Submitted:
12 February 2026
Posted:
13 February 2026
You are already at the latest version
Abstract
Keywords:
Results
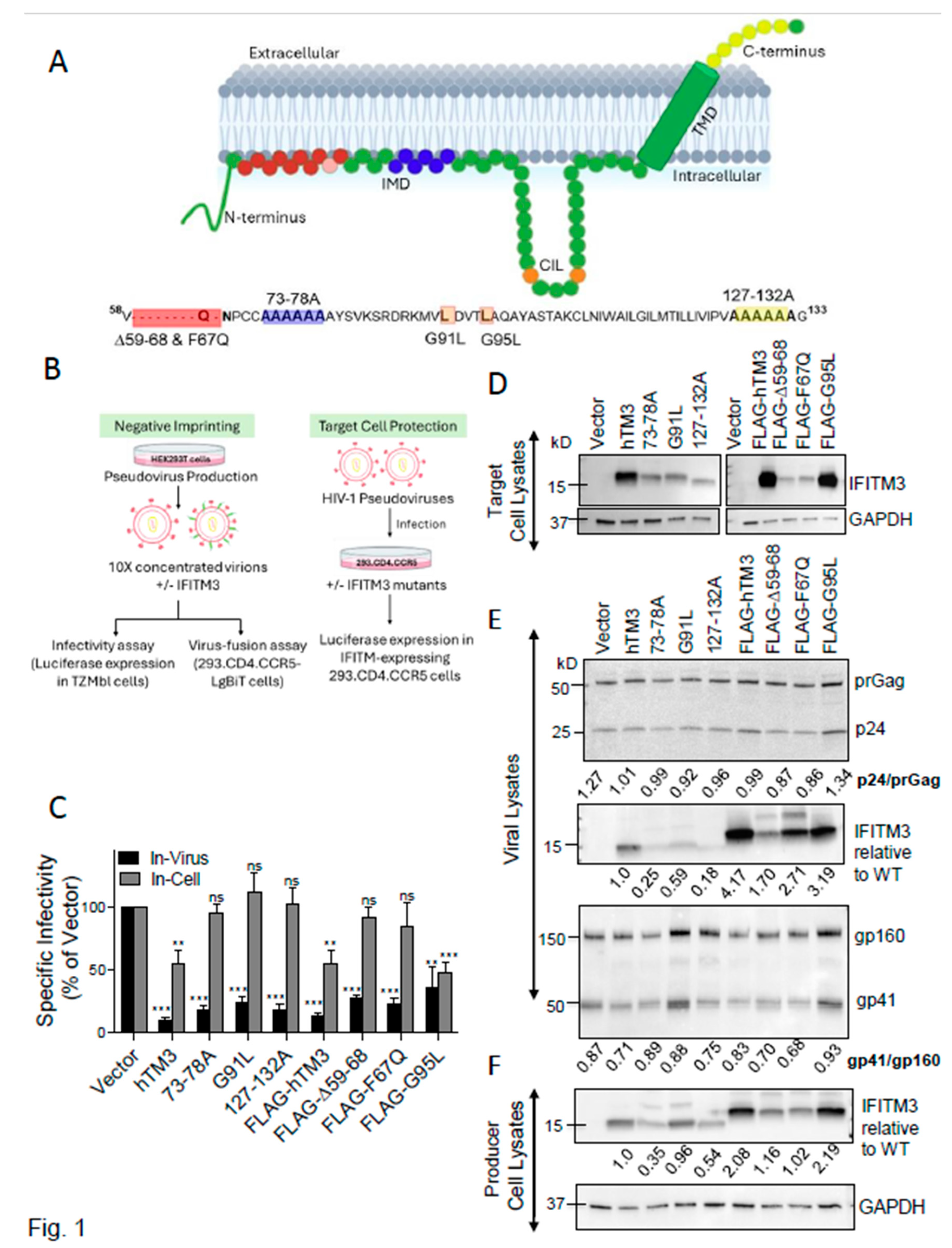
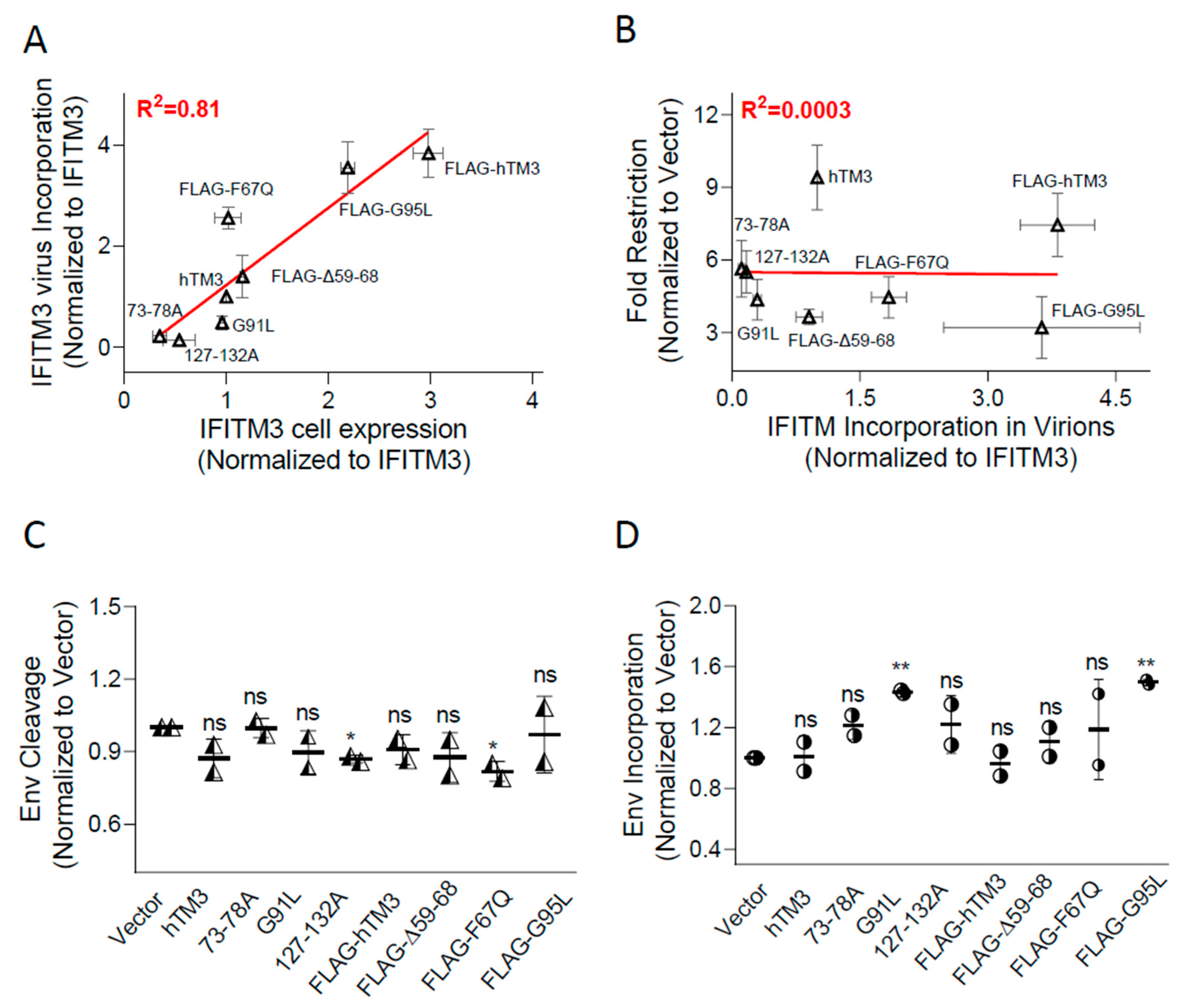
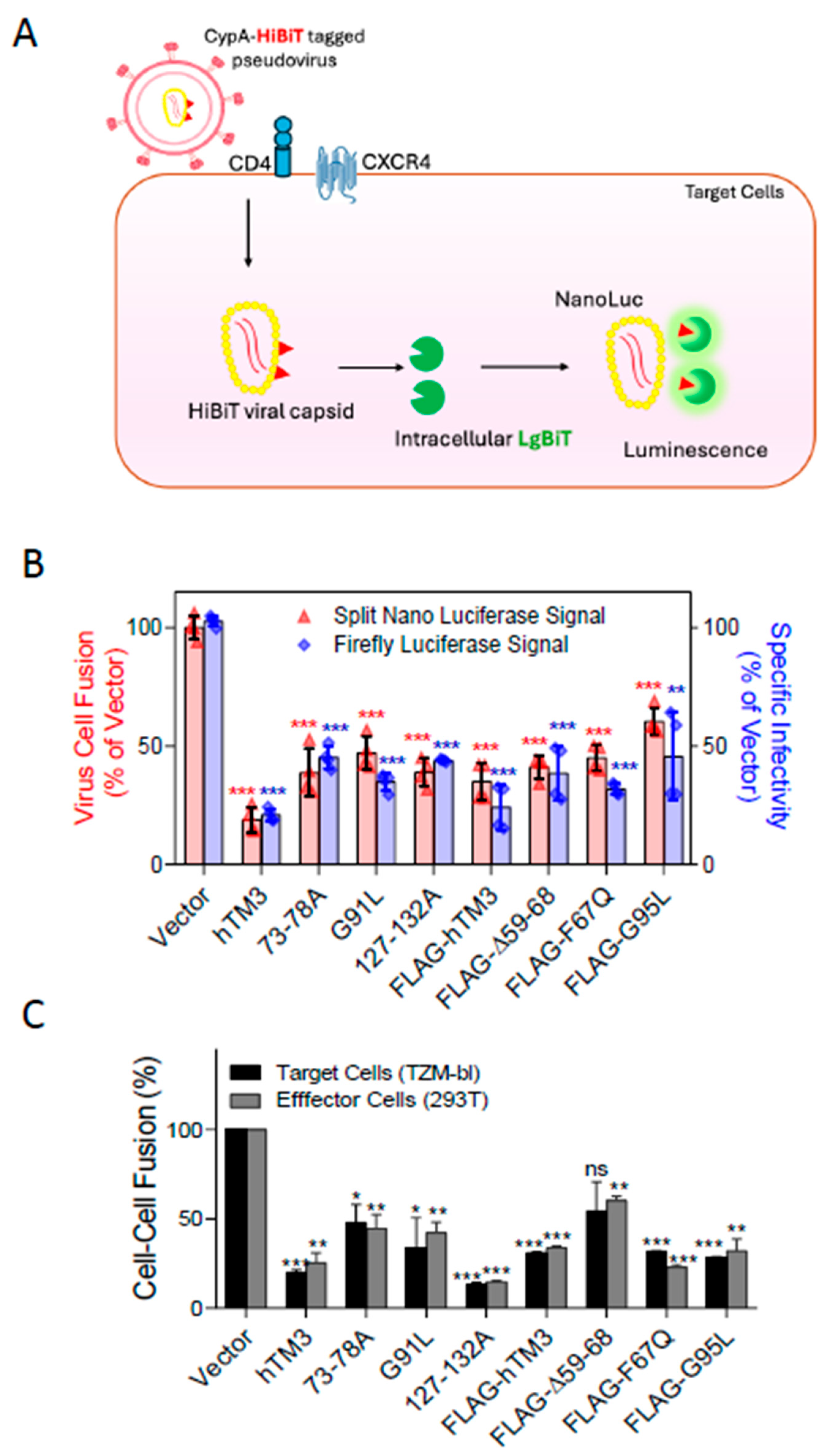
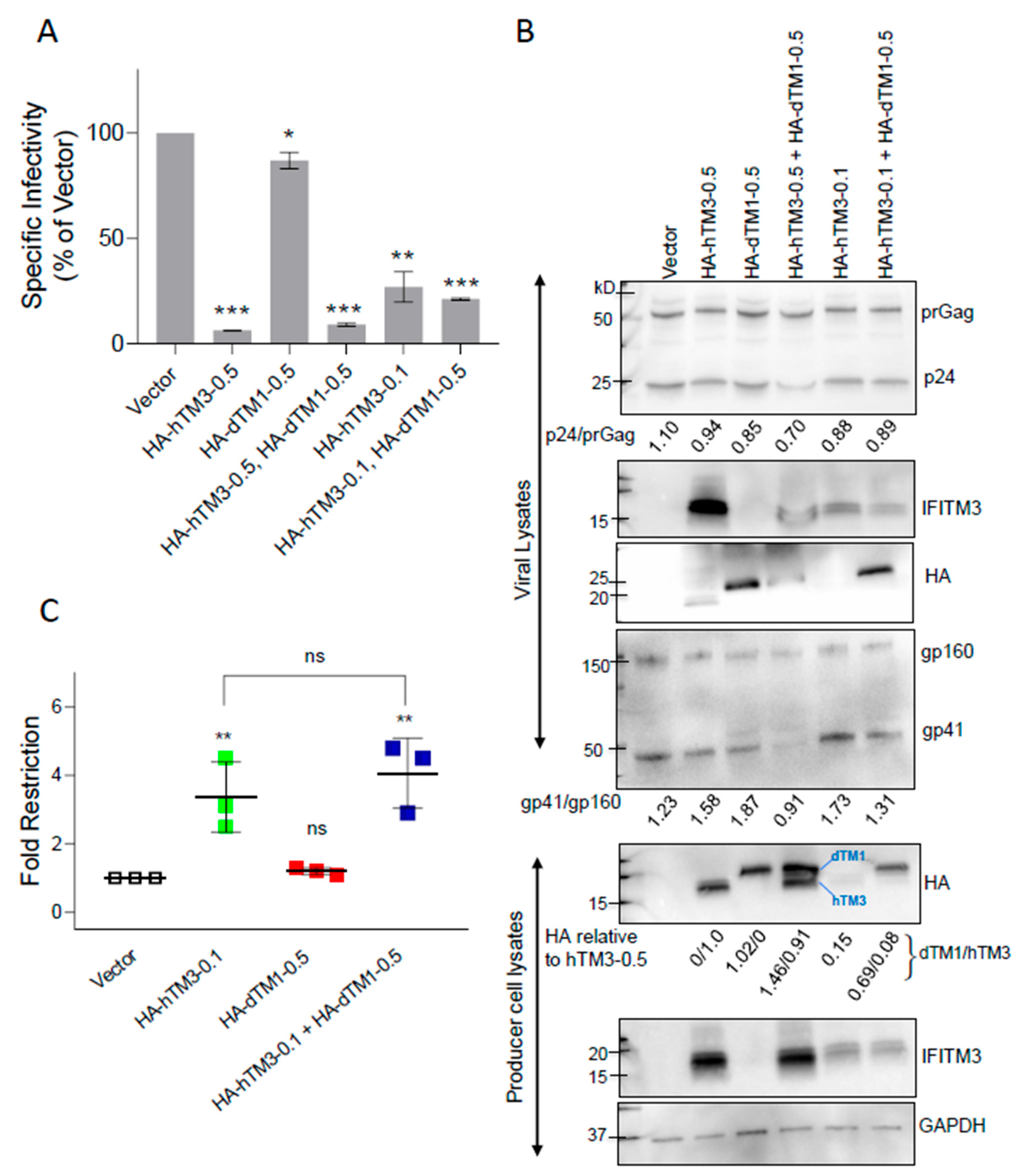
hTM3 Mutants Lacking Target Cell Protection Activity Negatively Imprint HIV-1
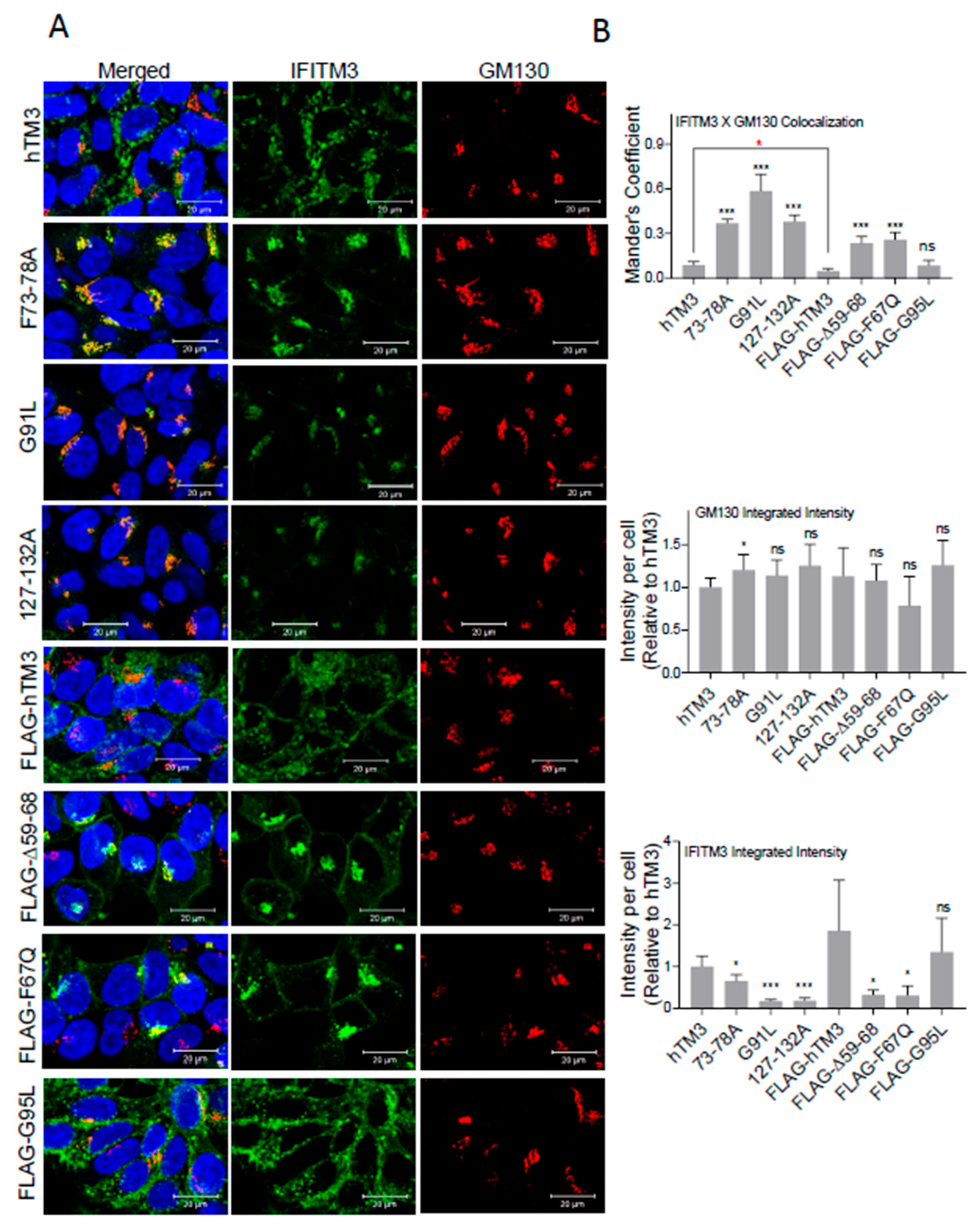
hTM3 Mutants Exhibit Altered Subcellular Localizations
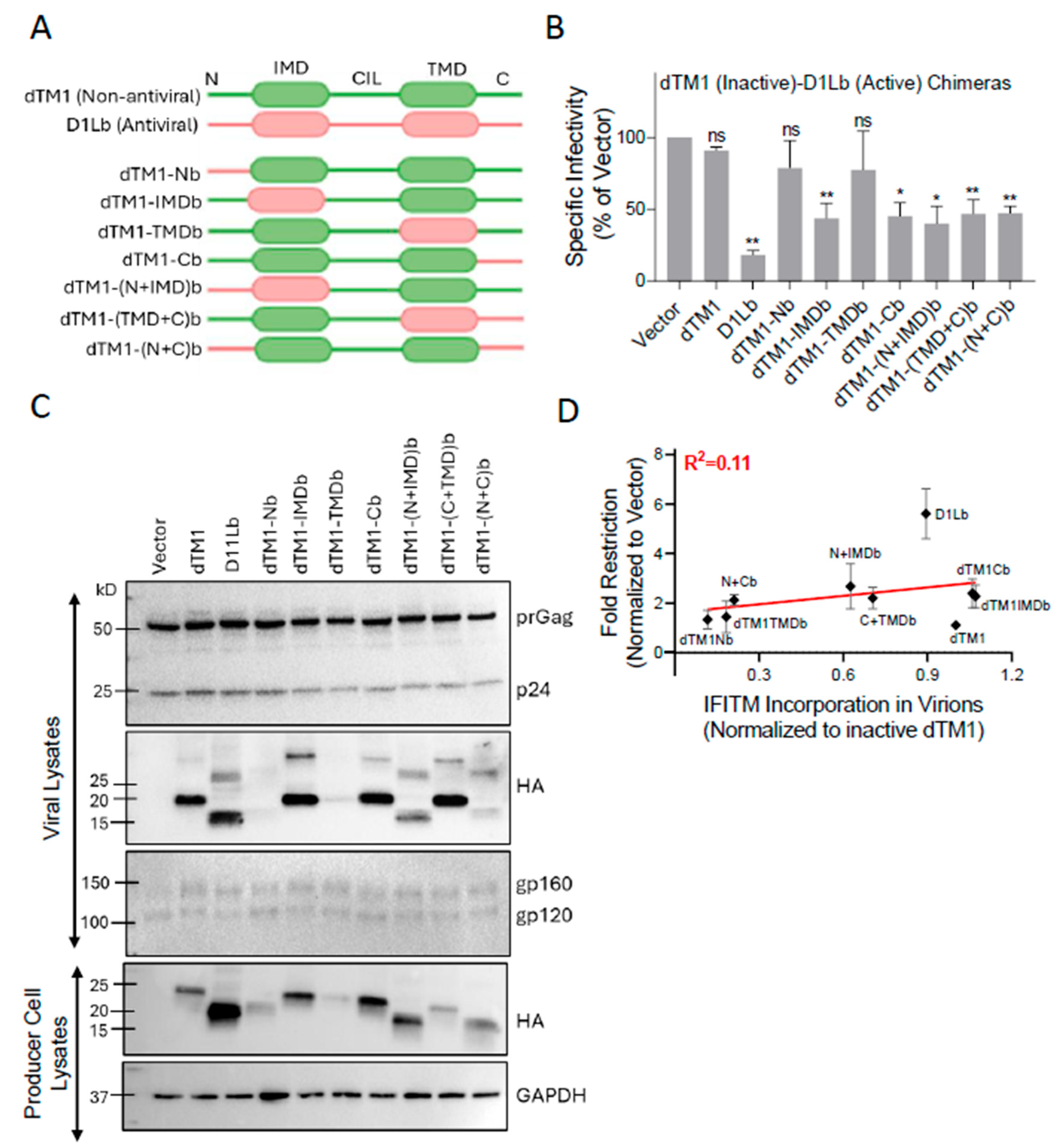
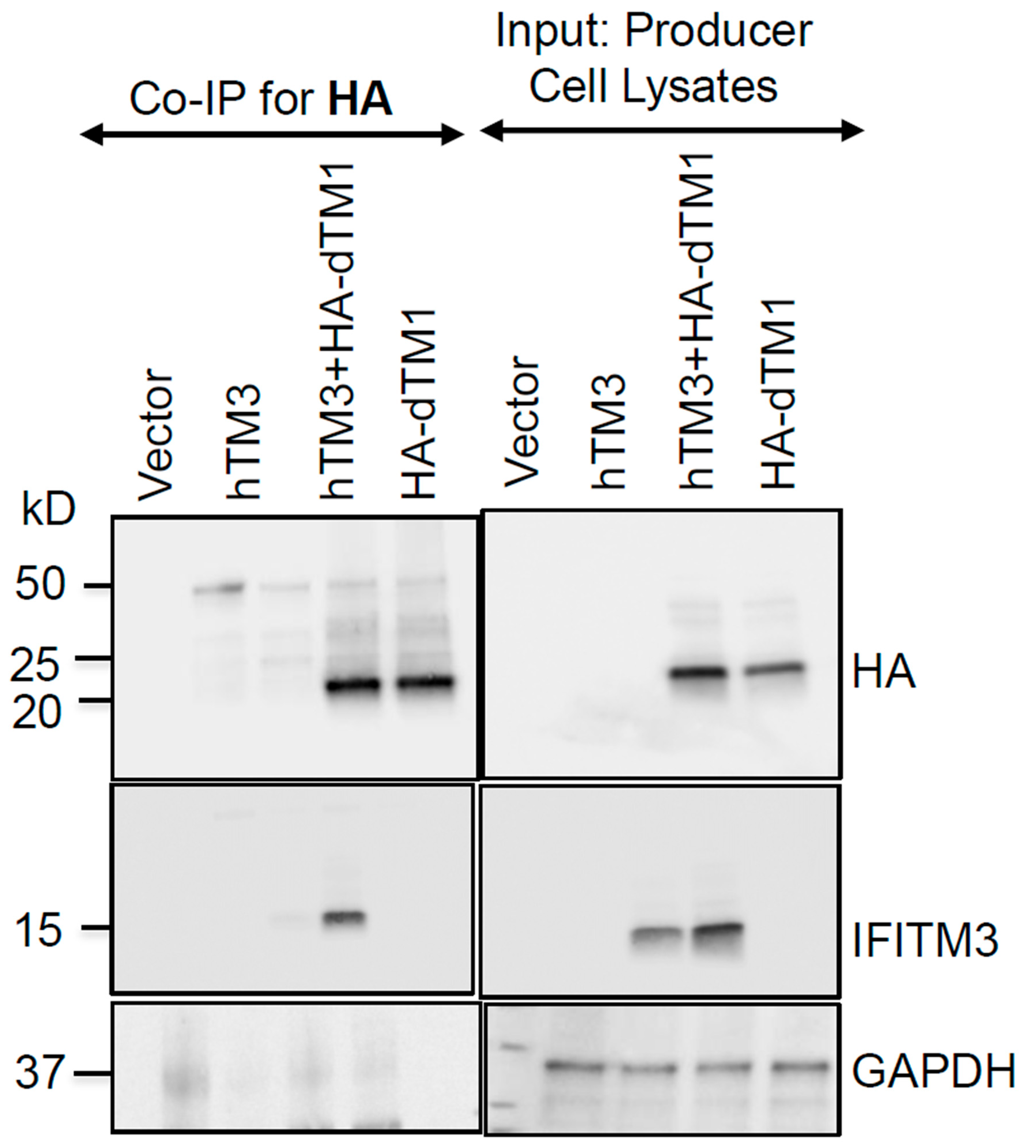
Multiple IFITM Domains Are Likely Required for Negative Imprinting of HIV-1
Discussion
Material and Methods
Cell Lines
Reagents, Plasmids, and Antibodies
Pseudovirus Production, ELISA and Infectivity Assay
Virus–Cell Fusion Assay Using Split NanoLuc Luciferase (NanoLuc) System
Western Blot and Immunoprecipitation
Cell-Cell Fusion assay
Flow Cytometry
Immunostaining, Microscopy, and Image Analysis
Statistical Analysis
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Agarwal, M.; Lai, K.K.; Wilt, I.; Majdoul, S.; Jolley, A.A.; Lewinski, M.; Compton, A.A. Restriction of HIV-1 infectivity by interferon and IFITM3 is counteracted by Nef. Science Advances 11 eadz7083. [CrossRef]
- Brass, A.L.; Huang, I.C.; Benita, Y.; John, S.P.; Krishnan, M.N.; Feeley, E.M.; Ryan, B.J.; Weyer, J.L.; van der Weyden, L.; Fikrig, E.; et al. The IFITM Proteins Mediate Cellular Resistance to Influenza A H1N1 Virus, West Nile Virus, and Dengue Virus. Cell 2009, 139, 1243–1254. [Google Scholar] [CrossRef] [PubMed]
- Perreira, J.M.; Chin, C.R.; Feeley, E.M.; Brass, A.L. IFITMs Restrict the Replication of Multiple Pathogenic Viruses. Journal of Molecular Biology 2013, 425, 4937–4955. [Google Scholar] [CrossRef]
- Wang, J.; Luo, Y.; Katiyar, H.; Liang, C.; Liu, Q. The Antiviral Activity of Interferon-Induced Transmembrane Proteins and Virus Evasion Strategies. Viruses 2024, 16. [Google Scholar] [CrossRef] [PubMed]
- Denz Parker, J.; Yount Jacob, S. IFITM3 variants point to a critical role in emergent virus infections. mBio 2025, 16, e03347-03324. [Google Scholar] [CrossRef] [PubMed]
- Chesarino, N.M.; Compton, A.A.; McMichael, T.M.; Kenney, A.D.; Zhang, L.; Soewarna, V.; Davis, M.; Schwartz, O.; Yount, J.S. IFITM3 requires an amphipathic helix for antiviral activity. EMBO reports 2017, 18, 1740-1751-1751. [Google Scholar] [CrossRef]
- Guo, X.; Steinkühler, J.; Marin, M.; Li, X.; Lu, W.; Dimova, R.; Melikyan, G.B. Interferon-Induced Transmembrane Protein 3 Blocks Fusion of Diverse Enveloped Viruses by Altering Mechanical Properties of Cell Membranes. ACS Nano 2021, 15, 8155–8170. [Google Scholar] [CrossRef]
- John Sinu, P.; Chin Christopher, R.; Perreira Jill, M.; Feeley Eric, M.; Aker Aaron, M.; Savidis, G.; Smith Sarah, E.; Elia Andrew, E.H.; Everitt Aaron, R.; Vora, M.; et al. The CD225 Domain of IFITM3 Is Required for both IFITM Protein Association and Inhibition of Influenza A Virus and Dengue Virus Replication. Journal of Virology 2013, 87, 7837–7852. [Google Scholar] [CrossRef]
- Rahman, K.; Coomer, C.A.; Majdoul, S.; Ding, S.Y.; Padilla-Parra, S.; Compton, A.A. Homology-guided identification of a conserved motif linking the antiviral functions of IFITM3 to its oligomeric state. eLife 2020, 9, e58537. [Google Scholar] [CrossRef] [PubMed]
- Huang, I.C.; Bailey, C.C.; Weyer, J.L.; Radoshitzky, S.R.; Becker, M.M.; Chiang, J.J.; Brass, A.L.; Ahmed, A.A.; Chi, X.; Dong, L.; et al. Distinct Patterns of IFITM-Mediated Restriction of Filoviruses, SARS Coronavirus, and Influenza A Virus. PLOS Pathogens 2011, 7, e1001258. [Google Scholar] [CrossRef] [PubMed]
- Savidis, G.; Perreira, Jill M.; Portmann, Jocelyn M.; Meraner, P.; Guo, Z.; Green, S.; Brass, Abraham L. The IFITMs Inhibit Zika Virus Replication. Cell Reports 2016, 15, 2323–2330. [Google Scholar] [CrossRef] [PubMed]
- Foster, Toshana L.; Wilson, H.; Iyer, Shilpa S.; Coss, K.; Doores, K.; Smith, S.; Kellam, P.; Finzi, A.; Borrow, P.; Hahn, Beatrice H.; et al. Resistance of Transmitted Founder HIV-1 to IFITM-Mediated Restriction. Cell Host & Microbe 2016, 20, 429–442. [Google Scholar] [CrossRef] [PubMed]
- Hussein, H.A.M.; Briestenska, K.; Mistrikova, J.; Akula, S.M. IFITM1 expression is crucial to gammaherpesvirus infection, in vivo. Sci Rep 2018, 8, 14105. [Google Scholar] [CrossRef]
- Everitt, A.R.; Clare, S.; McDonald, J.U.; Kane, L.; Harcourt, K.; Ahras, M.; Lall, A.; Hale, C.; Rodgers, A.; Young, D.B.; et al. Defining the Range of Pathogens Susceptible to Ifitm3 Restriction Using a Knockout Mouse Model. PLOS ONE 2013, 8, e80723. [Google Scholar] [CrossRef]
- Smith, S.E.; Busse, D.C.; Binter, S.; Weston, S.; Diaz Soria, C.; Laksono, B.M.; Clare, S.; Van Nieuwkoop, S.; Van den Hoogen, B.G.; Clement, M.; et al. Interferon-Induced Transmembrane Protein 1 Restricts Replication of Viruses That Enter Cells via the Plasma Membrane. Journal of Virology 2019, 93. [Google Scholar] [CrossRef]
- Aref, S.; Castleton, A.Z.; Bailey, K.; Burt, R.; Dey, A.; Leongamornlert, D.; Mitchell, R.J.; Okasha, D.; Fielding, A.K. Type 1 Interferon Responses Underlie Tumor-Selective Replication of Oncolytic Measles Virus. Molecular Therapy 2020, 28, 1043–1055. [Google Scholar] [CrossRef]
- Tartour, K.; Appourchaux, R.; Gaillard, J.; Nguyen, X.-N.; Durand, S.; Turpin, J.; Beaumont, E.; Roch, E.; Berger, G.; Mahieux, R.; et al. IFITM proteins are incorporated onto HIV-1 virion particles and negatively imprint their infectivity. Retrovirology 2014, 11, 103. [Google Scholar] [CrossRef]
- Li, K.; Markosyan, R.M.; Zheng, Y.-M.; Golfetto, O.; Bungart, B.; Li, M.; Ding, S.; He, Y.; Liang, C.; Lee, J.C.; et al. IFITM Proteins Restrict Viral Membrane Hemifusion. PLOS Pathogens 2013, 9, e1003124. [Google Scholar] [CrossRef]
- Desai, T.M.; Marin, M.; Chin, C.R.; Savidis, G.; Brass, A.L.; Melikyan, G.B. IFITM3 Restricts Influenza A Virus Entry by Blocking the Formation of Fusion Pores following Virus-Endosome Hemifusion. PLOS Pathogens 2014, 10, e1004048. [Google Scholar] [CrossRef]
- Compton, Alex A.; Bruel, T.; Porrot, F.; Mallet, A.; Sachse, M.; Euvrard, M.; Liang, C.; Casartelli, N.; Schwartz, O. IFITM Proteins Incorporated into HIV-1 Virions Impair Viral Fusion and Spread. Cell Host & Microbe 2014, 16, 736–747. [Google Scholar] [CrossRef]
- Yu, J.; Li, M.; Wilkins, J.; Ding, S.; Swartz, Talia H.; Esposito, Anthony M.; Zheng, Y.-M.; Freed, Eric O.; Liang, C.; Chen, Benjamin K.; et al. IFITM Proteins Restrict HIV-1 Infection by Antagonizing the Envelope Glycoprotein. Cell Reports 2015, 13, 145–156. [Google Scholar] [CrossRef]
- Tartour, K.; Nguyen, X.-N.; Appourchaux, R.; Assil, S.; Barateau, V.; Bloyet, L.-M.; Burlaud Gaillard, J.; Confort, M.-P.; Escudero-Perez, B.; Gruffat, H.; et al. Interference with the production of infectious viral particles and bimodal inhibition of replication are broadly conserved antiviral properties of IFITMs. PLOS Pathogens 2017, 13, e1006610. [Google Scholar] [CrossRef]
- Drouin, A.; Migraine, J.; Durand, M.-A.; Moreau, A.; Burlaud-Gaillard, J.; Beretta, M.; Roingeard, P.; Bouvin-Pley, M.; Braibant, M. Escape of HIV-1 Envelope Glycoprotein from Restriction of Infection by IFITM3. Journal of Virology 2021, 95, 10.1128/jvi.01994–01920. [Google Scholar] [CrossRef]
- Wang, Y.; Pan, Q.; Ding, S.; Wang, Z.; Yu, J.; Finzi, A.; Liu, S.-L.; Liang, C. The V3 Loop of HIV-1 Env Determines Viral Susceptibility to IFITM3 Impairment of Viral Infectivity. Journal of Virology 2017, 91, 10.1128/jvi.02441–02416. [Google Scholar] [CrossRef]
- Compton, A.A.; Roy, N.; Porrot, F.; Billet, A.; Casartelli, N.; Yount, J.S.; Liang, C.; Schwartz, O. Natural mutations in IFITM3 modulate post-translational regulation and toggle antiviral specificity. EMBO reports 2016, 17, 1657-1671-1671. [Google Scholar] [CrossRef] [PubMed]
- Appourchaux, R.; Delpeuch, M.; Zhong, L.; Burlaud-Gaillard, J.; Tartour, K.; Savidis, G.; Brass, A.; Etienne, L.; Roingeard, P.; Cimarelli, A. Functional Mapping of Regions Involved in the Negative Imprinting of Virion Particle Infectivity and in Target Cell Protection by Interferon-Induced Transmembrane Protein 3 against HIV-1. Journal of Virology 2019, 93. [Google Scholar] [CrossRef]
- Marziali, F.; Delpeuch, M.; Kumar, A.; Appourchaux, R.; Dufloo, J.; Tartour, K.; Etienne, L.; Cimarelli, A. Functional Heterogeneity of Mammalian IFITM Proteins against HIV-1. Journal of Virology 2021, 95, 10.1128/jvi.00439–00421. [Google Scholar] [CrossRef]
- Rahman, K.; Datta, S.A.K.; Beaven, A.H.; Jolley, A.A.; Sodt, A.J.; Compton, A.A. Cholesterol Binds the Amphipathic Helix of IFITM3 and Regulates Antiviral Activity. Journal of Molecular Biology 2022, 434, 167759. [Google Scholar] [CrossRef] [PubMed]
- Beitari, S.; Pan, Q.; Finzi, A.; Liang, C. Differential Pressures of SERINC5 and IFITM3 on HIV-1 Envelope Glycoprotein over the Course of HIV-1 Infection. Journal of Virology 2020, 94, 10.1128/jvi.00514–00520. [Google Scholar] [CrossRef] [PubMed]
- Verma, S.; Chen, Y.-C.; Marin, M.; Gillespie, S.E.; Melikyan, G.B. IFITM1 and IFITM3 Proteins Inhibit the Infectivity of Progeny HIV-1 without Disrupting Envelope Glycoprotein Clusters. Viruses 2023, 15. [Google Scholar] [CrossRef] [PubMed]
- Hall, M.P.; Unch, J.; Binkowski, B.F.; Valley, M.P.; Butler, B.L.; Wood, M.G.; Otto, P.; Zimmerman, K.; Vidugiris, G.; Machleidt, T.; et al. Engineered Luciferase Reporter from a Deep Sea Shrimp Utilizing a Novel Imidazopyrazinone Substrate. ACS Chemical Biology 2012, 7, 1848–1857. [Google Scholar] [CrossRef] [PubMed]
- Dixon, A.S.; Schwinn, M.K.; Hall, M.P.; Zimmerman, K.; Otto, P.; Lubben, T.H.; Butler, B.L.; Binkowski, B.F.; Machleidt, T.; Kirkland, T.A.; et al. NanoLuc Complementation Reporter Optimized for Accurate Measurement of Protein Interactions in Cells. ACS Chemical Biology 2016, 11, 400–408. [Google Scholar] [CrossRef]
- Nakamura, N.; Lowe, M.; Levine, T.P.; Rabouille, C.; Warren, G. The Vesicle Docking Protein p115 Binds GM130, a cis-Golgi Matrix Protein, in a Mitotically Regulated Manner. Cell 1997, 89, 445–455. [Google Scholar] [CrossRef]
- Marra, P.; Maffucci, T.; Daniele, T.; Tullio, G.D.; Ikehara, Y.; Chan, E.K.L.; Luini, A.; Beznoussenko, G.; Mironov, A.; De Matteis, M.A. The GM130 and GRASP65 Golgi proteins cycle through and define a subdomain of the intermediate compartment. Nature Cell Biology 2001, 3, 1101–1113. [Google Scholar] [CrossRef]
- Zhong, L.; Song, Y.; Marziali, F.; Uzbekov, R.; Nguyen, X.-N.; Journo, C.; Roingeard, P.; Cimarelli, A. A novel domain within the CIL regulates egress of IFITM3 from the Golgi and reveals a regulatory role of IFITM3 on the secretory pathway. Life Science Alliance 2022, 5, e202101174. [Google Scholar] [CrossRef]
- Prikryl, D.; Zhang, Y.; Verma, S.; Melikyan, G.B. Attenuation of IFITM proteins’ antiviral activity through sequestration into intraluminal vesicles of late endosomes. Frontiers in Immunology 2025, 16–2025. [Google Scholar] [CrossRef]
- Shi, G.; Schwartz, O.; Compton, A.A. More than meets the I: the diverse antiviral and cellular functions of interferon-induced transmembrane proteins. Retrovirology 2017, 14, 53. [Google Scholar] [CrossRef] [PubMed]
- Wilkins, J.; Zheng, Y.-M.; Yu, J.; Liang, C.; Liu, S.-L. Nonhuman Primate IFITM Proteins Are Potent Inhibitors of HIV and SIV. PLOS ONE 2016, 11, e0156739. [Google Scholar] [CrossRef]
- Wei, X.; Decker Julie, M.; Liu, H.; Zhang, Z.; Arani Ramin, B.; Kilby, J.M.; Saag Michael, S.; Wu, X.; Shaw George, M.; Kappes John, C. Emergence of Resistant Human Immunodeficiency Virus Type 1 in Patients Receiving Fusion Inhibitor (T-20) Monotherapy. Antimicrobial Agents and Chemotherapy 2002, 46, 1896–1905. [Google Scholar] [CrossRef] [PubMed]
- Johnston Samantha, H.; Lobritz Michael, A.; Nguyen, S.; Lassen, K.; Delair, S.; Posta, F.; Bryson Yvonne, J.; Arts Eric, J.; Chou, T.; Lee, B. A Quantitative Affinity-Profiling System That Reveals Distinct CD4/CCR5 Usage Patterns among Human Immunodeficiency Virus Type 1 and Simian Immunodeficiency Virus Strains. Journal of Virology 2009, 83, 11016–11026. [Google Scholar] [CrossRef]
- Sood, C.; Marin, M.; Chande, A.; Pizzato, M.; Melikyan, G.B. SERINC5 protein inhibits HIV-1 fusion pore formation by promoting functional inactivation of envelope glycoproteins. Journal of Biological Chemistry 2017, 292, 6014–6026. [Google Scholar] [CrossRef] [PubMed]
- Yamamoto, M.; Du, Q.; Song, J.; Wang, H.; Watanabe, A.; Tanaka, Y.; Kawaguchi, Y.; Inoue, J.-i.; Matsuda, Z. Cell–cell and virus–cell fusion assay–based analyses of alanine insertion mutants in the distal α9 portion of the JRFL gp41 subunit from HIV-1. Journal of Biological Chemistry 2019, 294, 5677–5687. [Google Scholar] [CrossRef] [PubMed]
- Markosyan, R.M.; Cohen, F.S.; Melikyan, G.B. HIV-1 envelope proteins complete their folding into six-helix bundles immediately after fusion pore formation. Mol Biol Cell 2003, 14, 926–938. [Google Scholar] [CrossRef] [PubMed]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.



