Submitted:
15 February 2026
Posted:
27 February 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
Sequence Alignment and Chimeric Construct Design
Structural Prediction and Refinement
Molecular Dynamics (MD) Simulations
Trajectory Analysis
3. Results
Predicted Structural Features and Model Validation
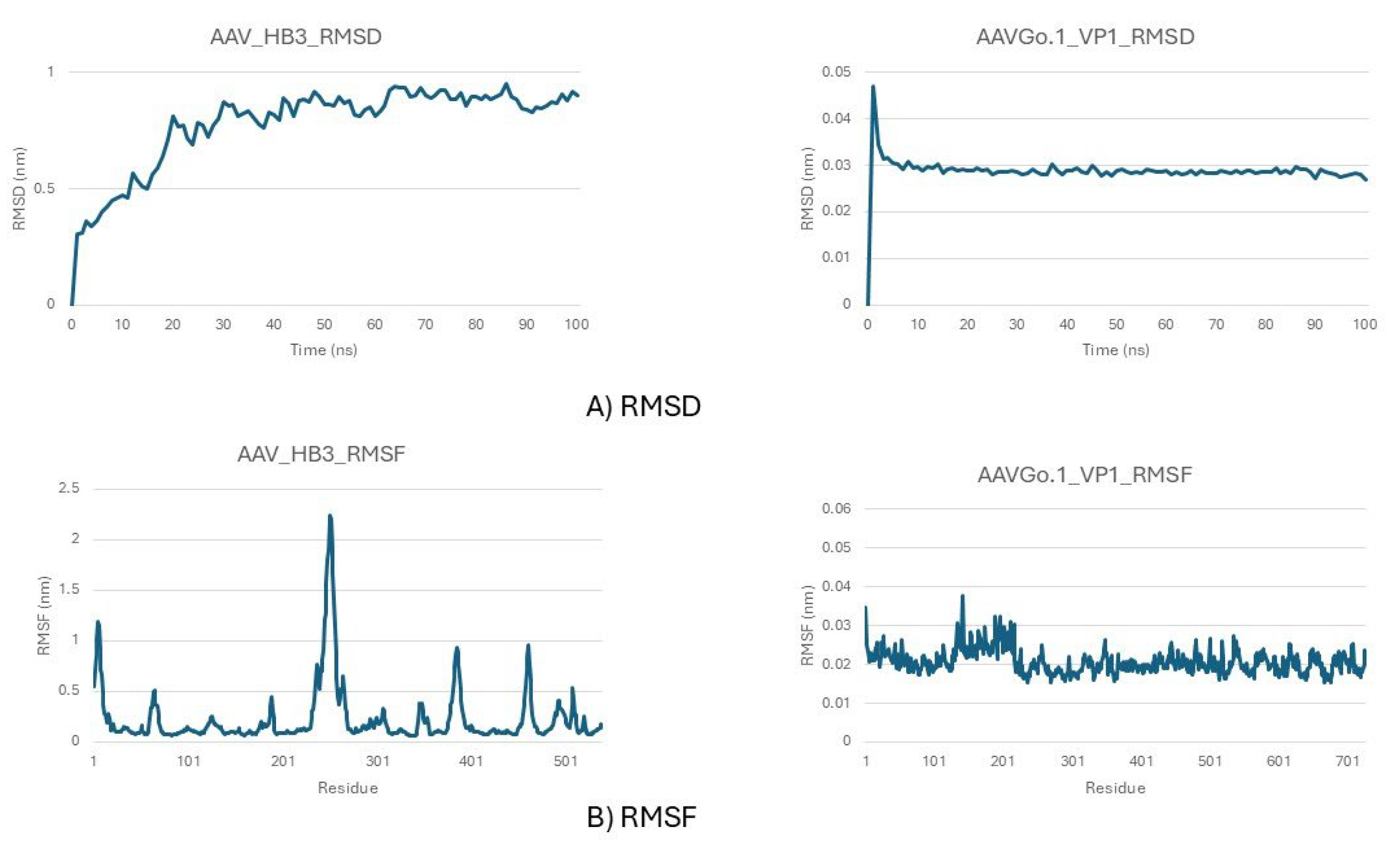
Stability Profiles from Molecular Dynamics
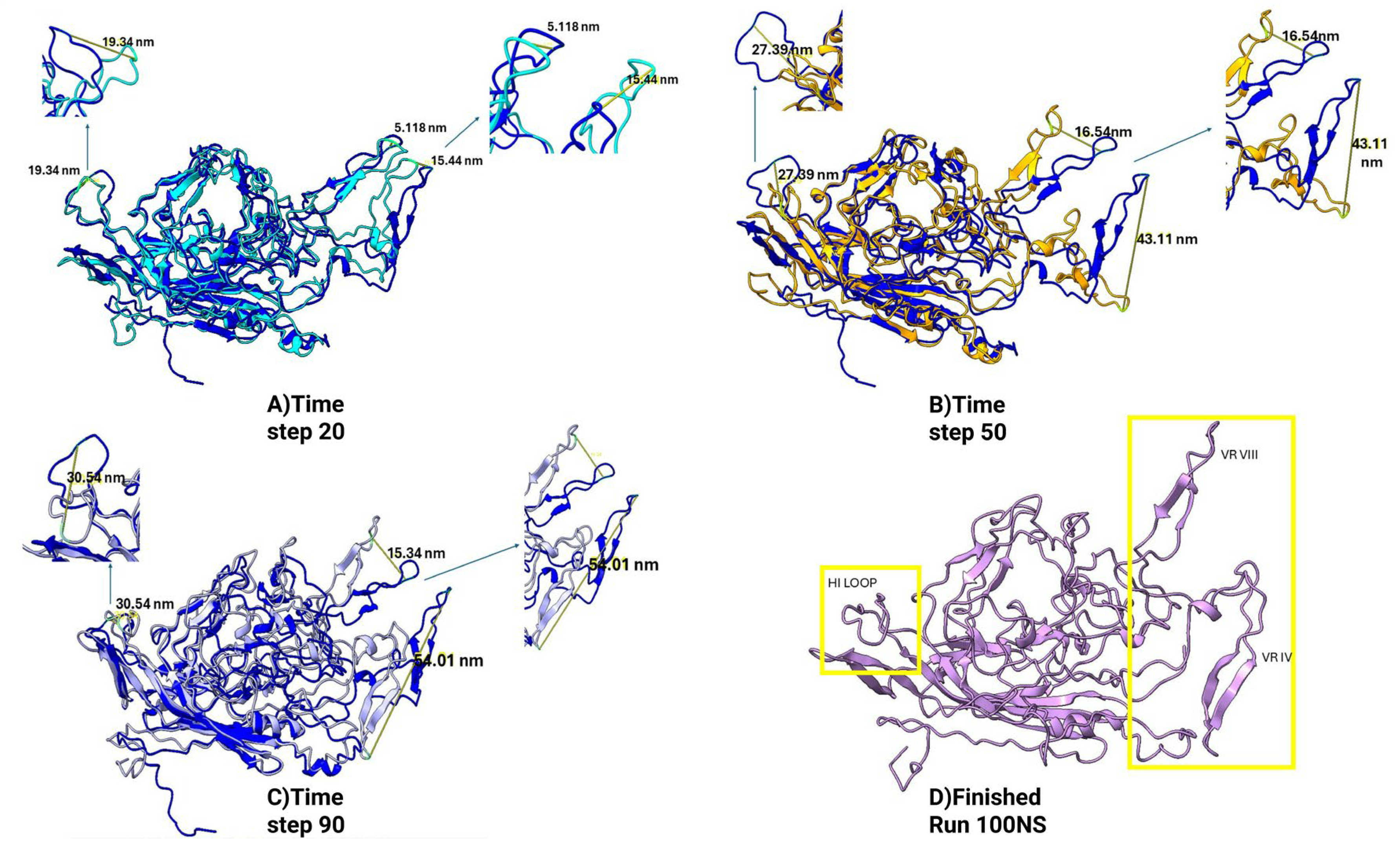
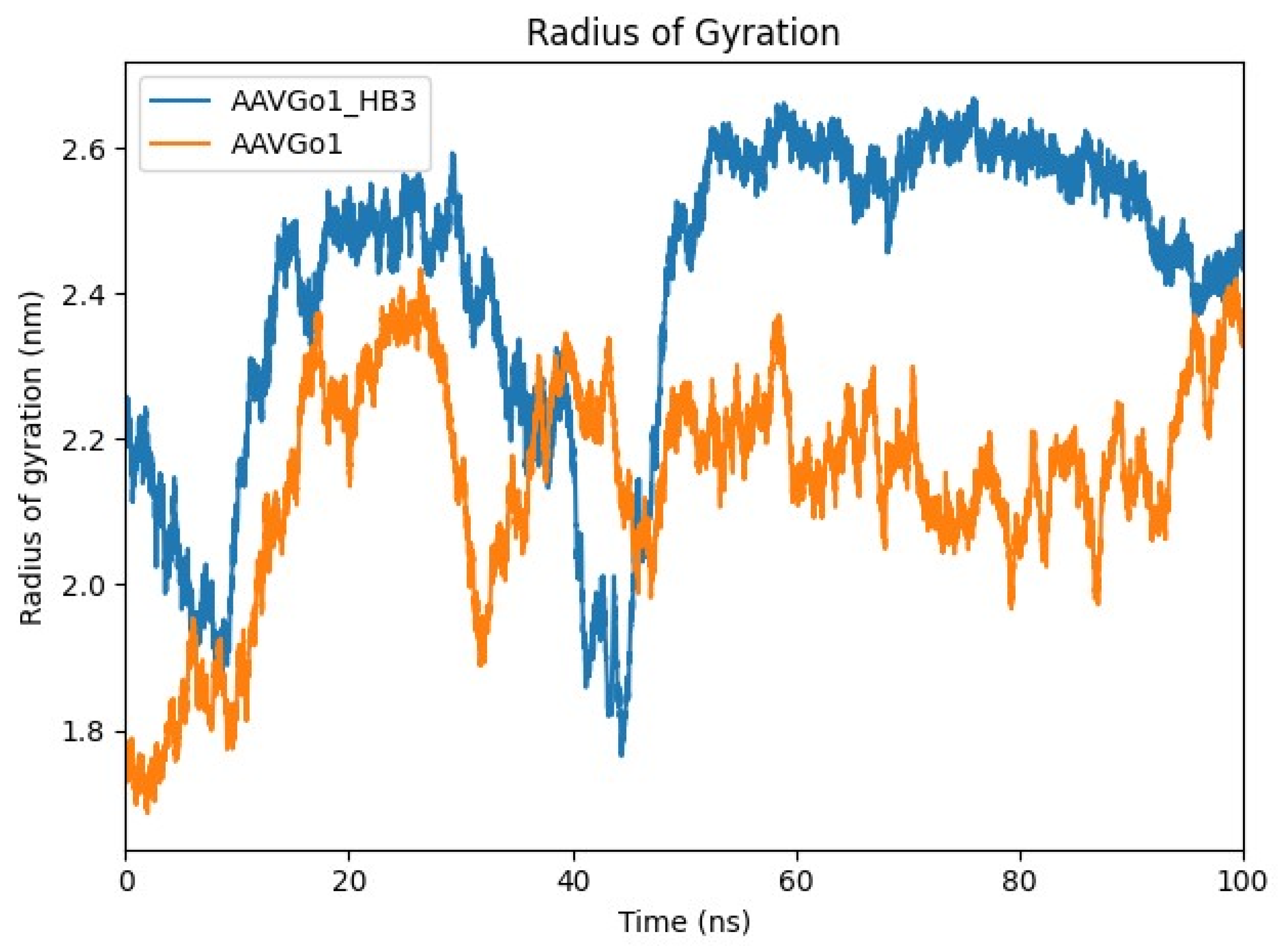
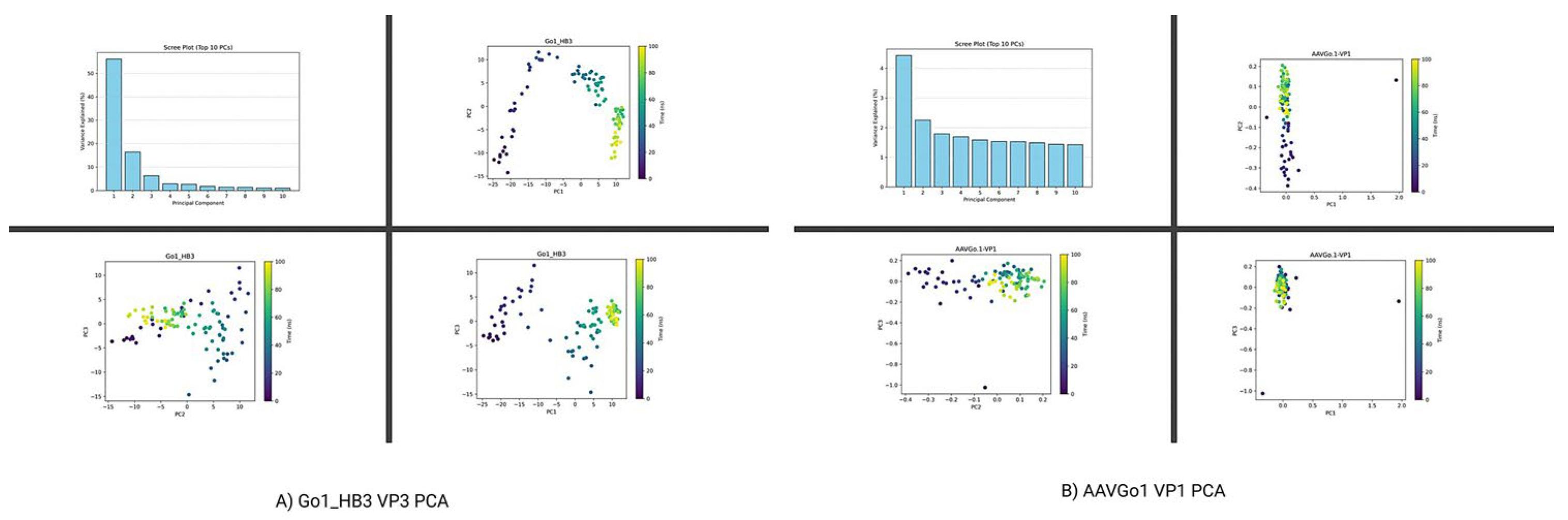
Comparative Structural Dynamics
4. Discussion
6. Conclusion
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
References
- Cotmore, S.F.; Agbandje-McKenna, M.; Canuti, M.; Chiorini, J.A.; Eis-Hubinger, A.-M.; Hughes, J.; et al. ICTV virus taxonomy profile: Parvoviridae. J. Gen. Virol. 2019, 100, 367–368. [Google Scholar] [CrossRef]
- Pénzes, J.J.; Söderlund-Venermo, M.; Canuti, M.; Eis-Hübinger, A.M.; Hughes, J.; Cotmore, S.F.; et al. Reorganizing the family Parvoviridae: A revised taxonomy independent of the canonical approach based on host association. Arch. Virol. 2020, 165, 2133–2146. [Google Scholar] [CrossRef]
- Mohammadi, M. HBoV-1: Virus structure, genomic features, life cycle, pathogenesis, epidemiology, diagnosis and clinical manifestations. Front. Cell. Infect. Microbiol. 2023, 13, 1198127. [Google Scholar] [CrossRef] [PubMed]
- Liu, D.; Li, T.; Liu, L.; Che, X.; Li, X.; Liu, C.; et al. Adeno-associated virus therapies: Pioneering solutions for human genetic diseases. Cytokine Growth Factor Rev. 2024, 80, 109–120. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.-H.; Gessler, D.J.; Zhan, W.; Gallagher, T.L.; Gao, G. Adeno-associated virus as a delivery vector for gene therapy of human diseases. Signal Transduct. Target. Ther. 2024, 9, 78. [Google Scholar] [CrossRef] [PubMed]
- Araujo, N.M. A dual role for adeno-associated virus in human health. Virol. J. 2023, 20, 228. [Google Scholar] [CrossRef]
- Hastie, E.; Samulski, R.J. Adeno-associated virus at 50: A golden anniversary of discovery, research, and gene therapy success—A personal perspective. Hum. Gene Ther. 2015, 26, 257–265. [Google Scholar] [CrossRef]
- Issa, S.S.; Shaimardanova, A.A.; Solovyeva, V.V.; Rizvanov, A.A. Various AAV serotypes and their applications in gene therapy: An overview. Cells 2023, 12, 785. [Google Scholar] [CrossRef]
- Arbetman, A.E.; Lochrie, M.; Zhou, S.; Wellman, J.; Scallan, C.; Doroudchi, M.M.; et al. Novel caprine adeno-associated virus (AAV-Go.1) capsid (AAV-Go.1) is closely related to the primate AAV-5 and has unique tropism and neutralization properties. J. Virol. 2005, 79, 15238–15245. [Google Scholar] [CrossRef] [PubMed]
- Large, E.E.; Silveria, M.A.; Weerakoon, O.; White, T.A.; Chapman, M.S. Cross-species permissivity: Structure of a goat adeno-associated virus and its complex with the human receptor AAVR. J. Virol. 2022, 96, e01484-22. [Google Scholar] [CrossRef]
- Irigoyen, C.; Amenabar Alonso, A.; Sanchez-Molina, J.; Rodríguez-Hidalgo, M.; Lara-López, A.; Ruiz-Ederra, J. Subretinal injection techniques for retinal disease: A review. J. Clin. Med. 2022, 11, 4717. [Google Scholar] [CrossRef]
- U.S. Food and Drug Administration. Approved cellular and gene therapy products. U.S. Department of Health and Human Services. 2025. Available online: https://www.fda.gov/vaccines-blood-biologics/cellular-gene-therapy-products/approved-cellular-and-gene-therapy-products.
- Ylä-Herttuala, S. Endgame: Glybera finally recommended for approval as the first gene therapy drug in the European Union. Mol. Ther. 2012, 20, 1831–1832. [Google Scholar] [CrossRef]
- Watanabe, N.; Yano, K.; Tsuyuki, K.; Okano, T.; Yamato, M. Re-examination of regulatory opinions in Europe: Possible contribution for the approval of the first gene therapy product Glybera. Mol. Ther. Methods Clin. Dev. 2015, 2, 14066. [Google Scholar] [CrossRef]
- Schulz, M.; Levy, D.I.; Petropoulos, C.J.; Bashirians, G.; Winburn, I.; Mahn, M.; et al. Binding and neutralizing anti-AAV antibodies: Detection and implications for rAAV-mediated gene therapy. Mol. Ther. 2023, 31, 616–630. [Google Scholar] [CrossRef]
- Zwi-Dantsis, L.; Mohamed, S.; Massaro, G.; Moeendarbary, E. Adeno-associated virus vectors: Principles, practices, and prospects in gene therapy. Viruses 2025, 17, 239. [Google Scholar] [CrossRef]
- Prasad, S.; Dimmock, D.P.; Greenberg, B.; Walia, J.S.; Sadhu, C.; Tavakkoli, F.; et al. Immune responses and immunosuppressive strategies for adeno-associated virus-based gene therapy for treatment of central nervous system disorders: Current knowledge and approaches. Hum. Gene Ther. 2022, 33, 1228–1245. [Google Scholar] [CrossRef] [PubMed]
- Yan, Z.; Keiser, N.W.; Song, Y.; Deng, X.; Cheng, F.; Qiu, J.; Engelhardt, J.F. A novel chimeric adeno-associated virus 2/human bocavirus 1 parvovirus vector efficiently transduces human airway epithelia. Mol. Ther. 2013, 21, 2181–2194. [Google Scholar] [CrossRef] [PubMed]
- Guido, M.; Tumolo, M.R.; Verri, T.; Romano, A.; Serio, F.; De Giorgi, M.; et al. Human bocavirus: Current knowledge and future challenges. World J. Gastroenterol. 2016, 22, 8684–8697. [Google Scholar] [CrossRef] [PubMed]
- Jartti, T.; Hedman, K.; Jartti, L.; Ruuskanen, O.; Allander, T.; Söderlund-Venermo, M. Human bocavirus—the first 5 years. Rev. Med. Virol. 2012, 22, 46–64. [Google Scholar] [CrossRef]
- Yan, Z.; Zou, W.; Feng, Z.; Shen, W.; Park, S.Y.; Deng, X.; et al. Establishment of a high-yield recombinant adeno-associated virus/human bocavirus vector production system independent of bocavirus nonstructural proteins. Hum. Gene Ther. 2019, 30, 556–570. [Google Scholar] [CrossRef]
- Fakhiri, J.; Schneider, M.A.; Puschhof, J.; Stanifer, M.; Schildgen, V.; Holderbach, S.; et al. Novel chimeric gene therapy vectors based on adeno-associated virus and four different mammalian bocaviruses. Mol. Ther. Methods Clin. Dev. 2019, 12, 202–222. [Google Scholar] [CrossRef] [PubMed]
- Wang, Z.; Deng, X.; Zou, W.; Engelhardt, J.F.; Yan, Z.; Qiu, J. Human bocavirus 1 is a novel helper for adeno-associated virus replication. J. Virol. 2017, 91, e00710-17. [Google Scholar] [CrossRef] [PubMed]
- Catalán-Tatjer, D.; Tzimou, K.; Nielsen, L.K.; Lavado-García, J. Unravelling the essential elements for recombinant adeno-associated virus (rAAV) production in animal cell-based platforms. Biotechnol. Adv. 2024, 72, 108370. [Google Scholar] [CrossRef] [PubMed]
- Nisanov, A.M.; Rivera de Jesús, J.A.; Schaffer, D.V. Advances in AAV capsid engineering: Integrating rational design, directed evolution and machine learning. Mol. Ther. 2025, 33, 1937–1945. [Google Scholar] [CrossRef] [PubMed]
- Guo, J.; Lin, L.F.; Oraskovich, S.V.; Rivera de Jesús, J.A.; Listgarten, J.; Schaffer, D.V.; et al. Computationally guided AAV engineering for enhanced gene delivery. Trends Biochem. Sci. 2024, 49, 457–469. [Google Scholar] [CrossRef]
- Vu Hong, A.; Suel, L.; Petat, E.; Dubois, A.; Le Brun, P.-R.; Guerchet, N.; Veron, P.; Poupiot, J.; Richard, I. An engineered AAV targeting integrin alpha V beta 6 presents improved myotropism across species. Nat. Commun. 2024, 15, 7965. [Google Scholar] [CrossRef]
- Klein, A.H.; Kuiper, M.J.; Burgess, M.; Wickramarachchi, A.; Jain, Y.; Bauer, D.C.; et al. CapBuild: A cloud-native tool for adeno-associated virus capsid engineering. Nucleic Acids Res. 2025, 53, W110–W117. [Google Scholar] [CrossRef]
- Brittain, T.J.; Jang, S.; Coughlin, G.M.; Barcelona, B.H.; Giriat, I.; Ristic, F.; et al. Structural basis of liver de-targeting and neuronal tropism of CNS-targeted AAV capsids. bioRxiv 2025, preprint. [Google Scholar] [CrossRef]
- Tarasova, E.; Farafonov, V.; Khayat, R.; Okimoto, N.; Komatsu, T.S.; Taiji, M.; et al. All-atom molecular dynamics simulations of entire virus capsid reveal the role of ion distribution in capsid’s stability. J. Phys. Chem. Lett. 2017, 8, 779–784. [Google Scholar] [CrossRef]
- Pipatpadungsin, N.; Chao, K.; Rouse, S.L. Coarse-grained simulations of adeno-associated virus and its receptor reveal influences on membrane lipid organization and curvature. J. Phys. Chem. B 2024, 128, 10139–10153. [Google Scholar] [CrossRef]
- Sen, D. Improving clinical efficacy of adeno-associated vectors by rational capsid bioengineering. J. Biomed. Sci. 2014, 21, 103. [Google Scholar] [CrossRef]
- Lee, E.J.; Guenther, C.M.; Suh, J. Adeno-associated virus (AAV) vectors: Rational design strategies for capsid engineering. Curr. Opin. Biomed. Eng. 2018, 7, 58–63. [Google Scholar] [CrossRef]
- Kanaan, N.M.; Sellnow, R.C.; Boye, S.L.; Coberly, B.; Bennett, A.; Agbandje-McKenna, M.; et al. Rationally engineered AAV capsids improve transduction and volumetric spread in the CNS. Mol. Ther. Nucleic Acids 2017, 8, 184–197. [Google Scholar] [CrossRef]
- Zhong, L.; Li, B.; Mah, C.S.; Govindasamy, L.; Agbandje-McKenna, M.; Cooper, M.; et al. Next generation of adeno-associated virus 2 vectors: Point mutations in tyrosines lead to high-efficiency transduction at lower doses. Mol. Ther. 2008, 16, 1717–1722. [Google Scholar] [CrossRef]
- Han, J.; Zhu, L.; Zhang, J.; Guo, L.; Sun, X.; Huang, C.; et al. Rational engineering of adeno-associated virus capsid enhances human hepatocyte tropism and reduces immunogenicity. Cell Prolif. 2022, 55, e13339. [Google Scholar] [CrossRef]
- Li, W.; Feng, S.L.; Herrschaft, L.; Samulski, R.J.; Li, C. Rationally engineered novel AAV capsids for intra-articular gene delivery. Mol. Ther. Methods Clin. Dev. 2024, 32, 101211. [Google Scholar] [CrossRef] [PubMed]
- Suarez-Amaran, L.; Song, L.; Tretiakova, A.P.; Mikhail, S.A.; Samulski, R.J. AAV vector development, back to the future. Mol. Ther. 2025, 33, 1903–1936. [Google Scholar] [CrossRef] [PubMed]
- Tabebordbar, M.; Lagerborg, K.A.; Stanton, A.; King, E.M.; Ye, S.; Tellez, L.; et al. Directed evolution of a family of AAV capsid variants enabling potent muscle-directed gene delivery across species. Cell 2021, 184, 4919–4938.e22. [Google Scholar] [CrossRef]
- Tan, F.; Dong, Y.; Qi, J.; Yu, W.; Chai, R. Artificial intelligence-based approaches for AAV vector engineering. Adv. Sci. 2025, 12, 2411062. [Google Scholar] [CrossRef] [PubMed]
- Eid, F.-E.; Chen, A.T.; Chan, K.Y.; Huang, Q.; Zheng, Q.; Tobey, I.G.; et al. Systematic multi-trait AAV capsid engineering for efficient gene delivery. Nat. Commun. 2024, 15, 6602. [Google Scholar] [CrossRef]
- Wec, A.Z.; Lin, K.S.; Kwasnieski, J.C.; Sinai, S.; Gerold, J.; Kelsic, E.D. Overcoming immunological challenges limiting capsid-mediated gene therapy with machine learning. Front. Immunol. 2021, 12, 674021. [Google Scholar] [CrossRef]
- Zhu, D.; Brookes, D.H.; Busia, A.; Carneiro, A.; Fannjiang, C.; Popova, G.; et al. Optimal trade-off control in machine learning-based library design, with application to adeno-associated virus (AAV) for gene therapy. Sci. Adv. 2024, 10, eadj3786. [Google Scholar] [CrossRef]
- Jang, M.J.; Coughlin, G.M.; Jackson, C.R.; Chen, X.; Chuapoco, M.R.; Vendemiatti, J.L.; et al. Spatial transcriptomics for profiling the tropism of viral vectors in tissues. Nat. Biotechnol. 2023, 41, 1272–1286. [Google Scholar] [CrossRef]
- Jumper, J.; Evans, R.; Pritzel, A.; Green, T.; Figurnov, M.; Ronneberger, O.; et al. Highly accurate protein structure prediction with AlphaFold. Nature 2021, 596, 583–589. [Google Scholar] [CrossRef] [PubMed]
- Noriega, H.A.; Wang, Q.; Yu, D.; Wang, X.S. Structural studies of Parvoviridae capsid assembly and evolution: Implications for novel AAV vector design. Front. Artif. Intell. 2025, 8, 1559461. [Google Scholar] [CrossRef] [PubMed]
- Abramson, J.; Adler, J.; Dunger, J.; Evans, R.; Green, T.; Pritzel, A.; et al. Accurate structure prediction of biomolecular interactions with AlphaFold 3. Nature 2024, 630, 493–500. [Google Scholar] [CrossRef]
- Baek, M.; Anishchenko, I.; Humphreys, I.R.; Cong, Q.; Baker, D.; DiMaio, F. Efficient and accurate prediction of protein structure using RoseTTAFold2. bioRxiv 2023, preprint. [Google Scholar] [CrossRef]
- Weber, S.; Ramirez, C.; Doerfler, W. Signal hotspot mutations in SARS-CoV-2 genomes evolve as the virus spreads and actively replicates in different parts of the world. Virus Res. 2020, 289, 198170. [Google Scholar] [CrossRef]
- Berman, H.M.; Westbrook, J.; Feng, Z.; Gilliland, G.; Bhat, T.N.; Weissig, H.; et al. The Protein Data Bank. Nucleic Acids Res. 2000, 28, 235–242. [Google Scholar] [CrossRef]
- Meng, E.C.; Goddard, T.D.; Pettersen, E.F.; Couch, G.S.; Pearson, Z.J.; Morris, J.H.; et al. UCSF ChimeraX: Tools for structure building and analysis. Protein Sci. 2023, 32, e4792. [Google Scholar] [CrossRef]
- Pettersen, E.F.; Goddard, T.D.; Huang, C.C.; Meng, E.C.; Couch, G.S.; Croll, T.I.; Morris, J.H.; Ferrin, T.E. UCSF ChimeraX: Structure visualization for researchers, educators, and developers. Protein Sci. 2021, 30, 70–82. [Google Scholar] [CrossRef]
- Goddard, T.D.; Huang, C.C.; Meng, E.C.; Pettersen, E.F.; Couch, G.S.; Morris, J.H.; Ferrin, T.E. UCSF ChimeraX: Meeting modern challenges in visualization and analysis. Protein Sci. 2018, 27, 14–25. [Google Scholar] [CrossRef]
- Abraham, M.J.; Murtola, T.; Schulz, R.; Páll, S.; Smith, J.C.; Hess, B.; et al. GROMACS: High performance molecular simulations through multi-level parallelism from laptops to supercomputers. SoftwareX 2015, 1–2, 19–25. [Google Scholar] [CrossRef]
- Bjelkmar, P.; Larsson, P.; Cuendet, M.A.; Hess, B.; Lindahl, E. GROMACS: Analysis of protein stability effects from correction maps, virtual interaction sites, and water models. J. Chem. Theory Comput. 2010, 6, 459–466. [Google Scholar] [CrossRef] [PubMed]
- Berendsen, H.J.C.; Postma, J.P.M.; van Gunsteren, W.F.; DiNola, A.; Haak, J.R. Molecular dynamics with coupling to an external bath. J. Chem. Phys. 1984, 81, 3684–3690. [Google Scholar] [CrossRef]
- Parrinello, M.; Rahman, A. Polymorphic transitions in single crystals: A new molecular dynamics method. J. Appl. Phys. 1981, 52, 7182–7190. [Google Scholar] [CrossRef]
- Hess, B.; Bekker, H.; Berendsen, H.J.C.; Fraaije, J.G.E.M. LINCS: A linear constraint solver for molecular simulations. J. Comput. Chem. 1997, 18, 1463–1472. [Google Scholar] [CrossRef]
- Darden, T.; York, D.; Pedersen, L. Particle mesh Ewald: An N·log(N) method for Ewald sums in large systems. J. Chem. Phys. 1993, 98, 10089–10092. [Google Scholar] [CrossRef]
- Essmann, U.; Perera, L.; Berkowitz, M.L.; Darden, T.; Lee, H.; Pedersen, L.G. A smooth particle mesh Ewald method. J. Chem. Phys. 1995, 103, 8577–8593. [Google Scholar] [CrossRef]
- Benson, N.C.; Daggett, V. A comparison of multiscale methods for the analysis of molecular dynamics simulations. J. Phys. Chem. B 2012, 116, 8722–8731. [Google Scholar] [CrossRef]
- May, E.R. Recent developments in molecular simulation approaches to study spherical virus capsids. Mol. Simul. 2014, 40, 878–888. [Google Scholar] [CrossRef] [PubMed]
- Jones, P.E.; Pérez-Segura, C.; Bryer, A.J.; Perilla, J.R.; Hadden-Perilla, J.A. Molecular dynamics of the viral life cycle: Progress and prospects. Curr. Opin. Virol. 2021, 50, 128–138. [Google Scholar] [CrossRef]
- Ito, M.; Takino, N.; Nomura, T.; Kan, A.; Muramatsu, S.-i. Engineered adeno-associated virus 3 vector with reduced reactivity to serum antibodies. Sci. Rep. 2021, 11, 9322. [Google Scholar] [CrossRef]
- DiPrimio, N.; Asokan, A. Surface loop dynamics in adeno-associated virus capsid assembly. J. Virol. 2008, 82, 5178–5189. [Google Scholar] [CrossRef]
- Mietzsch, M.; Barnes, C.; Hull, J.A.; Chipman, P.; Xie, J.; Bhattacharya, N.; et al. Comparative analysis of the capsid structures of AAVrh.10, AAVrh.39, and AAV8. J. Virol. 2020, 94, e01769-19. [Google Scholar] [CrossRef] [PubMed]
- Huang, J.; MacKerell, A.D., Jr. CHARMM36 all-atom additive protein force field: Validation based on comparison to NMR data. J. Comput. Chem. 2013, 34, 2135–2145. [Google Scholar] [CrossRef] [PubMed]

Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.



