Submitted:
20 February 2026
Posted:
26 February 2026
You are already at the latest version
Abstract
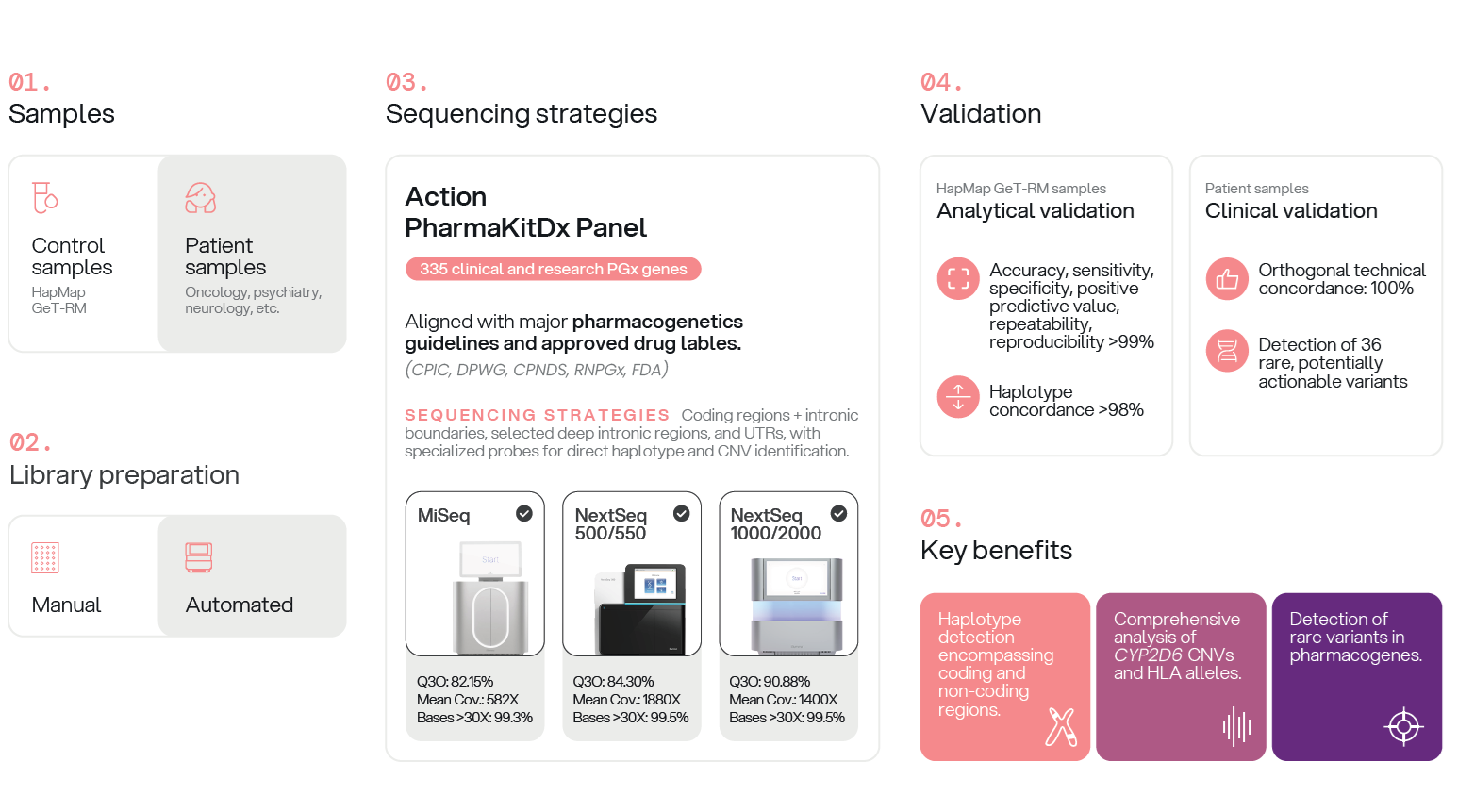
Keywords:
1. Introduction
2. Results
2.1. Sequencing Performance
2.2. Analytical Performance
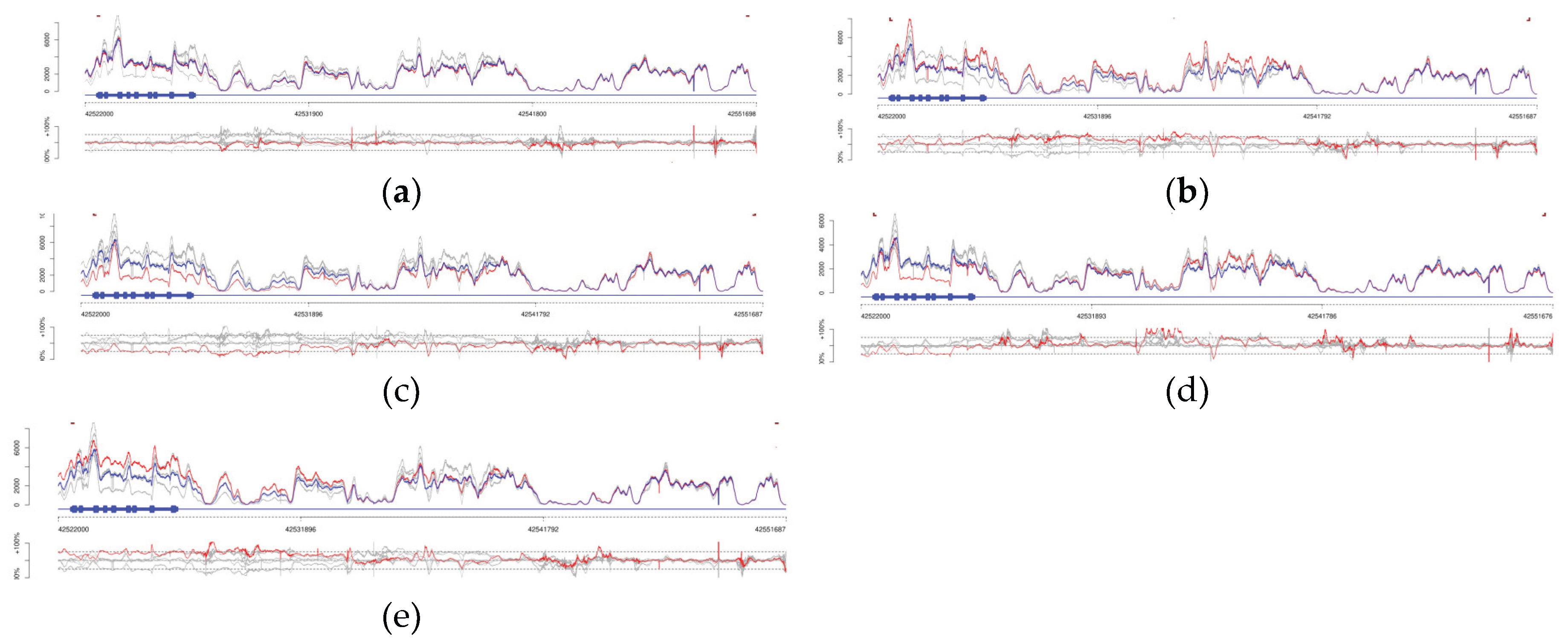
2.3. Pharmacogenetic Haplotypes
2.4. Clinical Feasibility
3. Discussion
Genomic Coverage and Design Strategies
Sequencing Performance and Coverage
Analytical Accuracy and Validation
Bioinformatics and Clinical Interpretation
Clinical Validation with Patient Samples
Detection of Rare and Additional Variants
Clinical Implications and Future Perspectives
4. Materials and Methods
Panel Design
Sample Selection
DNA Extraction
Bioinformatic Pipeline
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
Abbreviations
| ADME | Absorption, Distribution, Metabolism, and Excretion |
| AEMPS | Agencia Española de Medicamentos y Productos Sanitarios |
| CNV | Copy Number Variation |
| CPNDS | Canadian Pharmacogenomics Network for Drug Safety |
| CPIC | Clinical Pharmacogenetics Implementation Consortium |
| DPWG | Dutch Pharmacogenetics Working Group |
| EMA | European Medicines Agency |
| FDA | U.S. Food and Drug Administration |
| GeT-RM | Genetic Testing Reference Materials Coordination Program |
| HLA | Human Leukocyte Antigen |
| IVDD | In Vitro Diagnostic Directive |
| NGS | Next-Generation Sequencing |
| NPA | Negative Percent Agreement |
| PCR | Polymerase Chain Reaction |
| PGx | Pharmacogenomics |
| PPA | Positive Percent Agreement |
| PPV | Positive Predictive Value |
| Q30 | Phred Quality Score ≥30 |
| SNV | Single Nucleotide Variant |
| STR | Short Tandem Repeat |
| SV | Structural Variant |
| UTR | Untranslated Region |
| WES | Whole-Exome Sequencing |
| WGS | Whole-Genome Sequencing |
Appendix A. Supplementary Methods
Appendix A.1. Target Enrichment and Capture Design
Appendix A.1.1. Full Coding Regions with Extended Intronic and Regulatory Coverage (23 Genes)
- All coding exons
- ±50 bp of flanking intronic sequence
- Selected deep intronic regions containing known pathogenic variants
- Relevant 5′ and 3′ untranslated regions (UTRs)
Appendix A.1.2. Direct Haplotype Capture for Highly Polymorphic Genes (6 Genes)
- CYP2D6
- HLA-B, HLA-C, HLA-DRB1, HLA-E, HLA-G
Appendix A.1.3. Targeted Capture of Clinically Relevant Loci (307 Genes)
- Coding exons and selected noncoding regions containing well-established pharmacogenetic variants
- Specific loci documented in curated databases or medical literature
Appendix A.1.4. Total Gene Content
ABCC6, ABCC8, ABCC9, ABCG1, ABCG2, ACE, ACKR1, ACYP2, ADA,
ADD1, ADGRB3, ADGRL2, ADH1A, ADH1B, ADH1C, ADORA2A, ADRB1,
ADRB2, AFF3, AHR, AKR1E2, ALDH1A1, ALDH2, ALOX5, ALPL, AMPD1,
ANKK1, APOA5, APOE, ARL14, ATIC, ATP7A, BDNF, BMP2, BMP7,
BRINP1, BTG1, C11orf65, C1orf167, C8orf34, CACNA1S, CADM2, CALU,
CARD8, CBR3, CCDC179, CCHCR1, CCR5, CD2, CD226, CD69, CD80, CD84,
CD86, CDA, CEP72, CES1, CES2, CETP, CHRNA3, CHRNB2, CHST7,
CHUK, CLEC2D, CLSTN2, CNTN5, COL22A1, COMT, COQ2, CREB1,
CRHR1, CRHR2, CRP, CSMD1, CSMD3, CUL1, CYP17A1, CYP19A1,
CYP1A1, CYP1A2, CYP1B1, CYP2A13, CYP2A6, CYP2B6, CYP2C18,
CYP2C19, CYP2C8, CYP2C9, CYP2D6, CYP2E1, CYP2F1, CYP2J2, CYP2S1,
CYP3A4, CYP3A43, CYP3A5, CYP3A7, CYP4B1, CYP4F2, DCK, DHFR,
DHODH, DLG2, DOCK1, DPP6, DPYD, DRD1, DRD2, DRD3, DRD4,
DYNC2H1, EGF, EGLN3, EMCN, ENOX1, EPHX1, ERCC1, ESR1, ESR2,
ETV1, EYA4, F5, FAAH, FAR1, FCGR2A, FCGR2B, FCGR3A, FDPS, FKBP5,
FLT3, FMO2, FMO3, FOXP3, FPGS, FTO, G6PD, GABBR2, GALNT18,
GATM, GFRA1, GFRA2, GGCX, GGH, GNB3, GP1BA, GRIK4, GSTM1,
GSTP1, GSTT1, HAS3, HLA-A, HLA-B, HLA-C, HLA-DRB1, HLA-E, HLA-G,
HMGCR, HTR1A, HTR2A, HTR2C, HTR3A, IFNG, IFNGR2, IFNL3, IFNL4,
IGHM, IKBKB, IL10, IL12B, IL17A, IL17F, IL18, IL1B, IL1R1, IL2, IL4R, IL6,
IL6R, IRAK3, ITGB7, ITPA, KCNE1, KCNH2, KCNIP1, KCNJ11, KIAA0391,
KIF6, KLRC1, KLRD1, LMO3, LMX1A, LPA, LRPAP1, LRRC55, LTC4S,
MAP2K6, MAP3K1, MAP3K14, MAP3K7, MAPK14, MAPKAPK2, MAPT,
MC4R, MED15, MICA, MMP20, MRPL36, MS4A1, MTHFD1, MTHFR,
MTR, MTRR, MYD88, NAT1, NAT2, NAV2, NEBL, NEDD4L, NFIB,
NFKBIA, NFKBIB, NFKBIE, NLRP3, NQO1, NR1I2, NR2F2, NR3C1, NT5C2,
NTRK2, NUBPL, NUDT15, OPRD1, OPRM1, OR2B11, OSR1, P2RY1,
P2RY12, PDZD2, PLA2G4A, PNPLA3, PRELID3B, PRKCA, PRSS53, PTGFR,
PTGIS, PTGS1, PTGS2, PTPRC, PTPRM, QKI, RAD51B, RHOBTB1,
RPS6KA5, RYR1, SCN1A, SCN5A, SCNN1A, SEMA3C, SHMT1, SLC15A1,
SLC15A2, SLC16A7, SLC19A1, SLC22A1, SLC22A11, SLC22A2, SLC22A6,
SLC28A3, SLC46A1, SLC6A4, SLC9A7, SLCO1A2, SLCO1B1, SLCO1B3,
SLCO1C1, SLCO2B1, SOD2, SP3, SRY, STAT4, STRBP, SULT1A1, SULT1A2,
SYNM, TANC1, TBXAS1, TCF7L2, TEC, TGFB1, TGFBR2, TLR1, TLR10,
TLR2, TLR4, TLR5, TNF, TNFAIP3, TNFRSF1A, TNFRSF1B, TNFSF13B,
TP53, TPMT, TRAF1, TUBB2A, TYMS, UGT1A1, UGT1A10, UGT1A3,
UGT1A4, UGT1A5, UGT1A6, UGT1A7, UGT1A8, UGT1A9, UGT2B10,
UGT2B15, UGT2B17, UGT2B7, UMPS, VDR, VKORC1, WDR27, XPC,
XRCC1, YEATS4, ZNF595.
Appendix A.2. Pharmacogenetic Alleles Included in the Panel
ABCG2 [rs2231142]
CACNA1S [c.1589G>A/p.Arg530His, c.1598G>A/p.Arg533His]
CALU [c.606+133A>G (rs339097)]
COMT [c.615+310C>T (rs4646316)]
CYP1A2 [*1A, *1C, *1D, *1F, *1K, *1L, *2, *3, *4, *5, *6, *7, *8, *11, *15, *16]
CYP2B6 [*2, *3, *4, *5, *6, *7, *8, *9, *11, *12, *13, *14, *15, *16, *18, *19, *20,
*21, *22, *26, *27, *28]
CYP2C18 [rs12777823]
CYP2C19 [*2A, *2B, *3, *4, *5, *6, *7, *8, *9, *10, *12, *13, *14, *15, *17]
CYP2C9 [*2, *3, *4, *5, *6, *9, *10, *11, *12, *13, *15, *16, *25]
CYP2D6 [*2, *3, *4, *5, *6, *7, *8, *9, *10A, *11, *12, *14A, *14B, *15, *17, *18,
*19, *20, *21A, *29, *38, *40, *41, *42, *44, *56A, *56B, *64, x2, xN, hyb]
CYP3A4 [*1B, *2, *3, *4, *5, *6, *7, *8, *10, *11, *12, *13, *14, *15A, *15B, *16,
*17, *18, *19, *20, *22]
CYP3A5 [*1A, *2, *3B, *3C, *3D, *3F, *3G, *3K, *3L, *4, *5, *6, *7, *8, *9]
CYP4F2 [*1, *2, *3]
DPYD [c.1024G>A (rs183385770), c.1057C>T (rs143154602), c.1129-5923C>G,
c.1236G>A (HapB3), c.1156G>T (*12), c.1314T>G (rs186169810),
c.1484A>G (rs111858276), c.1679T>G (*13), c.1774C>T (rs59086055),
c.1775G>A (rs138616379), c.1777G>A (rs145773863), c.1898delC (*3),
c.1905+1G>A (*2A), c.2021G>A (rs137999090), c.2279C>T (rs112766203),
c.2639G>T (rs55674432), c.2656C>T (rs147545709), c.2846A>T (rs67376798),
c.2872A>G (rs141044036), c.2933A>G (rs72547601), c.295_298delTCAT (*7),
c.2983G>T (*10), c.557A>G (rs115232898), c.601A>C (rs72549308),
c.61C>T (rs72549310), c.632A>G (rs72549307), c.703C>T (*8),
c.868A>G (rs146356975)]
F5 [Factor V Leiden]
G6PD [202G>A_376A>G_1264C>G, A, A- 202A_376G, Aachen, Abeno,
Acrokorinthos, Alhambra, Amazonia, Amiens, Amsterdam, Anadia, Ananindeua,
Andalus, Arakawa, Asahi, Asahikawa, Aures, Aveiro, B (wildtype),
Bajo Maumere, Bangkok, Bangkok Noi, Bao Loc, Bari, Belem, Beverly Hills,
Genova, Iwate, Niigata, Yamaguchi, Brighton, Buenos Aires, Cairo,
Calvo Mackenna, Campinas, Canton, Taiwan-Hakka, Gifu-like, Agrigento-like,
Cassano, Chatham, Chikugo, Chinese-1, Chinese-5, Cincinnati, Cleveland Corum,
Clinic, Coimbra Shunde, Cosenza, Costanzo, Covao do Lobo, Crispim, Dagua,
Durham, Farroupilha, Figuera da Foz, Flores, Fukaya, Fushan,
G6PD A- 680T_376G, G6PD A- 968C_376G, G6PDNice, Gaohe, Georgia, Gidra,
Gond, Guadalajara, Guangzhou, Haikou, Hammersmith, Harilaou, Harima,
Hartford, Hechi, Hermoupolis, Honiara, Ierapetra, Ilesha, Insuli, Iowa,
Walter Reed, Springfield, Iwatsuki, Japan, Shinagawa, Kaiping, Anant, Dhon,
Sapporo-like, Wosera, Kalyan-Kerala, Jamnaga, Rohini, Kambos, Kamiube,
Keelung, Kamogawa, Kawasaki, Kozukata, Krakow, La Jolla, Lages, Lagosanto,
Laibin, Lille, Liuzhou, Loma Linda, Ludhiana, Lynwood, Madrid, Mahidol,
Malaga, Manhattan, Mediterranean Haplotype, Mediterranean, Dallas, Panama‚
Sassari, Cagliari, Birmingham, Metaponto, Mexico City, Miaoli, Minnesota,
Marion, Gastonia, LeJeune, Mira d’Aire, Mizushima, Montalbano, Montpellier,
Mt Sinai, Munich, Murcia Oristano, Musashino, Namouru, Nankang, Nanning,
Naone, Nara, Nashville, Anaheim, Portici, Neapolis, Nilgiri, No name,
North Dallas, Olomouc, Omiya, Orissa, Osaka, Palestrina, Papua, Partenope,
Pawnee, Pedoplis-Ckaro, Piotrkow, Plymouth, Praha, Puerto Limon, Quing Yan,
Radlowo, Rehevot, Rignano, Riley, Riverside, Roubaix, S. Antioco,
Salerno Pyrgos, Santa Maria, Santiago, Santiago de Cuba, Morioka, Sao Borja,
Seattle, Lodi, Modena, Ferrara II, Athens-like, Seoul, Serres, Shenzen, Shinshu,
Sibari, Sierra Leone, Sinnai, Songklanagarind, Split, Stonybrook, Sugao, Sumare,
Sunderland, Surabaya, Suwalki, Swansea, Taipei‚ Chinese-3, Telti/Kobe,
Tenri, Tokyo, Fukushima, Toledo, Tomah, Tondela, Torun, Tsukui, Ube Konan,
Union, Maewo, Chinese-2, Kalo, Urayasu, Utrecht, Valladolid, Vancouver,
Vanua Lava, Viangchan, Jammu, Villeurbanne, Volendam, Wayne, West Virginia,
Wexham, Wisconsin, Yunan]
GGCX [c.2084+45G>C (rs11676382))]
HLA-A [c.*66A>T (rs1061235-T) (*31:01)]
HLA-B [base de datos IMGT/HLA v 3.12 compuesta 2932 alelos]
IFNL4 [c.151-152G>A (rs12979860)]
NAT1 [*4, *5, *11, *11C, *14, *15, *17, *19A, *19B, *22, *23, *27, *30]
NAT2 [*4, *5A, *5E, *6A, *6J, *7A, *7D, *10, *12D, *14A, *14D, *14F, *17, *18, *19]
NUDT15 [*2, *3, *4, *5, *6, *7, *8, *9, *10, *11, *12, *13, *14, *15, *16, *17, *18, *19]
RYR1 [p.Ala2350Thr, p.Ala2428Thr, p.Arg163Cys, p.Arg163Leu, p.Arg2163Cys,
p.Arg2163His, p.Arg2336His, p.Arg2355Trp, p.Arg2435His, p.Arg2452Trp,
p.Arg2454Cys, p.Arg2454His, p.Arg2458Cys, p.Arg2458Leu, p.Arg2508Cys,
p.Arg2508Gly, p.Arg2508His, p.Arg328Trp, p.Arg401Gly, p.Arg44Cys,
p.Arg4861His, p.Arg530His, p.Arg533His, p.Arg533Ser, p.Arg552Trp,
p.Arg614Cys, p.Arg614Leu, p.Glu2348del, p.Glu3104Lys, p.Gly2375Ala,
p.Gly2434Arg, p.Gly248Arg, p.Gly341Arg, p.Gly3990Val, p.His4833Tyr,
p.Ile403Met, p.Ile4898Thr, p.Leu4838Val, p.Thr2206Arg, p.Thr2206Met,
p.Thr4826Ile, p.Tyr4796Cys, p.Tyr522Ser, p.Val2168Met, p.Val4849Ile]
SLC28A3 [c.1381C>T, (rs7853758)]
SLCO1B1 [*1B, *2, *3, *4, *5, *6, *7, *8, *9, *11, *13, *14, *15, *16, *17, *18, *21, *31]
TPMT [*1, *2, *3A, *3B, *3C, *3D, *4, *8, *24]
UGT1A1 [*1, *27, *28, *36, *37, *6, *60, *93]
UGT1A6 [*1, *2, *3A, *4A, *4b, *5]
VKORC1 [c.-1639G>A (rs9923231)].
Appendix A.3. Analytical Performance Metrics
Appendix A.4. Rare Variant Filtering and Prioritization Workflow
Appendix A.4.1. Quality-Based Filtering
- Depth of coverage ≥ 30
- Base quality ≥ 100
-
Allele fraction thresholds adapted to zygosity:
- a.
- Heterozygous variants ≥ 0.25
- b.
- Homozygous variants ≥ 0.80
- c.
- Variants with undefined zygosity were conservatively required to meet ≥ 0.25
Appendix A.4.2. Population Frequency Filtering
- gnomAD
- ExAC
- 1000 Genomes
- 5000 Exomes
- dbSNP frequency annotations
Appendix A.4.3. Functional Annotation Filtering
- Located in coding regions or exons
- Caused a non-synonymous amino acid change, including missense, nonsense, frameshift, indels, or disruptions affecting the reading frame
-
Annotated as splicing-relevant, either through:
- d.
- Explicit splicing flags in the annotation, or
- e.
- NNSPLICE predictions indicating ≥10% reduction in splice-site strength
Appendix A.4.4. Pathogenicity Predictor Filtering
- DANN score ≥ 0.9
- FATHMM coding-group classification containing “damaging” or “deleterious”
- MutationTaster prediction = “D” (disease causing)
Appendix A.4.5. ClinVar-Based Filtering
- Variants classified as pathogenic, likely pathogenic, conflicting, or VUS were retained.
- Variants labelled benign or likely benign were excluded unless at least one in silico predictor suggested deleteriousness (as defined above).
- Variants with no ClinVar annotation passed this step by default.
Appendix A.4.6. Final Variant Set
References
- Kabbani, D.; Akika, R.; Wahid, A.; Daly, A.K.; Cascorbi, I.; Zgheib, N.K. Pharmacogenomics in Practice: A Review and Implementation Guide. Front. Pharmacol. 2023, 14. [Google Scholar] [CrossRef] [PubMed]
- Russell, L.; Zhou, Y.; Almousa, A.A.; Sodhi, J.K.; Nwabufo, C.K.; Lauschke, V.M. Pharmacogenomics in the Era of next Generation Sequencing – from Byte to Bedside. Drug Metab. Rev. 2021, 53, 253–278. [Google Scholar] [CrossRef]
- Schwarz, U.I.; Gulilat, M.; Kim, R.B. The Role of Next-Generation Sequencing in Pharmacogenetics and Pharmacogenomics. Cold Spring Harb. Perspect. Med. 2018, 9. [Google Scholar] [CrossRef] [PubMed]
- Klein, K.; Tremmel, R.; Winter, S.; Fehr, S.; Battke, F.; Scheurenbrand, T.; Schaeffeler, E.; Biskup, S.; Schwab, M.; Zanger, U.M. A New Panel-Based Next-Generation Sequencing Method for ADME Genes Reveals Novel Associations of Common and Rare Variants With Expression in a Human Liver Cohort. Front. Genet. 2019, 10. [Google Scholar] [CrossRef]
- McInnes, G.; Lavertu, A.; Sangkuhl, K.; Klein, T.E.; Whirl-Carrillo, M.; Altman, R.B. Pharmacogenetics at Scale: An Analysis of the UK Biobank. Clin. Pharmacol. Ther. 2020, 109, 1528–1537. [Google Scholar] [CrossRef]
- Chen, X.; Shen, F.; Gonzaludo, N.; Malhotra, A.; Rogert, C.; Taft, R.J.; Bentley, D.; Eberle, M.A. Cyrius: Accurate CYP2D6 Genotyping Using Whole-Genome Sequencing Data. Pharmacogenomics J. 2021, 21, 251–261. [Google Scholar] [CrossRef]
- Luo, S.; Jiang, R.; Grzymski, J.J.; Lee, W.; Lu, J.T.; Washington, N.L. Comprehensive Allele Genotyping in Critical Pharmacogenes Reduces Residual Clinical Risk in Diverse Populations. Clin. Pharmacol. Ther. 2021, 110, 759–767. [Google Scholar] [CrossRef]
- Turner, A.; Derezinski, A.D.; Gaedigk, A.; Berres, M.E.; Gregornik, D.; Brown, K.; Broeckel, U.; Scharer, G. Characterization of Complex Structural Variation in the CYP2D6-CYP2D7-CYP2D8 Gene Loci Using Single-Molecule Long-Read Sequencing. Front. Pharmacol. 2023, 14. [Google Scholar] [CrossRef]
- Caspar, S.M.; Schneider, T.; Meienberg, J.; Matyas, G. Added Value of Clinical Sequencing: WGS-Based Profiling of Pharmacogenes. Int. J. Mol. Sci. 2020, 21. [Google Scholar] [CrossRef]
- Caspar, S.M.; Schneider, T.; Stoll, P.; Meienberg, J.; Mátyás, G. Potential of Whole-Genome Sequencing-Based Pharmacogenetic Profiling. Pharmacogenomics 2021, 22, 177–190. [Google Scholar] [CrossRef] [PubMed]
- Gulilat, M.; Lamb, T.; Teft, W.A.; Wang, J.; Dron, J.S.; Robinson, J.F.; Tirona, R.G.; Hegele, R.A.; Kim, R.B.; Schwarz, U.I. Targeted next Generation Sequencing as a Tool for Precision Medicine. BMC Med. Genomics 2019, 12, 81. [Google Scholar] [CrossRef]
- Lee, S.B.; Shin, J.Y.; Kwon, N.J.; Kim, C.; Seo, J.S. ClinPharmSeq: A Targeted Sequencing Panel for Clinical Pharmacogenetics Implementation. PLoS ONE 2022, 17. [Google Scholar] [CrossRef]
- van der Lee, M.; Kriek, M.; Guchelaar, H.-J.; Swen, J.J. Technologies for Pharmacogenomics: A Review. Genes 2020, 11, 1456. [Google Scholar] [CrossRef]
- David, V.; Fylan, B.; Bryant, E.; Smith, H.; Sagoo, G.S.; Rattray, M. An Analysis of Pharmacogenomic-Guided Pathways and Their Effect on Medication Changes and Hospital Admissions: A Systematic Review and Meta-Analysis. Front. Genet. 2021, 12. [Google Scholar] [CrossRef] [PubMed]
- Morris, S.A.; Alsaidi, A.T.; Verbyla, A.; Cruz, A.; Macfarlane, C.; Bauer, J.; Patel, J.N. Cost Effectiveness of Pharmacogenetic Testing for Drugs with Clinical Pharmacogenetics Implementation Consortium (CPIC) Guidelines: A Systematic Review. Clin. Pharmacol. Ther. 2022, 112, 1318–1328. [Google Scholar] [CrossRef]
- Abdullah-Koolmees, H.; van Keulen, A.M.; Nijenhuis, M.; Deneer, V.H.M. Pharmacogenetics Guidelines: Overview and Comparison of the DPWG, CPIC, CPNDS, and RNPGx Guidelines. Front. Pharmacol. 2020, 11, 595219. [Google Scholar] [CrossRef] [PubMed]
- Pritchard, D.; Patel, J.N.; Stephens, L.E.; McLeod, H.L. Comparison of FDA Table of Pharmacogenetic Associations and Clinical Pharmacogenetics Implementation Consortium Guidelines. Am. J. Health-Syst. Pharm. AJHP Off. J. Am. Soc. Health-Syst. Pharm. 2022, 79, 993–1005. [Google Scholar] [CrossRef]
- Action PharmaKit Manual-RUO. Health Code.
- Ramudo-Cela, L.; López-Martí, J.M.; Colmeiro-Echeberría, D.; De-Uña-Iglesias, D.; Santomé-Collazo, J.L.; Monserrat-Iglesias, L. Development and Validation of a Next-Generation Sequencing Panel for Clinical Pharmacogenetics. Farm. Hosp. Organo Of. Expresion Cient. Soc. Espanola Farm. Hosp. 2020, 44, 243–253. [Google Scholar] [CrossRef]
- Gaedigk, A.; Sangkuhl, K.; Whirl-Carrillo, M.; Twist, G.P.; Klein, T.E.; Miller, N.A. The Evolution of PharmVar. Clin. Pharmacol. Ther. 2019, 105, 29–32. [Google Scholar] [CrossRef]
- Whirl-Carrillo, M.; Huddart, R.; Gong, L.; Sangkuhl, K.; Thorn, C.F.; Whaley, R.; Klein, T.E. An Evidence-Based Framework for Evaluating Pharmacogenomics Knowledge for Personalized Medicine. Clin. Pharmacol. Ther. 2021, 110, 563–572. [Google Scholar] [CrossRef] [PubMed]
- De-Uña, D. System and Method to Detect Structural Genetic Variants 2019.
- Ramudo-Cela, L.; López-Martí, J.M.; Colmeiro-Echeberría, D.; De-Uña-Iglesias, D.; Santomé-Collazo, J.L.; Monserrat-Iglesias, L. Development and Validation of a Next-Generation Sequencing Panel for Clinical Pharmacogenetics. Farm. Hosp. Organo Of. Expresion Cient. Soc. Espanola Farm. Hosp. 2020, 44, 243–253. [Google Scholar] [CrossRef]
- Wittig, M.; Anmarkrud, J.A.; Kassens, J.C.; Koch, S.; Forster, M.; Ellinghaus, E.; Hov, J.R.; Sauer, S.; Schimmler, M.; Ziemann, M.; et al. Development of a High-Resolution NGS-Based HLA-Typing and Analysis Pipeline. Nucleic Acids Res. 2015, 43, e70. [Google Scholar] [CrossRef]
| Parameters | MiSeq (Manual workflow) |
MiSeq (Automated workflow) |
NextSeq 500/550 (Automated workflow) |
NextSeq 1000/2000 (Automated workflow) | |
|---|---|---|---|---|---|
| Sequencing quality metrics | Q30 (%) | 82.2 | 93.4 | 84.3 | 90.9 |
| Clusters passing filter (%) | 82.2 | 89.1 | 89.4 | NA | |
| Mean coverage (×) | 582 | 926 | 1,88 | 1,4 | |
| Bases >30× (%) | 99.3 | 99.4 | >99.5 | >99.5 | |
| Analytical performance metrics | Analytical accuracy (%) | 99.9 | >99.9 | 99.9 | 99.9 |
|
Analytical sensitivity (PPA%) |
99.4 | >99.9 | 99.8 | 99.9 | |
|
Analytical specificity (NPA%) |
>99.9 | >99.9 | >99.9 | >99.9 | |
|
Positive predictive value (PPV %) |
>99.9 | >99.9 | >99.9 | >99.9 | |
| Repeatability (%) | 99.1 | >99.9 | 99.8 | 99.9 | |
| Reproducibility (%) | 99.1 | >99.9 | 99.8 | 99.9 | |
| Gene | Detected Allele(s) | Sample ID(s) | Reference method(s) |
| CYP2D6 | *3, *4, *5, *6, *9, *10, *17, *41, xN | 23V15551, 23T15552, 23Y15555, 23Q15562, 23Z15563, 23P15568, 23R15570, 23V11743 | Allelic discrimination PCR; fluorescent PCR for copy number determination; Sanger sequencing |
| CYP2C19 | *2, *17 | 23R15553, 23Y15555, 23X15564, 23P15568, 23R15570, 23Y11750 | Allelic discrimination PCR |
| CYP2C9 | *2, *3 | 23P15554, 23Y15555, 23R15570, 23R11745, 23Z11738, 23Q11740, 23W11751, 23Q11754, 24U11012, 24S11013, 24X11016 | Sanger sequencing (rs1799853, rs1057910); allelic discrimination PCR |
| CYP1A2 | *1C, *1F | 23Q15562, 23X15564, 23P15568, 23R15570 | Allelic discrimination PCR |
| CYP3A4 | *22 | 23R15570 | Sanger sequencing |
| CYP3A5 | *3, *7 | 23Q15559, 23V15565 | Allelic discrimination PCR |
| DPYD | rs3918290 (*2A), rs75017182+ rs56038477 (HapB3), rs67376798 (D949V), rs55886062 (*13) | 23U15557, 23S15558, 23U15560, 23U11752, 23P11746, 23W11748, 24R11022, 24P11023, 24W11025 | Sanger sequencing; allelic discrimination PCR |
| UGT1A1 | *28, *37, c.1220_1221insG | 23U15557, 23S15558, 23T11744, 23U11749, 23U15560, 23U11752, 24V11017, 24T11018, 24R11019, 24V11020 | STR analysis of (TA)n promoter repeat; Sanger sequencing; whole-exome sequencing |
| TPMT | *2, *3A, *3B, *3C | 23T15566, 23R15567, 23Y15569 | Allelic discrimination PCR |
| NUDT15 | *3 | 23R15567, 23Y15569 | Allelic discrimination PCR |
| RYR1 | c.7858C>T (p.Gln2620*) | 24S11027 | Whole-exome sequencing |
| Clinical Category | N (%) | Genes tested | Sample IDs |
|---|---|---|---|
| Oncology (fluoropyrimidines, irinotecan, tamoxifen) | 16 (39%) | DPYD, UGT1A1, CYP2D6 | 23U15557, 23S15558, 23U15560, 23T11744, 23U11749, 23U11746, 23W11748, 23R15570*, 23U11752, 24R11022, 24P11023, 24W11025, 24V11017, 24T11018, 24R11019, 24V11020 |
| Psychiatry (psychotropic therapy optimization) | 10 (24%) | CYP2D6, CYP2C19, CYP2C9, CYP1A2, CYP3A4 | 23V15555, 23Q15562, 23Z15563, 23R15570*, 23V15551, 23T15552, 23R15553, 23P15554, 23P15568, 23X15564 |
| Neurology (siponimod therapy) | 7 (17%) | CYP2C9 | 23R11745, 23Z11738, 23Q11740, 23W11751, 24U11012, 24S11013, 24X11016 |
| Gastroenterology (thiopurines) | 3 (7%) | TPMT, NUDT15 | 23T15566, 23R15567, 23Y15569 |
| Transplantation (tacrolimus therapy) | 2 (5%) | CYP3A5 | 23Q15559, 23V15565 |
| Other individual cases | 4 (10% | CYP2D6, CYP2C19, CYP2C9, RYR1 | 23Y11750, 23Q11754, 23V11743, 24S11027 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.




