Submitted:
18 February 2026
Posted:
26 February 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
| Gene name | Primer forward 5’-3’ | Primer reverse 5’-3’ |
| Atg9a | GCGTTGCTGGTCACTCTATG | CAGGATAGAGCCAGCGAAGA |
| Atg16l1 | TCGCCTCAATGCAGAGAATG | GAGGTTCCTTTGCTGCTTCT |
| Atp6v0c | CAGAGCTGATCATGAAGTCCATC | GAAAACTCCTGTAGATGATG |
| Atp6v1d | GGATTGAACGCACCCTTG | GCTGCTCTCCGCCGCTC |
| Bdnf | AAGTCTGCATTACATTCCTCGA | GTTTTCTGAAAGAGCGACAGTT |
| C-fos | AGAGCGCCCCATCCTTAC | GCAGCCATCTTATTCCGTTC |
| Clcn3 | CCTGGCTGCTGATGTTATGA | TGAGGGCAAATCCCACTAAT |
| Ctsa | TTCTGATCCAGCCAGATGGTG | TACAGCACGTTGGCAATCAGG |
| Ctsd | CGTCCTTTGACATCCACTACGG | TGGAACCGATACAGTGTCCTGG |
| Cyclophilin | GGCAAATGCTGGACCAAACACAA | GTAAAATGCCCGCAAGTCAAAAG |
| Egr1 | CCTTCAATCCTCAAGGGAGC | AACCGAGTCGTTTGGCTGGGA |
| Foxo1 | GTGGATGGTGAAGAGCGTGCCC | GCTGTGAAGGGACAGATTGTGGCG |
| Foxo3 | AAGGGCGACAGCAACAGCTCTG | CATGAAGCGGCTGTGCAGGGAC |
| Gabarapl1 | CAAAGAGGAGCATCCGTTCGAG | TTGTCCAGGTCTCCTATCCGAG |
| Glb1 | AAATGGCTGGCAGTCCTTCTG | ACCTGCACGGTTATGATCGGT |
| Hexa | CTACATCCAGACGCTGCTGGAC | TACTGGCATTTCTTCCCGCCAC |
| Hif1-α | GCTGAAGACACAGAGGCAAA | TACTTGGAGGGCTTGGAGAA |
| Lamp1 | CCTACGAGACTGCGAATGGT | CCACAAGAACTGCCATTTTTC |
| Map1lc3b | GCTTGCAGCTCAATGCTAAC | CCTGCGAGGCATAAACCATGTA |
| Mcol1 | GCGCCTATGACACCATCAA | TATCCTGGCACTGCTCGAT |
| Naglu | TCCAACAGCACGAGTTTGAG | CTGCGATGGCTAATCTGTCA |
| Pgc1-α | GAATCAAGCCACTACAGACACGC | CATCCCTCTTGAGCCTTTCGTG |
| Slc38a9 | TTGAAAGCGAGGGAAATGATGGTC | ATGGGAATGAGGGTCACTGAGAAG |
| Sqstm1 | GAAGCTGCCCTATACCCACA | TGGGAGAGGGACTCAATCAG |
| Tfe3 | AGGATCAAAGAGCTGGGCAC | CCGGCTCTCCAGGTCTTTG |
| Tfeb | GTCATTGACAACATTATGCGCC | GCGTGTTAGGCATCTTGCATCT |
| Tpp1 | CCCCTCATGTGGATTTTGTGG | TGGTTCTGGACGTTGTCTTGG |
| Ulk1 | TGGAGACCTGGCTGACTACCTGC | TGGATGATGCCCTTGCTGTGCA |
3. Results
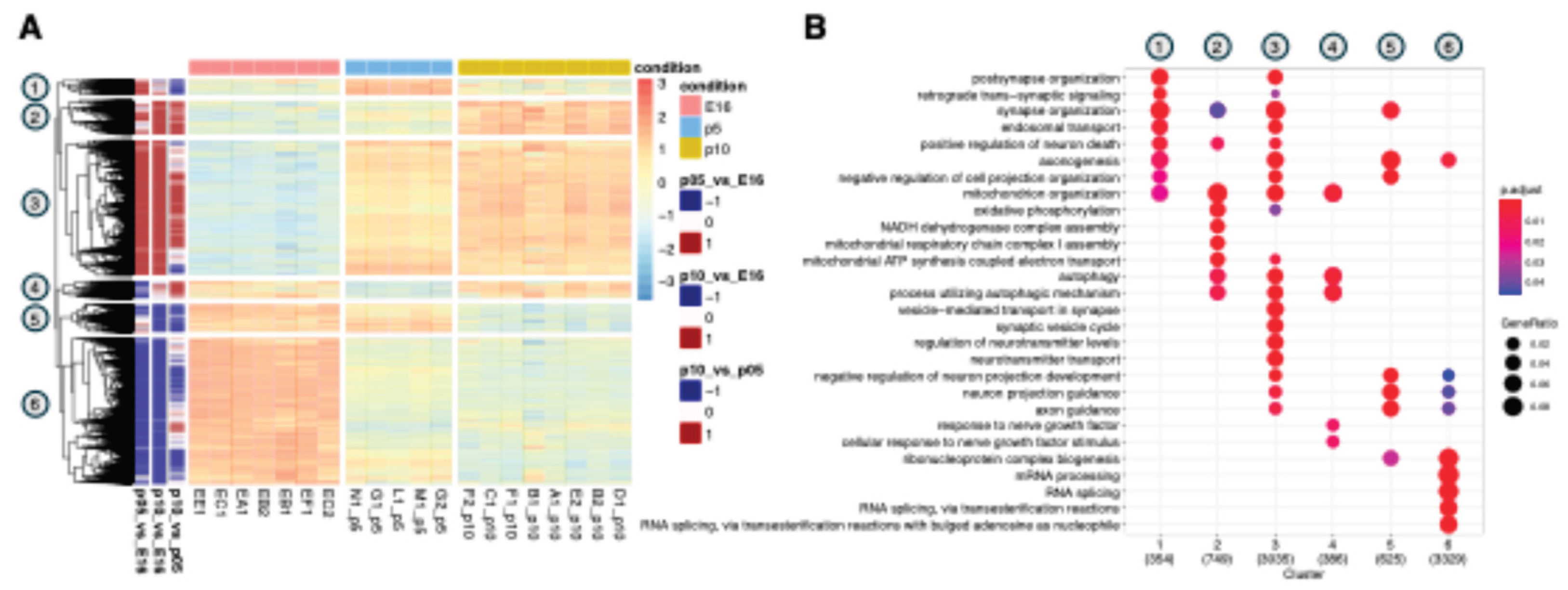
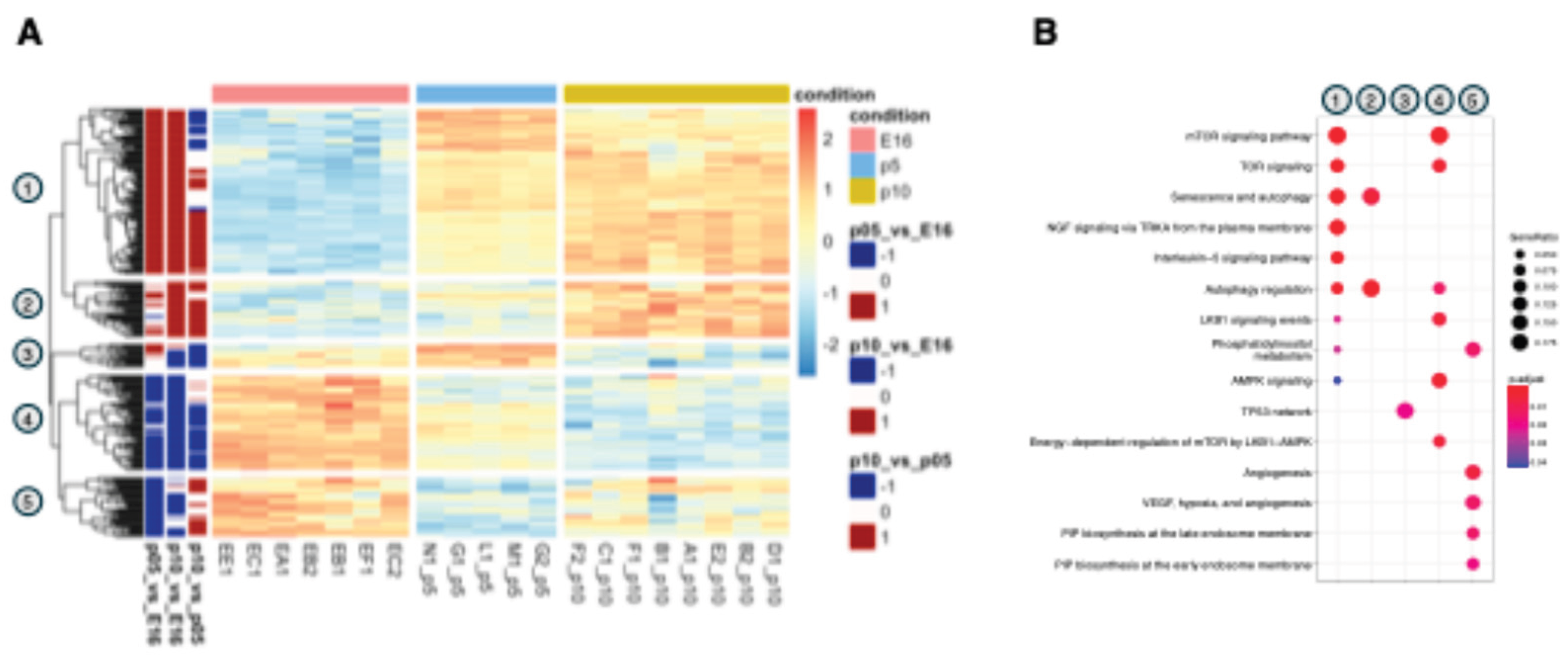
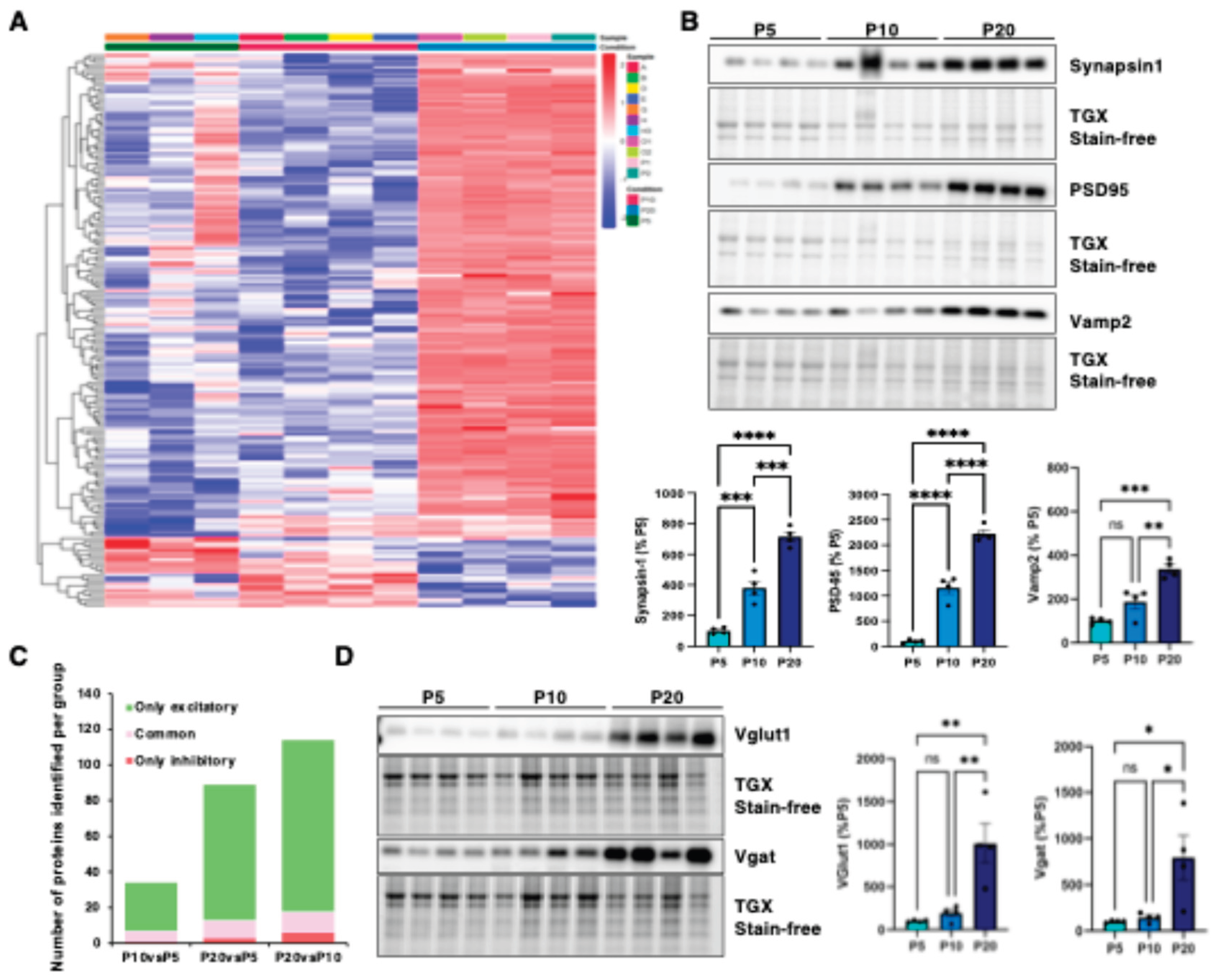
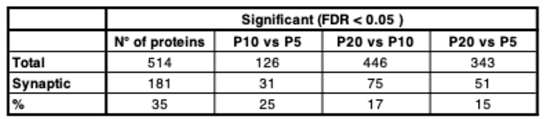
3.1. Identification of Distinct Differential Expression Patterns and Functional Gene Clusters in the Developing Cortex
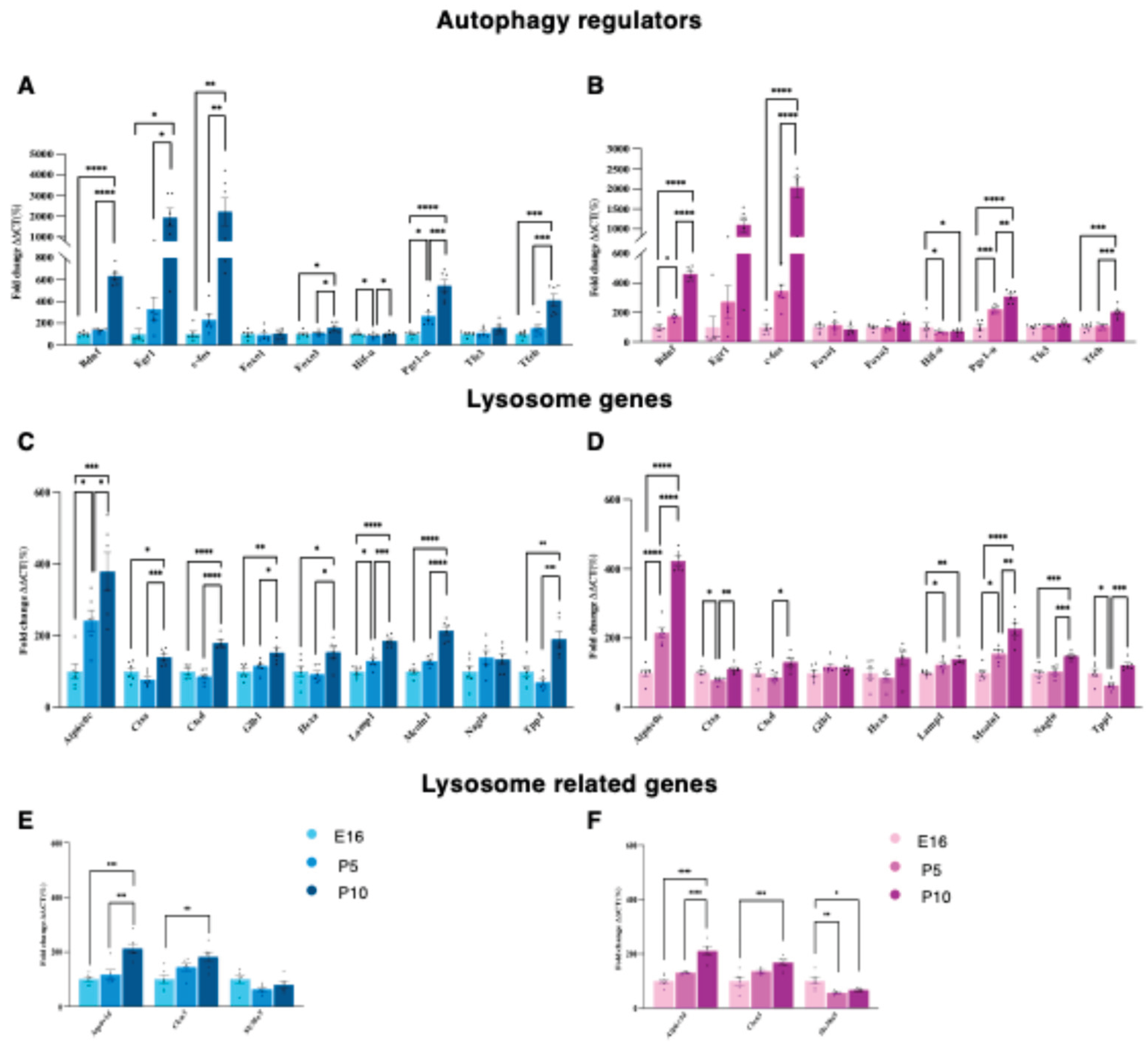
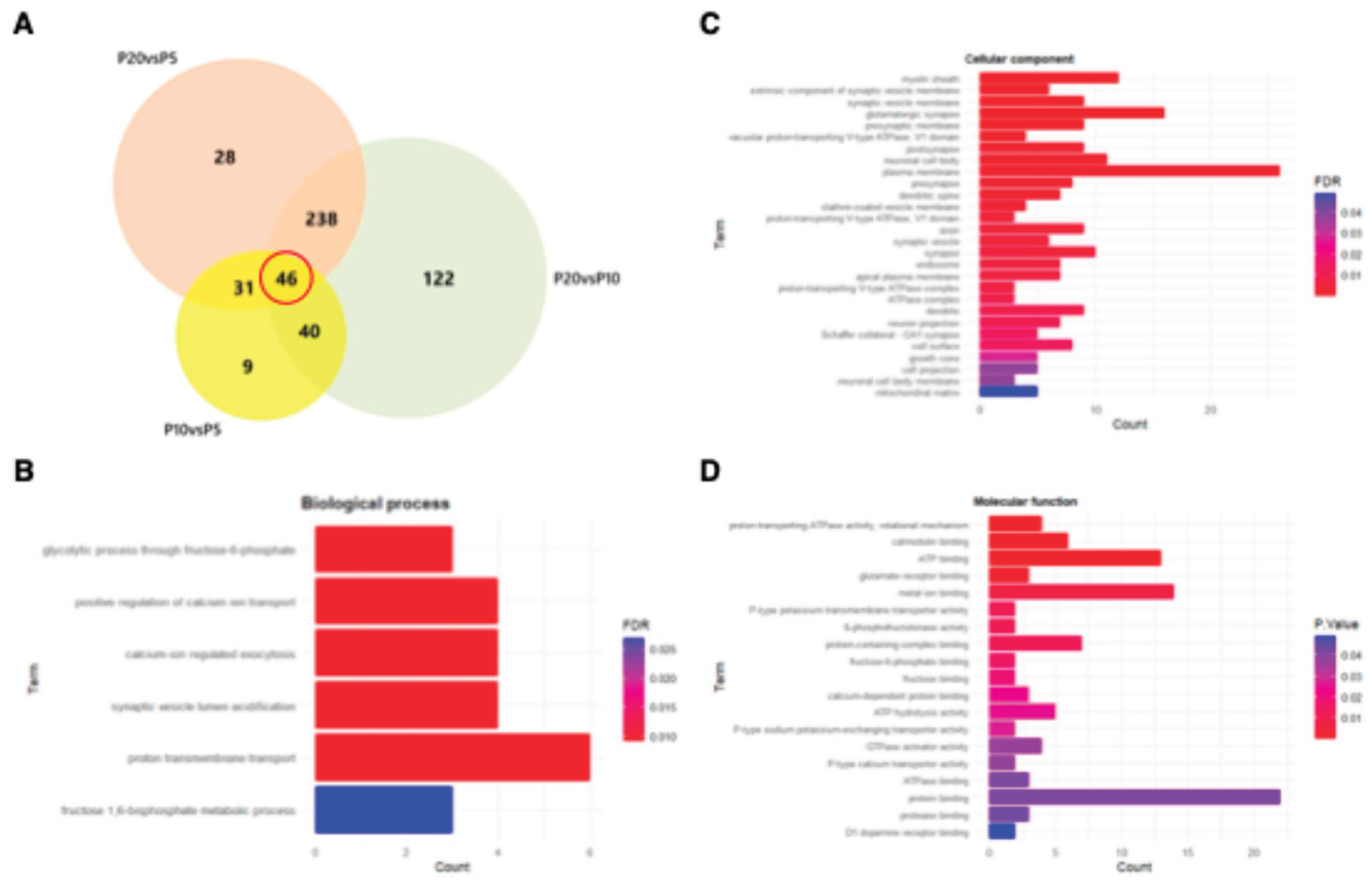
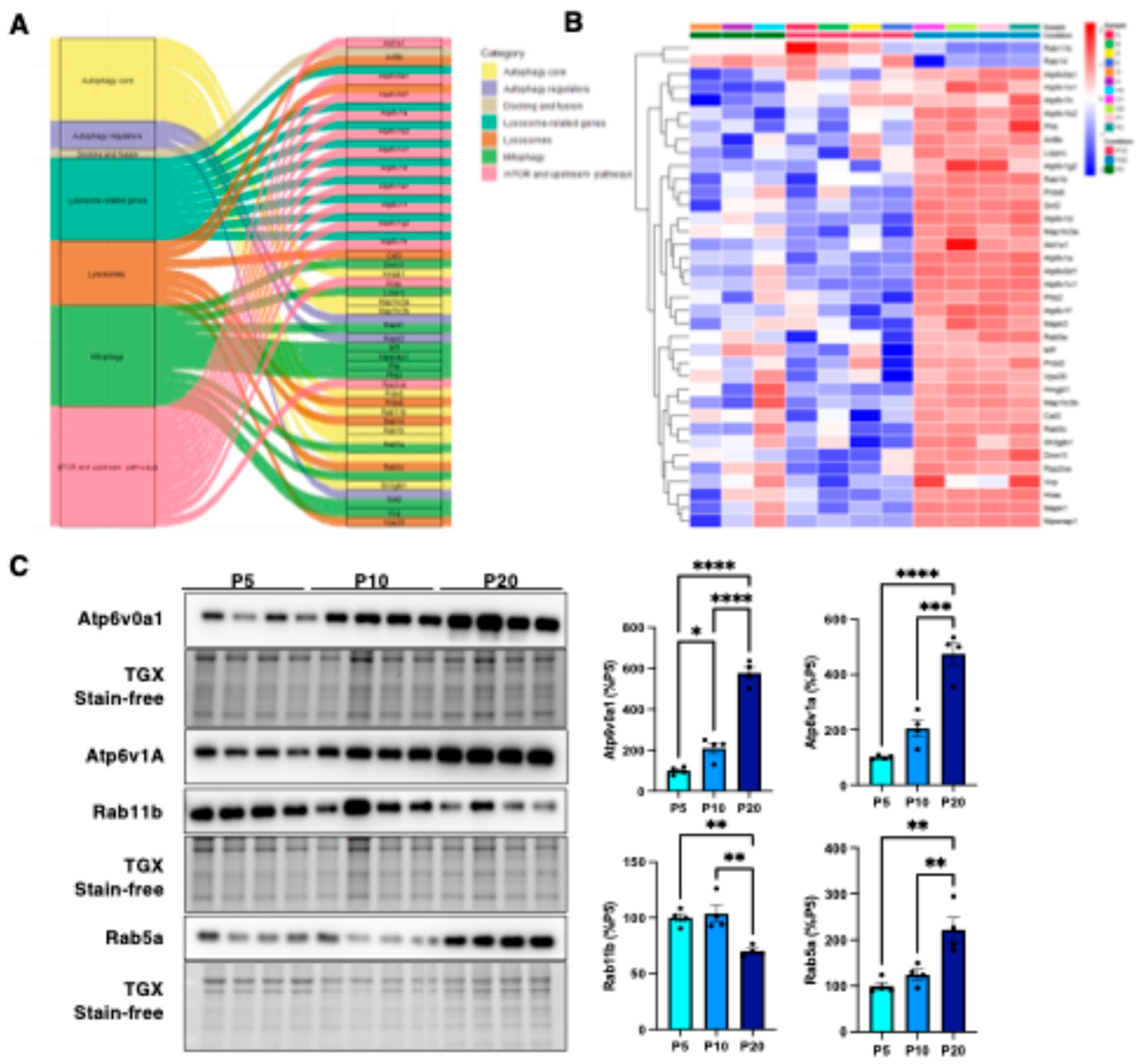
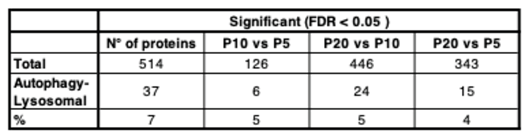
3.2. Mapping the Correlation Between Autophagy and Synaptic Architecture in the Postnatal Cortex.
4. Discussion
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| mTOR | (mechanistic target of rapamycin) |
| AMPK | (AMP-activated protein kinase) |
| ULK1 | (Unc-51 Like Autophagy Activating Kinase 1) |
| LKB1 | (Liver Kinase B1) |
| SEM | (Standard Error of the Mean) |
| NGF | (Nerve Growth Factor) |
| TRKA | (Tyrosine kinase A) |
References
- Dikic, I.; Elazar, Z. Mechanism and medical implications of mammalian autophagy. Nat. Rev. Mol. Cell Biol. 2018, 19, 349–364. [Google Scholar] [CrossRef]
- Stavoe, A.K.H.; Holzbaur, E.L.F. Autophagy in Neurons. Annu. Rev. Cell Dev. Biol. 2019, 35, 477–500. [Google Scholar] [CrossRef]
- Palmer, J.E.; Wilson, N.; Son, S.M.; Obrocki, P.; Wrobel, L.; Rob, M.; Takla, M.; Korolchuk, V.I.; Rubinsztein, D.C. Autophagy, aging, and age-related neurodegeneration. Neuron 2025, 113, 29–48. [Google Scholar] [CrossRef]
- Ventruti, A.; Cuervo, A.M. Autophagy and neurodegeneration. Curr. Neurol. Neurosci. Rep. 2007, 7, 443–451. [Google Scholar] [CrossRef]
- Son, J.H.; Shim, J.H.; Kim, K.H.; Ha, J.Y.; Han, J.Y. Neuronal autophagy and neurodegenerative diseases. Exp. Mol. Med. 2012, 44, 89–98. [Google Scholar] [CrossRef] [PubMed]
- Xi, Y.; Dhaliwal, J.S.; Ceizar, M.; Vaculik, M.; Kumar, K.L.; Lagace, D.C. Knockout of Atg5 delays the maturation and reduces the survival of adult-generated neurons in the hippocampus. Cell Death Dis. 2016, 7, e2127. [Google Scholar] [CrossRef]
- Coupé, B.; Ishii, Y.; Dietrich, M.O.; Komatsu, M.; Horvath, T.L.; Bouret, S.G. Loss of autophagy in pro-opiomelanocortin neurons perturbs axon growth and causes metabolic dysregulation. Cell Metab. 2012, 15, 247–255. [Google Scholar] [CrossRef] [PubMed]
- Nikoletopoulou, V.; Sidiropoulou, K.; Kallergi, E.; Dalezios, Y.; Tavernarakis, N. Modulation of autophagy by BDNF underlies synaptic plasticity. Cell Metab. 2017, 26, 230–242.e5. [Google Scholar] [CrossRef] [PubMed]
- Torres, C.A.; Sulzer, D. Macroautophagy can press a brake on presynaptic neurotransmission. Autophagy 2012, 8, 1540–1541. [Google Scholar] [CrossRef]
- Tang, G.; Gudsnuk, K.; Kuo, S.-H.; Cotrina, M.L.; Rosoklija, G.; Sosunov, A.; Sonders, M.S.; Kanter, E.; Castagna, C.; Yamamoto, A.; Yue, Z.; Arancio, O.; Peterson, B.S.; Champagne, F.; Dwork, A.J.; Goldman, J.; Sulzer, D. Loss of mTOR-dependent macroautophagy causes autistic-like synaptic pruning deficits. Neuron 2014, 83, 1131–1143. [Google Scholar] [CrossRef]
- Ebrahimi-Fakhari, D.; Saffari, A.; Wahlster, L.; Lu, J.; Byrne, S.; Hoffmann, G.F.; Jungbluth, H.; Sahin, M. Congenital disorders of autophagy: an emerging novel class of inborn errors of neuro-metabolism. Brain 2016, 139, 317–337. [Google Scholar] [CrossRef] [PubMed]
- Fassio, A.; Falace, A.; Esposito, A.; Aprile, D.; Guerrini, R.; Benfenati, F. Emerging role of the autophagy/lysosomal degradative pathway in neurodevelopmental disorders with epilepsy. Front. Cell. Neurosci. 2020, 14, 39. [Google Scholar] [CrossRef]
- Esposito, A.; Seri, T.; Breccia, M.; Indrigo, M.; De Rocco, G.; Nuzzolillo, F.; Denti, V.; Pappacena, F.; Tartaglione, G.; Serrao, S.; Paglia, G.; Murru, L.; de Pretis, S.; Cioni, J.-M.; Landsberger, N.; Guarnieri, F.C.; Palmieri, M. Unraveling autophagic imbalances and therapeutic insights in Mecp2-deficient models. EMBO Mol. Med. 2024, 16, 2795–2826. [Google Scholar] [CrossRef] [PubMed]
- Zhu, Y.; Runwal, G.; Obrocki, P.; Rubinsztein, D.C. Autophagy in childhood neurological disorders. Dev. Med. Child Neurol. 2019, 61, 639–645. [Google Scholar] [CrossRef]
- Palmieri, M.; Impey, S.; Kang, H.; di Ronza, A.; Pelz, C.; Sardiello, M.; Ballabio, A. Characterization of the CLEAR network reveals an integrated control of cellular clearance pathways. Hum. Mol. Genet. 2011, 20, 3852–3866. [Google Scholar] [CrossRef]
- Sardiello, M.; Palmieri, M.; di Ronza, A.; Medina, D.L.; Valenza, M.; Gennarino, V.A.; Di Malta, C.; Donaudy, F.; Embrione, V.; Polishchuk, R.S.; Banfi, S.; Parenti, G.; Cattaneo, E.; Ballabio, A. A gene network regulating lysosomal biogenesis and function. Science 2009, 325, 473–477. [Google Scholar] [CrossRef]
- Raben, N.; Puertollano, R. TFEB and TFE3: linking lysosomes to cellular adaptation to stress. Annu. Rev. Cell Dev. Biol. 2016, 32, 255–278. [Google Scholar] [CrossRef]
- Bordi, M.; De Cegli, R.; Testa, B.; Nixon, R.A.; Ballabio, A.; Cecconi, F. A gene toolbox for monitoring autophagy transcription. Cell Death Dis. 2021, 12, 1044. [Google Scholar] [CrossRef]
- Picone, P.; Porcelli, G.; Bavisotto, C.C.; Nuzzo, D.; Galizzi, G.; Biagio, P.L.S.; Bulone, D.; Di Carlo, M. Synaptosomes: new vesicles for neuronal mitochondrial transplantation. J. Nanobiotechnology 2021, 19, 6. [Google Scholar] [CrossRef] [PubMed]
- Plum, S.; Eggers, B.; Helling, S.; Stepath, M.; Theiss, C.; Leite, R.E.P.; Molina, M.; Grinberg, L.T.; Riederer, P.; Gerlach, M.; May, C.; Marcus, K. Proteomic Characterization of Synaptosomes from Human Substantia Nigra Indicates Altered Mitochondrial Translation in Parkinson’s Disease. Cells 2020, 9. [Google Scholar] [CrossRef]
- Gonzalez-Lozano, M.A.; Klemmer, P.; Gebuis, T.; Hassan, C.; van Nierop, P.; van Kesteren, R.E.; Smit, A.B.; Li, K.W. Dynamics of the mouse brain cortical synaptic proteome during postnatal brain development. Sci. Rep. 2016, 6, 35456. [Google Scholar] [CrossRef]
- van Oostrum, M.; Blok, T.M.; Giandomenico, S.L.; Tom Dieck, S.; Tushev, G.; Fürst, N.; Langer, J.D.; Schuman, E.M. The proteomic landscape of synaptic diversity across brain regions and cell types. Cell 2023, 186, 5411–5427.e23. [Google Scholar] [CrossRef]
- Dahlhaus, M.; Li, K.W.; van der Schors, R.C.; Saiepour, M.H.; van Nierop, P.; Heimel, J.A.; Hermans, J.M.; Loos, M.; Smit, A.B.; Levelt, C.N. The synaptic proteome during development and plasticity of the mouse visual cortex. Mol. Cell. Proteomics 2011, 10, M110.005413. [Google Scholar] [CrossRef]
- Tau, G.Z.; Peterson, B.S. Normal development of brain circuits. Neuropsychopharmacology 2010, 35, 147–168. [Google Scholar] [CrossRef]
- Zhang, Z.; Jiao, Y.Y.; Sun, Q.Q. Developmental maturation of excitation and inhibition balance in principal neurons across four layers of somatosensory cortex. Neuroscience 2011, 174, 10–25. [Google Scholar] [CrossRef] [PubMed]
- Chen, J.L.; Villa, K.L.; Cha, J.W.; So, P.T.C.; Kubota, Y.; Nedivi, E. Clustered dynamics of inhibitory synapses and dendritic spines in the adult neocortex. Neuron 2012, 74, 361–373. [Google Scholar] [CrossRef] [PubMed]
- Gowrisankaran, S.; Milosevic, I. Regulation of synaptic vesicle acidification at the neuronal synapse. IUBMB Life 2020, 72, 568–576. [Google Scholar] [CrossRef] [PubMed]
- Maestro, I.; Boya, P.; Martinez, A. Serum- and glucocorticoid-induced kinase 1, a new therapeutic target for autophagy modulation in chronic diseases. Expert Opin. Ther. Targets 2020, 24, 231–243. [Google Scholar] [CrossRef]
- Galluzzi, L.; Pietrocola, F.; Levine, B.; Kroemer, G. Metabolic control of autophagy. Cell 2014, 159, 1263–1276. [Google Scholar] [CrossRef]
- Bolger, A.M.; Lohse, M.; Usadel, B. Trimmomatic: A flexible trimmer for Illumina sequence data. Bioinformatics 2014, 30, 2114–2120. [Google Scholar] [CrossRef]
- Dobin, A.; Davis, C.A.; Schlesinger, F.; Drenkow, J.; Zaleski, C.; Jha, S.; Batut, P.; Chaisson, M.; Gingeras, T.R. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 2013, 29, 15–21. [Google Scholar] [CrossRef]
- Liao, Y.; Smyth, G.K.; Shi, W. featureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 2014, 30, 923–930. [Google Scholar] [CrossRef]
- Love, M.I.; Huber, W.; Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 2014, 15, 550. [Google Scholar] [CrossRef] [PubMed]
- Xu, S.; Hu, E.; Cai, Y.; Xie, Z.; Luo, X.; Zhan, L.; Tang, W.; Wang, Q.; Liu, B.; Wang, R.; Xie, W.; Wu, T.; Xie, L.; Yu, G. Using clusterProfiler to characterize multiomics data. Nat. Protoc. 2024, 19, 3292–3320. [Google Scholar] [CrossRef] [PubMed]
- Wiśniewski, J.R.; Zougman, A.; Nagaraj, N.; Mann, M. Universal sample preparation method for proteome analysis. Nat. Methods 2009, 6, 359–362. [Google Scholar] [CrossRef]
- Cox, J.; Neuhauser, N.; Michalski, A.; Scheltema, R.A.; Olsen, J.V.; Mann, M. Andromeda: a peptide search engine integrated into the MaxQuant environment. J. Proteome Res. 2011, 10, 1794–1805. [Google Scholar] [CrossRef] [PubMed]
- Sherman, B.T.; Hao, M.; Qiu, J.; Jiao, X.; Baseler, M.W.; Lane, H.C.; Imamichi, T.; Chang, W. DAVID: a web server for functional enrichment analysis and functional annotation of gene lists (2021 update). Nucleic Acids Res. 2022, 50, W216–W221. [Google Scholar] [CrossRef]
 |
 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




