Submitted:
12 February 2026
Posted:
13 February 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
2.1. Study Population
2.2. Sample Preparation
2.2.1. Milk Collection
2.2.2. Milk Serum Separation
2.2.3. Sample Preparation for LC-MS/MS Analysis
2.2.4. Liquid Chromatography – Tandem Mass Spectrometry
2.2.5. Mass Spectrometric Raw Data Analysis, Spectral Library Generation and DIA Quantitation
2.3. Statistical Analysis
3. Results
3.1. Study Population and Maternal Characteristics
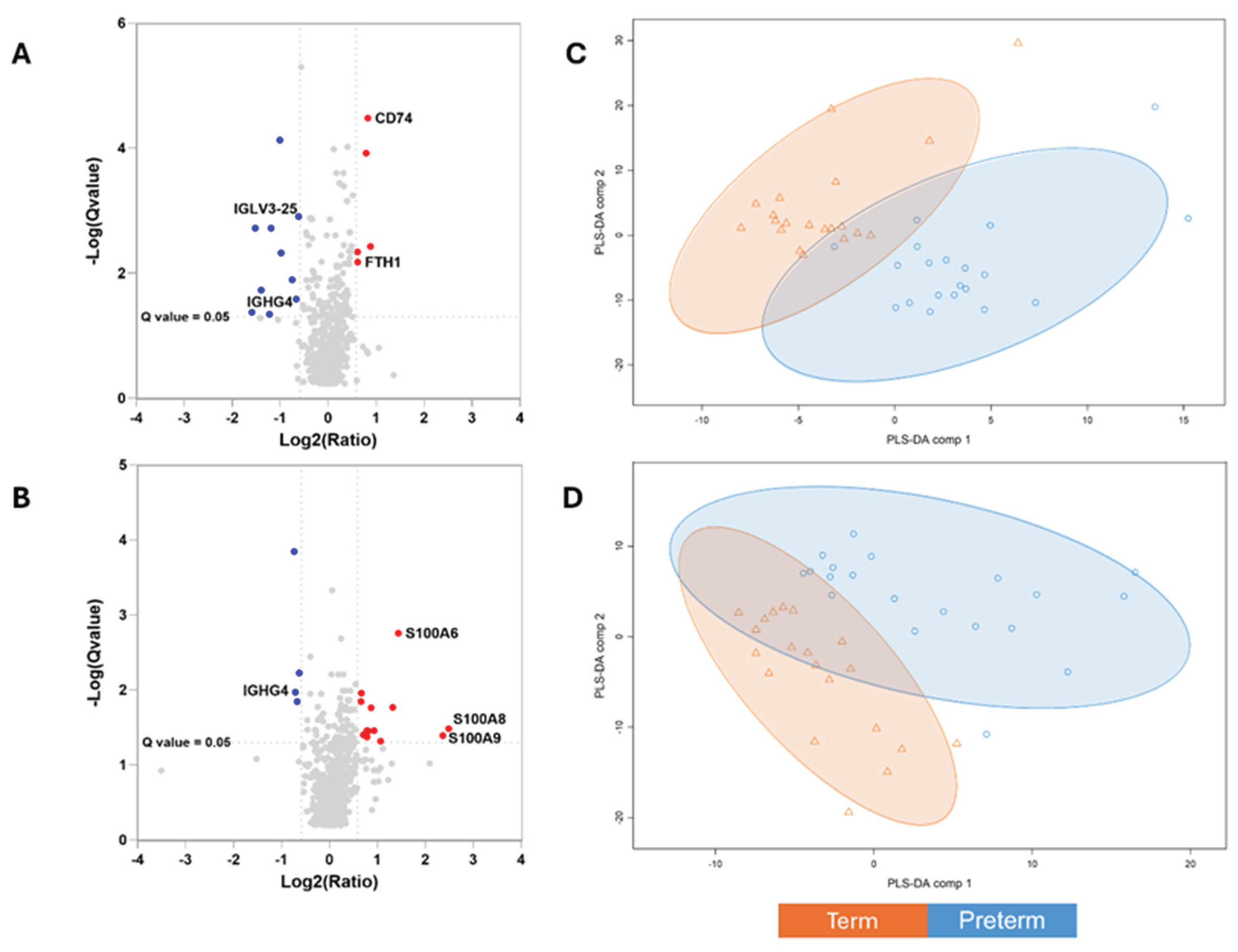
3.2. Differential Milk Proteome in Early Lactation
3.3. Differential Milk Proteome in Later Lactation
4. Discussion
4.1. Summary of Principal Findings
4.2. Biological Interpretation of Key Proteins
4.2.1. Immune-Related Proteins and Immunoglobulins
4.2.2. Proteins Related to Innate Immunity and Inflammatory Response
4.2.3. Iron Metabolism and Adaptive Functions
4.3. Lactation Stage as Contextual Framework Rather than Primary Driver
4.4. Novelty and Significance of the Study
4.5. Limitations
4.6. Implications and Future Directions
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Victora, C.G.; Bahl, R.; Barros, A.J.; França, G.V.; Horton, S.; Krasevec, J.; et al. Breastfeeding in the 21st century: Epidemiology, mechanisms, and lifelong effect. Lancet 2016, 387(10017), 475–490. [Google Scholar] [CrossRef]
- Ballard, O.; Morrow, A.L. Human milk composition: Nutrients and bioactive factors. Pediatr Clin North Am. 2013, 60(1), 49–74. [Google Scholar] [CrossRef]
- Andreas, N.J.; Kampmann, B.; Mehring Le-Doare, K. Human breast milk: A review on its composition and bioactivity. Early Hum Dev. 2015, 91(11), 629–635. [Google Scholar] [CrossRef]
- Castellote, C.; Casillas, R.; Ramírez-Santana, C.; Pérez-Cano, F.J.; Castell, M.; Moretones, M.G.; et al. Premature delivery influences the immunological composition of colostrum and transitional and mature human milk. J Nutr. 2011, 141(6), 1181–1187. [Google Scholar] [CrossRef]
- Lönnerdal, B. Bioactive proteins in human milk: Mechanisms of action. J Pediatr. 2010, 156((2) Suppl, S26–S30. [Google Scholar] [CrossRef]
- Zasada, M.; Suski, M.; Bokiniec, R.; Szwarc-Duma, M.; Borszewska-Kornacka, M.K.; Madej, J.; et al. Comparative two time-point proteome analysis of the plasma from preterm infants with and without bronchopulmonary dysplasia. Ital J Pediatr. 2019, 45(1), 112. [Google Scholar] [CrossRef] [PubMed]
- Dekker, P.M.; Boeren, S.; van Goudoever, J.B.; Vervoort, J.J.M.; Hettinga, K.A. Exploring Human Milk Dynamics: Interindividual Variation in Milk Proteome, Peptidome, and Metabolome. J Proteome Res. 2022, 21(4), 1002–1016. [Google Scholar] [CrossRef] [PubMed]
- Molinari, C.E.; Casadio, Y.S.; Hartmann, B.T.; Livk, A.; Bringans, S.; Arthur, P.G.; et al. Proteome mapping of human skim milk proteins in term and preterm milk. J Proteome Res. 2012, 11(3), 1696–1714. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Zhang, X.; Mi, L.; Li, C.; Zhang, Y.; Bi, R.; et al. Comparative Proteomic Analysis of Proteins in Breast Milk during Different Lactation Periods. Nutrients. 2022, 14(17), 3648. [Google Scholar] [CrossRef]
- Wiśniewski, J.R.; Gaugaz, F.Z. Fast and sensitive total protein and Peptide assays for proteomic analysis. Anal Chem. 2015, 87(8), 4110–4116. [Google Scholar] [CrossRef]
- Wiśniewski, J.R.; Zougman, A.; Nagaraj, N.; Mann, M. Universal sample preparation method for proteome analysis. Nat Methods. 2009, 6(5), 359–362. [Google Scholar] [CrossRef] [PubMed]
- Bruderer, R.; Bernhardt, O.M.; Gandhi, T.; Miladinović, S.M.; Cheng, L.Y.; Messner, S.; et al. Extending the limits of quantitative proteome profiling with data-independent acquisition and application to acetaminophen-treated three-dimensional liver microtissues. Mol Cell Proteomics. 2015, 14(5), 1400–1410. [Google Scholar] [CrossRef] [PubMed]
- Zhang, B.; Chambers, M.C.; Tabb, D.L. Proteomic parsimony through bipartite graph analysis improves accuracy and transparency. J Proteome Res. 2007, 6(9), 3549–3557. [Google Scholar] [CrossRef] [PubMed]
- Storey, J.D. A Direct Approach to False Discovery Rates. Journal of the Royal Statistical Society Series B: Statistical Methodology. 2002, 64(3), 479–498. [Google Scholar] [CrossRef]
- Rohart, F.; Gautier, B.; Singh, A.; Le Cao, K.A. mixOmics: An R package for ‘omics feature selection and multiple data integration. PLoS Comput Biol. 2017, 13(11), e1005752. [Google Scholar] [CrossRef]
- Huang da, W.; Sherman, B.T.; Lempicki, R.A. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat Protoc. 2009, 4(1), 44–57. [Google Scholar] [CrossRef]
- Sherman, B.T.; Hao, M.; Qiu, J.; Jiao, X.; Baseler, M.W.; Lane, H.C.; et al. DAVID: A web server for functional enrichment analysis and functional annotation of gene lists (2021 update). Nucleic Acids Res. 2022, 50(W1), W216–W221. [Google Scholar] [CrossRef]
- Vidarsson, G.; Dekkers, G.; Rispens, T. IgG subclasses and allotypes: From structure to effector functions. Front Immunol. 2014, 5, 520. [Google Scholar] [CrossRef]
- Stríz, I.; Trebichavský, I. Calprotectin - a pleiotropic molecule in acute and chronic inflammation. Physiol Res. 2004, 53(3), 245–253. [Google Scholar] [CrossRef]
- Burri, E.; Beglinger, C. Faecal calprotectin -- a useful tool in the management of inflammatory bowel disease. Swiss Med Wkly. 2012, 142, w13557. [Google Scholar] [CrossRef]
- Wang, S.; Song, R.; Wang, Z.; Jing, Z.; Ma, J. S100A8/A9 in Inflammation. Front Immunol. 2018, 9, 1298. [Google Scholar] [CrossRef]
- Lönnerdal, B.; Keen, C.L.; Hurley, L.S. Iron, copper, zinc, and manganese in milk. Annu Rev Nutr. 1981, 1, 149–174. [Google Scholar] [CrossRef]
- Lönnerdal, B. Excess iron intake as a factor in growth, infections, and development of infants and young children. Am J Clin Nutr. 2017, 106 (Suppl 6), 1681s–1687s. [Google Scholar] [CrossRef]
- Dempster, W.S.; Heese, H.D.; Pocock, F.H.; Kirsten, G.; Watermeyer, S. Ferritin levels in human milk. Ann Trop Paediatr. 1986, 6(3), 209–212. [Google Scholar] [CrossRef]
- Siimes, M.A.; Vuori, E.; Kuitunen, P. Breast milk iron--a declining concentration during the course of lactation. Acta Paediatr Scand. 1979, 68(1), 29–31. [Google Scholar] [CrossRef]
| Mothers of Full-Term Babies (n=20) | Mothers of Preterm Babies (n=20) | p-Value | |
|---|---|---|---|
| Characteristics of mothers | |||
| Gravidity; Me (Q1-Q3) | 1 (1-2) | 2 (1-3) | 0.1645W |
| Parity; Me (Q1-Q3) | 1 (1-2) | 2 (1-2) | 0.2057W |
| Maternal age [years]; mean (SD) | 30.2 (5.5) | 29.4 (6.1) | 0.7346T |
| Level of education; n (%) elementary secondary university vocational |
2 (10%) 9 (45%) 9 (45%) 0 (0%) |
1 (5%) 14 (70%) 4 (20%) 1 (5%) |
0.2356F |
| Maternal height [centimetres]; mean (SD) | 165.6 (6.0) | 162.4 (5.2) | 0.0839T |
| Maternal weight gain during pregnancy [kilograms]; Me (Q1-Q3) | 15.0 (12.6 – 17.0) | 9.5 (7.3 – 11.8) | 0.0003W |
| Maternal maximal weight before labour [kilograms]; Me (Q1-Q3) | 76.0 (60.8 – 85.8) | 70.0 (55.5 – 75.0) | 0.0933W |
| Maternal weight one week after labour [kilograms]; Me (Q1-Q3) | 72 (65 – 84) | 64 (58 – 85) | 0.0983W |
| Previously breastfeeding (whole group; n (%) Breastfeeding a previous child (only if there was a previous child); n (%) |
6 (30%) 6 (86%) |
10 (50%) 10 (91%) |
0.1967P 0.7324P |
| Interval between end of previous lactation [months]; mean (SD) | 74 (45) | 52 (32) | 0.3299T |
| Characteristics of the current pregnancy | |||
| Thyroid disease (any); n (%) | 5 (25%) | 2 (10%) | 0.2119P |
| Diabetes (any); n (%) | 0 (0%) | 0 (0%) | - |
| Heart problems; n (%) | 0 (0%) | 0 (0%) | - |
| Hypertension; n (%) | 3 (15%) | 4 (20%) | 0.6773P |
| Proteinuria; n (%) | 0 (0%) | 2 (10%) | 0.1468P |
| Swelling; n (%) | 2 (10%) | 5 (25%) | 0.2119P |
| Anaemia; n (%) | 4 (20%) | 3 (15%) | 0.6773P |
| Bacterial infection (any); n (%) | 4 (20%) | 4 (20%) | 0.9999P |
| Viral infection (any); n (%) | 6 (30%) | 2 (10%) | 0.1138P |
| Urinary tract infection; n (%) | 3 (15%) | 1 (5%) | 0.2918P |
| Vaginal rectal screening for GBS colonization; n (%) Positive Negative |
6 (32%) 13 (68%) |
1 (7%) 13 (93%) |
0.0897P |
| Smoking; n (%) | 1 (5%) | 2 (10%) | 0.5483P |
| Alcohol use; n (%) | 0 (0%) | 0 (0%) | - |
| Antenatal corticosteroid prophylaxis; n (%) No Yes |
20 (100%) 0 (0%) |
13 (65%) 7 (35%) |
0.0036P |
| Gestational age [weeks]; Me (Q1-Q3) | 39 (38-40) | 30 (29-31) | <0.0001W |
| Birth weight [grams], Me (Q1-Q3) | 3020 (2843 - 3515) | 1400 (1225 - 1575) | <0.0001W |
| Mode of delivery; n (%) Caesarean section Vaginal Vaginal + caesarean section (twin pregnancy) |
11 (55%) 9 (45%) 0 (0%) |
18 (90%) 1 (5%) 1 (5%) |
0.0084F |
| Newborn’s gender; n (%) Female Female + Male (twin pregnancy) Male |
12 (60%) 0 (0%) 8 (40%) |
7 (35%) 1 (5%) 12 (60%) |
0.2049F |
| UniProt Accession | Protein Name | Preterm vs Term (Fold Change) |
|---|---|---|
| Immunity | ||
| P02794 | Ferritin heavy chain | 1.5 |
| P04233 | HLA class II histocompatibility antigen gamma chain | 1.8 |
| P01717 | Immunoglobulin lambda variable 3-25 | -1.5 |
| P01861 | Immunoglobulin heavy constant gamma 4 | -1.6 |
| Signalling process | ||
| P0DTE7;P0DTE8;P0DUB6 | Alpha-amylase 1B;Alpha-amylase 1C;Alpha-amylase 1A | 1.5 |
| Q13444 |
Disintegrin and metalloproteinase domain-containing protein 15 | 1.8 |
| O00300 | Tumor necrosis factor receptor superfamily member 11B | -2.0 |
| Q6UX06 | Olfactomedin-4 | -2.3 |
| Other biological processes | ||
| P10451 | Osteopontin | 1.7 |
| Q10472 | Polypeptide N-acetylgalactosaminyltransferase 1 | -1.7 |
| UniProt Accession | Protein Name | Preterm vs Term (Fold Change) |
|---|---|---|
| Immunity | ||
| P05109 | Protein S100-A8 | 5.6 |
| P06702 | Protein S100-A9 | 5.2 |
| P00738 | Haptoglobin | 1.6 |
| P06703 | Protein S100-A6 | 2.7 |
| P01861 | Immunoglobulin heavy constant gamma 4 | -1.6 |
| P01780 | Immunoglobulin heavy variable 3-7 | 1.8 |
| A0A075B6R9;A0A0C4DH68 | Probable non-functional immunoglobulin kappa variable 2D-24;Immunoglobulin kappa variable 2-24 | 1.7 |
| A0A075B6I0 | Immunoglobulin lambda variable 8-61 | 1.9 |
| Signalling process | ||
| O00592 | Podocalyxin | 2.1 |
| Q96DA0 | Zymogen granule protein 16 homolog B | 1.8 |
| P04114 | Apolipoprotein B-100 | -1.7 |
| P12273 | Prolactin-inducible protein | 1.7 |
| Q06481 | Amyloid-like protein 2 | 1.7 |
| P0DTE7;P0DTE8;P0DUB6 | Alpha-amylase 1B;Alpha-amylase 1C;Alpha-amylase 1A | 1.6 |
| Q9UNW1 | Multiple inositol polyphosphate phosphatase 1 | -1.5 |
| Q6WN34 | Chordin-like protein 2 | -1.6 |
| Other | ||
| P13796 | Plastin-2 | 2.5 |
| P07437 | Tubulin beta chain | 1.7 |
| P16671 | Platelet glycoprotein 4 | 1.7 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.




