baracos, V.E. A practical and precise approach to quantification of body composition in cancer patients using computed tomography images acquired during routine care. Appl Physiol Nutr Metab 2008, 33, 997–1006. [Google Scholar] [CrossRef] [PubMed]
- Goodpaster, B.H.; Kelley, D.E.; Thaete, F.L.; He, J.; Ross, R. Skeletal muscle attenuation determined by computed tomography is associated with skeletal muscle lipid content. J Appl Physiol (1985) 2000, 89, 104–110. [Google Scholar] [CrossRef] [PubMed]
- Portal, D.; Hofstetter, L.; Eshed, I.; Dan-Lantsman, C.; Sella, T.; Urban, D.; Onn, A.; Bar, J.; Segal, G. L3 skeletal muscle index (L3SMI) is a surrogate marker of sarcopenia and frailty in non-small cell lung cancer patients. Cancer Manag Res 2019, 11, 2579–2588. [Google Scholar] [CrossRef]
- Ji, W.; Liu, X.; Zhang, Y.; Zhao, Y.; He, Y.; Cui, J.; Li, W. Development of Formulas for Calculating L3 Skeletal Muscle Mass Index and Visceral Fat Area Based on Anthropometric Parameters. Front Nutr 2022, 9, 910771. [Google Scholar] [CrossRef]
- Derstine, B.A.; Holcombe, S.A.; Goulson, R.L.; Ross, B.E.; Wang, N.C.; Sullivan, J.A.; Su, G.L.; Wang, S.C. Quantifying Sarcopenia Reference Values Using Lumbar and Thoracic Muscle Areas in a Healthy Population. J Nutr Health Aging 2017, 21, 180–185. [Google Scholar] [CrossRef]
- van der Werf, A.; Dekker, I.M.; Meijerink, M.R.; Wierdsma, N.J.; de van der Schueren, M.A.E.; Langius, JAE. Skeletal muscle analyses: agreement between non-contrast and contrast CT scan measurements of skeletal muscle area and mean muscle attenuation. Clin Physiol Funct Imaging 2018, 38, 366–372. [Google Scholar] [CrossRef] [PubMed]
- Rollins, K.E.; Gopinath, A.; Awwad, A.; Macdonald, I.A.; Lobo, D.N. Computed tomography-based psoas skeletal muscle area and radiodensity are poor sentinels for whole L3 skeletal muscle values. Clin Nutr 2020, 39, 2227–2232. [Google Scholar] [CrossRef]
- Onishi, S.; Kuwahara, T.; Tajika, M.; Tanaka, T.; Yamada, K.; Shimizu, M.; Niwa, Y.; Yamaguchi, R. Artificial intelligence for body composition assessment focusing on sarcopenia. Sci Rep 2025, 15, 1324. [Google Scholar] [CrossRef]
- Gomez-Perez, S.L.; Haus, J.M.; Sheean, P.; Patel, B.; Mar, W.; Chaudhry, V.; McKeever, L.; Braunschweig, C. Measuring Abdominal Circumference and Skeletal Muscle From a Single Cross-Sectional Computed Tomography Image: A Step-by-Step Guide for Clinicians Using National Institutes of Health ImageJ. JPEN J Parenter Enteral Nutr 2016, 40, 308–318. [Google Scholar] [CrossRef]
- Irving, B.A.; Weltman, J.Y.; Brock, D.W.; Davis, C.K.; Gaesser, G.A.; Weltman, A. NIH ImageJ and Slice-O-Matic computed tomography imaging software to quantify soft tissue. Obesity (Silver Spring) 2007, 15, 370–376. [Google Scholar] [CrossRef]
- Steele, S.; Lin, F.; Le, T.L.; Medline, A.; Higgins, M.; Sandberg, A.; Evans, S.; Hong, G.; Williams, M.A.; Bilen, M.A.; et al. Segmentation and Linear Measurement for Body Composition Analysis using Slice-O-Matic and Horos. J Vis Exp 2021. [Google Scholar] [CrossRef]
- Weston, A.D.; Korfiatis, P.; Kline, T.L.; Philbrick, K.A.; Kostandy, P.; Sakinis, T.; Sugimoto, M.; Takahashi, N.; Erickson, B.J. Automated Abdominal Segmentation of CT Scans for Body Composition Analysis Using Deep Learning. Radiology 2019, 290, 669–679. [Google Scholar] [CrossRef] [PubMed]
- Ackermans, L.; Volmer, L.; Wee, L.; Brecheisen, R.; Sanchez-Gonzalez, P.; Seiffert, A.P.; Gomez, E.J.; Dekker, A.; Ten Bosch, J.A.; Olde Damink, S.M.W.; et al. Deep Learning Automated Segmentation for Muscle and Adipose Tissue from Abdominal Computed Tomography in Polytrauma Patients. Sensors (Basel) 2021, 21. [Google Scholar] [CrossRef] [PubMed]
- Lin, J.; Zhang, W.; Chen, W.; Huang, Y.; Wu, R.; Chen, X.; Shen, X.; Zhu, G. Muscle Mass, Density, and Strength Are Necessary to Diagnose Sarcopenia in Patients With Gastric Cancer. J Surg Res 2019, 241, 141–148. [Google Scholar] [CrossRef]
- Woodward, A.J.; Wallen, M.P.; Ryan, J.; Ward, L.C.; Coombes, J.S.; Macdonald, G.A. Evaluation of techniques used to assess skeletal muscle quantity in patients with cirrhosis. Clin Nutr ESPEN 2021, 44, 287–296. [Google Scholar] [CrossRef]
- Kikuchi, N.; Uojima, H.; Hidaka, H.; Iwasaki, S.; Wada, N.; Kubota, K.; Nakazawa, T.; Shibuya, A.; Kako, M.; Take, A.; et al. Evaluation of Skeletal Muscle Mass in Patients with Chronic Liver Disease Shows Different Results Based on Bioelectric Impedance Analysis and Computed Tomography. Ann Nutr Metab 2022, 78, 336–344. [Google Scholar] [CrossRef]
- Ying, T.; Borrelli, P.; Edenbrandt, L.; Enqvist, O.; Kaboteh, R.; Tragardh, E.; Ulen, J.; Kjolhede, H. AI-based fully automatic image analysis: Optimal abdominal and thoracic segmentation volumes for estimating total muscle volume on computed tomography scans. Osteoporos Sarcopenia 2024, 10, 78–83. [Google Scholar] [CrossRef] [PubMed]
- van Dijk, D.P.J.; Volmer, L.F.; Brecheisen, R.; Martens, B.; Dolan, R.D.; Bryce, A.S.; Chang, D.K.; McMillan, D.C.; Stoot, J.; West, M.A.; et al. External validation of a deep learning model for automatic segmentation of skeletal muscle and adipose tissue on abdominal CT images. Br J Radiol 2024, 97, 2015–2023. [Google Scholar] [CrossRef] [PubMed]
- Lee, Y.S.; Hong, N.; Witanto, J.N.; Choi, Y.R.; Park, J.; Decazes, P.; Eude, F.; Kim, C.O.; Chang Kim, H.; Goo, J.M.; et al. Deep neural network for automatic volumetric segmentation of whole-body CT images for body composition assessment. Clin Nutr 2021, 40, 5038–5046. [Google Scholar] [CrossRef] [PubMed]
- Nachit, M.; Horsmans, Y.; Summers, R.M.; Leclercq, I.A.; Pickhardt, P.J. AI-based CT Body Composition Identifies Myosteatosis as Key Mortality Predictor in Asymptomatic Adults. Radiology 2023, 307, e222008. [Google Scholar] [CrossRef]
- Koitka, S.; Kroll, L.; Malamutmann, E.; Oezcelik, A.; Nensa, F. Fully automated body composition analysis in routine CT imaging using 3D semantic segmentation convolutional neural networks. Eur Radiol 2021, 31, 1795–1804. [Google Scholar] [CrossRef]
- Isensee, F.; Jaeger, P.F.; Kohl, S.A.A.; Petersen, J.; Maier-Hein, K.H. nnU-Net: a self-configuring method for deep learning-based biomedical image segmentation. Nat Methods 2021, 18, 203–211. [Google Scholar] [CrossRef]
- Antonelli, M.; Reinke, A.; Bakas, S.; Farahani, K.; Kopp-Schneider, A.; Landman, B.A.; Litjens, G.; Menze, B.; Ronneberger, O.; Summers, R.M.; et al. The Medical Segmentation Decathlon. Nat Commun 2022, 13, 4128. [Google Scholar] [CrossRef]
- Zhang, G.; Yang, Z.; Huo, B.; Chai, S.; Jiang, S. Multiorgan segmentation from partially labeled datasets with conditional nnU-Net. Comput Biol Med 2021, 136, 104658. [Google Scholar] [CrossRef]
- Wasserthal, J.; Breit, H.C.; Meyer, M.T.; Pradella, M.; Hinck, D.; Sauter, A.W.; Heye, T.; Boll, D.T.; Cyriac, J.; Yang, S.; et al. TotalSegmentator: Robust Segmentation of 104 Anatomic Structures in CT Images. Radiol Artif Intell 2023, 5, e230024. [Google Scholar] [CrossRef]
- Hesamian, M.H.; Jia, W.; He, X.; Kennedy, P. Deep Learning Techniques for Medical Image Segmentation: Achievements and Challenges. J Digit Imaging 2019, 32, 582–596. [Google Scholar] [CrossRef]
- Kelly, C.J.; Karthikesalingam, A.; Suleyman, M.; Corrado, G.; King, D. Key challenges for delivering clinical impact with artificial intelligence. BMC Med 2019, 17, 195. [Google Scholar] [CrossRef]
- Bae, K.T. Intravenous contrast medium administration and scan timing at CT: considerations and approaches. Radiology 2010, 256, 32–61. [Google Scholar] [CrossRef] [PubMed]
- Lell, M.M.; Wildberger, J.E.; Alkadhi, H.; Damilakis, J.; Kachelriess, M. Evolution in Computed Tomography: The Battle for Speed and Dose. Invest Radiol 2015, 50, 629–644. [Google Scholar] [CrossRef]
- Willemink, M.J.; Koszek, W.A.; Hardell, C.; Wu, J.; Fleischmann, D.; Harvey, H.; Folio, L.R.; Summers, R.M.; Rubin, D.L.; Lungren, M.P. Preparing Medical Imaging Data for Machine Learning. Radiology 2020, 295, 4–15. [Google Scholar] [CrossRef]
- Bedrikovetski, S.; Seow, W.; Kroon, H.M.; Traeger, L.; Moore, J.W.; Sammour, T. Artificial intelligence for body composition and sarcopenia evaluation on computed tomography: A systematic review and meta-analysis. Eur J Radiol 2022, 149, 110218. [Google Scholar] [CrossRef]
- Zech, J.R.; Badgeley, M.A.; Liu, M.; Costa, A.B.; Titano, J.J.; Oermann, E.K. Variable generalization performance of a deep learning model to detect pneumonia in chest radiographs: A cross-sectional study. PLoS Med 2018, 15, e1002683. [Google Scholar] [CrossRef] [PubMed]
- Ha, J.; Park, T.; Kim, H.K.; Shin, Y.; Ko, Y.; Kim, D.W.; Sung, Y.S.; Lee, J.; Ham, S.J.; Khang, S.; et al. Development of a fully automatic deep learning system for L3 selection and body composition assessment on computed tomography. Sci Rep 2021, 11, 21656. [Google Scholar] [CrossRef] [PubMed]
- Finlayson, S.G.; Subbaswamy, A.; Singh, K.; Bowers, J.; Kupke, A.; Zittrain, J.; Kohane, I.S.; Saria, S. The Clinician and Dataset Shift in Artificial Intelligence. N Engl J Med 2021, 385, 283–286. [Google Scholar] [CrossRef] [PubMed]
- Borrelli, P.; Kaboteh, R.; Enqvist, O.; Ulen, J.; Tragardh, E.; Kjolhede, H.; Edenbrandt, L. Artificial intelligence-aided CT segmentation for body composition analysis: a validation study. Eur Radiol Exp 2021, 5, 11. [Google Scholar] [CrossRef] [PubMed]
- Greco, F.; Mallio, C.A. Artificial intelligence and abdominal adipose tissue analysis: a literature review. Quant Imaging Med Surg 2021, 11, 4461–4474. [Google Scholar] [CrossRef] [PubMed]
- Magudia, K.; Bridge, C.P.; Bay, C.P.; Babic, A.; Fintelmann, F.J.; Troschel, F.M.; Miskin, N.; Wrobel, W.C.; Brais, L.K.; Andriole, K.P.; et al. Population-Scale CT-based Body Composition Analysis of a Large Outpatient Population Using Deep Learning to Derive Age-, Sex-, and Race-specific Reference Curves. Radiology 2021, 298, 319–329. [Google Scholar] [CrossRef]
- Urooj, B.; Ko, Y.; Na, S.; Kim, I.O.; Lee, E.H.; Cho, S.; Jeong, H.; Khang, S.; Lee, J.; Kim, K.W. Implementation of Fully Automated AI-Integrated System for Body Composition Assessment on Computed Tomography for Opportunistic Sarcopenia Screening: Multicenter Prospective Study. JMIR Form Res 2025, 9, e69940. [Google Scholar] [CrossRef]
| Characteristic | Value |
|---|---|
| Total Cases (n) | 50 |
| Age (years) | 63 ± 15 |
| Sex (Male : Female) | 18 : 32 |
| Primary Clinical Indications (n, %) | |
| Liver cirrhosis | 11 (22%) |
| Routine evaluation for TIPS | 10 (20%) |
| Refractory ascites | 9 (18%) |
| Hepatocellular carcinoma | 8 (16%) |
| Portosystemic shunt | 6 (12%) |
| PV thrombosis evaluation | 5 (10%) |
| Hepatic hydrothorax | 3 (6%) |
| Axial CT Findings at L3 | |
| Ascites | 44 (88%) |
| Subcutaneous edema | 3 (6%) |
| Liver masses | 1 (2%) |
| No ascites | 6 (12%) |
| Fold | Psoas | Paraspinal | Abdominal wall | Mean DSC |
|---|---|---|---|---|
| 0 | 0.932 | 0.940 | 0.898 | 0.924 |
| 1 | 0.939 | 0.956 | 0.923 | 0.939 |
| 2 | 0.941 | 0.952 | 0.908 | 0.934 |
| 3 | 0.925 | 0.935 | 0.894 | 0.918 |
| 4 | 0.946 | 0.958 | 0.919 | 0.941 |
| Mean | 0.937 | 0.948 | 0.908 | 0.931 ± 0.010 |
| Metric | Single fold (Fold 4) | 5-fold ensemble | P-value |
|---|---|---|---|
| Overall mean DSC | 0.937 | 0.937 | 0.736 |
| Psoas DSC | 0.939 | 0.941 | 0.448 |
| Paraspinal DSC | 0.960 | 0.960 | 0.810 |
| Abdominal wall DSC | 0.912 | 0.911 | 0.858 |
| Minimum observed DSC | 0.720 | 0.822 | — |
| Metric | Pearson r | MAE | Mean bias (nnU-Net - manual) |
|---|---|---|---|
| Total muscle area (cm²) | 0.955 | 8.00 | +7.17 |
| Total mean attenuation (HU) | 0.968 | 2.33 | -1.67 |
| Metric | Pearson r | MAE | Mean bias (round 2 - round 1) |
|---|---|---|---|
| Total muscle area (cm²) | 0.995 | 1.98 | -1.31 |
| Total mean attenuation (HU) | 0.995 | 0.74 | +0.56 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).





