Submitted:
11 February 2026
Posted:
12 February 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Results
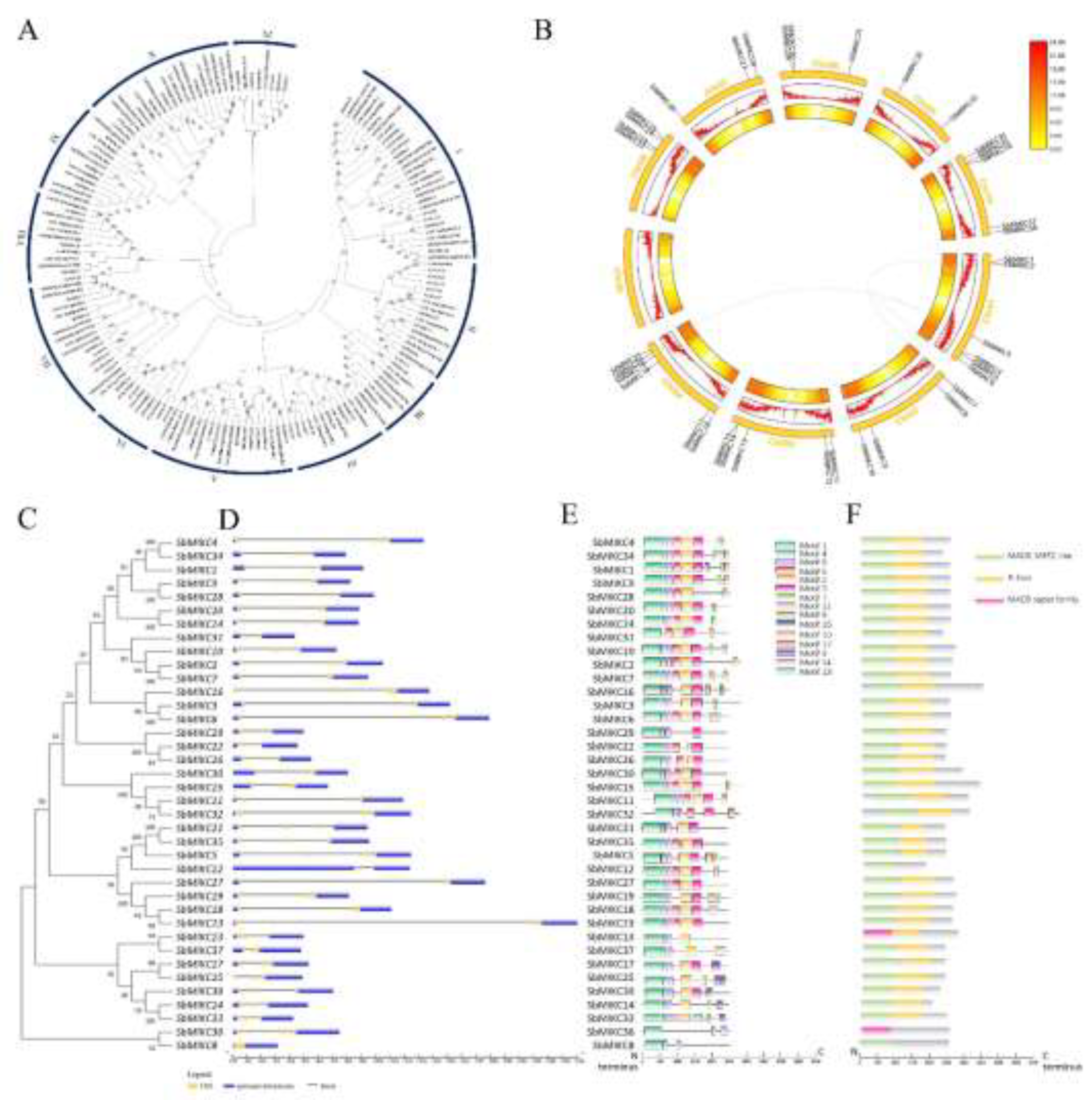
2.1. Identification of the MIKC-Type MADS Factors in Sorghum
2.2. Sequence Characterization of SbMIKCs
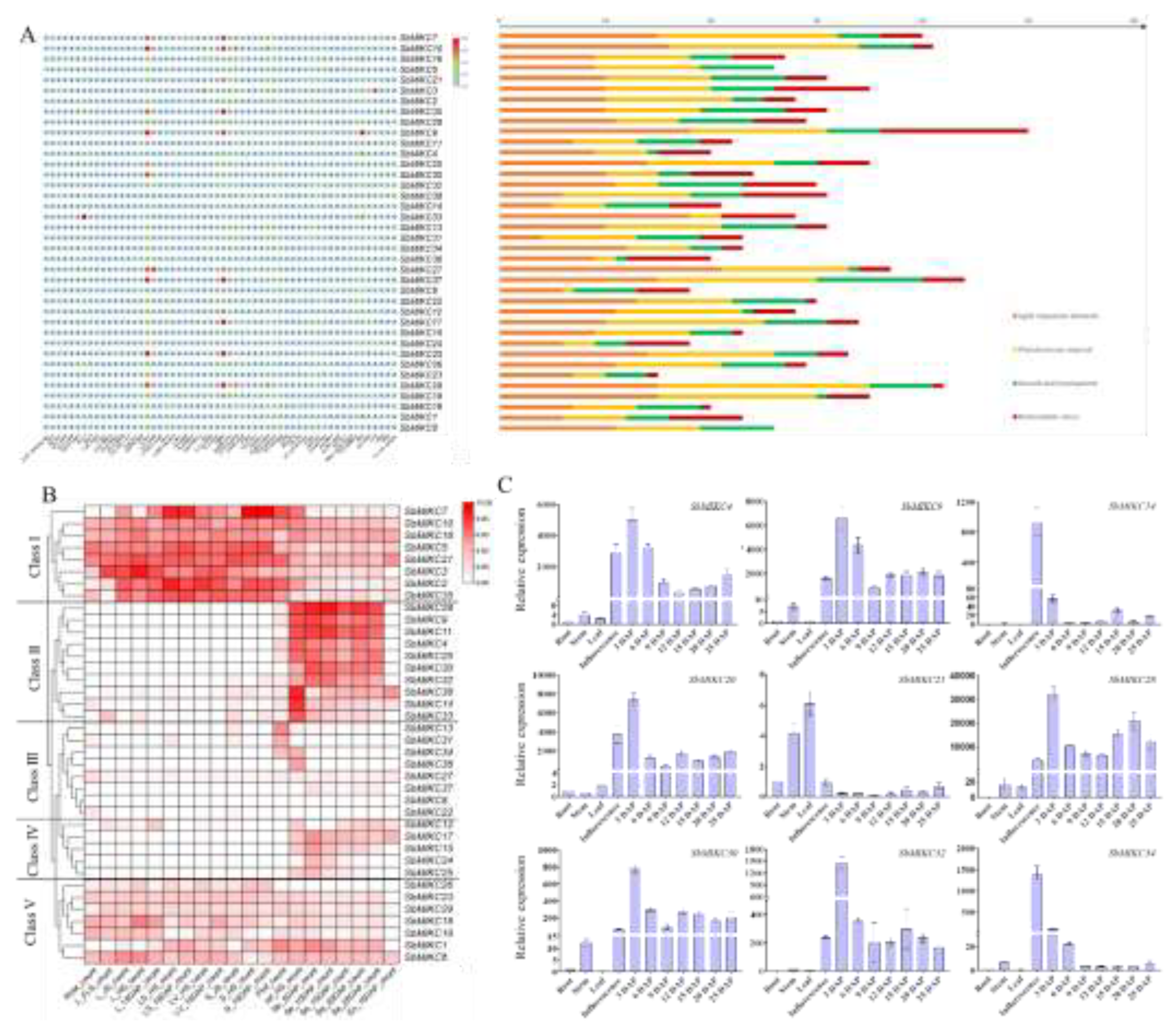
2.3. Cis-Element Architecture and Expression of SbMIKCs
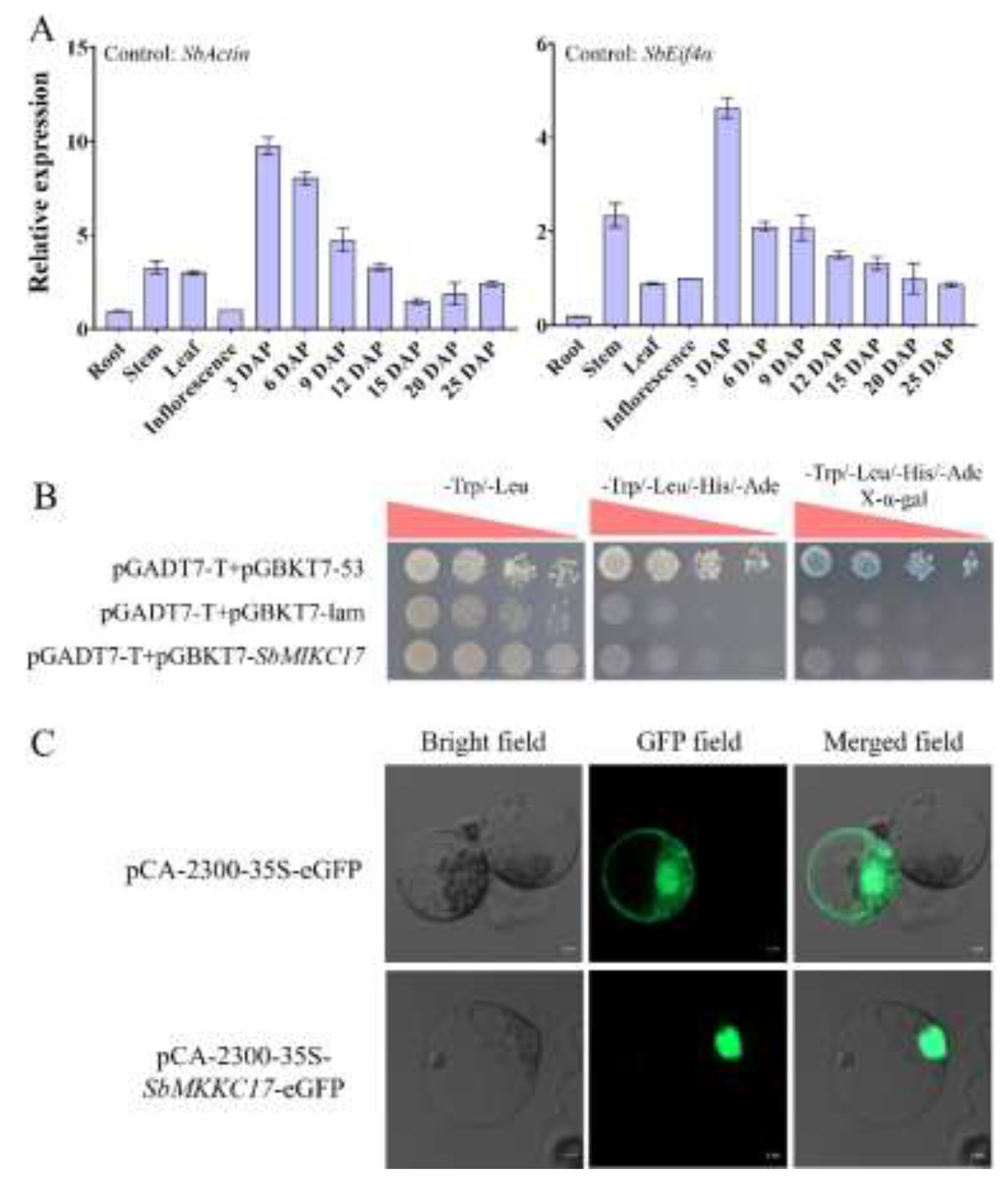
2.4. Functional Properties Analysis of SbMIKC17
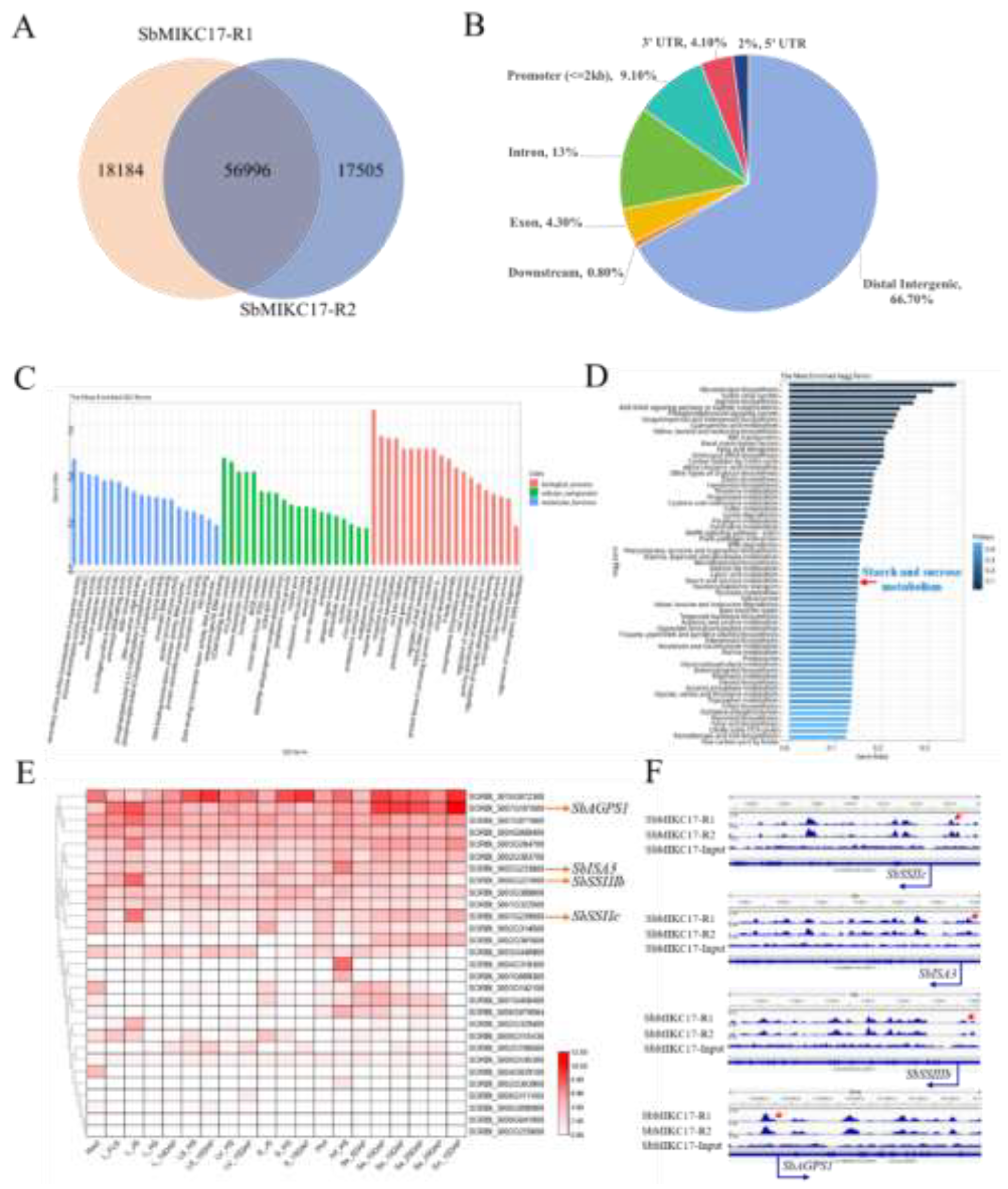
2.5. Identification of the Target Gene Regulated by SbMIKC17
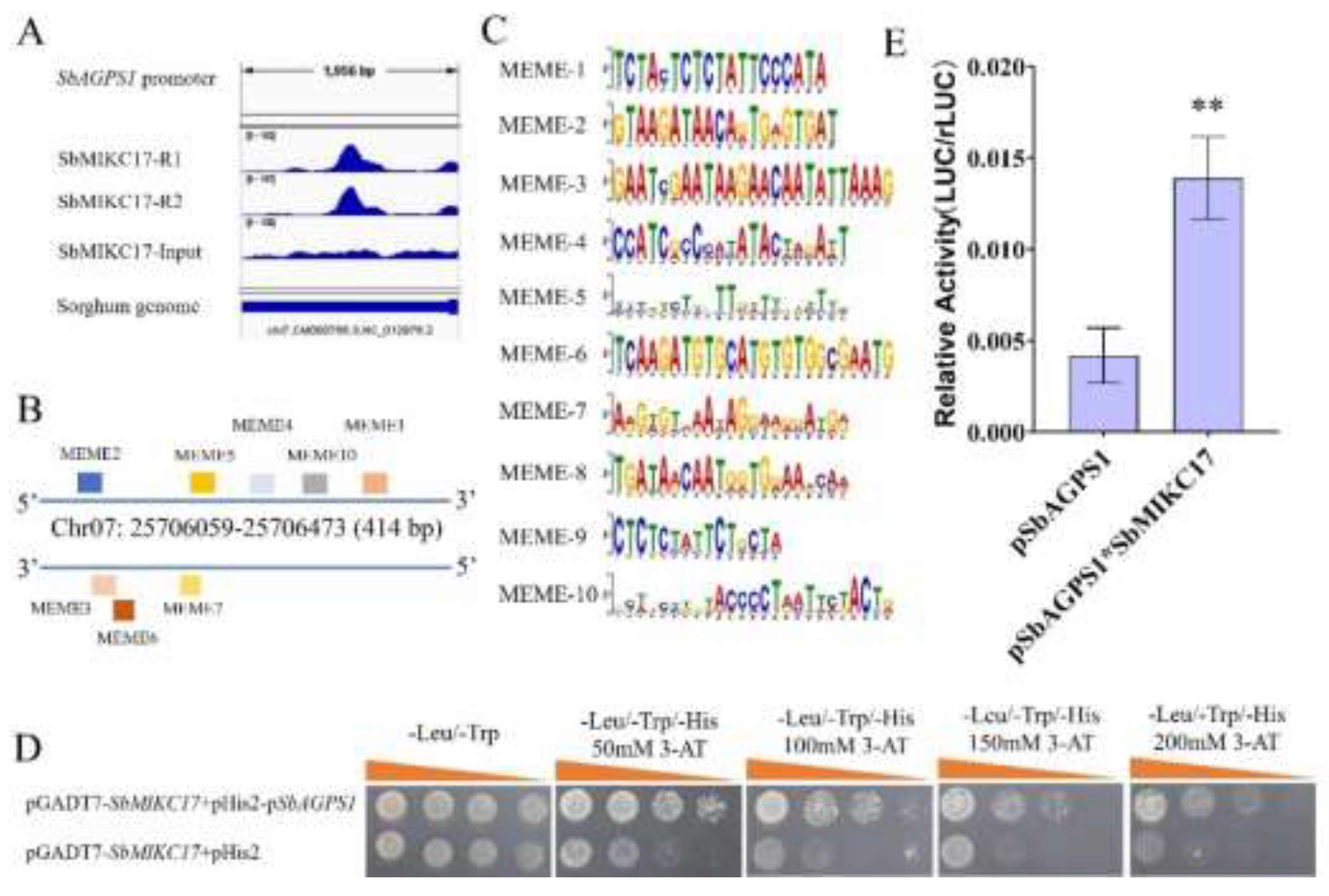
2.6. SbMIKC17 Directly Regulates the Transcription of SbAGPS1
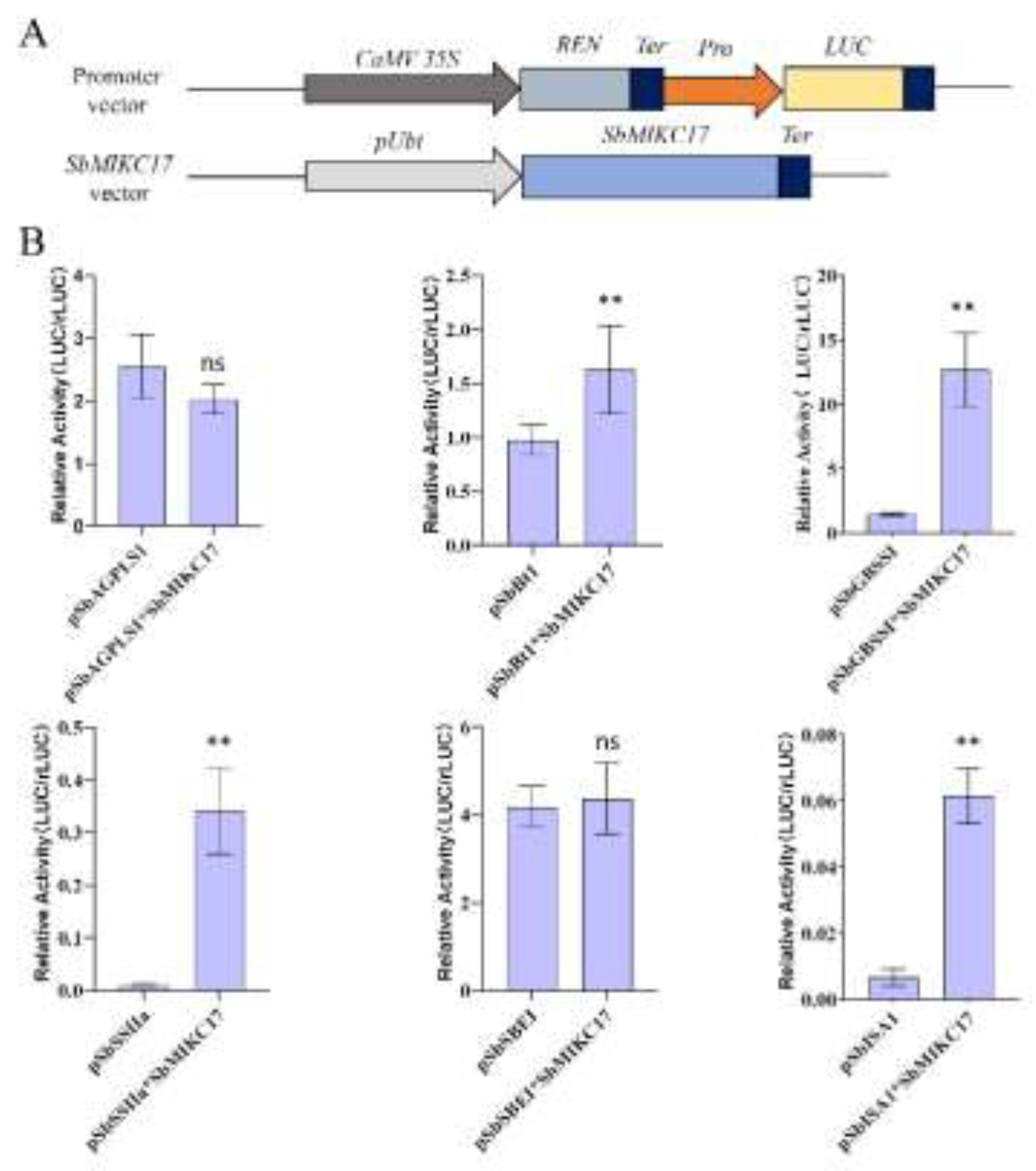
2.7. SbMIKC17 Regulates the Promoter Activities of Other Key SBRGs
3. Discussion
4. Materials and Methods
4.1. Plant Materials and Growth Conditions
4.2. Identification of MIKC-Type TFs from the Sorghum Genome
4.3. Bioinformatic Analysis of Sorghum MIKC-Type TFs
4.4. Transcriptome Data Analysis
4.5. RNA Extraction and Quantitative Real-Time PCR
4.6. Gene Cloning and Vector Constructions
4.7. Functional Property Analysis of SbMIKC17
4.8. DAP-Seq Library Preparation and Data Analysis
4.9. Dual-Luciferase Reporting Assay
4.10. Yeast One-Hybrid Assay
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
References
- Xin, Z.; Wang, M.; Cuevas, H.E.; Chen, J.; Harrison, M.; Pugh, N.A.; Morris, G. Sorghum genetic, genomic, and breeding resources. Planta 2021, 254, 114. [Google Scholar] [CrossRef]
- Hao, H.; Li, Z.; Leng, C.; Lu, C.; Luo, H.; Liu, Y.; Wu, X.; Liu, Z.; Shang, L.; Jing, H.C. Sorghum breeding in the genomic era: opportunities and challenges. Theor Appl Genet 2021, 134, 1899–1924. [Google Scholar] [CrossRef]
- Zahid, M.K.; Ahmad, D.; Amin, R.; Bao, J. Sorghum starch: Composition, structure, functionality, and strategies for its improvement. Compr Rev Food Sci Food Saf 2025, 24, e70101. [Google Scholar] [CrossRef]
- Khoddami, A.; Messina, V.; Vadabalija Venkata, K.; Farahnaky, A.; Blanchard, C.L.; Roberts, T.H. Sorghum in foods: Functionality and potential in innovative products. Crit Rev Food Sci Nutr 2023, 63, 1170–1186. [Google Scholar] [CrossRef]
- Haziman, M.L.; Ishaq, M.I.; Qonit, M.A.H.; Lestari, E.G.; Susilawati, P.N.; Widarsih, W.; Syukur, C.; Herawati, H.; Arief, R.; Santosa, B.; Purba, R.; Andoyo, R.; Yursak, Z.; Tan, S.S.; Musfal, M.; Mubarok, S. Sorghum starch review: Structural properties, interactions with proteins and polyphenols, and modification of physicochemical properties. Food Chem 2025, 463 Pt 1, 139810. [Google Scholar] [CrossRef] [PubMed]
- Cai, T.; Sun, H.; Qiao, J.; Zhu, L.; Zhang, F.; Zhang, J.; Tang, Z.; Wei, X.; Yang, J.; Yuan, Q.; Wang, W.; Yang, X.; Chu, H.; Wang, Q.; You, C.; Ma, H.; Sun, Y.; Li, Y.; Li, C.; Jiang, H.; Wang, Q.; Ma, Y. Cell-free chemoenzymatic starch synthesis from carbon dioxide. Science 2021, 373, 1523–1527. [Google Scholar] [CrossRef]
- Bahaji, A.; Li, J.; Sánchez-López, Á.M.; Baroja-Fernández, E.; Muñoz, F.J.; Ovecka, M.; Almagro, G.; Montero, M.; Ezquer, I.; Etxeberria, E. Starch biosynthesis, its regulation and biotechnological approaches to improve crop yields. Biotechnol Advances 2014, 32, 87–106. [Google Scholar] [CrossRef]
- Zeeman, S.C.; Kossmann, J.; Smith, A.M. Starch: its metabolism, evolution, and biotechnological modification in plants. Annual Review in Plant Biology 2010, 61, 209–234. [Google Scholar] [CrossRef] [PubMed]
- Keeling, P.L.; Myers, A.M. Biochemistry and genetics of starch synthesis. Annual Review of Food Science and Technology 2010, 1, 271–303. [Google Scholar] [CrossRef]
- Jeon, J.S.; Ryoo, N.; Hahn, T.R.; Walia, H.; Nakamura, Y. Starch biosynthesis in cereal endosperm. Plant Physiology and Biochemistry 2010, 48, 383–392. [Google Scholar] [CrossRef] [PubMed]
- Giroux, M.J.; Hannah, L.C. ADP-glucose pyrophosphorylase in shrunken-2 and brittle-2 mutants of maize. Mol Gen Genet 1994, 243, 400–408. [Google Scholar] [CrossRef]
- Stark, D.M.; Timmerman, K.P.; Barry, G.F.; Preiss, J.; Kishore, G.M. Regulation of the amount of starch in plant tissues by ADP Glucose pyrophosphorylase. Science 1992, 258, 287–292. [Google Scholar] [CrossRef]
- Shannon, J.C.; Pien, F.M.; Cao, H.; Liu, K.C. Brittle-1, an adenylate translocator, facilitates transfer of extraplastidial synthesized ADP-glucose into amyloplasts of maize endosperms. Plant Physiology 1998, 117, 1235–1252. [Google Scholar] [CrossRef]
- Wang, Z.Y.; Zheng, F.Q.; Shen, G.Z.; Gao, J.P.; Snustad, D.P.; Li, M.G.; Zhang, J.L.; Hong, M.M. The amylose content in rice endosperm is related to the post-transcriptional regulation of the waxy gene. Plant J 1995, 7, 613–622. [Google Scholar] [CrossRef]
- Nakamura, T.; Yamamori, M.; Hirano, H.; Hidaka, S.; Nagamine, T. Production of Waxy (Amylose-Free) Wheats. Mol Gen Genet 1995, 248, 253–259. [Google Scholar] [CrossRef]
- Seo, B.S.; Kim, S.; Scott, M.P.; Singletary, G.W.; Wong, K.S.; James, M.G.; Myers, A.M. Functional interactions between heterologously expressed starch-branching enzymes of maize and the glycogen synthases of Brewer’s yeast. Plant Physiol 2002, 128, 1189–1199. [Google Scholar] [CrossRef]
- Hussain, H.; Mant, A.; Seale, R.; Zeeman, S.; Hinchliffe, E.; Edwards, A.; Hylton, C.; Bornemann, S.; Smith, A.M.; Martin, C. Three isoforms of isoamylase contribute different catalytic properties for the debranching of potato glucans. The Plant Cell 2003, 15, 133–149. [Google Scholar] [CrossRef] [PubMed]
- Sim, L.; Beeren, S.R.; Findinier, J.; Dauvillee, D.; Ball, S.G.; Henriksen, A.; Palcic, M.M. Crystal structure of the Chlamydomonas starch debranching enzyme isoamylase ISA1 reveals insights into the mechanism of branch trimming and complex assembly. J Biol Chem 2014, 289, 22991–23003. [Google Scholar] [CrossRef] [PubMed]
- Orawetz, T.; Malinova, I.; Orzechowski, S.; Fettke, J. Reduction of the plastidial phosphorylase in potato (Solanum tuberosum L.) reveals impact on storage starch structure during growth at low temperature. Plant Physiol Biochem 2016, 100, 141–149. [Google Scholar] [CrossRef]
- Subasinghe, R.M.; Liu, F.; Polack, U.C.; Lee, E.A.; Emes, M.J.; Tetlow, I.J. Multimeric states of starch phosphorylase determine protein-protein interactions with starch biosynthetic enzymes in amyloplasts. Plant Physiol Biochem 2014, 83, 168–179. [Google Scholar] [CrossRef] [PubMed]
- Huang, L.; Tan, H.; Zhang, C.; Li, Q.; Liu, Q. Starch biosynthesis in cereal endosperms: An updated review over the last decade. Plant Commun 2021, 2, 100237. [Google Scholar] [CrossRef]
- Ahmad, D.; Ying, Y.; Bao, J. Understanding starch biosynthesis in potatoes for metabolic engineering to improve starch quality: A detailed review. Carbohydr Polym 2024, 346, 122592. [Google Scholar] [CrossRef]
- Fu, F.; Xue, H. Coexpression analysis identifies Rice Starch Regulator1, a rice AP2/EREBP family transcription factor, as a novel rice starch biosynthesis regulator. Plant Physiology 2010, 154, 927–938. [Google Scholar] [CrossRef]
- Wang, J.; Xu, H.; Zhu, Y.; Liu, Q.; Cai, X. OsbZIP58, a basic leucine zipper transcription factor, regulates starch biosynthesis in rice endosperm. Journal of Experimental Botany 2013, 64, 3453–3466. [Google Scholar] [CrossRef] [PubMed]
- Bello, B.K.; Hou, Y.; Zhao, J.; Jiao, G.; Jian, Z. NF-YB1-YC12-bHLH144 complex directly activates Wx to regulate grain quality in rice (Oryza sativa L.). Plant Biotechnol J 2019, 17, 1222–1235. [Google Scholar] [CrossRef]
- Wang, J.; Chen, Z.; Zhang, Q.; Meng, S.; Wei, C. The NAC transcription factors OsNAC20 and OsNAC26 regulate starch and storage protein synthesis. Plant physiol 2020, 184, 1775–1791. [Google Scholar] [CrossRef]
- Sun, Q.; Duan, E.; Jing, R.; Ren, Y.; Xu, H.; Gu, C.; Lv, W.; Jiang, X.; Chen, R.; Wang, Q.; Zhang, Y.; Zhang, R.; Xu, H.; Zhang, Y.; Chi, J.; Fu, Y.; Zhu, Y.; Zhang, Y.; Zhang, B.; Teng, X.; Dong, H.; Yang, X.; Zhou, L.; Tian, Y.; Liu, X.; Liu, S.; Guo, X.; Lei, C.; Jiang, L.; Wang, Y.; Wan, J. The bZIP transcription factor RISBZ1 balances grain filling and ER stress response in rice grains. Plant Commun 2025, 6, 101458. [Google Scholar] [CrossRef]
- Zhang, Z.; Dong, J.; Ji, C.; Wu, Y.; Messing, J. NAC-type transcription factors regulate accumulation of starch and protein in maize seeds. Proc Natl Acad Sci U S A 2019, 116, 11223–11228. [Google Scholar] [CrossRef] [PubMed]
- Xiao, Q.; Wang, Y.; Li, H.; Zhang, C.; Wei, B.; Wang, Y.; Huang, H.; Li, Y.; Yu, G.; Liu, H.; Zhang, J.; Liu, Y.; Hu, Y.; Huang, Y. Transcription factor ZmNAC126 plays an important role in transcriptional regulation of maize starch synthesis-related genes. Crop Journal 2021, 9, 192–203. [Google Scholar] [CrossRef]
- Xiao, Q.; Wang, Y.; Du, J.; Li, H.; Wei, B.; Wang, Y.; Li, Y.; Yu, G.; Liu, H.; Zhang, J.; Liu, Y.; Hu, Y.; Huang, Y. ZmMYB14 is an important transcription factor involved in the regulation of the activity of the ZmBT1 promoter in starch biosynthesis in maize. Febs J 2017, 284, 3079–3099. [Google Scholar] [CrossRef]
- Chen, J.; Yi, Q.; Cao, Y.; Wei, B.; Zheng, L.; Xiao, Q.; Xie, Y.; Gu, Y.; Li, Y.; Huang, H.; Wang, Y.; Hou, X.; Long, T.; Zhang, J.; Liu, H.; Liu, Y.; Yu, G.; Huang, Y. ZmbZIP91 regulates expression of starch synthesis-related genes by binding to ACTCAT elements in their promoters. J Exp Bot 2016, 67, 1327–1338. [Google Scholar] [CrossRef]
- Qing, X.; Li, J.; Lin, Z.; Wang, W.; Yi, F.; Chen, J.; Liu, Q.; Song, W.; Lai, J.; Chen, B.; Zhao, H.; Yang, Z. Maize transcription factor ZmEREB167 negatively regulates starch accumulation and kernel size. Journal of Genetics and Genomics 2025, 52, 411–421. [Google Scholar] [CrossRef]
- Gao, Y.; An, K.; Guo, W.; Chen, Y.; Zhang, R.; Zhang, X.; Chang, S.; Vincenzo, R.; Jin, F.; Cao, X.; Xin, M.; Peng, H.; Hu, Z.; Guo, W.; Du, J.; Ni, Z.; Sun, Q.; Yao, Y. The endosperm-specific transcription factor TaNAC019 regulates glutenin and starch accumulation and its elite allele improves wheat grain quality. Plant Cell 2021, 33, 603–622. [Google Scholar] [CrossRef] [PubMed]
- Liu, Z.; Gong, X.; Li, J.; Duan, H.; Dai, T.; Wei, W.; Yin, M.; Zheng, Y.; Gul, H.; Wang, J.; Liu, C.; Xiao, Q. Discovering of the transcriptional regulatory network involved in starch biosynthesis in sorghum grains. Plant Diversity 2025. [Google Scholar] [CrossRef]
- Yin, M.; Zheng, Y.; Zeng, H.; Yan, Z.; Sun, A.; Ma, K.; Gong, X.; Li, J.; Xiao, Q.; Liu, Z. SbNAC22 coordinates starch biosynthesis in sorghum grain by regulating pathway related functional genes. Journal of Integrative Agriculture 2025. [Google Scholar] [CrossRef]
- Shore, P.; Sharrocks, A.D. The MADS-Box Family of Transcription Factors. European Journal of Biochemistry 1995, 229, 1–13. [Google Scholar] [CrossRef]
- Adhikari, P.B.; Kasahara, R.D. An Overview on MADS Box Members in Plants: A Meta-Review. Int J Mol Sci 2024, 25, 8233. [Google Scholar] [CrossRef]
- Tilly, J.J.; Allen, D.W.; Jack, T. The CArG boxes in the promoter of the Arabidopsis floral organ identity gene APETALA3 mediate diverse regulatory effects. Development 1998, 125, 1647–1657. [Google Scholar] [CrossRef] [PubMed]
- Alvarez-Buylla, E.R.; Pelaz, S.; Liljegren, S.J.; Gold, S.E.; Burgeff, C.; Ditta, G.S.; Ribas de Pouplana, L.; Martínez-Castilla, L.; Yanofsky, M.F. An ancestral MADS-box gene duplication occurred before the divergence of plants and animals. Proc Natl Acad Sci U S A 2000, 97, 5328–5333. [Google Scholar] [CrossRef] [PubMed]
- Gramzow, L.; Theissen, G. A hitchhiker’s guide to the MADS world of plants. Genome Biol 2010, 11, 216. [Google Scholar] [CrossRef]
- Smaczniak, C.; Immink, R.G.H.; Angenent, G.C.; Kaufmann, K. Developmental and evolutionary diversity of plant MADS-domain factors: Insights from recent studies. Development 2012, 139, 3081–3098. [Google Scholar] [CrossRef]
- Han, J.P.; Wan, J.N.; Guan, Z.L.; Xu, H.; Wang, Q.F.; Wan, T. The origin, evolution, and diversification of MADS-box transcription factors in green plants. Plant Commun 2025, 6, 101462. [Google Scholar] [CrossRef]
- Kaufmann, K.; Melzer, R.; Theißen, G. MIKC-type MADS-domain proteins: structural modularity, protein interactions and network evolution in land plants. Gene 2005, 347, 183–198. [Google Scholar] [CrossRef]
- Henschel, K.; Kofuji, R.; Hasebe, M.; Saedler, H.; Munster, T.; Thei?En, G. Two Ancient Classes of MIKC-type MADS-box Genes are Present in the Moss Physcomitrella patens. Molecular Biology Evolution 2002, 19, 801–814. [Google Scholar] [CrossRef]
- Gramzow, L.; Theissen, G. Phylogenomics reveals surprising sets of essential and dispensable clades of MIKC(c)-group MADS-box genes in flowering plants. J Exp Zool B Mol Dev Evol 2015, 324, 353–362. [Google Scholar] [CrossRef]
- Schilling, S.; Pan, S.; Kennedy, A.; Melzer, R. MADS-box genes and crop domestication: the jack of all traits. J Exp Bot 2018, 69, 1447–1469. [Google Scholar] [CrossRef] [PubMed]
- Yamaguchi, T.; Hirano, H.Y. Function and diversification of MADS-box genes in rice. ScientificWorldJournal 2006, 6, 1923–1932. [Google Scholar] [CrossRef] [PubMed]
- Wu, Q.; Wu, Y.; Li, R.; Cao, H.; Li, Z.; Li, Q.; Zhou, L. Research Progress on the Regulation of Plant Floral Organ Development by the MADS-box Gene Family. Int J Mol Sci 2025, 26. [Google Scholar] [CrossRef]
- Colombo, M.; Masiero, S.; Vanzulli, S.; Lardelli, P.; Kater, M.M.; Colombo, L. AGL23, a type I MADS-box gene that controls female gametophyte and embryo development in Arabidopsis. Plant J 2008, 54, 1037–1048. [Google Scholar] [CrossRef] [PubMed]
- Bemer, M.; Heijmans, K.; Airoldi, C.; Davies, B.; Angenent, G.C. An atlas of type I MADS box gene expression during female gametophyte and seed development in Arabidopsis. Plant Physiol 2010, 154, 287–300. [Google Scholar] [CrossRef]
- Masiero, S.; Colombo, L.; Grini, P.E.; Schnittger, A.; Kater, M.M. The emerging importance of type I MADS box transcription factors for plant reproduction. Plant Cell 2011, 23, 865–872. [Google Scholar] [CrossRef]
- Bente, H.; Kohler, C. Molecular basis and evolutionary drivers of endosperm-based hybridization barriers. Plant Physiol 2024, 195, 155–169. [Google Scholar] [CrossRef] [PubMed]
- Malabarba, J.; Buffon, V.; Mariath, J.E.A.; Gaeta, M.L.; Dornelas, M.C.; Margis-Pinheiro, M.; Pasquali, G.; Revers, L.F. The MADS-box gene Agamous-like 11 is essential for seed morphogenesis in grapevine. J Exp Bot 2017, 68, 1493–1506. [Google Scholar] [CrossRef]
- Silva, C.S.; Sriharsha, P.; Adam, R.; Martha, B.; Agnès, J.; François, P.; Veronique, H.; Chloe, Z. Evolution of the Plant Reproduction Master Regulators LFY and the MADS Transcription Factors: The Role of Protein Structure in the Evolutionary Development of the Flower. Frontiers in Plant Science 2016, 6, 1193. [Google Scholar] [CrossRef]
- Tripathi, A.; Vishwakarma, K.; Tripathi, S.; Jadaun, J.S.; Nayak, A.K. Utilization of MADS-Box genes for agricultural advancement: current insights and future prospects. Mol Biol Rep 2025, 53, 20. [Google Scholar] [CrossRef]
- Liu, Y.; Cui, S.; Wu, F.; Yan, S.; Lin, X.; Du, X.; Chong, K.; Schilling, S.; Theissen, G.; Meng, Z. Functional conservation of MIKC*-Type MADS box genes in Arabidopsis and rice pollen maturation. Plant Cell 2013, 25, 1288–1303. [Google Scholar] [CrossRef] [PubMed]
- Lee, J.; Lee, I. Regulation and function of SOC1, a flowering pathway integrator. J Exp Bot 2010, 61, 2247–2254. [Google Scholar] [CrossRef]
- Adamczyk, B.J.; Lehti-Shiu, M.D.; Fernandez, D.E. The MADS domain factors AGL15 and AGL18 act redundantly as repressors of the floral transition in Arabidopsis. Plant J 2007, 50, 1007–1019. [Google Scholar] [CrossRef] [PubMed]
- Huang, Q.; Chen, X.; Zhong, S.; Wu, S.; Guo, J.; Wang, Q.; Li, J.; Li, D.; Xia, Y.; Zhang, J.; Wang, X. MIKC-Type MADS-Box Gene Analysis Reveals the Role of PlSOC1 in Bud Dormancy Transition in Herbaceous Peony. Plants (Basel) 2025, 14, 928. [Google Scholar] [CrossRef]
- Quesada-Traver, C.; Guerrero, B.I.; Badenes, M.L.; Rodrigo, J.; Rios, G.; Lloret, A. Structure and Expression of Bud Dormancy-Associated MADS-Box Genes (DAM) in European Plum. Front Plant Sci 2020, 11, 1288. [Google Scholar] [CrossRef]
- Ma, M.M.; Zhang, H.F.; Tian, Q.; Wang, H.C.; Zhang, F.Y.; Tian, X.; Zeng, R.F.; Huang, X.M. MIKC type MADS-box transcription factor LcSVP2 is involved in dormancy regulation of the terminal buds in evergreen perennial litchi (Litchi chinensis Sonn.). Hortic Res 2024, 11, uhae150. [Google Scholar] [CrossRef] [PubMed]
- Adamczyk, B.J.; Fernandez, D.E. MIKC* MADS domain heterodimers are required for pollen maturation and tube growth in Arabidopsis. Plant Physiol 2009, 149, 1713–1723. [Google Scholar] [CrossRef]
- Alvarez-Buylla, E.R.; Garcia-Ponce, B.; Sanchez, M.P.; Espinosa-Soto, C.; Garcia-Gomez, M.L.; Pineyro-Nelson, A.; Garay-Arroyo, A. MADS-box genes underground becoming mainstream: plant root developmental mechanisms. New Phytol 2019, 223, 1143–1158. [Google Scholar] [CrossRef]
- Griebel, T.; Zeier, J. Light regulation and daytime dependency of inducible plant defenses in Arabidopsis: phytochrome signaling controls systemic acquired resistance rather than local defense. Plant Physiol 2008, 147, 790–801. [Google Scholar] [CrossRef]
- Zhang, Z; Zou, W; Lin, P; Wang, Z; Chen, Y; Yang, X; Zhao, W; Zhang, Y; Wang, D; Que, Y; Wu, Q. Evolution and Function of MADS-Box Transcription Factors in Plants. International Journal of Molecular Sciences 2024b, 25. [Google Scholar] [CrossRef] [PubMed]
- Zha, K.; Xie, H.; Ge, M.; Wang, Z.; Wang, Y.; Si, W.; Gu, L. Expression of Maize MADS Transcription Factor ZmES22 Negatively Modulates Starch Accumulation in Rice Endosperm. Int J Mol Sci 2019, 20. [Google Scholar] [CrossRef]
- Dong, Q.; Wang, F.; Kong, J.; Xu, Q.; Li, T.; Chen, L.; Chen, H.; Jiang, H.; Li, C.; Cheng, B. Functional analysis of ZmMADS1a reveals its role in regulating starch biosynthesis in maize endosperm. Sci Rep 2019, 9, 3253. [Google Scholar] [CrossRef]
- Song, G.Q.; Han, X.; Ryner, J.T.; Thompson, A.; Wang, K. Utilizing MIKC-type MADS-box protein SOC1 for yield potential enhancement in maize. Plant Cell Rep 2021, 40, 1679–1693. [Google Scholar] [CrossRef]
- Feng, T.; Wang, L.; Li, L.; Liu, Y.; Chong, K.; Theissen, G.; Meng, Z. OsMADS14 and NF-YB1 cooperate in the direct activation of OsAGPL2 and Waxy during starch synthesis in rice endosperm. New Phytol 2022, 234, 77–92. [Google Scholar] [CrossRef] [PubMed]
- Li, R.; Wu, M.W.; Liu, J.; Xu, X.; Bao, Y.; Liu, C.M. NAC25 transcription factor regulates the degeneration of cytoplasmic membrane integrity and starch biosynthesis in rice endosperm through interacting with MADS29. Front Plant Sci 2025, 16, 1563065. [Google Scholar] [CrossRef] [PubMed]
- Zhang, H.; Xu, H.; Feng, M.; Zhu, Y. Suppression of OsMADS7 in rice endosperm stabilizes amylose content under high temperature stress. Plant Biotechnol J 2018, 16, 18–26. [Google Scholar] [CrossRef]
- Xiao, Q.; Huang, T.; Cao, W.; Ma, K.; Liu, T.; Xing, F.; Ma, Q.; Duan, H.; Ling, M.; Ni, X.; Liu, Z. Profiling of transcriptional regulators associated with starch biosynthesis in sorghum (Bicolor sorghum L.). Frontiers in Plant Science 2022, 13, 999747. [Google Scholar] [CrossRef] [PubMed]
- Zhao, D.; Chen, Z.; Xu, L.; Zhang, L.; Zou, Q. Genome-Wide Analysis of the MADS-Box Gene Family in Maize: Gene Structure, Evolution, and Relationships. Genes (Basel) 2021, 12. [Google Scholar] [CrossRef]
- Zhao, Y.; Li, X.; Chen, W.; Peng, X.; Cheng, X.; Zhu, S.; Cheng, B. Whole-genome survey and characterization of MADS-box gene family in maize and sorghum. Plant Cell, Tissue and Organ Culture (PCTOC) 2010, 105, 159–173. [Google Scholar] [CrossRef]
- Abdullah-Zawawi, M.R.; Ahmad-Nizammuddin, N.F.; Govender, N.; Harun, S.; Mohd-Assaad, N.; Mohamed-Hussein, Z.A. Comparative genome-wide analysis of WRKY, MADS-box and MYB transcription factor families in Arabidopsis and rice. Sci Rep 2021, 11, 19678. [Google Scholar] [CrossRef]
- Wang, F.; Zhou, Z.; Zhu, L.; Gu, Y.; Guo, B.; Lv, C.; Zhu, J.; Xu, R. Genome-wide analysis of the MADS-box gene family involved in salt and waterlogging tolerance in barley (Hordeum vulgare L.). Front Plant Sci 2023, 14, 1178065. [Google Scholar] [CrossRef]
- Zhang, L; Ma, F; Duan, G; Ju, Y; Yu, T; Zhang, Q; Sodmergen. MIKC*-type MADS transcription factors control JINGUBANG expression and the degree of pollen dormancy in Arabidopsis. Plant Physiol 2024a, 197. [Google Scholar] [CrossRef] [PubMed]
- Mou, Y.; Yuan, C.; Sun, Q.; Yan, C.; Zhao, X.; Wang, J.; Wang, Q.; Shan, S.; Li, C. MIKC-type MADS-box transcription factor gene family in peanut: Genome-wide characterization and expression analysis under abiotic stress. Front Plant Sci 2022, 13, 980933. [Google Scholar] [CrossRef]
- Paul, P.; Dhatt, B.K.; Miller, M.; Folsom, J.J.; Wang, Z.; Krassovskaya, I.; Liu, K.; Sandhu, J.; Yu, H.; Zhang, C.; Obata, T.; Staswick, P.; Walia, H. MADS78 and MADS79 Are Essential Regulators of Early Seed Development in Rice. Plant Physiol 2020, 182, 933–948. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Yang, T.; Li, Y.; Hou, J.; He, J.; Ma, N.; Zhou, X. Genome-wide identification and expression analysis of MIKC(C) genes in rose provide insight into their effects on flower development. Front Plant Sci 2022, 13, 1059925. [Google Scholar] [CrossRef]
- Yin, X.; Gao, Q.; Wang, F.; Liu, W.; Yu, S.; Zhong, S.; Feng, J.; Bai, R.; Luo, Y.; Chen, L.; Dai, X.; Liang, M. MIKC-type MADS-box transcription factor OsMADS31 positively regulates salinity tolerance in rice. Front Plant Sci 2025, 16, 1628305. [Google Scholar] [CrossRef]
- Li, Y.; Wang, J.; Zhong, S.; Huo, Q.; Wang, Q.; Shi, Y.; Liu, H.; Liu, J.; Song, Y.; Fang, X.; Lin, Z. MADS-box encoding gene Tunicate1 positively controls maize yield by increasing leaf number above the ear. Nat Commun 2024, 15, 9799. [Google Scholar] [CrossRef]
- Song, G.Q.; Chen, Q. Overexpression of the MADS-box gene K-domain increases the yield potential of blueberry. Plant Sci 2018, 276, 22–31. [Google Scholar] [CrossRef] [PubMed]
- Chen, C.; Wu, Y.; Li, J.; Wang, X.; Zeng, Z.; Xu, J.; Liu, Y.; Feng, J.; Chen, H.; He, Y.; Xia, R. TBtools-II: A “one for all, all for one” bioinformatics platform for biological big-data mining. Mol Plant 2023, 16, 1733–1742. [Google Scholar] [CrossRef] [PubMed]
- Tamura, K.; Stecher, G.; Kumar, S. MEGA11: Molecular Evolutionary Genetics Analysis Version 11. Mol Biol Evol 2021, 38, 3022–3027. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Liu, T.; Meyer, C.A.; Eeckhoute, J.; Johnson, D.S.; Bernstein, B.E.; Nusbaum, C.; Myers, R.M.; Brown, M.; Li, W.; Liu, X.S. Model-based analysis of ChIP-Seq (MACS). Genome Biol. 2008, 9, R137. [Google Scholar] [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).



