Submitted:
18 February 2026
Posted:
24 February 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
1.1. The Unmet Need for Individualized Chemotherapy
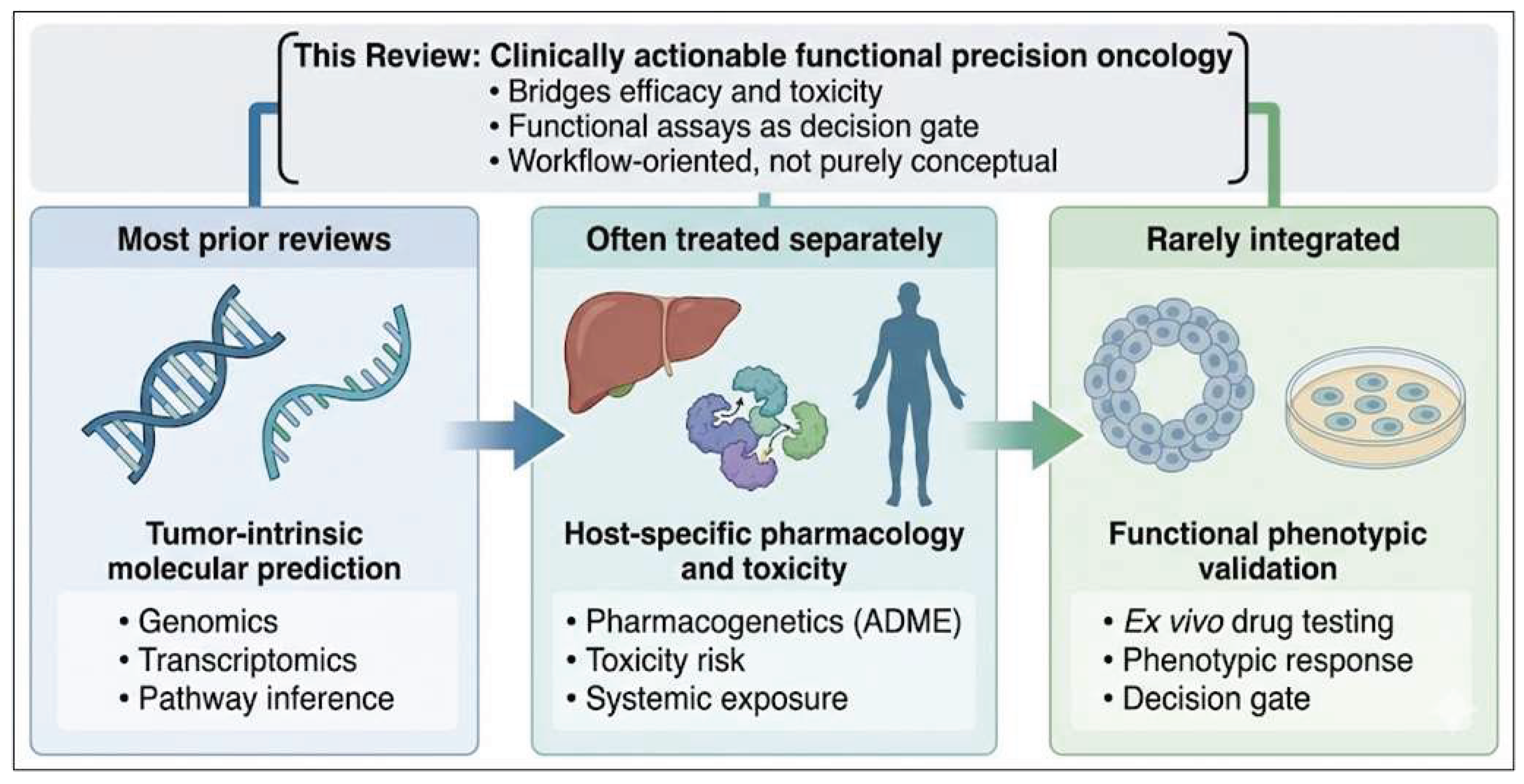
1.2. From Precision Oncology to Functional Precision Oncology
1.3. Aim and Scope of the Review
2. Molecular Oncodiagnostics: Conceptual and Technical Foundations
2.1. Definition and Components
2.2. Enabling Technologies
2.3. Clinical Implementation Models
3. Tumor Transcriptomics as a Predictor of Chemotherapy Response
3.1. Biological Rationale
3.2. Transcriptomic Signatures Associated with Drug Sensitivity and Resistance
3.3. Clinical Assays and Commercial Platforms
3.4. Limitations and Unresolved Challenges
4. Patient Pharmacogenetics in Oncology
4.1. Germline Pharmacogenetics: Principles and Relevance
4.2. Key Pharmacogenetic Pathways in Chemotherapy
4.3. Clinical Evidence and Guideline-Supported Applications
4.4. Limitations as a Standalone Tool
5. Integrating Tumor Transcriptomics and Patient Pharmacogenetics
5.1. Biological Rationale for Integration
5.2. Multi-Omics Integration Strategies
5.3. Emerging Evidence from Recent Studies
5.4. Clinical Decision-Making Implications
6. Ex Vivo Chemoresistance Testing Using Patient-Derived Tumor Cells
6.1. Current Technological Landscape (2026)
6.2. Clinical Utility and Feasibility
6.3. Emerging Enhancements
7. Predictive Modeling and Artificial Intelligence
7.1. From Molecular Findings to Clinical Decisions
7.2. Companion Diagnostics and Precision Therapeutics
7.3. Liquid Biopsy in Clinical Monitoring
7.4. Ethical, Regulatory, and Health System Considerations
7.5. Emerging Trends and Future Perspectives
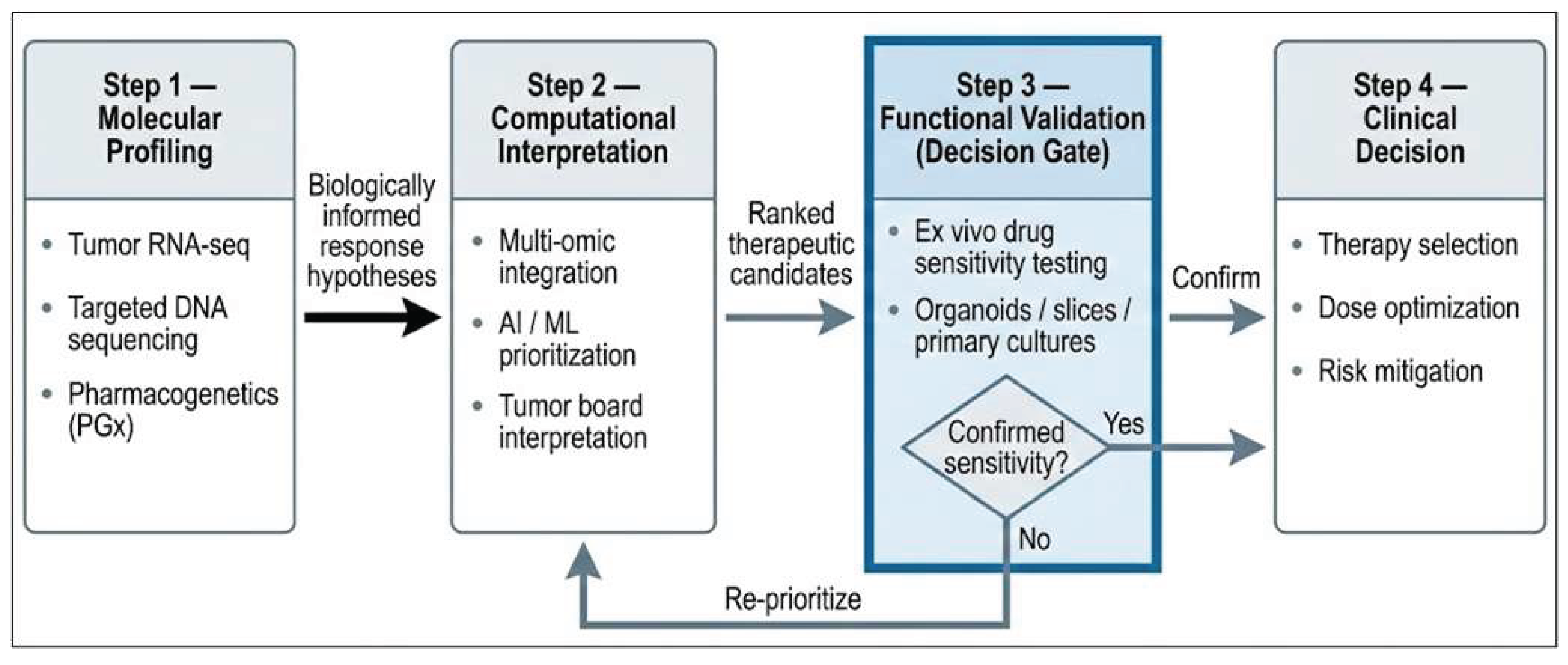
8. Functional Integration of Multi-Omic and Ex Vivo Data
8.1. Informing Drug Response Expectations with Multi-Omic Data
8.2. From Multi-Omic Prediction to Ex Vivo Validation
8.3. Functional Validation and Refinement of Therapeutic Strategies
9. Clinical Implementation: Opportunities and Barriers
9.1. Translation to Clinical Practice
9.2. Ethical and Regulatory Challenges
10. Final Considerations
10.1. Value Added by Functional Testing
10.2. Integration Challenges
10.3. Future Directions
11. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Akhoundova, D., & Rubin, M. A. (2022). Clinical application of advanced multi-omics tumor profiling: Shaping precision oncology of the future. Cancer Cell, 40(9), 920–938. [CrossRef]
- Al-Kabani, A., Huda, B., Haddad, J., Yousuf, M., Bhurka, F., Ajaz, F., et al. (2025). Exploring experimental models of colorectal cancer: A critical appraisal from 2D cell systems to organoids, humanized mouse avatars, organ-on-chip, CRISPR engineering, and AI-driven platforms—Challenges and opportunities for translational precision oncology. Cancers, 17(13), 2163. [CrossRef]
- Aman, L. A. O., Arfan, A., Asnaw, A., Putra, P. P., Hasan, H., & Papeo, D. R. P. (2025, December 17). Prediction of PLX-4720 sensitivity in cancer cell lines through multi-omics integration and attention-based fusion modeling. arXiv. https://arxiv.org/abs/2512.12113.
- Angulo, B., Lopez-Rios, F., & Gonzalez, D. (2014). A new generation of companion diagnostics: cobas BRAF, KRAS and EGFR mutation detection tests. Expert Review of Molecular Diagnostics, 14(5), 517–524. [CrossRef]
- Ayuda-Durán, P., et al. (2023). Standardized assays to monitor drug sensitivity in hematologic cancers. Cell Death Discovery, 9(1), 435. [CrossRef]
- Baranovskii, A., Gündüz, I. B., Franke, V., Uyar, B., & Akalin, A. (2022). Multi-omics alleviates the limitations of panel sequencing for cancer drug response prediction. Cancers, 14(22), 5604. [CrossRef]
- Benboubker, V., Ramzy, G. M., Jacobs, S., & Nowak-Sliwinska, P. (2024). Challenges in validation of combination treatment strategies for CRC using patient-derived organoids. Journal of Experimental & Clinical Cancer Research, 43(1), 259. [CrossRef]
- Bierlaagh, M. C., Ramalho, A. S., Silva, I. A. L., et al. (2024). Repeatability and reproducibility of the forskolin-induced swelling (FIS) assay on intestinal organoids from people with cystic fibrosis. Journal of Cystic Fibrosis, 23(4), 693–702. [CrossRef]
- Brufsky, A. M., Hoskins, K. F., Conter, H. J., et al. (2025). MammaPrint predicts chemotherapy benefit in HR+HER2− early breast cancer: FLEX Registry real-world data. JNCI Cancer Spectrum, 9(5), pkaf079. [CrossRef]
- Cartry, J., Bedja, S., Boilève, A., et al. (2023). Implementing patient-derived organoids in functional precision medicine for patients with advanced colorectal cancer. Journal of Experimental & Clinical Cancer Research, 42(1), 281. [CrossRef]
- Chanez, B., Delaye, M., Fraunhoffer, N., Iovanna, J., Neuzillet, C., & Dusetti, N. (2025). Toward precision chemotherapy for pancreatic cancer guided by transcriptomic signatures. United European Gastroenterology Journal, 13(10), 1905–1912. [CrossRef]
- ClinicalTrials.gov. (2024). Vemurafenib, cobimetinib, and atezolizumab in advanced melanoma with BRAF V600 mutations (Identifier NCT03860376). https://clinicaltrials.gov/study/NCT03860376.
- Creighton, C. J. (2023). Gene expression profiles in cancers and their therapeutic implications. Cancer Journal, 29(1), 9–14. [CrossRef]
- Deng, H., Liu, Q., Tian, Z., Liu, G., Wang, S., Liang, D., Wang, Y., Han, Y., & Xie, S. (2025). Spatial transcriptomics uncovers immune-cell plasticity and dedifferentiation signatures in aggressive lung adenocarcinoma subtypes. Frontiers in Immunology, 16, 1620886. [CrossRef]
- Dong, M., Wang, L., Hu, N., Rao, Y., Wang, Z., & Zhang, Y. (2025). Integration of multi-omics approaches in exploring intra-tumoral heterogeneity. Cancer Cell International, 25(1), 317. [CrossRef]
- El-Sayed, M. M., Bianco, J. R., Li, Y., & Fabian, Z. (2024). Tumor-agnostic therapy—The final step forward in the cure for human neoplasms? Cells, 13(12), 1071. [CrossRef]
- Giliberto, M., Thimiri Govinda Raj, D. B., et al. (2022). Ex vivo drug sensitivity screening in multiple myeloma identifies drug combinations that act synergistically. Molecular Oncology, 16(6), 1241–1258. [CrossRef]
- Goda, R. Y., & Abdel-Aziz, A. K. (2025). Exploiting artificial intelligence in precision oncology: An updated comprehensive review. Journal of Translational Medicine, 23(1), 1397. [CrossRef]
- Goh, L. S. H., Thng, D. K. H., Ang, Y. L. E., Ho, D., Toh, T. B., & Wong, A. L. A. (2026). The functional imperative in high-grade glioma. Experimental & Molecular Medicine. Advance online publication. [CrossRef]
- Gonçalves, I. M., Carvalho, V., Rodrigues, R. O., Pinho, D., Teixeira, S. F. C. F., Moita, A., Hori, T., Kaji, H., Lima, R., & Minas, G. (2022). Organ-on-a-chip platforms for drug screening and delivery in tumor cells: A systematic review. Cancers, 14(4), 935. [CrossRef]
- Gu, Z., Yao, Y., Yang, G., et al. (2022). Pharmacogenomic landscape of head and neck squamous cell carcinoma informs precision oncology therapy. Science Translational Medicine, 14(661), eabo5987. [CrossRef]
- Hennig, L., Monecke, A., Hoang, N. A., Thieme, R., Prill, S., Hoffmeister, A., Tuennemann, J., Bechmann, I., Lordick, F., & Kallendrusch, S. (2025). Personalized drug stratification using endoscopic samples to assess ex vivo gastric cancer tissue susceptibility to chemotherapy and immune checkpoint inhibitors. Clinical and Experimental Medicine, 25(1), 188. [CrossRef]
- Hertz, D. L., Lustberg, M. B., & Sonis, S. (2023). Evolution of predictive risk factor analysis for chemotherapy-related toxicity. Supportive Care in Cancer, 31(10), 601. [CrossRef]
- Hurkmans, E. G. E., Brand, A. C. A. M., Verdonschot, J. A. J., Te Loo, D. M. W. M., & Coenen, M. J. H. (2022). Pharmacogenetics of chemotherapy treatment response and toxicities in patients with osteosarcoma: A systematic review. BMC Cancer, 22(1), 1326. [CrossRef]
- Ivanisevic, T., & Sewduth, R. N. (2023). Multi-omics integration for the design of novel therapies and the identification of novel biomarkers. Proteomes, 11(4), 34. [CrossRef]
- Ji, G., Yang, Q., Wang, S., Yan, X., Ou, Q., Gong, L., et al. (2024). Single-cell profiling of response to neoadjuvant chemo-immunotherapy in surgically resectable esophageal squamous cell carcinoma. Genome Medicine, 16(1), 49. [CrossRef]
- Jørgensen, J. T. (2023). Twenty-five years with companion diagnostics. Chinese Clinical Oncology, 12(6), 65. [CrossRef]
- Jovanovic, A., Bright, F. K., Sadeghi, A., et al. (2026). Large-scale testing of antimicrobial lethality at single-cell resolution predicts mycobacterial infection outcomes. Nature Microbiology. Advance online publication. [CrossRef]
- Kenmogne, V. L., Takundwa, M. M., Nweke, E. E., Monchusi, B., Dube, P., Maher, H., Du Toit, J., Philip-Cherian, V., Fru, P. N., & Thimiri Govinda Raj, D. B. (2025). The first-in-Africa ex vivo drug sensitivity testing platform identifies novel drug combinations for South African leukaemia patient cohort. Scientific Reports, 15(1), 9160. [CrossRef]
- Kim, H. H., Im, S., Kim, J., et al. (2026). QAL333’s antitumor activity and predictive modeling: Integrated transcriptomic-bioinformatic analysis reveals selective cytotoxicity and sensitivity determinants. Scientific Reports. Advance online publication. [CrossRef]
- Kolodziej, M., Czarny, J., Dyla, H., Rekawek, D., Zlotek, M., Wlodarz, A., Baranowicz, M., Kwiatkowska-Borowczyk, E., Milbrandt, O., Ramlau, R., et al. (2026). Breast cancer multicellular spheroid models—A tool for studying cancer biology: A possible platform for drug screening and personalized medicine. International Journal of Molecular Sciences, 27(3), 1314. [CrossRef]
- Ladan, M. M., Meijer, T. G., Verkaik, N. S., de Monye, C., Koppert, L. B., Oomen-de Hoop, E., van Deurzen, C. H. M., Kanaar, R., Nonnekens, J., van Gent, D. C., & Jager, A. (2023). Proof-of-concept study linking ex vivo sensitivity testing to neoadjuvant anthracycline-based chemotherapy response in breast cancer patients. NPJ Breast Cancer, 9(1), 80. [CrossRef]
- Ladan, M. M., Meijer, T. G., Verkaik, N. S., Komar, Z. M., van Deurzen, C. H. M., den Bakker, M. A., Kanaar, R., van Gent, D. C., & Jager, A. (2022). Functional ex vivo tissue-based chemotherapy sensitivity testing for breast cancer. Cancers, 14(5), 1252. [CrossRef]
- Lazar, V., Zhang, B., Magidi, S., Le Tourneau, C., Raymond, E., Ducreux, M., Bresson, C., Raynaud, J., Wunder, F., Onn, A., Felip, E., Tabernero, J., Batist, G., Kurzrock, R., Rubin, E., & Schilsky, R. L. (2023). A transcriptomics approach to expand therapeutic options and optimize clinical trials in oncology. Therapeutic Advances in Medical Oncology, 15, 17588359231156382. [CrossRef]
- Leiva, M. C., Gustafsson, A., Garre, E., Ståhlberg, A., Kovács, A., Helou, K., & Landberg, G. (2023). Patient-derived scaffolds representing breast cancer microenvironments influence chemotherapy responses in adapted cancer cells consistent with clinical features. Journal of Translational Medicine, 21(1), 924. [CrossRef]
- Liebers, N., Bruch, P. M., Terzer, T., et al. (2023). Ex vivo drug response profiling for response and outcome prediction in hematologic malignancies: The prospective non-interventional SMARTrial. Nature Cancer, 4(12), 1648–1659. [CrossRef]
- Liu, L., Wang, H., Chen, R., Song, Y., Wei, W., Baek, D., Gillin, M., Kurabayashi, K., & Chen, W. (2025). Cancer-on-a-chip for precision cancer medicine. Lab on a Chip, 25(14), 3314–3347. [CrossRef]
- Liu, X., Fang, J., Huang, S., Wu, X., Xie, X., Wang, J., Liu, F., Zhang, M., & Peng, Z. (2021). Tumor-on-a-chip: From bioinspired design to biomedical application. Microsystems & Nanoengineering, 7, 50. [CrossRef]
- Lu, E. H., Rusyn, I., & Chiu, W. A. (2025). Incorporating new approach methods (NAMs) data in dose-response assessments: The future is now! Journal of Toxicology and Environmental Health, Part B: Critical Reviews, 28(1), 28–62. [CrossRef]
- Lu, Y., Huang, W., Li, Y., Xu, Y., Wei, Q., Sha, C., & Guo, P. (2025). Leveraging artificial intelligence in antibody-drug conjugate development: From target identification to clinical translation in oncology. NPJ Precision Oncology, 9(1), 374. [CrossRef]
- Meghdadi, B., Al-Holou, W. N., Scott, A. J., et al. (2026). Digital twins for in vivo metabolic flux estimations in patients with brain cancer. Cell Metabolism, 38(1), 228–246.e17. [CrossRef]
- Mhandire, D. Z., & Goey, A. K. L. (2022). The value of pharmacogenetics to reduce drug-related toxicity in cancer patients. Molecular Diagnosis & Therapy, 26(2), 137–151. [CrossRef]
- Miteva-Marcheva, N. N., Ivanov, H. Y., Dimitrov, D. K., & Stoyanova, V. K. (2020). Application of pharmacogenetics in oncology. Biomarker Research, 8, 32. [CrossRef]
- Motohashi, S., Katsuta, E., & Ban, D. (2025). Advances and challenges in drug screening for cancer therapy: A comprehensive review. Bioengineering, 12(12), 1315. [CrossRef]
- Mundi, P. S., Dela Cruz, F. S., Grunn, A., Diolaiti, D., Mauguen, A., Rainey, A. R., et al. (2023). A transcriptome-based precision oncology platform for patient-therapy alignment in a diverse set of treatment-resistant malignancies. Cancer Discovery, 13(6), 1386–1407. [CrossRef]
- Murciano-Goroff, Y. R., Suehnholz, S. P., Drilon, A., & Chakravarty, D. (2023). Precision oncology: 2023 in review. Cancer Discovery, 13(12), 2525–2531. [CrossRef]
- Nafchi, H. M., Solatzadeh, H., Hajimaghsoudi, E., & Babakhanzadeh, E. (2025). Personalizing cancer therapy: The role of pharmacogenetics in overcoming drug resistance and toxicity. Molecular Biology Reports, 52(1), 785. [CrossRef]
- O’Reilly, S., Gaynor, S., & Mayer, E. L. (2025). Oncology’s ongoing struggle with evaluating quality-of-life–limiting treatment-related toxicity. Supportive Care in Cancer, 33(11), 943. [CrossRef]
- Qiao, D., Wang, R. C., & Wang, Z. (2025). Precision oncology: Current landscape, emerging trends, challenges, and future perspectives. Cells, 14(22), 1804. [CrossRef]
- Qin, G., Knijnenburg, T. A., Gibbs, D. L., Moser, R., Monnat, R. J., Jr., Kemp, C. J., & Shmulevich, I. (2022). A functional module states framework reveals transcriptional states for drug and target prediction. Cell Reports, 38(3), 110269. [CrossRef]
- Qu, S., Xu, R., Yi, G., Li, Z., Zhang, H., Qi, S., & Huang, G. (2024). Patient-derived organoids in human cancer: A platform for fundamental research and precision medicine. Molecular Biomedicine, 5(1), 6. [CrossRef]
- Reizine, N. M., & O’Donnell, P. H. (2022). Modern developments in germline pharmacogenomics for oncology prescribing. CA: A Cancer Journal for Clinicians, 72(4), 315–332. [CrossRef]
- Ren, F., Wang, L., Wang, Y., Wang, J., Wang, Y., Song, X., Zhang, G., Nie, F., & Lin, S. (2024). Single-cell transcriptome profiles the heterogeneity of tumor cells and microenvironments for different pathological endometrial cancer and identifies specific sensitive drugs. Cell Death & Disease, 15(8), 571. [CrossRef]
- Saito, K., & Narukawa, M. (2025). Bridging gaps in oncology: Comparative analysis of development and approval pathways for anticancer drugs and companion diagnostics in the United States and Japan. Clinical and Translational Science, 18(2), e70162. [CrossRef]
- Sánchez-Bayona, R., Catalán, C., Cobos, M. A., & Bergamino, M. (2025). Pharmacogenomics in solid tumors: A comprehensive review of genetic variability and its clinical implications. Cancers, 17(6), 913. [CrossRef]
- Schmelz, K., Toedling, J., Huska, M., et al. (2021). Spatial and temporal intratumour heterogeneity has potential consequences for single biopsy-based neuroblastoma treatment decisions. Nature Communications, 12(1), 6804. [CrossRef]
- Schoonbeek, M. C., Gestraud, P., Vernooij, L., et al. (2026). Ex vivo drug sensitivity profiling to complement molecular profiling in pediatric precision oncology. NPJ Precision Oncology. Advance online publication. [CrossRef]
- Spiró, Z., El-Heliebi, A., Mair, M. J., Pieber, T. R., Prietl, B., Spiegl-Kreinecker, S., Stanzer, S., Wöhrer, A., Preusser, M., & Berghoff, A. S. (2025). Ex vivo drug screening on patient-derived tumor material to advance functional precision in oncology: An overview on current approaches and unresolved challenges. Cancer Treatment Reviews, 143, 103072. [CrossRef]
- Struyf, N., Österroos, A., Vesterlund, M., et al. (2024). Delineating functional and molecular impact of ex vivo sample handling in precision medicine. NPJ Precision Oncology, 8(1), 38. [CrossRef]
- Sundaresan, S., Lavanya, S. K., & Manickam, M. (2024). Emerging molecular technology in cancer testing. EJIFCC, 35(3), 142–153.
- Taurin, S., Alzahrani, R., Aloraibi, S., Ashi, L., Alharmi, R., & Hassani, N. (2025). Patient-derived tumor organoids: A preclinical platform for personalized cancer therapy. Translational Oncology, 51, 102226. [CrossRef]
- The, S., Schnepp, P. M., Shelley, G., Keller, J. M., Rao, A., & Keller, E. T. (2023). Integration of single-cell RNA-sequencing and network analysis to investigate mechanisms of drug resistance. Methods in Molecular Biology, 2660, 85–94. [CrossRef]
- Tilsed, C. M., Fisher, S. A., Nowak, A. K., Lake, R. A., & Lesterhuis, W. J. (2022). Cancer chemotherapy: Insights into cellular and tumor microenvironmental mechanisms of action. Frontiers in Oncology, 12, 960317. [CrossRef]
- Tomasik, B., Garbicz, F., Braun, M., Bieńkowski, M., & Jassem, J. (2023). Heterogeneity in precision oncology. Cambridge Prisms: Precision Medicine, 2, e2. [CrossRef]
- Tsimberidou, A. M., Fountzilas, E., Bleris, L., & Kurzrock, R. (2022). Transcriptomics and solid tumors: The next frontier in precision cancer medicine. Seminars in Cancer Biology, 84, 50–59. [CrossRef]
- Tsukamoto, Y., Hirashita, Y., Shibata, T., Fumoto, S., Kurogi, S., Nakada, C., Kinoshita, K., Fuchino, T., Murakami, K., Inomata, M., Moriyama, M., & Hijiya, N. (2023). Patient-derived ex vivo cultures and endpoint assays with surrogate biomarkers in functional testing for prediction of therapeutic response. Cancers, 15(16), 4104. [CrossRef]
- Valla, V., Alzabin, S., Koukoura, A., Lewis, A., Nielsen, A. A., & Vassiliadis, E. (2021). Companion diagnostics: State of the art and new regulations. Biomarker Insights, 16, 11772719211047763. [CrossRef]
- Waheed, Z., Bunka, M., Edwards, L., Bousman, C., Hoens, A., Austin, J. J., & Bryan, S. (2025). Characterizing models for delivery of pharmacogenomic testing: A scoping review. Pharmacogenomics, 26(13–14), 513–527. [CrossRef]
- Wang, Y., Chang, X., Deng, S., Tang, S., & Chen, P. (2026). Functional material probes and advanced technologies in organ-on-a-chip characterization. Theranostics, 16(5), 2488–2516. [CrossRef]
- Wegmann, R., et al. (2024). Single-cell landscape of innate and acquired drug resistance in acute myeloid leukemia. Nature Communications, 15(1), 9402. [CrossRef]
- Wu, Y., Chen, M., & Qin, Y. (2025). Anticancer drug response prediction integrating multi-omics pathway-based difference features and multiple deep learning techniques. PLoS Computational Biology, 21(3), e1012905. [CrossRef]
- Xu, R., Shao, J., Liu, J., Qu, B., & Liu, J. (2024). Single-cell analysis of chemotherapy-resistant microenvironment identifies a chemo-response biomarker for pancreatic cancer. Journal of Gastrointestinal Oncology, 15(4), 1836–1846. [CrossRef]
- Zhang, Y., Ji, M., Zhao, J. Y., Wang, H. F., Wang, C. W., Li, W., Ye, J. J., Lu, F., Lin, L. H., Gao, Y. T., Jin, J., Li, L., Ji, C. Y., Ballesteros, J., & Zhu, H. H. (2022). Ex vivo chemosensitivity profiling of acute myeloid leukemia and its correlation with clinical response and outcome to chemotherapy. Frontiers in Oncology, 11, 793773. [CrossRef]
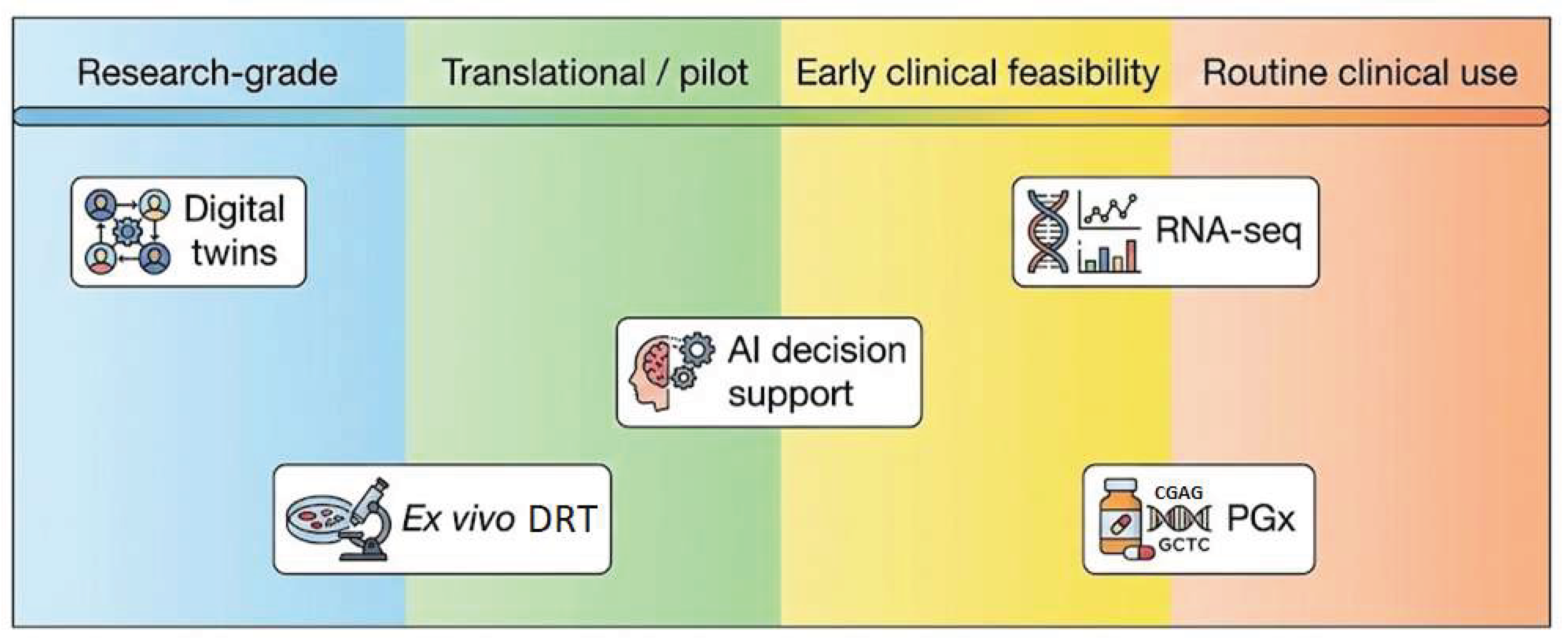
| Technology | Current Status (2026) | Clinical Use |
|---|---|---|
| PDO drug sensitivity testing | Pilot clinical feasibility | Select centers / trials |
| Tumor slice cultures | Research–pilot | Limited |
| Digital twins | Research-grade | Experimental |
| High-throughput DST platforms | Early clinical feasibility | Non-routine |
| Stage | Data Type | Functional Role in Integration |
|---|---|---|
| Prediction | RNA-seq | Identifies regulatory states and potential resistance pathways (e.g., efflux, bypass signaling) |
| Metabolic Context | PGx Profiling | Anticipates systemic processing of drugs (clearance/activation rates) based on genetic variants |
| Validation | Ex Vivo Assays | Confirms actual viability inhibition and drug hits in living cancer tissue within a defined window |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




