Submitted:
24 February 2026
Posted:
25 February 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
2.1. Sample Collection and Geographic Distribution
2.2. Serological Screening by Enzyme-Linked Immunosorbent Assay (ELISA)
2.2.1. Commercial ELISA
2.2.2. In-House ELISA
2.3. Statistical Analyses
3. Results
4. Discussion
Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Data Availability Statement
Conflicts of Interest
References
- Naqvi, A.A.T.; Fatima, K.; Mohammad, T.; Fatima, U.; Singh, I.K.; Singh, A.; Atif, S.M.; Hariprasad, G.; Hasan, G.M.; Hassan, M.I. Insights into SARS-CoV-2 genome, structure, evolution, pathogenesis and therapies: Structural genomics approach. Biochim. Biophys. Acta Mol. Basis Dis. 2020, 1866, 165878. [Google Scholar] [CrossRef] [PubMed]
- Decaro, N.; Buonavoglia, C. An update on canine coronaviruses: Viral evolution and pathobiology. Vet. Microbiol. 2008, 132, 221–234. [Google Scholar] [CrossRef]
- Whittaker, G.; Stout, A. Coronaviruses in wild canids: A review of the literature. Qeios 2022. [Google Scholar] [CrossRef]
- Coronaviridae Study Group of the International Committee on Taxonomy of Viruses. The species Severe acute respiratory syndrome-related coronavirus: Classifying 2019-nCoV and naming it SARS-CoV-2. Nat. Microbiol. 2020, 5, 536–544. [CrossRef]
- Harcourt, J.; Tamin, A.; Lu, X.; Kamili, S.; Sakthivel, S.K.; Murray, J.; et al. Severe acute respiratory syndrome coronavirus 2 from a patient with coronavirus disease, United States. Emerg. Infect. Dis. 2020, 26, 1266–1273. [Google Scholar] [CrossRef]
- Zhou, F.; Yu, T.; Du, R.; Fan, G.; Liu, Y.; Liu, Z.; et al. Clinical course and risk factors for mortality of adult in patients with COVID-19 in Wuhan, China: A retrospective cohort study. Lancet 2020, 395, 1054–1062. [Google Scholar] [CrossRef]
- World Health Organization. Coronavirus disease (COVID-2019) situation reports. Available online: https://www.who.int/emergencies/diseases/novel-coronavirus-2019/situation-reports/situation-reports-archive (accessed on 4 January 2026).
- LeDuc, J.W.; Barry, M.A. SARS, the first pandemic of the 21st century. Emerg. Infect. Dis. 2004, 10. [Google Scholar] [CrossRef]
- Zaki, A.M.; van Boheemen, S.; Bestebroer, T.M.; Osterhaus, A.D.; Fouchier, R.A. Isolation of a novel coronavirus from a man with pneumonia in Saudi Arabia. N. Engl. J. Med. 2012, 367, 1814–1820. [Google Scholar] [CrossRef]
- Lytras, S.; et al. The animal origin of SARS-CoV-2. Science 2021, 373, 968–970. [Google Scholar] [CrossRef] [PubMed]
- Fischhoff, I.R.; Castellanos, A.A.; Rodrigues, J.P.G.L.M.; Varsani, A.; Han, B.A. Predicting the zoonotic capacity of mammals to transmit SARS-CoV-2. Proc. R. Soc. B 2021, 288, 20211651. [Google Scholar] [CrossRef] [PubMed]
- Porter, S.M.; Hartwig, A.E.; Bielefeldt-Ohmann, H.; Bosco-Lauth, A.M.; Root, J.J. Susceptibility of wild canids to SARS-CoV-2. Emerg. Infect. Dis. 2022, 28, 1852–1855. [Google Scholar] [CrossRef]
- Andre, M.; Lau, L.S.; Pokharel, M.D.; Ramelow, J.; Owens, F.; Souchak, J.; Akkaoui, J.; Ales, E.; Brown, H.; Shil, R.; Nazaire, V.; Manevski, M.; Paul, N.P.; Esteban-Lopez, M.; Ceyhan, Y.; El-Hage, N. From Alpha to Omicron: How different variants of concern of the SARS-coronavirus-2 impacted the world. Biology 2023, 12, 1267. [Google Scholar] [CrossRef]
- Decaro, N.; Balboni, A.; Bertolotti, L.; Martino, P.A.; Mazzei, M.; Mira, F.; Pagnini, U. SARS-CoV-2 infection in dogs and cats: Facts and speculations. Front. Vet. Sci. 2021, 8, 619207. [Google Scholar] [CrossRef] [PubMed]
- Munir, K.; Ashraf, S.; Munir, I.; Khalid, H.; Muneer, M.A.; Mukhtar, N.; Amin, S.; Ashraf, S.; Imran, M.A.; Chaudhry, U.; et al. Zoonotic and reverse zoonotic events of SARS-CoV-2 and their impact on global health. Emerg. Microbes Infect. 2020, 9, 2222–2235. [Google Scholar] [CrossRef]
- Pickering, B.; Lung, O.; Maguire, F.; Kruczkiewicz, P.; Kotwa, J.D.; Buchanan, T.; Gagnier, M.; Guthrie, J.L.; Jardine, C.M.; Marchand-Austin, A.; Massé, A.; McClinchey, H.; Nirmalarajah, K.; Aftanas, P.; Blais-Savoie, J.; Chee, H.Y.; Chien, E.; Yim, W.; Banete, A.; Griffin, B.D.; Yip, L.; Goolia, M.; Suderman, M.; Pinette, M.; Smith, G.; Sullivan, D.; Rudar, J.; Vernygora, O.; Adey, E.; Nebroski, M.; Goyette, G.; Finzi, A.; Laroche, G.; Ariana, A.; Vahkal, B.; Côté, M.; McGeer, A.J.; Nituch, L.; Mubareka, S.; Bowman, J. Divergent SARS-CoV-2 variant emerges in white-tailed deer with deer-to-human transmission. Nat. Microbiol. 2022, 7, 2011–2024. [Google Scholar] [CrossRef]
- Larsen, H.D.; Fonager, J.; Lomholt, F.K.; Dalby, T.; Benedetti, G.; Kristensen, B.; Urth, T.R.; Rasmussen, M.; Lassaunière, R.; Rasmussen, T.B.; Strandbygaard, B.; Lohse, L.; Chaine, M.; Møller, K.L.; Berthelsen, A.N.; Nørgaard, S.K.; Sönksen, U.W.; Boklund, A.E.; Hammer, A.S.; Belsham, G.J.; Krause, T.G.; Mortensen, S.; Bøtner, A.; Fomsgaard, A.; Mølbak, K. Preliminary report of an outbreak of SARS-CoV-2 in mink and mink farmers associated with community spread, Denmark, June to November 2020. Euro Surveill. 2021, 26, 2100009. [Google Scholar] [CrossRef]
- World Organisation for Animal Health (WOAH). Cases of SARS-CoV-2 infection in animals reported to WOAH since March 2020. Available online: https://www.woah.org/en/disease/sars-cov-2/#ui-id-2 (accessed on 21 February 2026).
- Shi, J.; Wen, Z.; Zhong, G.; Yang, H.; Wang, C.; Huang, B.; et al. Susceptibility of ferrets, cats, dogs, and other domesticated animals to SARS-coronavirus 2. Science 2020, 368, 1016–1020. [Google Scholar] [CrossRef]
- Jemeršić, L.; Lojkić, I.; Krešić, N.; Keros, T.; Zelenika, T.A.; Jurinović, L.; Skok, D.; Bata, I.; Boras, J.; Habrun, B.; Brnić, D. Investigating the presence of SARS-CoV-2 in free-living and captive animals. Pathogens 2021, 10, 635. [Google Scholar] [CrossRef]
- Miller, M.R.; Braun, E.; Ip, H.S.; Tyson, G.H. Domestic and wild animal samples and diagnostic testing for SARS-CoV-2. Vet. Q. 2023, 43, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Hamer, S.A.; Pauvolid-Corrêa, A.; Zecca, I.B.; Davila, E.; Auckland, L.D.; Roundy, C.M.; Tang, W.; Torchetti, M.K.; Killian, M.L.; Jenkins-Moore, M.; et al. SARS-CoV-2 infections and viral isolations among serially tested cats and dogs in households with infected owners in Texas, USA. Viruses 2021, 13, 938. [Google Scholar] [CrossRef] [PubMed]
- Institute of Public Health of the Republic of Serbia “Dr Milan Jovanović Batut”. Health Statistical Yearbook of the Republic of Serbia 2023. 2024. Available online: https://www.batut.org.rs/download/aktuelno/04012023.pdf (accessed on 10 February 2026).
- Stanojević, S.; Radojičić, S.; Mišić, D.; Srejić, D.; Vasiljević, D.V.; Prokić, K.; Ilić, N. Frequency of SARS-CoV-2 infection in dogs and cats: Results of a retrospective serological survey in Šumadija District, Serbia. Prev. Vet. Med. 2022, 208, 105755. [Google Scholar] [CrossRef]
- Petrović, T.; Videnović, D.; Savić, S.; Vidanović, D.; Gajdov, V.; Lazić, G.; Samojlović, M.; Lupulović, D.; Lazić, S. Study on presence of SARS-CoV-2 in pet dogs and cats – Results from Serbia. In Proceedings of the 14th Annual Meeting EPIZONE, Barcelona, Spain, 18–20 May 2022. [Google Scholar]
- Statistical Office of the Republic of Serbia. Statistical Yearbook of the Republic of Serbia 2023. Available online: https://publikacije.stat.gov.rs/G2023/pdf/G20232056.pdf (accessed on 12 December 2025).
- Official Gazette of the Republic of Serbia. No. 21/2024. Available online: http://demo.paragraf.rs/demo/combined/Old/t/t2024_03/SG_021_2024_005.htm (accessed on 21 February 2026).
- Baralić, M.; Laušević, M.; Ćujić, D.; Bontić, A.; Pavlović, J.; Brković, V.; Kezić, A.; Mihajlovski, K.; Hadži Tanović, L.; Assi Milošević, I.; Lukić, J.; Gnjatović, M.; Todorović, A.; Stojanović, N.M.; Jovanović, D.; Radović, M. The importance of natural and acquired immunity to SARS-CoV-2 infection in patients on peritoneal dialysis. Vaccines 2024, 12, 135. [Google Scholar] [CrossRef]
- Kotur-Stevuljević, J.; Roksandić Milenković, M.; Vekić, J.; Dimić, N.; Dimić, D.; Ćujić, D.; Guzonjić, A.; Gnjatović, M.; Todorović, A.; Bogavac-Stanojević, N. Leukocyte telomere dynamic change in patients with mild to moderate COVID-19 during three weeks of follow-up: Relation with therapy. J. Med. Biochem. 2026, 45, 1–11. [Google Scholar] [CrossRef]
- Cohen, J. A coefficient of agreement for nominal scales. Educ. Psychol. Meas. 1960, 20, 37–46. [Google Scholar] [CrossRef]
- EFSA AHAW Panel (EFSA Panel on Animal Health and Animal Welfare); Nielsen, S.S.; Alvarez, J.; Bicout, D.J.; Calistri, P.; Canali, E.; Drewe, J.A.; Garin-Bastuji, B.; Gonzales Rojas, J.L.; Gortázar, C.; Herskin, M.; Michel, V.; Miranda Chueca, M.A.; Padalino, B.; Pasquali, P.; Roberts, H.C.; Spoolder, H.; Velarde, A.; Viltrop, A.; Winckler, C.; Adlhoch, C.; Aznar, I.; Baldinelli, F.; Boklund, A.; Broglia, A.; Gerhards, N.; Mur, L.; Nannapaneni, P.; Stahl, K. SARS-CoV-2 in animals: Susceptibility of animal species, risk for animal and public health, monitoring, prevention and control. EFSA J. 2023, 21, 7822. [Google Scholar] [CrossRef]
- Abay, Z.; Sadikaliyeva, S.; Nurpeisova, A.; Jekebekov, K.; Shorayeva, K.; Yespembetov, B.; Nurabayev, S.; Kerimbayev, A.; Khairullin, B.; Yoo, H.; et al. Breaking the Barrier: SARS-CoV-2 Infections in Wild and Companion Animals and Their Implications for Public Health. Viruses 2024, 16, 956. [Google Scholar] [CrossRef]
- Kuhn, J.; Marti, I.; Ryser-Degiorgis, M.-P.; Wernike, K.; Jones, S.; Tyson, G.; Delalay, G.; Scherrer, P.; Borel, S.; Hosie, M.J.; et al. Investigations on the Potential Role of Free-Ranging Wildlife as a Reservoir of SARS-CoV-2 in Switzerland. Viruses 2024, 16, 1407. [Google Scholar] [CrossRef] [PubMed]
- Chan, T.; Ginders, J.; Kuhlmeier, E.; Meli, M.L.; Bönzli, E.; Meili, T.; Hüttl, J.; Hatt, J.-M.; Hindenlang Clerc, K.; Kipar, A.; et al. Detection of SARS-CoV-2 RNA in a Zoo-Kept Red Fox (Vulpes vulpes). Viruses 2024, 16, 521. [Google Scholar] [CrossRef] [PubMed]
- Goll, A.; Krupińska, M.; Nowicka, J.; Baranowicz, K.; Rabalski, L.; Lass, A.; Gorska, A.; Sironen, T.; Kant, R.; Grzybek, M. Wild Red Foxes (Vulpes vulpes) Do Not Participate in SARS-CoV-2 Circulation in Poland. One Health 2024, 19, 100845. [Google Scholar] [CrossRef]
- Delahay, R.J.; de la Fuente, J.; Smith, G.C.; Sharun, K.; Snary, E.L.; Flores Girón, L.; Nziza, J.; Fooks, A.R.; Brookes, S.M.; Lean, F.Z.X.; et al. Assessing the Risks of SARS-CoV-2 in Wildlife. One Health Outlook 2021, 3, 7. [Google Scholar] [CrossRef] [PubMed]
- Yang, X.Y.; Huang, J.S.; Gong, Q.L.; Sun, J.M.; Li, Y.J.; Liu, B.; Zhang, Y.M.; Shi, C.W.; Yang, G.L.; Yang, W.T.; Wang, C.F. SARS-CoV-2 Prevalence in Wildlife 2020–2022: A Worldwide Systematic Review and Meta-Analysis. Microbes Infect. 2024, 26, 105350. [Google Scholar] [CrossRef]
- Goldberg, A.R.; Langwig, K.E.; Brown, K.L.; Marano, J.M.; Rai, P.; King, K.M.; Sharp, A.K.; Ceci, A.; Kailing, C.D.; Kailing, M.J.; et al. Widespread Exposure to SARS-CoV-2 in Wildlife Communities. Nat. Commun. 2024, 15, 6210. [Google Scholar] [CrossRef]
- Tinto, B.; Revel, J.; Virolle, L.; Chenet, B.; Reboul Salze, F.; Ortega, A.; Beltrame, M.; Simonin, Y. Monitoring SARS-CoV-2 Seroprevalence in Domestic and Exotic Animals in Southern France. Trop. Med. Infect. Dis. 2023, 8, 426. [Google Scholar] [CrossRef]
- Greenhorn, J.E.; Kotwa, J.D.; Bowman, J.; Bruce, L.; Buchanan, T.; Buck, P.A.; Davy, C.M.; Dibernardo, A.; Flockhart, L.; Gagnier, M.; et al. SARS-CoV-2 Wildlife Surveillance in Ontario and Québec. Can. Commun. Dis. Rep. 2022, 48, 243–251. [Google Scholar] [CrossRef]
- Ristić, M.; et al. Sentinel Surveillance of Acute Respiratory Infections and Distribution of SARS-CoV-2, Influenza A and B, and Respiratory Syncytial Virus in the Post-COVID-19 Period. Vojnosanit. Pregl. 2025, 82, 490–503. [Google Scholar] [CrossRef]
- Campbell, S.J.; Ashley, W.; Gil-Fernandez, M.; Newsome, T.M.; Di Giallonardo, F.; Ortiz-Baez, A.S.; Mahar, J.E.; Towerton, A.L.; Gillings, M.; Holmes, E.C.; Carthey, A.J.R.; Geoghegan, J.L. Red Fox Viromes in Urban and Rural Landscapes. Virus Evol. 2020, 6, veaa065. [Google Scholar] [CrossRef] [PubMed]
- Davidson, W.R.; Appel, M.J.; Doster, G.L.; Baker, O.E.; Brown, J.F. Diseases and Parasites of Red Foxes, Gray Foxes, and Coyotes from Commercial Sources Selling to Fox-Chasing Enclosures. J. Wildl. Dis. 1992, 28, 581–589. [Google Scholar] [CrossRef] [PubMed]
- Luan, J.; Lu, Y.; Jin, X.; Zhang, L. Spike Protein Recognition of Mammalian ACE2 Predicts the Host Range and an Optimized ACE2 for SARS-CoV-2 Infection. Biochem. Biophys. Res. Commun. 2020, 526, 165–169. [Google Scholar] [CrossRef] [PubMed]
- Tindih Sheltone, H.; Obara, I.; Alessandro, M.; et al. First Detection of SARS-CoV-2-like Antibodies in Domestic and Wild Animals in Kenya. Authorea 2022, May 25.
- Liu, W.; Liu, L.; Kou, G.; Zheng, Y.; Ding, Y.; Ni, W.; Wang, Q.; Tan, L.; Wu, W.; Tang, S.; Xiong, Z. Evaluation of Nucleocapsid and Spike Protein-Based Enzyme-Linked Immunosorbent Assays for Detecting Antibodies against SARS-CoV-2. J. Clin. Microbiol. 2020, 58, e00461-20. [Google Scholar] [CrossRef]
- Kontou, P.I.; Braliou, G.G.; Dimou, N.L.; Nikolopoulos, G.; Bagos, P.G. Antibody Tests in Detecting SARS-CoV-2 Infection: A Meta-Analysis. Diagnostics 2020, 10, 319. [Google Scholar] [CrossRef]
- Barua, S.; Hoque, M.; Adekanmbi, F.; Kelly, P.; Jenkins-Moore, M.; Torchetti, M.K.; Chenoweth, K.; Wood, T.; Wang, C.M. Antibodies to SARS-CoV-2 in Dogs and Cats, USA. Emerg. Microbes Infect. 2021, 10, 1669–1674. [Google Scholar] [CrossRef] [PubMed]
- Dias, J.M.; Costa, M.O.; de Souza, A.O.; Faria, M.L.; Golcman, D.H.; Selim, M.B.; Medici, E.P.; Haga, G.S.I.; Romaldini, A.H.C.N.; Okuda, L.H. Serological Assay for SARS-CoV-2 in Domestic and Wild Animals in Middle and Southeast Regions in Brazil. Fortune J. Health Sci. 2024, 7, 299–306. [Google Scholar]
- Pomorska Mól, M.; Turlewicz Podbielska, H.; Gogulski, M.; Ruszkowski, J.R.; Kubiak, M.; Kuriga, A.; Barket, P.; Postrzech, M. A Cross-Sectional Retrospective Study of SARS-CoV-2 Seroprevalence in Domestic Cats, Dogs and Rabbits in Poland. BMC Vet. Res. 2021, 17, 322. [Google Scholar] [CrossRef] [PubMed]
- Fernández-Bastit, L.; Marfil, S.; Pradenas, E.; Blanco, J.; Vergara-Alert, J.; Segalés, J. Comparison of Three Commercial ELISA Kits for Detection of Antibodies Against SARS-CoV-2 in Serum Samples from Different Animal Species. Viruses 2025, 17, 716. [Google Scholar] [CrossRef] [PubMed]
- Udom, K.; Jairak, W.; Chamsai, E.; Charoenkul, K.; Boonyapisitsopa, S.; Bunpapong, N.; Techakriengkrai, N.; Amonsin, A. Serological Survey of Antibodies Against SARS-CoV-2 in Dogs and Cats, Thailand. Transbound. Emerg. Dis. 2022, 69, 2140–2147. [Google Scholar] [CrossRef]
- Whitman, J.D.; Hiatt, J.; Mowery, C.T.; Shy, B.R.; Yu, R.; Yamamoto, T.N.; Rathore, U.; Goldgof, G.M.; Whitty, C.; Woo, J.M.; et al. Test Performance Evaluation of SARS-CoV-2 Serological Assays. medRxiv [Preprint] 2020, 2020.04.25.20074856. Update in: Nat. Biotechnol. 2020, 38, 1174–1183. https://doi.org/10.1038/s41587-020-0659-0. [CrossRef]
- Gobena, D.; Gudina, E.K.; Degfie, T.T.; et al. Diagnostic Performance Evaluation of Elecsys Anti-SARS-CoV-2 Assay Against RT-PCR for SARS-CoV-2 Detection and Surveillance in Ethiopian Referral Hospitals: Cross-Sectional Study. Sci. Rep. 2026, 16, 5207. [Google Scholar] [CrossRef]
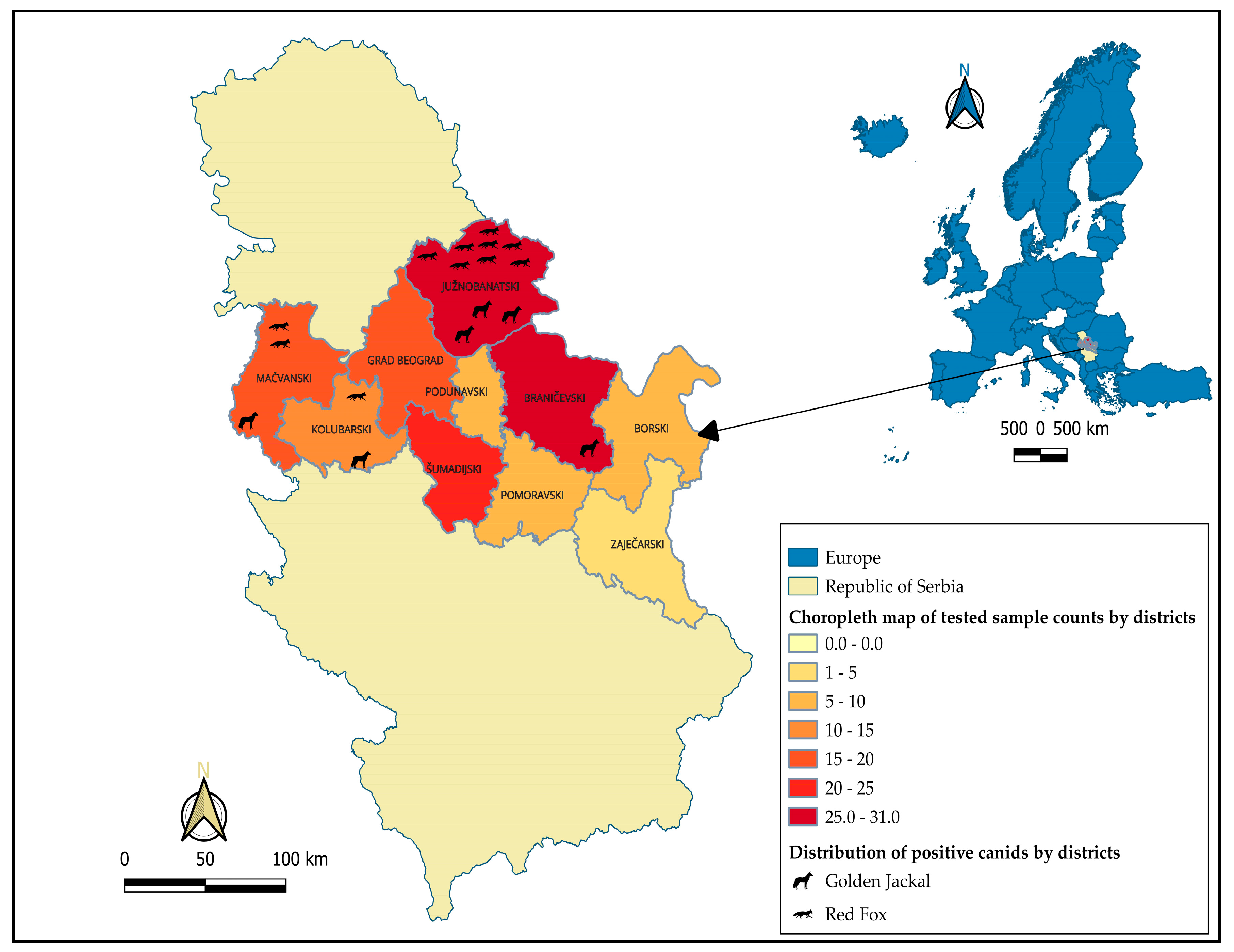
| District | Species | Total | |
|---|---|---|---|
| Red fox | Golden jackal | ||
| Borski | 6 | 4 | 10 |
| Braničevski | 7 | 21 | 28 |
| Južnobanatski | 16 | 15 | 31 |
| Kolubarski | 13 | 2 | 15 |
| Mačvanski | 16 | 3 | 19 |
| Podunavski | 0 | 7 | 7 |
| Pomoravski | 1 | 5 | 6 |
| Šumadijski | 16 | 9 | 25 |
| Zaječarski | 1 | 3 | 4 |
| City of Belgrade | 7 | 13 | 20 |
| Total | 83 | 82 | 165 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




