Submitted:
16 February 2026
Posted:
26 February 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
2.1. Study Groups and Ethics Statement
2.2. Blood Collection and Sample Preparations
2.3. Viscosity Measurements
2.4. Description of Microfluidic System
2.5. Design of the Experiments
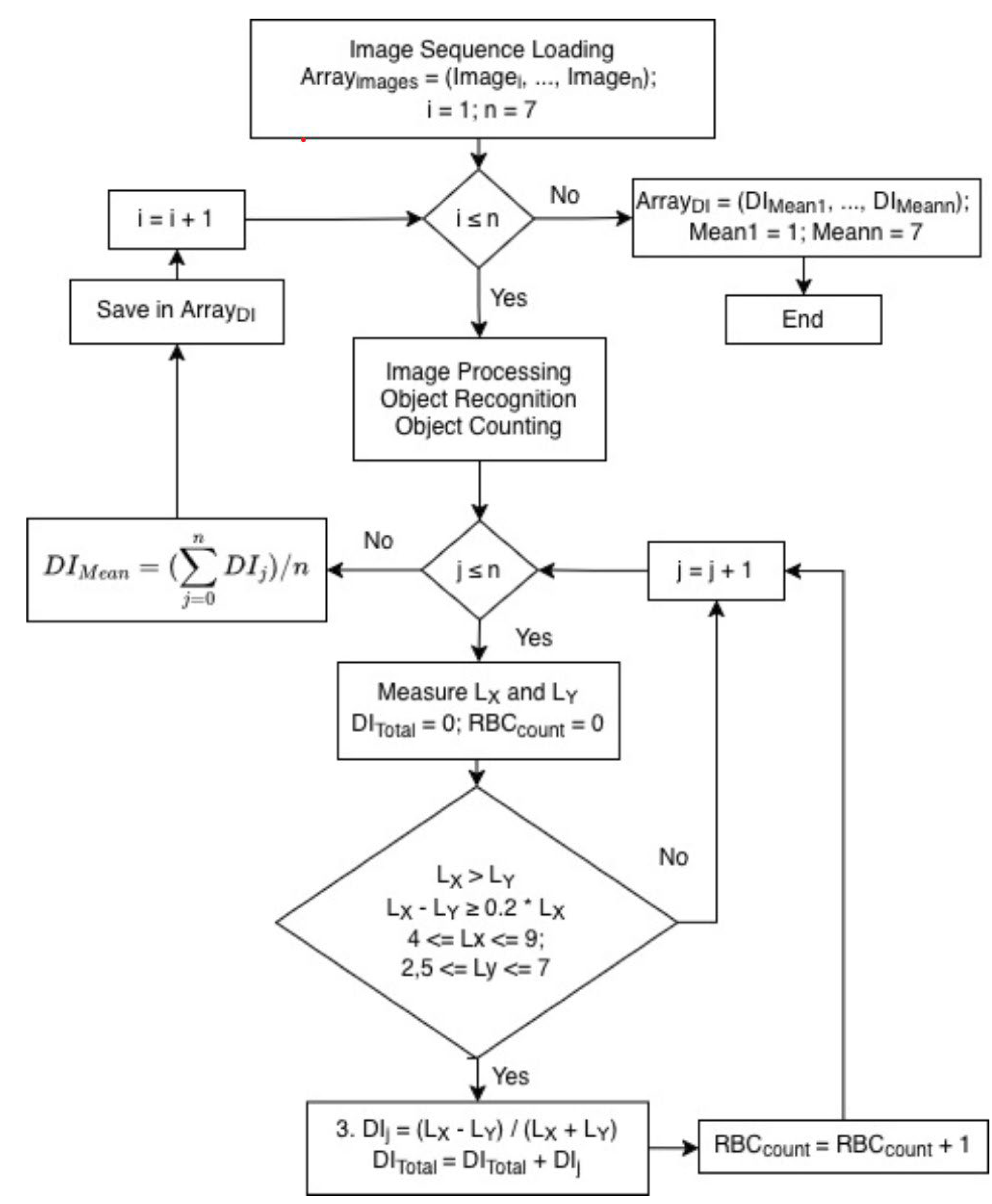
2.6. Algorithm for Software Image Flow Analysis of RBC Deformability
2.7. Statistics
3. Results
3.1. Clinical and Hematological Characteristics of the CLL Patients and Healthy Individuals
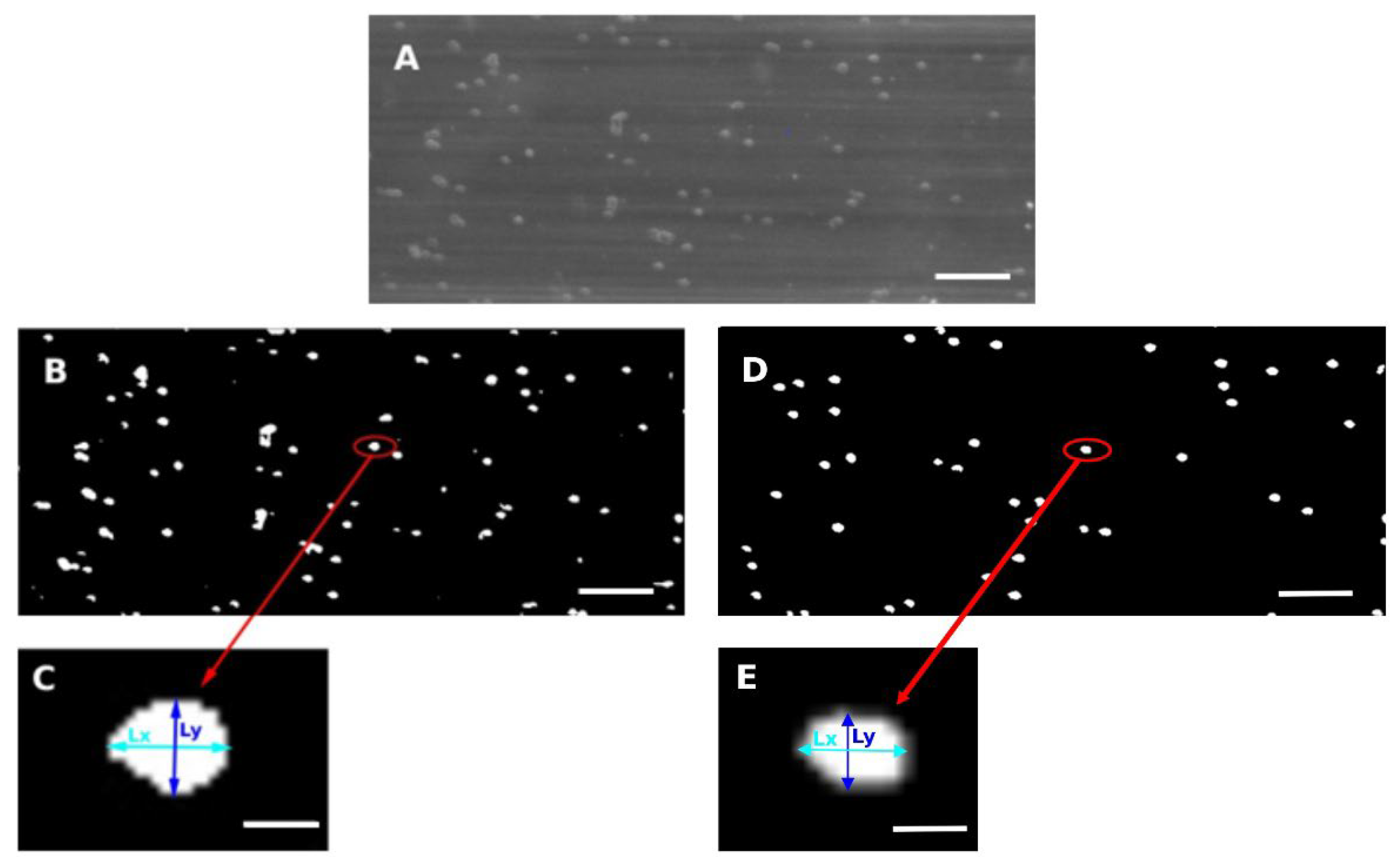
3.2. Development of a Software Application for the Analysis of Red Blood Cell Deformation under Flow Conditions
3.2.1. Software Application using LabVIEW
- a)
- Lₓ > Lᵧ
- b)
- Lₓ − Lᵧ ≥ 0.2 × Lₓ
- c)
- Lx and Ly within the ranges 4 ≤ Lx ≤ 9 and 2.5 ≤ Ly ≤ 7 pixels
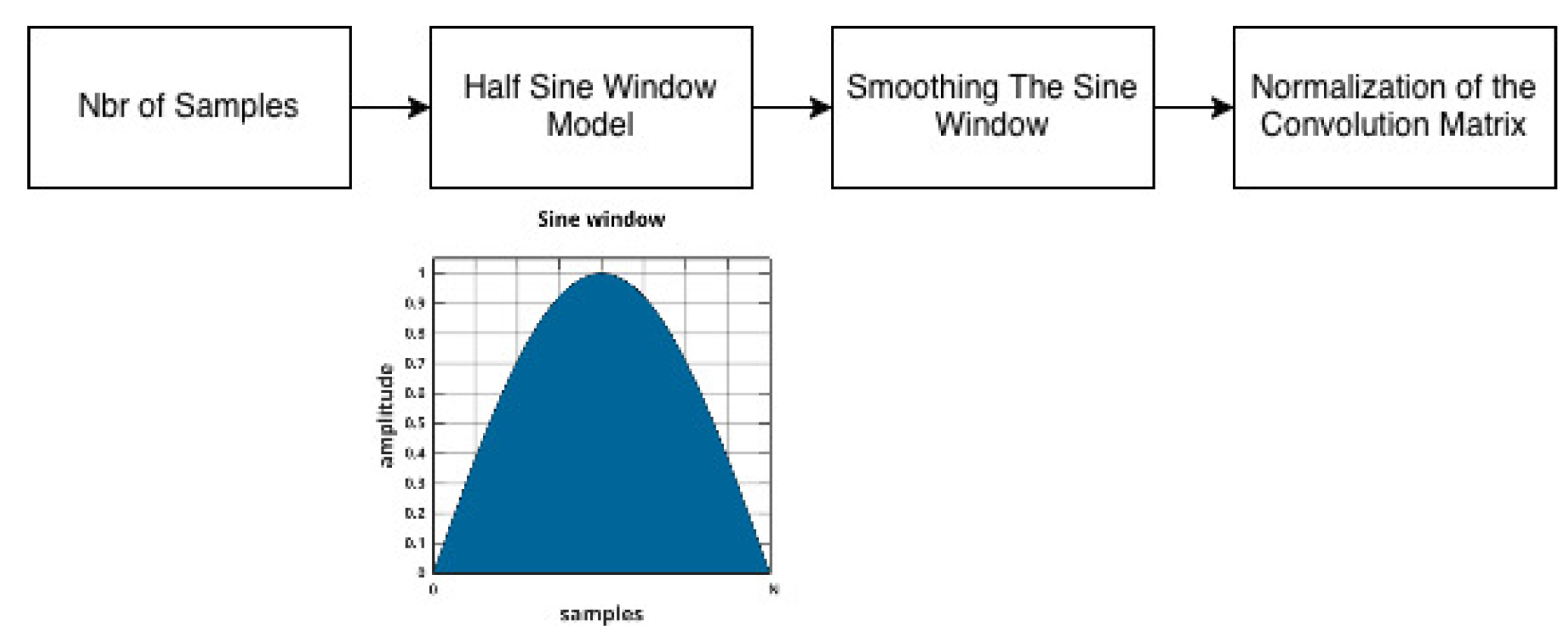
3.1.2. Implementation of the Sine Window Filter
3.2.3. Software Application using Python Programming Language
3.3. Limitations and Sources of Inter-Platform Variability
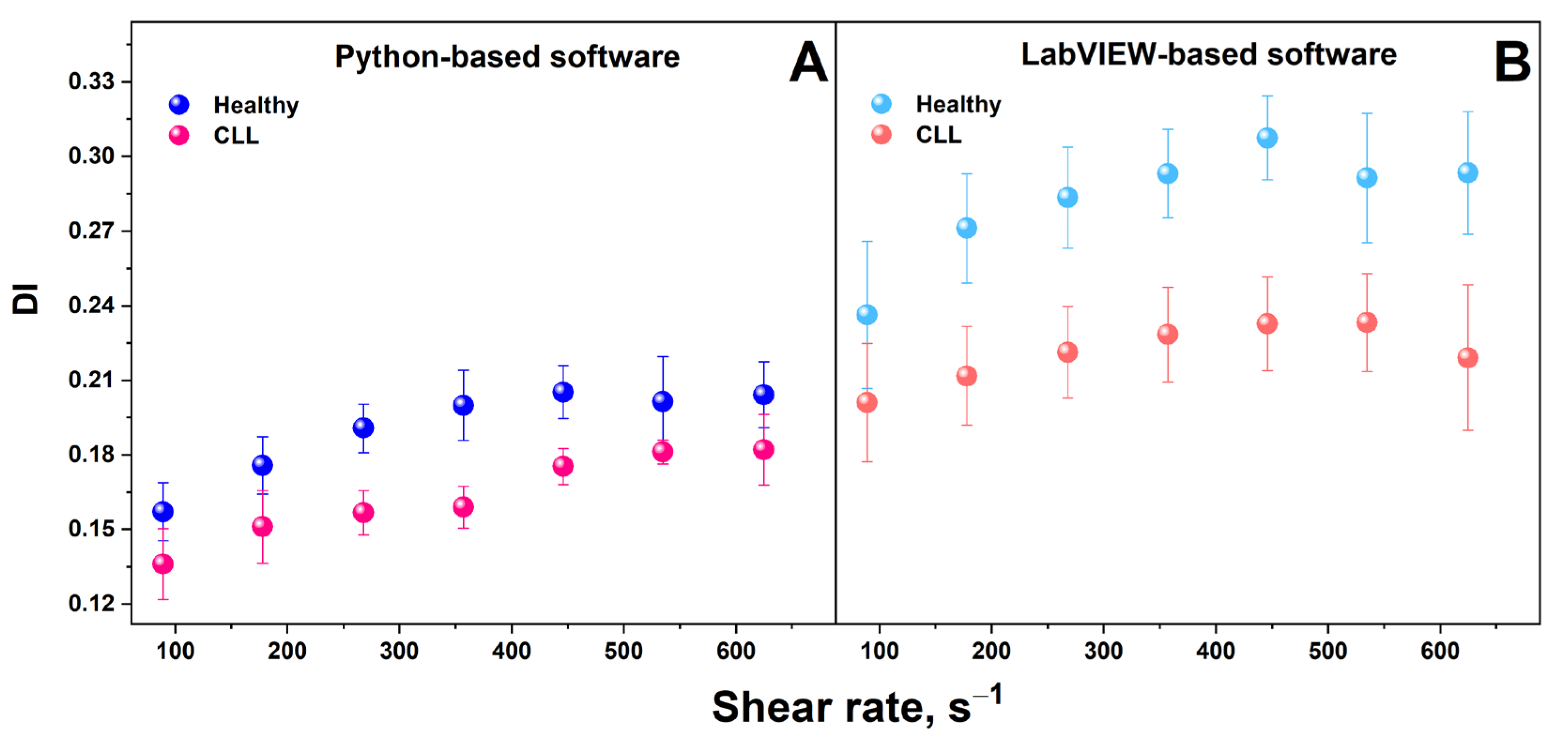
3.4. Comparison between Deformability Index Values for Healthy Individuals and Patients with CLL
3.4.1. Python-Based Analysis
3.4.2. LabVIEW-Based Analysis
3.4.3. Comparison between Python- and LabVIEW-Based Calculations
4. Discussion
4.1. RBC Deformability in Healthy Donors
4.2. Altered Deformability Patterns in CLL Patients
4.3. Cross-Platform Comparison and Methodological Implications
4.4. Clinical and Methodological Relevance
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Chen, M.; Leng, Y.; He, C.; et al. Red blood cells: a potential delivery system. J. Nanobiotechnol. 2023, 21, 288. [Google Scholar] [CrossRef] [PubMed]
- Jiang, H.; Li, X.; Liu, Z.; Luo, S.; Huang, J.; Chen, C.; Chen, R.; Li, F. A microfluidic approach for assessing the rheological properties of healthy versus thalassemic red blood cells. Micromachines 2025, 16, 957. [Google Scholar] [CrossRef] [PubMed]
- Mehendale, N.; Mitra, D.; Paul, D. A fast microfluidic device to measure the deformability of red blood cells. bioRxiv 2019, 644161. [Google Scholar] [CrossRef]
- Liang, M.; Ming, D.; Zhong, J.; Shannon, C.S.; Rojas-Carabali, W.; Agrawal, K.; Ai, Y.; Agrawal, R. Pathophysiological associations and measurement techniques of red blood cell deformability. Biosensors 2025, 15, 566. [Google Scholar] [CrossRef]
- Moutzouri, A.G.; Athanassiou, G.A.; Dimitropoulou, D.; Skoutelis, A.T.; Gogos, C.A. Severe sepsis and diabetes mellitus have additive effects on red blood cell deformability. J. Infect. 2008, 57, 147–151. [Google Scholar] [CrossRef]
- Shin, S.; Ku, Y.; Park, M.S.; et al. Deformability of red blood cells: A determinant of blood viscosity. J. Mech. Sci. Technol. 2005, 19, 216–223. [Google Scholar] [CrossRef]
- Alexy, T.; Detterich, J.; Connes, P.; Toth, K.; Nader, E.; Kenyeres, P.; Arriola-Montenegro, J.; Ulker, P.; Simmonds, M.J. Physical properties of blood and their relationship to clinical conditions. Front. Physiol. 2022, 13, 906768. [Google Scholar] [CrossRef]
- Ortín, X.; Giralt, M.; Romeu, M.; Lejeune, M.; Nogues, M.R.; Sanchez-Martos, V.; Rodriguez-Luaces, M.; Sansa, T.; Font, L. Oxidative stress in patients with early stage chronic lymphocytic leukemia: Assessment and correlation with prognostic factors. J. Hematol. 2012, 1, 77–88. [Google Scholar] [CrossRef]
- D'Alessandro, A.; Zolla, L. Proteomic analysis of red blood cells and the potential for the clinic: What have we learned so far? Expert Rev. Proteomics 2017, 14, 243–252. [Google Scholar] [CrossRef]
- Alexandrova-Watanabe, A.; Abadjieva, E.; Gartcheva, L.; Langari, A.; Ivanova, M.; Guenova, M.; Tiankov, T.; Strijkova, V.; Krumova, S.; Todinova, S. The impact of targeted therapies on red blood cell aggregation in patients with chronic lymphocytic leukemia evaluated using software image flow analysis. Micromachines 2025, 16, 95. [Google Scholar] [CrossRef]
- Zheng, Y.; Wen, J.; Nguyen, J.; Cachia, M.A.; Wang, C.; Sun, Y. Decreased deformability of lymphocytes in chronic lymphocytic leukemia. Sci. Rep. 2015, 5, 7613. [Google Scholar] [CrossRef]
- Alapan, Y.; Little, J.; Gurkan, U. Heterogeneous red blood cell adhesion and deformability in sickle cell disease. Sci. Rep. 2014, 4, 7173. [Google Scholar] [CrossRef] [PubMed]
- Artmann, G.M. Microscopic photometric quantification of stiffness and relaxation time of red blood cells in a flow chamber. Biorheology 1995, 32, 553–570. [Google Scholar] [PubMed]
- Boas, L.V.; Faustino, V.; Lima, R.; Miranda, J.M.; Minas, G.; Fernandes, C.S.V.; Catarino, S.O. Assessment of the deformability and velocity of healthy and artificially impaired red blood cells in narrow PDMS microchannels. Micromachines 2018, 9, 384. [Google Scholar] [CrossRef] [PubMed]
- Cluitmans, J.C.; Chokkalingam, V.; Janssen, A.M.; Brock, R.; Huck, W.T.; Bosman, G.J. Alterations in red blood cell deformability during storage: A microfluidic approach. Biomed. Res. Int. 2014, 764268. [Google Scholar] [CrossRef]
- Edwin, C.E.; Masnok, K.; Sawa, M.; Ibrahim, M.D.; Roslan, L.; Watanabe, N. Comparison between actual and theoretical ellipsoidal shape of human erythrocyte in modulated shear flow. J. Biorheol. 2025, 39, 10–24. [Google Scholar] [CrossRef]
- Žura, N.; Vince, S.; Perić, P.; Vilić, M.; Malarić, K.; Rimac, V.; Golubić Ćepulić, B.; Vajdić, M.; Jurak, I.; Milinković Tur, S.; et al. Short-Term In Vitro Exposure of Human Blood to 5G Network Frequencies: Do Sex and Frequency Additionally Affect Erythrocyte Morphometry? Biomedicines 2025, 13, 478. [Google Scholar] [CrossRef]
- Guo, Q.; Duffy, S.P.; Matthews, K.; Santoso, A.T.; Scott, M.D.; Ma, H. Microfluidic analysis of red blood cell deformability. J. Biomech. 2014, 47, 1767–1776. [Google Scholar] [CrossRef]
- Vdovin, A.; Muravyov, A.V.; Pevzner, A.A. Aggregometer-deformometer of human blood cells. South Sib. Sci. Bull. 2012, 2, 99–103. [Google Scholar]
- Lamoureux, E.S.; Islamzada, E.; Wiens, M.V.J.; Matthews, K.; Duffy, S.P.; Ma, H. Assessing red blood cell deformability from microscopy images using deep learning. Lab Chip, 2022. [Google Scholar]
- Hallek, M.; Cheson, B.D.; Catovsky, D.; et al. iwCLL guidelines for diagnosis, indications for treatment, response assessment, and supportive management of CLL. Blood 2018, 131, 2745–2760. [Google Scholar] [CrossRef]
- Hallek, M.; Cheson, B.D.; Catovsky, D.; et al. Guidelines for the diagnosis and treatment of chronic lymphocytic leukemia. Blood 2008, 111, 5446–5456. [Google Scholar] [CrossRef]
- Alexandrova-Watanabe, A.; Abadjieva, E.; Giosheva, I.; Langari, A.; Tiankov, T.; Gartchev, E.; Komsa-Penkova, R.; Todinova, S. Assessment of red blood cell aggregation in preeclampsia by microfluidic image flow analysis. Int. J. Mol. Sci. 2024, 25, 3732. [Google Scholar] [CrossRef] [PubMed]
- Mohandas, N.; Chasis, J.A.; Shohet, S.B. The influence of membrane skeleton on red cell deformability. Semin. Hematol. 1983, 20, 225–242. [Google Scholar] [PubMed]
- Meram, E.; Yilmaz, B.D.; Bas, C.; Atac, N.; Yalcin, O.; Meiselman, H.J.; Baskurt, O.K. Shear stress-induced improvement of red blood cell deformability. Biorheology 2013, 50, 165–176. [Google Scholar] [CrossRef] [PubMed]
- Yan, X.-J.; Dozmorov, I.; Li, W.; et al. Identification of outcome-correlated cytokine clusters in chronic lymphocytic leukemia. Blood 2011, 118, 5201–5210. [Google Scholar] [CrossRef]
- Rozovski, U.; Keating, M.J.; Estrov, Z. Targeting inflammatory pathways in chronic lymphocytic leukemia. Crit. Rev. Oncol. Hematol. 2013, 88, 655–666. [Google Scholar] [CrossRef]
- Strijkova, V.; Katrova, V.; Ivanova, M.; Langari, A.; Gartcheva, L.; Guenova, M.; Alexandrova-Watanabe, A.; Taneva, S.G.; Krumova, S.; Todinova, S. Nanomechanical and thermodynamic alterations of red blood cells in chronic lymphocytic leukemia. Int. J. Mol. Sci. 2026, 27, 353. [Google Scholar] [CrossRef]
- Baskurt, O.K.; Meiselman, H.J. Blood rheology and hemodynamics. Semin. Thromb. Hemost. 2003, 29, 435–450. [Google Scholar] [CrossRef]
- Parrow, N.L.; Violet, P.C.; Tu, H.; et al. Measuring deformability and red cell heterogeneity in blood by ektacytometry. J. Vis. Exp. 2018, 56910. [Google Scholar]
- Heo, Y.; Jung, H.; Shin, S. Osmotic deformability of erythrocytes at various shear stresses. Clin. Hemorheol. Microcirc. 2015, 59, 211–218. [Google Scholar] [CrossRef]
- Baskurt, O.K.; Hardeman, M.R.; Uyuklu, M.; et al. Parameterization of red blood cell elongation index–shear stress curves obtained by ektacytometry. Scand. J. Clin. Lab. Invest. 2009, 69, 777–788. [Google Scholar] [CrossRef]

| Parameters | Reference Value | Studied Groups | |
|
Healthy controls (n = 13) |
Untreated CLL Patients (n = 9) |
||
| Age (years) | - | 58.00 ± 7.4 | 63.89 ± 12.01 |
| Gender (F/M) | 8/5 | 4/5 | |
| Rai stage | 0 – 1 | ||
| RBC count (T/L) | 4.60–6.20 | 4.97 ± 0.23 | 4.98 ± 0.39 |
| Hb (g/L) | 140.00–180.00 | 161.40 ± 8.09 | 146.44± 11.86 |
| Ht (L/L) | 0.40–0.54 | 0.48 ± 0.01 | 0.44 ± 0.03 |
| MCV (fl) | 80.00–95.00 | 89.10 ± 3.74 | 87.93 ± 5.01 |
| MCH (pg/L) | 27.00–32.00 | 30.55 ± 1.33 | 29.43 ± 1.69 |
| MCHC (g/L) | 320.00–360.00 | 344.75 ± 4.15 | 335.00 ± 8.62 |
| RDW % | 11.60–14.80 | 13.88 ± 0.80 | 14.50 ± 1.32 |
| WBC | 3.50–10.50 | 6.3 ± 1.1 | 11.7 ÷ 157.5 * |
| Lymphocytes (ABS) | 1.10–3.80 | 1.91 ± 0.17 | 6.70 ÷ 144.28 * |
| Shear rate (s⁻¹) | Python-based software | LabVIEW-based software | ||||
| Healthy controls | Untreated CLL patients | p | Healthy controls | Untreated CLL patients | p | |
| 89 | 0.157 ± 0.011 | 0.136 ± 0.014 | 0.18 | 0.236 ± 0.029 | 0.201 ± 0.023 | 0.074 |
| 178 | 0.176 ± 0.011 | 0.151 ± 0.014 | 0.20 | 0.271 ± 0.022 | 0.212 ± 0.020* | 0.046 |
| 268 | 0.191 ± 0.010 | 0.156 ± 0.009* | 0.015 | 0.284 ± 0.020 | 0.221 ± 0.018* | 0.016 |
| 357 | 0.200 ± 0.014 | 0.159 ± 0.008* | 0.031 | 0.293 ± 0.017 | 0.228 ± 0.019* | 0.012 |
| 446 | 0.205 ± 0.010 | 0.175 ± 0.007* | 0.039 | 0.307 ± 0.016 | 0.233 ± 0.019* | 0.014 |
| 535 | 0.202 ± 0.018 | 0.181 ± 0.005 | 0.19 | 0.291 ± 0.025 | 0.233 ± 0.020* | 0.022 |
| 625 | 0.204 ± 0.013 | 0.182 ± 0.014 | 0.14 | 0.293 ± 0.024 | 0.219 ± 0.029* | 0.035 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




