Submitted:
15 February 2026
Posted:
26 February 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Results
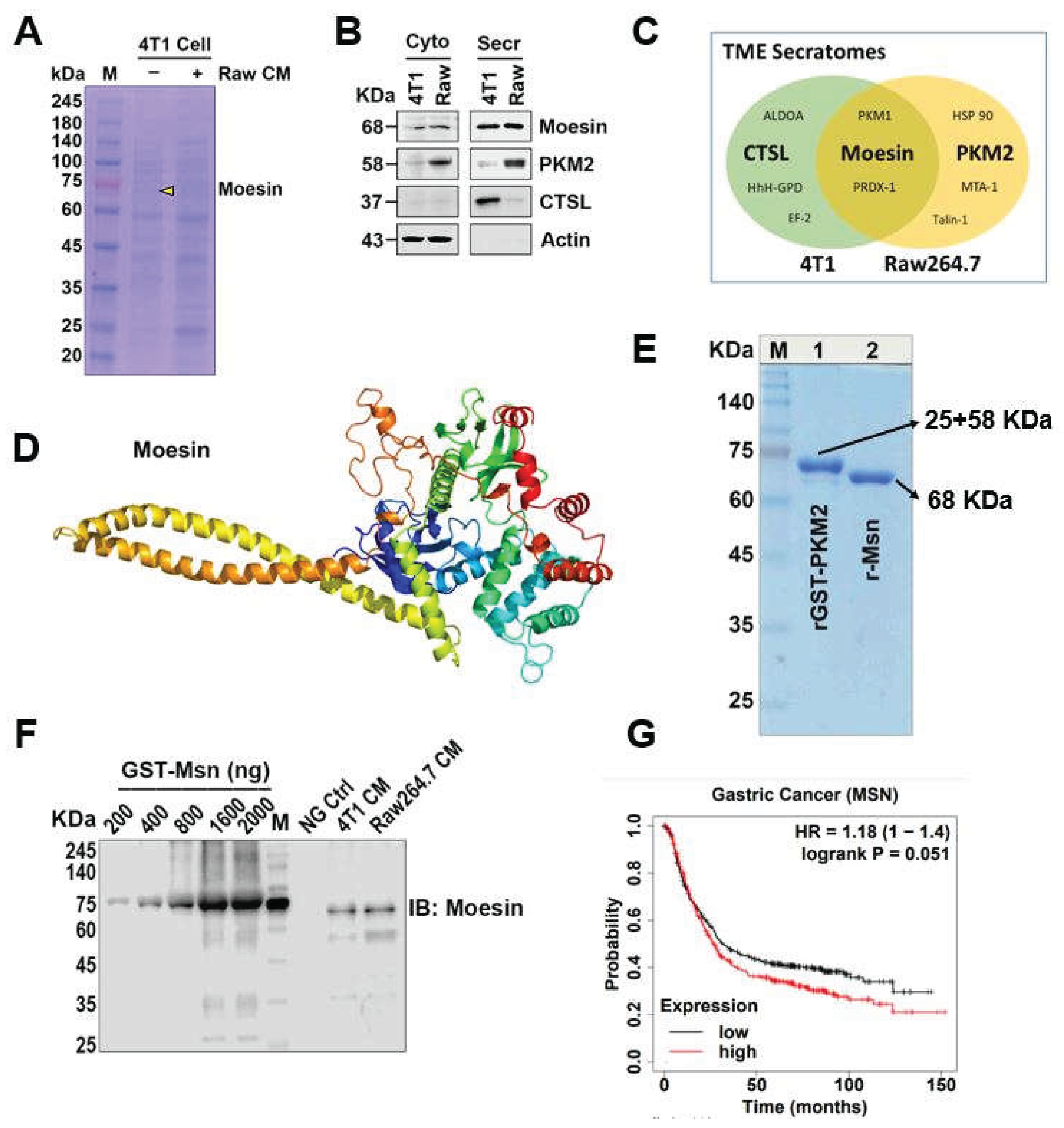
2.1. Msn Was Identified as Being Secreted from Cancer (4T1) and Macrophage (Raw264.7) Cells
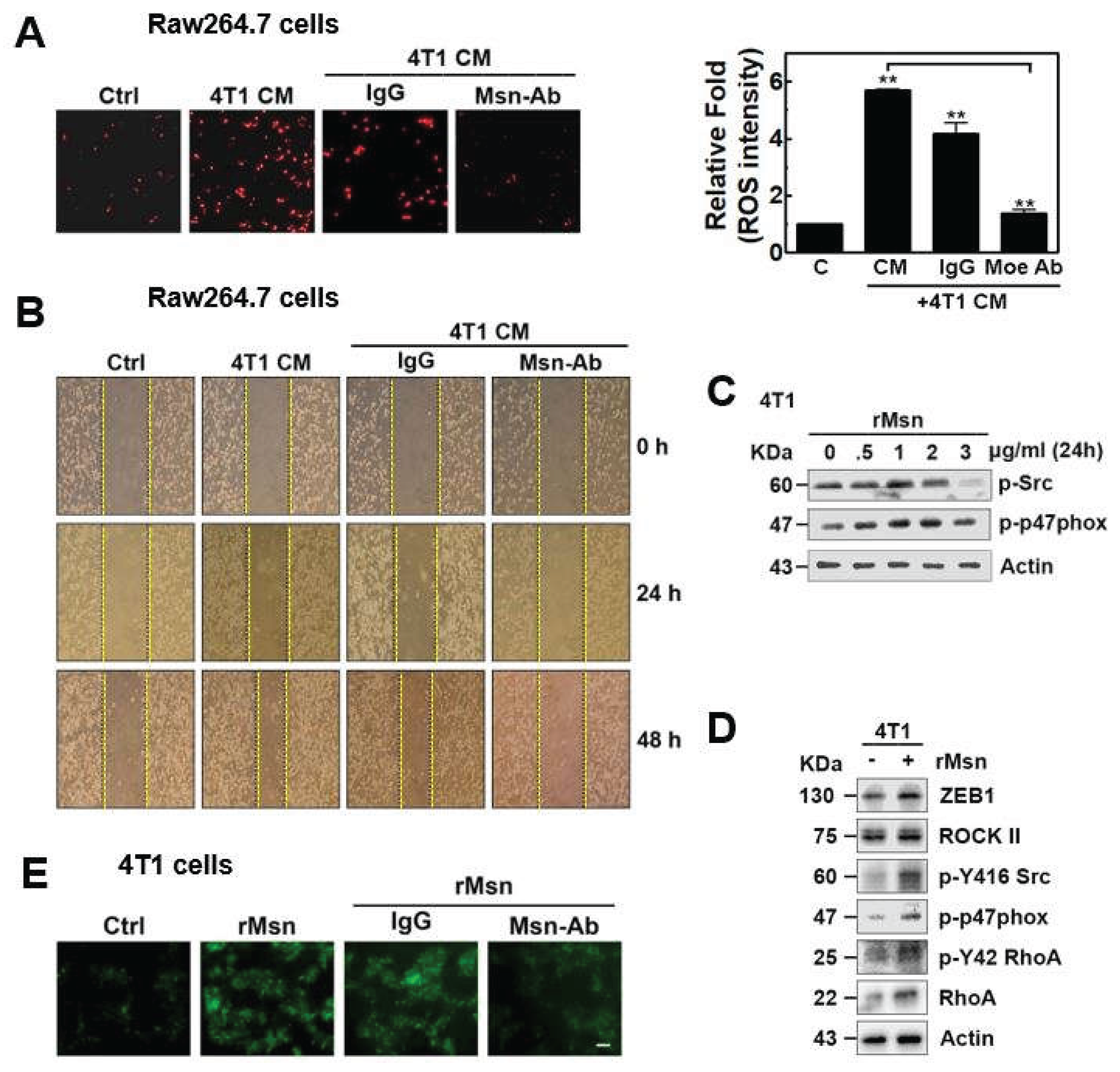
2.2. Intracellular Signaling Through Extracellular Msn and Cell Physiology in TME
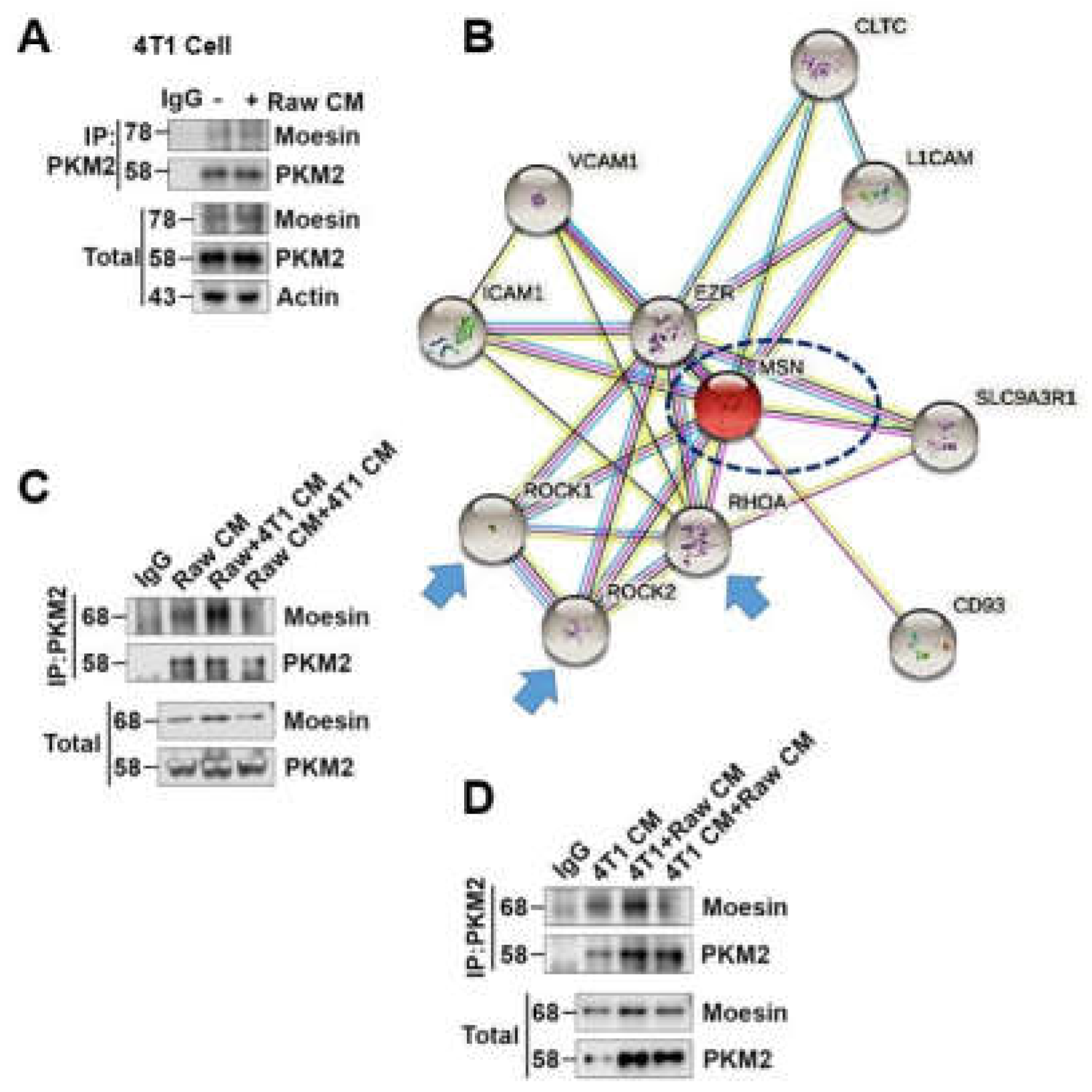
2.3. Extracellular Msn (exMsn) Interacts with Extracellular PKM2 (exPKM2)
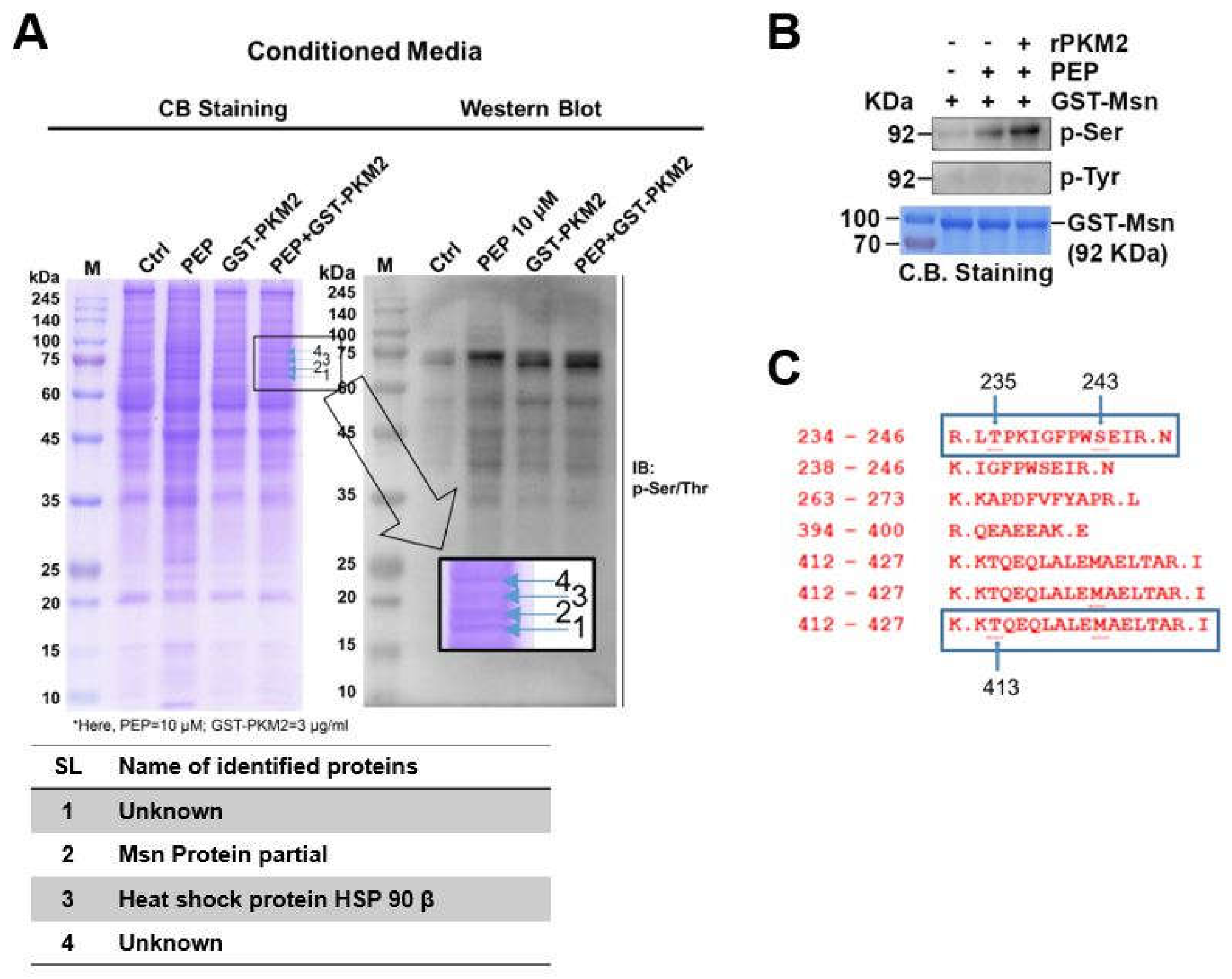
2.4. exMsn Can Be Phosphorylated by exPKM2 in the Presence of PEP
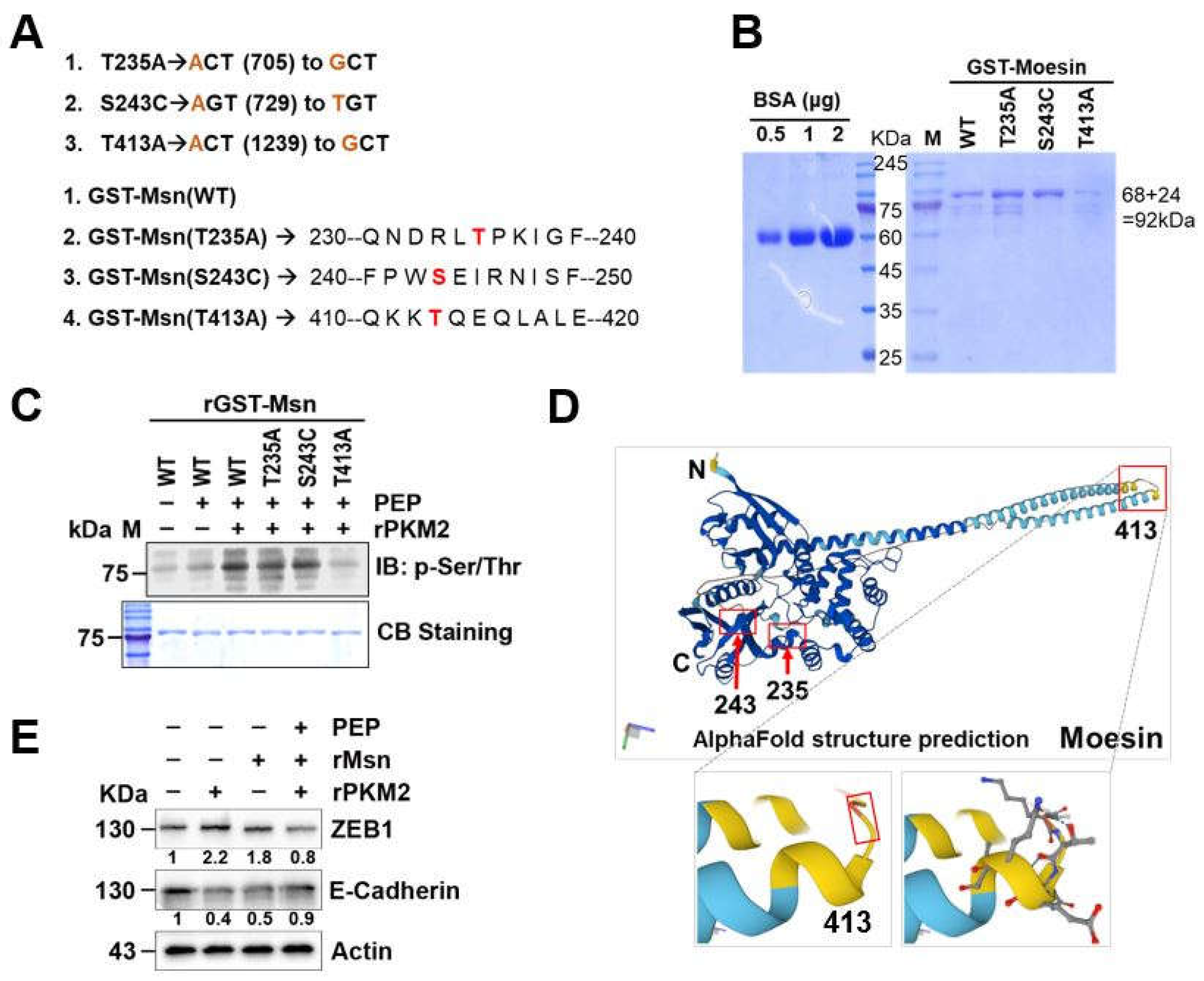
2.5. N-Terminal Extended α-Helical Globular Domain of Msn Was Phosphorylated by exPKM2
3. Discussion
4. Materials and Methods
4.1. Cell Culture
4.2. Preparation of Conditioned Media (CM)
4.3. Recombinant Msn Preparation
4.4. Cell Migration by Wound Healing Assay
4.5. In Vitro Kinase Assay
4.6. Site-Directed Mutagenesis
4.7. Western Blotting
4.8. Immunoprecipitation
4.9. Superoxide Measurement Assay
4.10. Protein Identification by Mass Spectrometry (PMF-MALDI-TOF)
4.11. Bioinformatics Analysis
4.12. Statistical Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
Abbreviations
| TME | Tumor Microenviromnment |
| exMsn | Extracellular Moesin |
| rMsn | Recombinant Moesin |
| exPKM2 | Extracellular Pyruvate Kinase |
| CM | conditioned media |
| EMT | epithelial-mesenchymal transition |
References
- da Cunha, B.R.; Domingos, C.; Stefanini, A.C.B.; Henrique, T.; Polachini, G.M.; Castelo-Branco, P.; Tajara, E.H. Cellular Interactions in the Tumor Microenvironment: The Role of Secretome. J Cancer 2019, 10, 4574–4587. [Google Scholar] [CrossRef]
- Morimoto, H.; Ito, Y.; Yoden, E.; Horie, M.; Tanaka, N.; Komurasaki, Y.; Yamamoto, R.; Mihara, K.; Minami, K.; Hirato, T. Non-clinical evaluation of JR-051 as a biosimilar to agalsidase beta for the treatment of Fabry disease. Mol Genet Metab 2018, 125, 153–160. [Google Scholar] [CrossRef]
- Winkler, J.K.; Haenssle, H.A.; Enk, A.; Toberer, F. Orange Verrucous Pitted Lesion on Lateral Foot of a 9-year-old Girl: A Quiz. Acta Derm Venereol 2020, 100, adv00201. [Google Scholar] [CrossRef]
- Xue, H.; Lu, B.; Lai, M. The cancer secretome: a reservoir of biomarkers. J Transl Med 2008, 6, 52. [Google Scholar] [CrossRef]
- Hamza, A.; Cho, J.Y.; Cap, K.C.; Hossain, A.J.; Kim, J.G.; Park, J.B. Extracellular pyruvate kinase M2 induces cell migration through p-Tyr42 RhoA-mediated superoxide generation and epithelial-mesenchymal transition. Free Radic Biol Med 2023, 208, 614–629. [Google Scholar] [CrossRef]
- Kessenbrock, K.; Plaks, V.; Werb, Z. Matrix metalloproteinases: regulators of the tumor microenvironment. Cell 2010, 141, 52–67. [Google Scholar] [CrossRef] [PubMed]
- Fehon, R.G.; McClatchey, A.I.; Bretscher, A. Organizing the cell cortex: the role of ERM proteins. Nat Rev Mol Cell Biol 2010, 11, 276–287. [Google Scholar] [CrossRef]
- Eustace, B.K.; Sakurai, T.; Stewart, J.K.; Yimlamai, D.; Unger, C.; Zehetmeier, C.; Lain, B.; Torella, C.; Henning, S.W.; Beste, G.; et al. Functional proteomic screens reveal an essential extracellular role for hsp90 alpha in cancer cell invasiveness. Nat Cell Biol 2004, 6, 507–514. [Google Scholar] [CrossRef]
- Lankes, W.T.; Furthmayr, H. Moesin: a member of the protein 4.1-talin-ezrin family of proteins. Proc Natl Acad Sci U S A 1991, 88, 8297–8301. [Google Scholar] [CrossRef]
- Chen, Q.Y.; Xu, W.; Jiao, D.M.; Wu, L.J.; Song, J.; Yan, J.; Shi, J.G. Silence of ezrin modifies migration and actin cytoskeleton rearrangements and enhances chemosensitivity of lung cancer cells in vitro. Mol Cell Biochem 2013, 377, 207–218. [Google Scholar] [CrossRef]
- Kunda, P.; Pelling, A.E.; Liu, T.; Baum, B. Moesin controls cortical rigidity, cell rounding, and spindle morphogenesis during mitosis. Curr Biol 2008, 18, 91–101. [Google Scholar] [CrossRef]
- Neisch, A.L.; Fehon, R.G. Ezrin, Radixin and Moesin: key regulators of membrane-cortex interactions and signaling. Curr Opin Cell Biol 2011, 23, 377–382. [Google Scholar] [CrossRef]
- Adyshev, D.M.; Dudek, S.M.; Moldobaeva, N.; Kim, K.M.; Ma, S.F.; Kasa, A.; Garcia, J.G.; Verin, A.D. Ezrin/radixin/moesin proteins differentially regulate endothelial hyperpermeability after thrombin. Am J Physiol Lung Cell Mol Physiol 2013, 305, L240–255. [Google Scholar] [CrossRef] [PubMed]
- Wei, X.; He, S.; Wang, Z.; Wu, J.; Zhang, J.; Cheng, Y.; Yang, J.; Xu, X.; Chen, Z.; Ye, J.; et al. Fibroblast growth factor 1attenuates 6-hydroxydopamine-induced neurotoxicity: an in vitro and in vivo investigation in experimental models of parkinson’s disease. Am J Transl Res 2014, 6, 664–677. [Google Scholar] [PubMed]
- Wang, Q.; Fan, A.; Yuan, Y.; Chen, L.; Guo, X.; Huang, X.; Huang, Q. Role of Moesin in Advanced Glycation End Products-Induced Angiogenesis of Human Umbilical Vein Endothelial Cells. Sci Rep 2016, 6, 22749. [Google Scholar] [CrossRef] [PubMed]
- Li, Q.; Liu, H.; Du, J.; Chen, B.; Li, Q.; Guo, X.; Huang, X.; Huang, Q. Advanced glycation end products induce moesin phosphorylation in murine brain endothelium. Brain Res 2011, 1373, 1–10. [Google Scholar] [CrossRef]
- Wang, Y.; Kaiser, M.S.; Larson, J.D.; Nasevicius, A.; Clark, K.J.; Wadman, S.A.; Roberg-Perez, S.E.; Ekker, S.C.; Hackett, P.B.; McGrail, M.; et al. Moesin1 and Ve-cadherin are required in endothelial cells during in vivo tubulogenesis. Development 2010, 137, 3119–3128. [Google Scholar] [CrossRef]
- Li, Y.Q.; Zheng, Z.; Liu, Q.X.; Lu, X.; Zhou, D.; Zhang, J.; Zheng, H.; Dai, J.G. Moesin as a prognostic indicator of lung adenocarcinoma improves prognosis by enhancing immune lymphocyte infiltration. World J Surg Oncol 2021, 19, 109. [Google Scholar] [CrossRef]
- Siu, L.L. Clinical trials in the elderly--a concept comes of age. N Engl J Med 2007, 356, 1575–1576. [Google Scholar] [CrossRef]
- Kwon, O.K.; Lee, W.; Kim, S.J.; Lee, Y.M.; Lee, J.Y.; Kim, J.Y.; Bae, J.S.; Lee, S. In-depth proteomics approach of secretome to identify novel biomarker for sepsis in LPS-stimulated endothelial cells. Electrophoresis 2015, 36, 2851–2858. [Google Scholar] [CrossRef]
- Ahandoust, S.; Li, K.; Sun, X.; Li, B.Y.; Yokota, H.; Na, S. Intracellular and extracellular moesins differentially regulate Src activity and beta-catenin translocation to the nucleus in breast cancer cells. Biochem Biophys Res Commun 2023, 639, 62–69. [Google Scholar] [CrossRef]
- Gao, X.; Wang, H.; Yang, J.J.; Liu, X.; Liu, Z.R. Pyruvate kinase M2 regulates gene transcription by acting as a protein kinase. Mol Cell 2012, 45, 598–609. [Google Scholar] [CrossRef]
- Jiang, Y.; Wang, Y.; Wang, T.; Hawke, D.H.; Zheng, Y.; Li, X.; Zhou, Q.; Majumder, S.; Bi, E.; Liu, D.X.; et al. PKM2 phosphorylates MLC2 and regulates cytokinesis of tumour cells. Nat Commun 2014, 5, 5566. [Google Scholar] [CrossRef] [PubMed]
- Koppenol, W.H.; Bounds, P.L.; Dang, C.V. Otto Warburg’s contributions to current concepts of cancer metabolism. Nat Rev Cancer 2011, 11, 325–337. [Google Scholar] [CrossRef] [PubMed]
- Yang, W.; Xia, Y.; Ji, H.; Zheng, Y.; Liang, J.; Huang, W.; Gao, X.; Aldape, K.; Lu, Z. Nuclear PKM2 regulates β-catenin transactivation upon EGFR activation. Nature 2011, 480, 118–122. [Google Scholar] [CrossRef] [PubMed]
- Wang, C.; Zhang, S.; Liu, J.; Tian, Y.; Ma, B.; Xu, S.; Fu, Y.; Luo, Y. Secreted Pyruvate Kinase M2 Promotes Lung Cancer Metastasis through Activating the Integrin Beta1/FAK Signaling Pathway. Cell Rep 2020, 30, 1780–1797 e1786. [Google Scholar] [CrossRef]
- Hou, P.P.; Luo, L.J.; Chen, H.Z.; Chen, Q.T.; Bian, X.L.; Wu, S.F.; Zhou, J.X.; Zhao, W.X.; Liu, J.M.; Wang, X.M.; et al. Ectosomal PKM2 Promotes HCC by Inducing Macrophage Differentiation and Remodeling the Tumor Microenvironment. Mol Cell 2020, 78, 1192–1206 e1110. [Google Scholar] [CrossRef]
- Whitesell, L.; Lindquist, S.L. HSP90 and the chaperoning of cancer. Nat Rev Cancer 2005, 5, 761–772. [Google Scholar] [CrossRef]
- Wang, X.; Song, X.; Zhuo, W.; Fu, Y.; Shi, H.; Liang, Y.; Tong, M.; Chang, G.; Luo, Y. The regulatory mechanism of Hsp90alpha secretion and its function in tumor malignancy. Proc Natl Acad Sci U S A 2009, 106, 21288–21293. [Google Scholar] [CrossRef]
- Zhang, S.; Wang, C.; Ju, J.; Wang, C. Extracellular Hsp90alpha Supports the ePKM2-GRP78-AKT Axis to Promote Tumor Metastasis. Front Oncol 2022, 12, 906080. [Google Scholar] [CrossRef]
- Li, Q.; Nance, M.R.; Kulikauskas, R.; Nyberg, K.; Fehon, R.; Karplus, P.A.; Bretscher, A.; Tesmer, J.J. Self-masking in an intact ERM-merlin protein: an active role for the central alpha-helical domain. J Mol Biol 2007, 365, 1446–1459. [Google Scholar] [CrossRef] [PubMed]
- Vilmos, P.; Kristo, I.; Szikora, S.; Jankovics, F.; Lukacsovich, T.; Kari, B.; Erdelyi, M. The actin-binding ERM protein Moesin directly regulates spindle assembly and function during mitosis. Cell Biol Int 2016, 40, 696–707. [Google Scholar] [CrossRef] [PubMed]
- Kim, J.G.; Mahmud, S.; Min, J.K.; Lee, Y.B.; Kim, H.; Kang, D.C.; Park, H.S.; Seong, J.; Park, J.B. RhoA GTPase phosphorylated at tyrosine 42 by src kinase binds to beta-catenin and contributes transcriptional regulation of vimentin upon Wnt3A. Redox Biol 2021, 40, 101842. [Google Scholar] [CrossRef]
- Cap, K.C.; Kim, J.G.; Hamza, A.; Park, J.B. P-Tyr42 RhoA GTPase amplifies superoxide formation through p47phox, phosphorylated by ROCK. Biochem Biophys Res Commun 2020, 523, 972–978. [Google Scholar] [CrossRef] [PubMed]
- Hu, S.; Shi, X.; Liu, Y.; He, Y.; Du, Y.; Zhang, G.; Yang, C.; Gao, F. CD44 cross-linking increases malignancy of breast cancer via upregulation of p-Moesin. Cancer Cell Int 2020, 20, 563. [Google Scholar] [CrossRef]
- Sanchez-Sanchez, B.J.; Marcotti, S.; Salvador-Garcia, D.; Diaz-de-la-Loza, M.D.; Burki, M.; Davidson, A.J.; Wood, W.; Stramer, B.M. Moesin integrates cortical and lamellar actin networks during Drosophila macrophage migration. Nat Commun 2025, 16, 1414. [Google Scholar] [CrossRef]
- Barros, F.B.A.; Assao, A.; Garcia, N.G.; Nonogaki, S.; Carvalho, A.L.; Soares, F.A.; Kowalski, L.P.; Oliveira, D.T. Moesin expression by tumor cells is an unfavorable prognostic biomarker for oral cancer. BMC Cancer 2018, 18, 53. [Google Scholar] [CrossRef]
- Haynes, J.; Srivastava, J.; Madson, N.; Wittmann, T.; Barber, D.L. Dynamic actin remodeling during epithelial-mesenchymal transition depends on increased moesin expression. Mol Biol Cell 2011, 22, 4750–4764. [Google Scholar] [CrossRef]
- Tlili, A.; Pintard, C.; Hurtado-Nedelec, M.; Liu, D.; Marzaioli, V.; Thieblemont, N.; Dang, P.M.; El-Benna, J. ROCK2 interacts with p22phox to phosphorylate p47phox and to control NADPH oxidase activation in human monocytes. Proc Natl Acad Sci U S A 2023, 120, e2209184120. [Google Scholar] [CrossRef]
- Senbanjo, L.T.; Chellaiah, M.A. CD44: A Multifunctional Cell Surface Adhesion Receptor Is a Regulator of Progression and Metastasis of Cancer Cells. Front Cell Dev Biol 2017, 5, 18. [Google Scholar] [CrossRef]
| SL | Identity | Primer sequence(5’ to 3’) |
|---|---|---|
| 1 | MSN- T235A_F | GAATGACAGACTAGCTCCCAAGATAG |
| 2 | MSN- T235A_R | CTATCTTGGGAGCTAGTCTGTCATTC |
| 3 | MSN- T243C_F | GGCTTCCCCTGGAGTGAAATCAGGAACATC |
| 4 | MSN- T243C_R | GGCTTCCCCTGGTCTGAAATCAGGAACATC |
| 5 | MSN- T413A_F | CAGAAAAAGGCTCAGGAACAG |
| 6 | MSN- T413A_R | CTGTTCCTGAGCCTTTTTCTG |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




