Submitted:
10 February 2026
Posted:
12 February 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Materials and Methods
2.1. Retrieval of the FIV Sequences and the Multiple Sequence Alignment (MSA)
2.2. Prediction and Mapping of Some Common T Cell Epitopes Across Some Key Target FIV Proteins
2.3. Prediction and Mapping of Some Common B Cell Epitopes of the Target Viral Proteins
2.3. The Immunological and Safety Profiling of the Mapped FIV Epitopes Across the Key Proteins
2.4. Prediction of the Ability of Mapped T- Cell Epitopes for the Feline Cytokine Production and Cross Reactivity
2.5. In Silico Construction of the FIV Multiepitope DNA Vaccine
2.5. Analysis of the Physicochemical and the Peptide Solubility Properties of the FIV Vaccine Construct
2.6. Prediction of the Secondary and Tertiary Structures of the Potentially Expressed Proteins
2.7. Molecular Docking and Simulation
2.8. In Silico Prediction of the Immune Stimulation Simulation of the Designed FIV Vaccine Construct
2.9. In Silico Cloning of the Multiepitope FIV Vaccine Construct
2.10. Software
| Software | Reference | Software | Reference |
|---|---|---|---|
| Geneious | - | Alphafold | [44,45] |
| ABCpred | [38] | Colabfold | [46] |
| IEDB | [47] | ProSA | [48,49] |
| NCBI Blastp | - | MolProbity | [50] |
| Vaxijenv2.0 | [51] | GalaxyRefine | [52,53] |
| Allertop v2.1 | [54] | HDock | [55] |
| Toxinpred | [56] | PDBePISA | [57] |
| TMHMM-2.0 | [58] | C-Immsim | [59] |
| NETPHOS-3.1 | [60,61] | Addgene | - |
| Expasy [62] | [62] | Snapgene | - |
| NetNGlyc 1.0 | [63] | IL4Pred | [64] |
| NPS | - | IFNEpitope | [65] |
3. Results
3.1. FIV Multiple Sequence Analysis
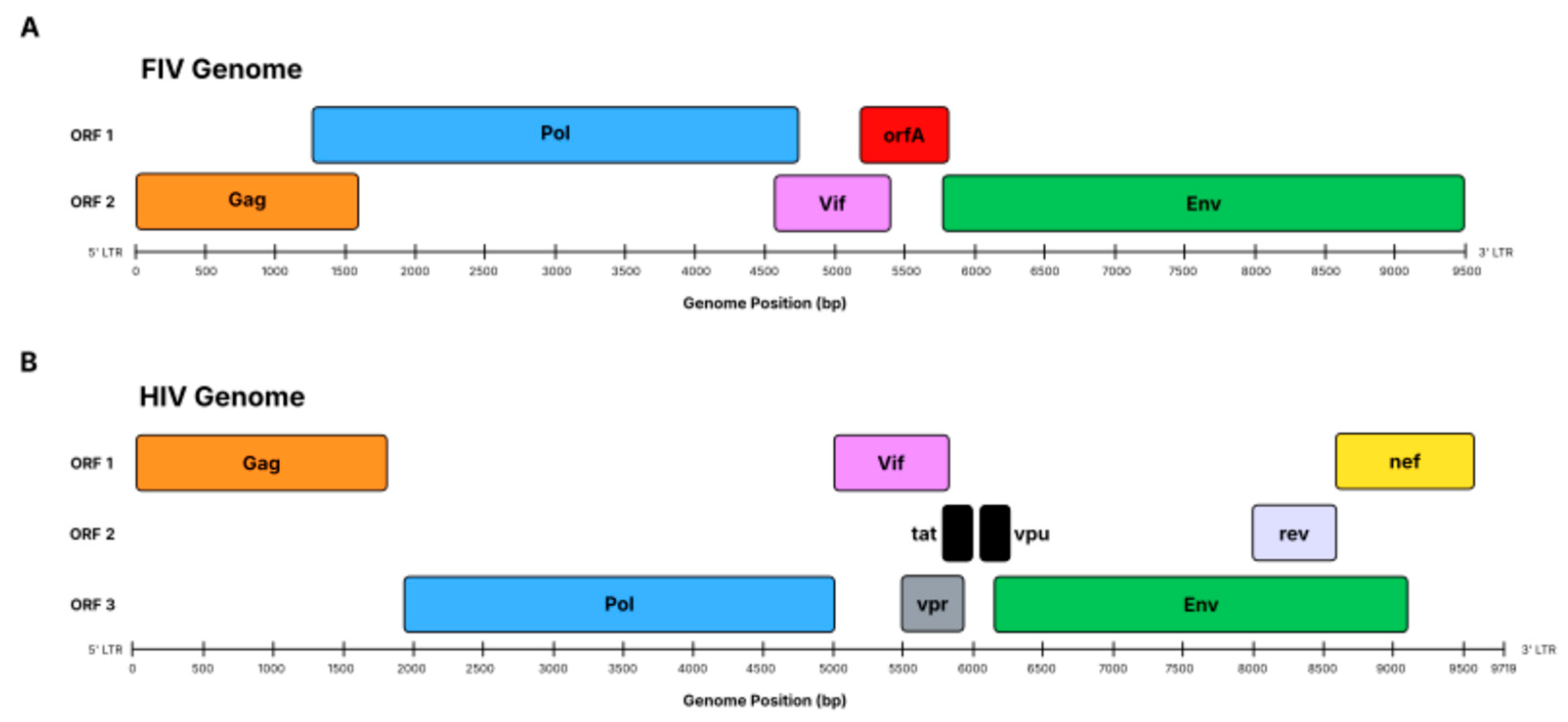
3.2. Mapping Some B-Cell Epitope Across the Key Proteins of the FIV Genomes
| Gene | Epitope (aa) | Antigenicity | Allergenicity | Toxicity | Surface exposed | Cross- Reactivity |
|---|---|---|---|---|---|---|
| Gag | 161IQTVNGAPQYVALDPK176 | 1.266 | Non- allergen |
Non-toxin | Yes | No |
| Gag | 332VKLYLKQSLSIANANP347 | 0.847 | Non- allergen |
Non-toxin | Yes | No |
| Pol | 744GEGILDKRAEDAGYDL759 | 1.15 | Non- allergen |
Non-toxin | Yes | No |
| Pol | 513GPHQICYQVYQKEGNP528 | 0.742 | Non- allergen |
Non-toxin | Yes | No |
| Env | 298KVNISLCLTGGKMLYN313 | 0.873 | Non- allergen |
Non-toxin | Yes | No |
| Env | 523KAVEMYNIAGNWSCTS538 | 0.813 | Non- allergen |
Non-toxin | Yes | No |
3.3. Mapping and Selection of Some Cytotoxic T Lymphocyte (CD8+) Epitopes Across the Key Proteins of the FIV Genomes
| Gene | Epitope (aa) | Restricting MHC allele | Antigenicity | Allergenicity | Toxicity | Cross- Reactivity |
|---|---|---|---|---|---|---|
| Gag | 55DLQERREKF63 | DLA-8803401 | 1.229 | Non- allergen |
Non-toxin | No |
| Gag | 138RMANVSTGR146 | DLA-8803401 | 1.108 | Non- allergen |
Non-toxin | No |
| Pol | 267SLAVHSLNF275 | DLA-8803401 | 1.053 | Non- allergen |
Non-toxin | No |
| Pol | 397RMLIDFREL405 | DLA-8803401 | 0.967 | Non- allergen |
Non-toxin | No |
| Env | 379RTQSQPGSW387 | DLA-8803401 | 1.365 | Non- allergen |
Non-toxin | No |
| Env | 512YTAFAMQEL520 | DLA-8803401 | 1.085 | Non- allergen |
Non-toxin | No |
3.4. Mapping and Selection of Some Helper T Lymphocyte (CD4+) Epitopes Across the Key Proteins of the FIV Genomes
| Gene | Epitope (aa) | Restricting MHC allele | Antigenicity | Allergenicity | Toxicity | IL-4 | IFN-γ | Cross- Reactivity |
|---|---|---|---|---|---|---|---|---|
| Gag | 134RWAIRMANVTTGREP148 | HLA-DRB1*01:01 | 1.082 | Non- allergen |
Non-toxin | Yes | No | No |
| Gag | 269LTQEQQAEPRFAPAR283 | HLA-DRB1*04:01 | 0.656 | Non- allergen |
Non-toxin | Yes | No | No |
| Gag | 217QLWFTAFSANLTPTD231 | HLA-DRB1*01:01 | 0.843 | Non- allergen |
Non-toxin | Yes | Yes | No |
| Pol | 312FTQNQQWIGPEEAEE326 | HLA-DRB1*07:01 | 0.613 | Non- allergen |
Non-toxin | Yes | No | No |
| Pol | 401DPDYAPYTAFTLPRK415 | HLA-DRB1*11:01 | 0.933 | Non- allergen |
Non-toxin | Yes | No | No |
| Env | 386GSWFRAISSWKQRNR400 | HLA-DRB1*15:01 | 0.459 | Non-allergen | Non-toxin | Yes | Yes | No |
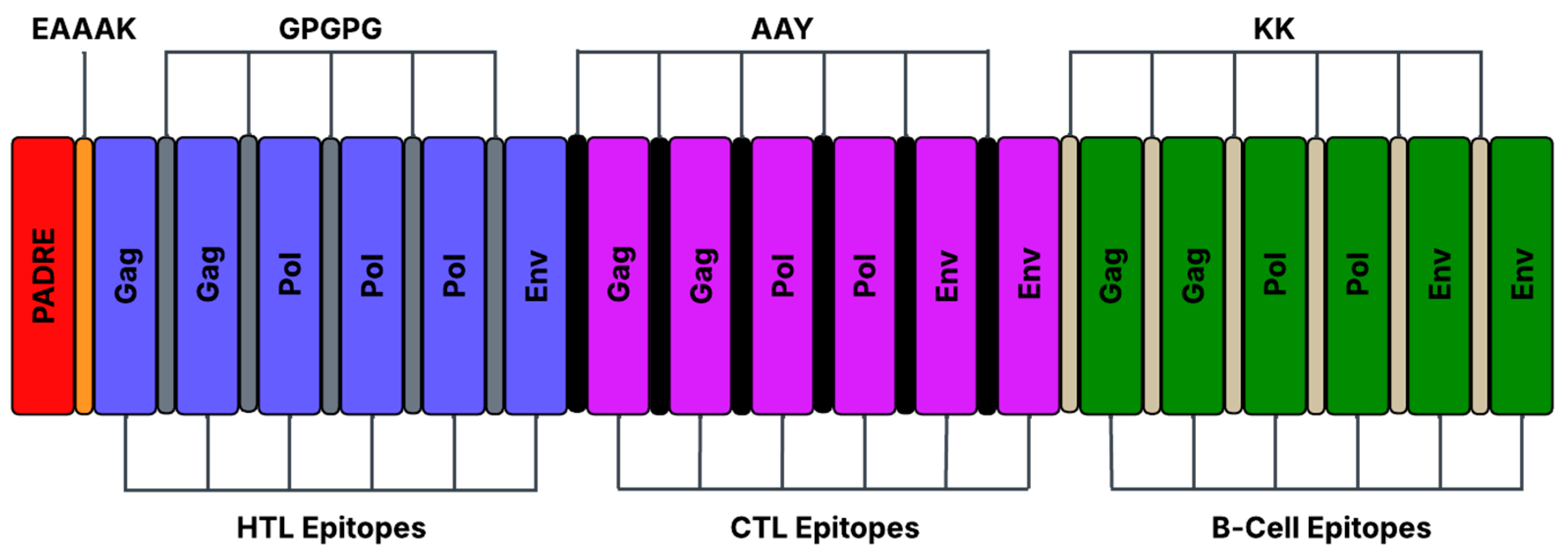
3.5. Selection of the Top Ranked Epitope and the Design of the FIV-DNA Vaccine Construct
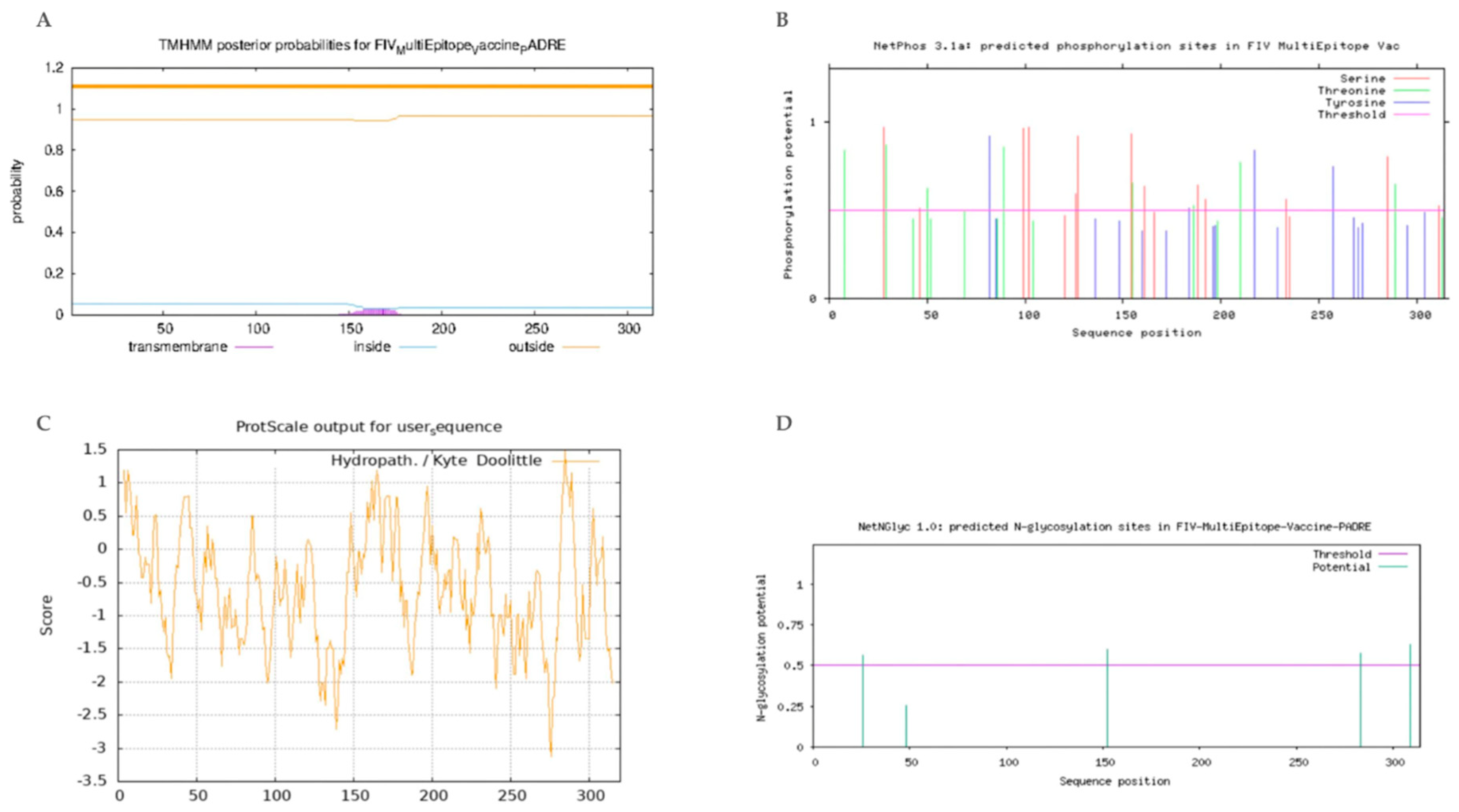
3.6. The Physicochemical Properties of the Designed FIV-DNA Vaccine Construct
3.7. Results of the Predicted Antigenicity and Allergenicity of the FIV-DNA Vaccine Construct
3.8. The Secondary Structure of the Multiepitope FIV-DNA Vaccine Construct
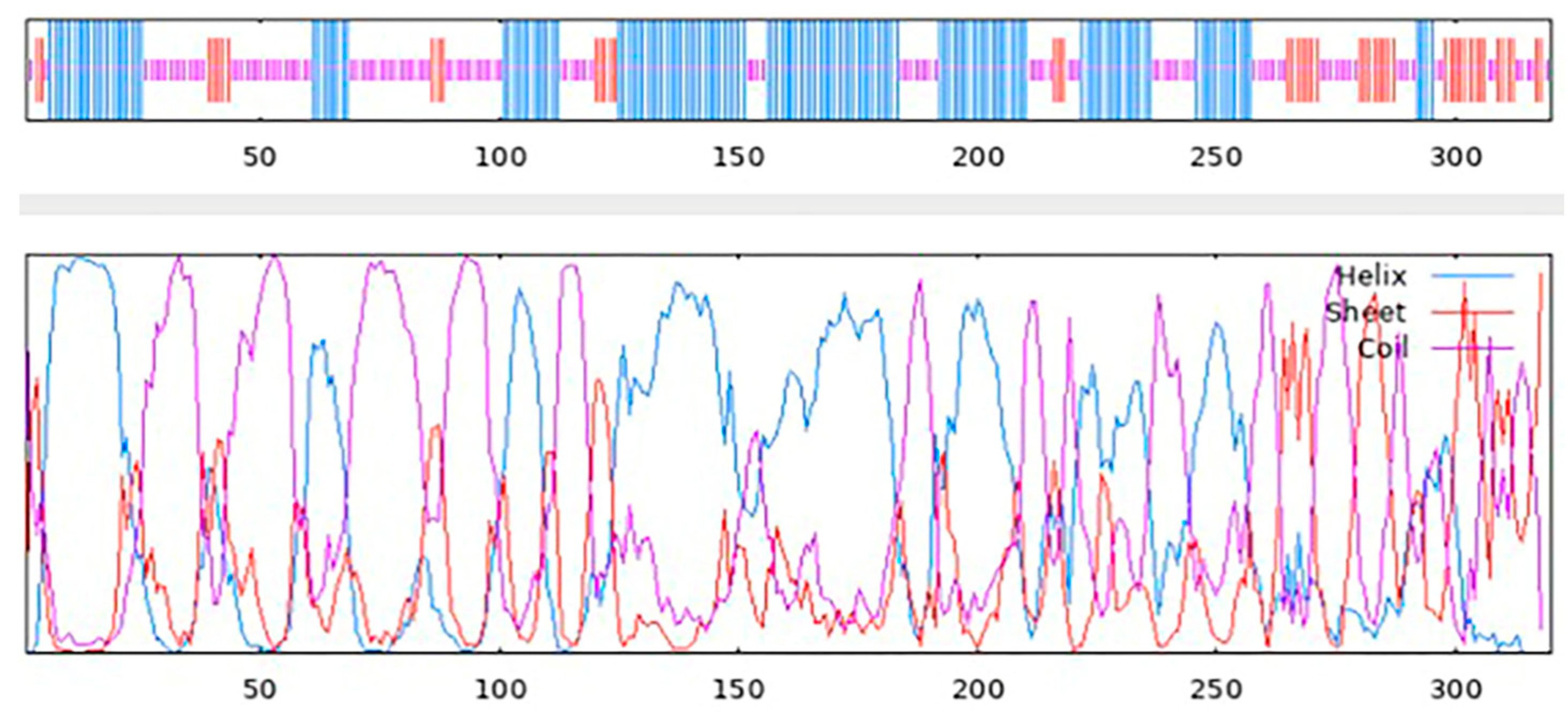
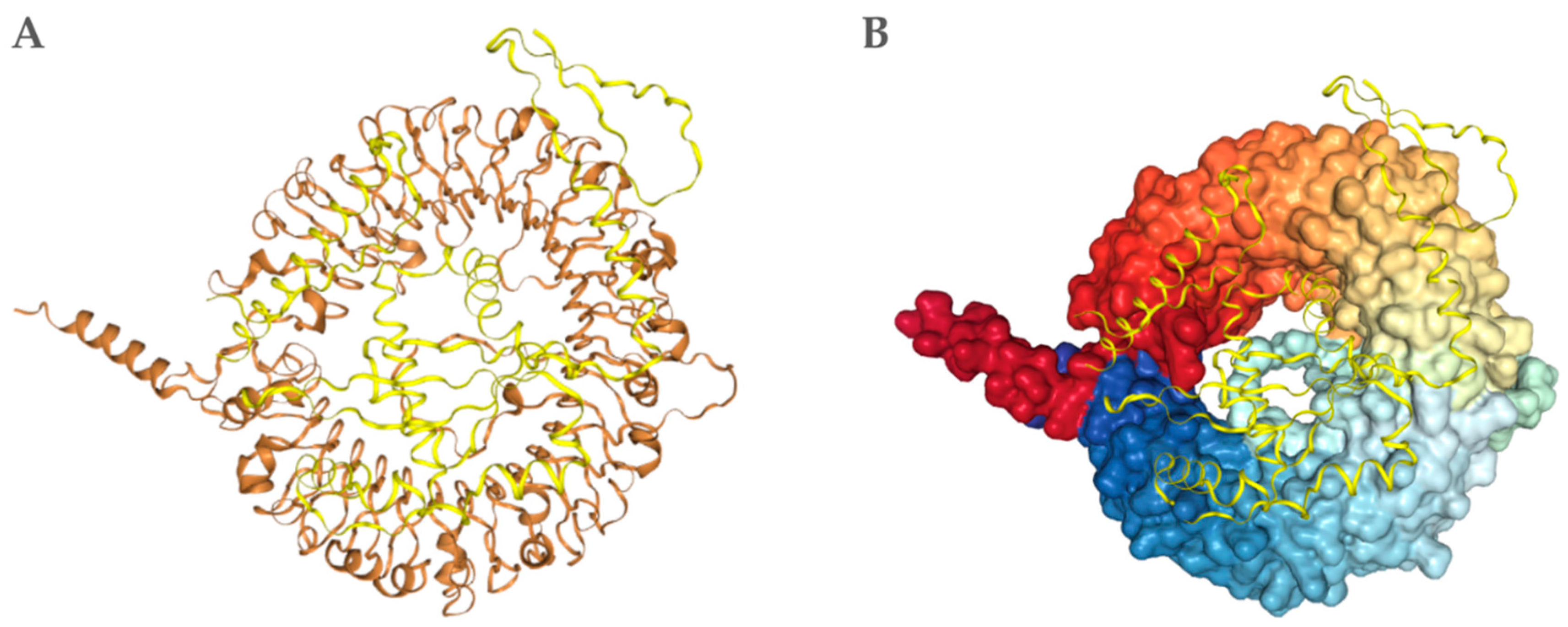
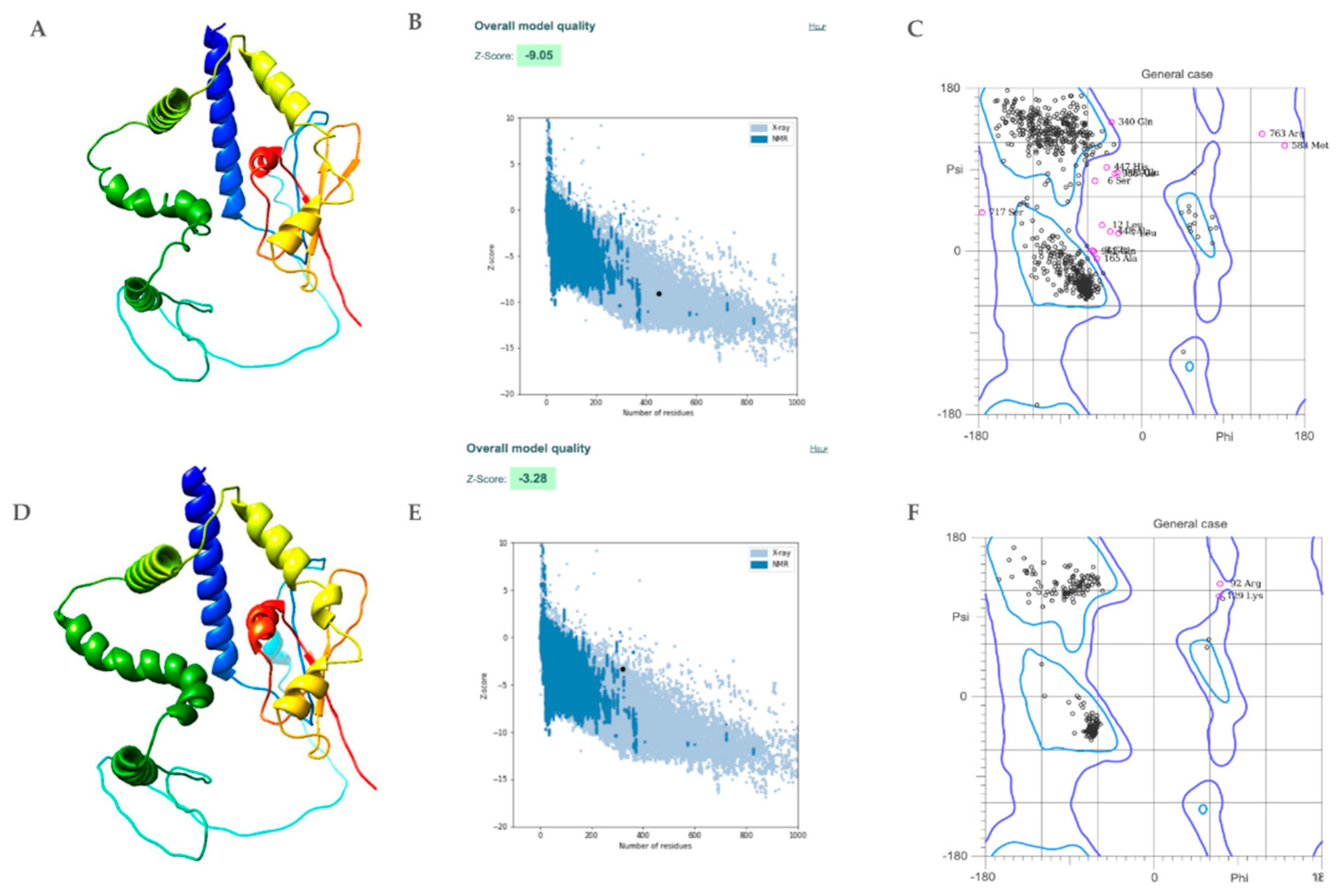
3.9. The Tertiary Structure of the Multiepitope FIV-DNA Vaccine Construct
3.10. Results of the Molecular Docking with Ligand-Binding Domain of the Feline TLR9
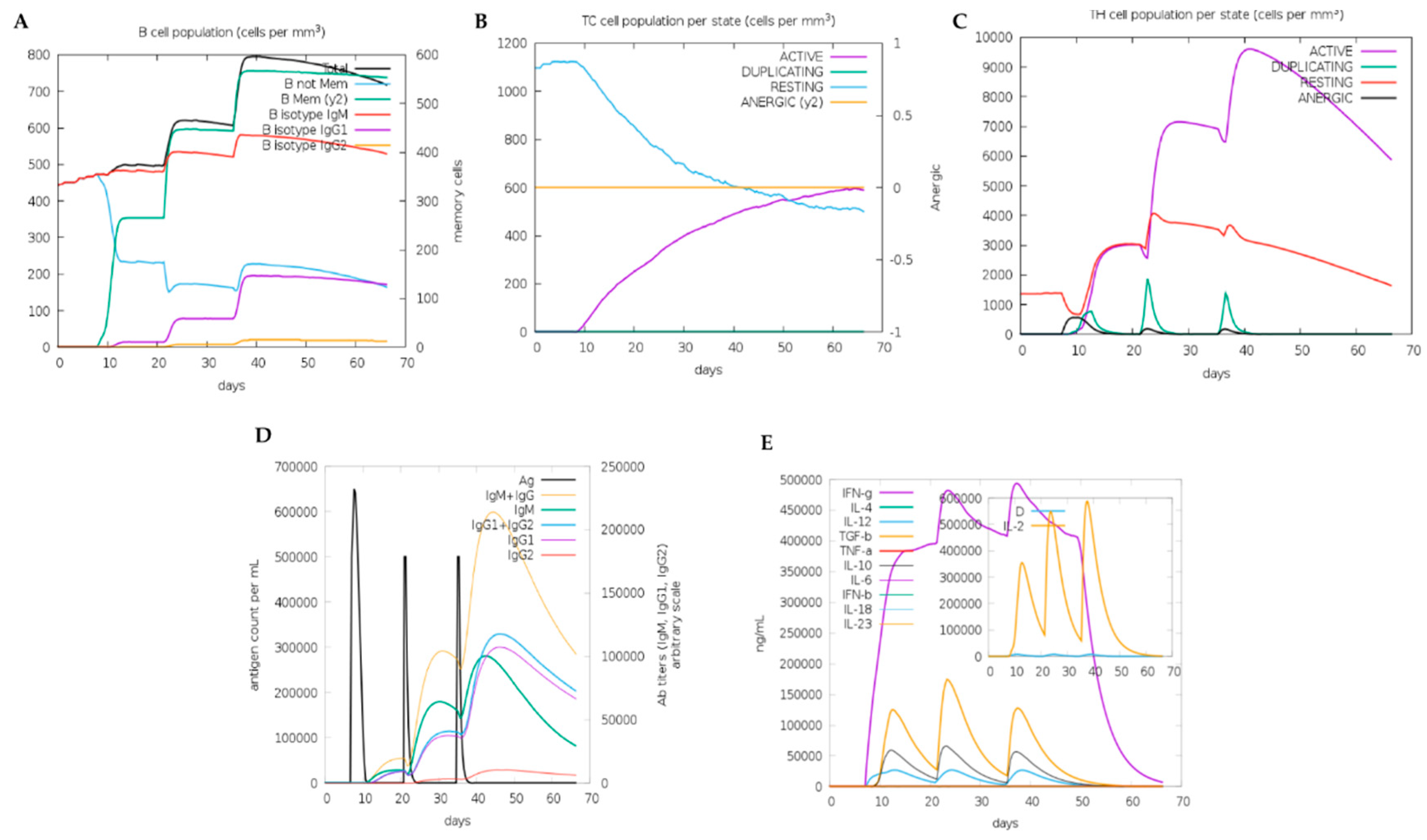
3.11. The Immune Stimulation Results of the Designed Multiepitope FIV-DNA Vaccine Construct
3.9. In Silico Cloning Results of the Final Recombinant Vaccine Construct
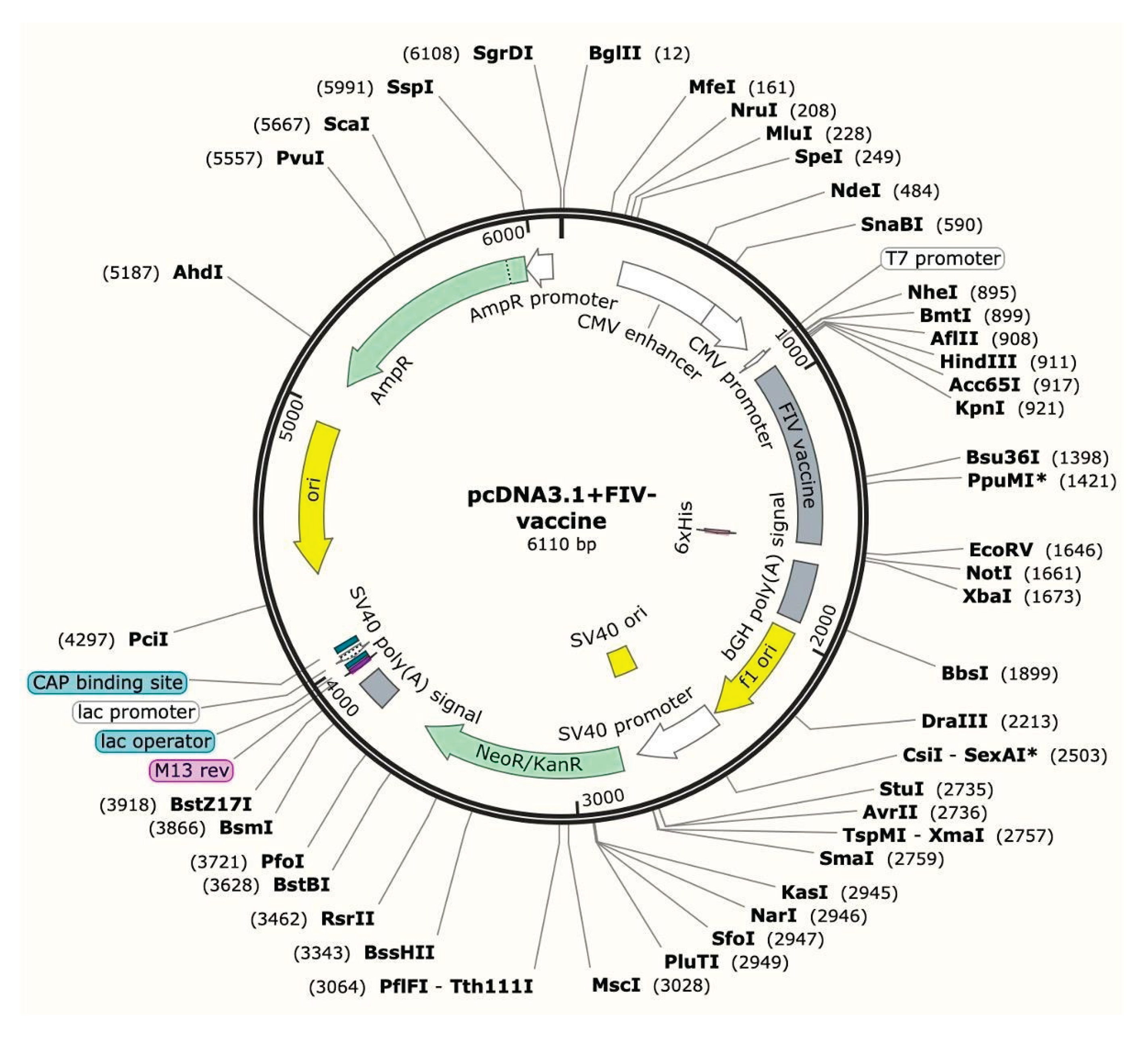
4. Discussion
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Pennings, P.S., HIV drug resistance: problems and perspectives. Infectious Disease Reports 2013, 5(11), e5.
- Hill, A.L.; Rosenbloom, D.I.S.; Nowak, M.A. Evolutionary dynamics of HIV at multiple spatial and temporal scales . Journal of Molecular Medicine 2012, 90(5), 543–561. [Google Scholar] [CrossRef] [PubMed]
- Pang, X. Emergence of HIV-1 drug resistance mutations among children and adolescents undergoing prolonged antiretroviral therapy in Guangxi . Journal of Global Antimicrobial Resistance 2024, 37, 208–213. [Google Scholar] [CrossRef] [PubMed]
- Barouch, D.H. Challenges in the development of an HIV-1 vaccine . Nature 2008, 455(7213), 613–619. [Google Scholar] [CrossRef] [PubMed]
- Cedeño-Muñoz, J.S. Antibiotic resistant bacteria and antibiotic resistance genes as contaminants of emerging concern: Occurrences, impacts, mitigations and future guidelines . Science of The Total Environment 2024, 952, 175906. [Google Scholar] [CrossRef]
- Hussaini, I.M. Unveiling the arsenal against antibiotic resistance: Antibacterial peptides as broad-spectrum weapons targeting multidrug-resistant bacteria . The Microbe 2024. 5, 100169. [Google Scholar] [CrossRef]
- Asquith, C.R.M. Synthesis and comparison of substituted 1,2,3-dithiazole and 1,2,3-thiaselenazole as inhibitors of the feline immunodeficiency virus (FIV) nucleocapsid protein as a model for HIV infection . Bioorganic & Medicinal Chemistry Letters 2019, 29(14), 1765–1768. [Google Scholar] [CrossRef]
- Laitinen, T. Synthesis and evaluation of 1,2,3-dithiazole inhibitors of the nucleocapsid protein of feline immunodeficiency virus (FIV) as a model for HIV infection . Bioorganic & Medicinal Chemistry 2022, 68, 116834. [Google Scholar] [CrossRef]
- Evermann, J.F.; Kennedy, M.A. Chapter 16 - Viral Infections, in Small Animal Pediatrics; Peterson, M.E., Kutzler, M.A., Eds.; W.B. Saunders: Saint Louis, 2011; pp. 119–129. [Google Scholar]
- Weaver, J.L. 11.32 - Immunomodulation and the Risk for Neoplasia, in Comprehensive Toxicology (Third Edition); McQueen, C.A., Ed.; Elsevier: Oxford, 2018; pp. 717–731. [Google Scholar]
- Miller, C. Applications of the FIV Model to Study HIV Pathogenesis . Viruses 2018, 10(4). [Google Scholar] [CrossRef]
- Burkhard, M.J.; Dean, G.A. Transmission and immunopathogenesis of FIV in cats as a model for HIV . Curr HIV Res 2003, 1(1), 15–29. [Google Scholar] [CrossRef]
- Mary Jo, B.; Gregg, A.D. Transmission and Immunopathogenesis of FIV in Cats as a Model for HIV . Current HIV Research 2003, 1(1), 15–29. [Google Scholar] [CrossRef]
- Bhattacharya, M. A novel mutation-proof, next-generation vaccine to fight against upcoming SARS-CoV-2 variants and subvariants, designed through AI enabled approaches and tools, along with the machine learning based immune simulation: A vaccine breakthrough . International Journal of Biological Macromolecules 2023, 242, 124893. [Google Scholar] [CrossRef]
- Vaghasiya, J.; Khan, M.; Bakhda, T. Milan. A meta-analysis of AI and machine learning in project management: Optimizing vaccine development for emerging viral threats in biotechnology . International Journal of Medical Informatics 2025, 195, 105768. [Google Scholar] [CrossRef] [PubMed]
- Sarkar, B. Chapter Five - The power of AI in viral vaccine production: A paradigm shift in efficiency and costs . In Methods in Microbiology; Srivastava, A., Mishra, V., Eds.; Academic Press, 2025; pp. 125–146. [Google Scholar]
- Jorge, S.; Dellagostin, O.A. The development of veterinary vaccines: a review of traditional methods and modern biotechnology approaches . Biotechnology Research and Innovation 2017, 1(1), 6–13. [Google Scholar] [CrossRef]
- Khalaf, W.S.; Morgan, R.N.; Elkhatib, W.F. Clinical microbiology and artificial intelligence: Different applications, challenges, and future prospects . Journal of Microbiological Methods 2025. 232-234, 107125. [Google Scholar] [CrossRef] [PubMed]
- Olawade, D.B. Leveraging artificial intelligence in vaccine development: A narrative review . Journal of Microbiological Methods 2024, 224, 106998. [Google Scholar] [CrossRef]
- Kannan, S.; Subbaram, K.; Faiyazuddin, M. Chapter 17 - Artificial intelligence in vaccine development: Significance and challenges ahead, in A Handbook of Artificial Intelligence in Drug Delivery; Philip, A., et al., Eds.; Academic Press, 2023; pp. 467–486. [Google Scholar]
- Malhotra, Y. Chapter Eight - AI in infectious disease diagnosis and vaccine development, in Methods in Microbiology; Srivastava, A., Mishra, V., Eds.; Academic Press, 2025; pp. 191–218. [Google Scholar]
- Al-Kubati, A.A.G. Immunoinformatic prediction of the pathogenicity of bovine viral diarrhea virus genotypes: implications for viral virulence determinants, designing novel diagnostic assays and vaccines development . Front Vet Sci 2023, 10, 1130147. [Google Scholar] [CrossRef] [PubMed]
- Duraisamy, N. Machine learning tools used for mapping some immunogenic epitopes within the major structural proteins of the bovine coronavirus (BCoV) and for the in silico design of the multiepitope-based vaccines . Front Vet Sci 2024, 11, 1468890. [Google Scholar] [CrossRef]
- Goud, M.D. Artificial Intelligence Driven Framework for the Design and Development of Next-Generation Avian Viral Vaccines . Microorganisms 2025, 13(10). [Google Scholar] [CrossRef]
- Hemida, M.G. The next-generation coronavirus diagnostic techniques with particular emphasis on the SARS-CoV-2 . J Med Virol 2021, 93(7), 4219–4241. [Google Scholar] [CrossRef]
- Khan, M.Y. Repurposing of Some Nucleoside Analogs Targeting Some Key Proteins of the Avian H5N1 Clade 2.3.4.4b to Combat the Circulating HPAI in Birds: An In Silico Approach . Viruses 2025, 17(7). [Google Scholar] [CrossRef]
- Khan, M.Y. Leveraging Artificial Intelligence and Gene Expression Analysis to Identify Some Potential Bovine Coronavirus (BCoV) Receptors and Host Cell Enzymes Potentially Involved in the Viral Replication and Tissue Tropism . Int J Mol Sci 2025, 26(3). [Google Scholar] [CrossRef]
- Malik, Y.S. Advances in Diagnostic Approaches for Viral Etiologies of Diarrhea: From the Lab to the Field . Front Microbiol 2019, 10, 1957. [Google Scholar] [CrossRef]
- Al-Mubarak, A.I.A. Detection of Avian Orthoavulavirus-1 genotypes VI.2.1 and VII.1.1 with neuro-viscerotropic tropism in some backyard pigeons (Columbidae) in Eastern Saudi Arabia . Front Vet Sci 2024, 11, 1352636. [Google Scholar] [CrossRef] [PubMed]
- Alturaiki, W. The pivotal roles of the host immune response in the fine-tuning the infection and the development of the vaccines for SARS-CoV-2 . Hum Vaccin Immunother 2021, 17(10), 3297–3309. [Google Scholar] [CrossRef]
- Hemida, M.G.; Al-Ghadeer, H.M. Evidence of Peste des petits Ruminants' Virus in Dromedary Camels in the Kingdom of Saudi Arabia between 2014 and 2016 . In Vet Med Int; 2019; p. 4756404. [Google Scholar]
- Hemida, M.G. Molecular characterization and phylogenetic analyses of virulent infectious bronchitis viruses isolated from chickens in Eastern Saudi Arabia . Virusdisease 2017, 28(2), 189–199. [Google Scholar] [CrossRef]
- Hemida, M.G. Prevalence and molecular characterization of some circulating strains of the peste-des-petits-ruminants virus in Saudi Arabia between 2014-2016 . PeerJ 2020. 8, p. e9035. [Google Scholar] [CrossRef]
- Hemida, M.G. Longitudinal study of Middle East Respiratory Syndrome coronavirus infection in dromedary camel herds in Saudi Arabia, 2014-2015 . Emerg Microbes Infect 2017, 6(6), p. e56. [Google Scholar] [CrossRef] [PubMed]
- Hemida, M.G. Molecular characterisation of an avian influenza (H5N8) outbreak in backyard flocks in Al Ahsa, Eastern Saudi Arabia, 2017-2018 . Vet Rec Open 2019, 6(1), p. e000362. [Google Scholar] [CrossRef] [PubMed]
- Hemida, M.G. Evidence of equine influenza A (H3N8) activity in horses from Eastern and Central Saudi Arabia: 2013-2015 . Equine Vet J 2019, 51(2), 218–221. [Google Scholar] [CrossRef]
- Hemida, M.G. Foot-and-mouth disease virus O/ME-SA/Ind 2001 lineage outbreak in vaccinated Holstein Friesian cattle in Saudi Arabia in 2016 . Vet Q 2018, 38(1), 88–98. [Google Scholar] [CrossRef] [PubMed]
- Saha, S.; Raghava, G.P. Prediction of continuous B-cell epitopes in an antigen using recurrent neural network . Proteins 2006, 65(1), 40–8. [Google Scholar] [CrossRef]
- Shah, A.U.; Gauger, P.; Hemida, M.G. Isolation and molecular characterization of an enteric isolate of the genotype-Ia bovine coronavirus with notable mutations in the receptor binding domain of the spike glycoprotein . Virology 2025. 603, 110313. [Google Scholar] [CrossRef]
- Shah, A.U. Some novel field isolates belonging to lineage-1 of the genotype GI-avian infectious bronchitis virus (AIBV) show strong evidence of recombination with field/vaccinal strains . Infect Genet Evol 2025. 129, 105723. [Google Scholar] [CrossRef]
- Singh, K.; Malik, Y.S.; Hemida, M.G. Comprehensive Evolutionary and Structural Analysis of the H5N1 Clade 2.4.3.4b Influenza a Virus Based on the Sequences and Data Mining of the Hemagglutinin, Nucleoprotein and Neuraminidase Genes Across Multiple Hosts . Pathogens 2025. 14, 9. [Google Scholar] [CrossRef]
- Liu, H. Design and development of a novel multi-epitope DNA vaccine candidate against infectious bronchitis virus: an immunoinformatic approach . Archives of Microbiology 2025, 207(4), p. 84. [Google Scholar] [CrossRef]
- Sher, H. Employing computational tools to design a multi-epitope vaccine targeting human immunodeficiency virus-1 (HIV-1) . BMC Genomics 2023, 24(1), 276. [Google Scholar] [CrossRef] [PubMed]
- Fleming, J. AlphaFold Protein Structure Database and 3D-Beacons: New Data and Capabilities . Journal of Molecular Biology 2025, 437(15), p. 168967. [Google Scholar] [CrossRef]
- Jumper, J. Highly accurate protein structure prediction with AlphaFold . Nature 2021, 596(7873), 583–589. [Google Scholar] [CrossRef]
- Mirdita, M. ColabFold: making protein folding accessible to all . Nature Methods 2022, 19(6), 679–682. [Google Scholar] [CrossRef]
- Vita, R. The Immune Epitope Database (IEDB): 2018 update . Nucleic Acids Res 2019, 47(D1), D339–d343. [Google Scholar] [CrossRef]
- Wiederstein, M.; Sippl, M.J. ProSA-web: interactive web service for the recognition of errors in three-dimensional structures of proteins . Nucleic Acids Res 2007, 35(Web Server issue), W407–10. [Google Scholar] [CrossRef]
- Sippl, M.J. Recognition of errors in three-dimensional structures of proteins . Proteins 1993, 17(4), 355–62. [Google Scholar] [CrossRef]
- Chen, V.B. MolProbity: all-atom structure validation for macromolecular crystallography . Acta Crystallogr D Biol Crystallogr 2010, 66 Pt 1, 12–21. [Google Scholar] [CrossRef]
- Doytchinova, I.A.; Flower, D.R. VaxiJen: a server for prediction of protective antigens, tumour antigens and subunit vaccines . BMC Bioinformatics 2007, 8, 4. [Google Scholar] [CrossRef]
- Heo, L.; Park, H.; Seok, C. GalaxyRefine: Protein structure refinement driven by side-chain repacking . Nucleic Acids Res 2013, 41(Web Server issue), W384–8. [Google Scholar] [CrossRef]
- Lee, G.R.; Heo, L.; Seok, C. Effective protein model structure refinement by loop modeling and overall relaxation . Proteins 2016, 84 Suppl 1, 293–301. [Google Scholar] [CrossRef] [PubMed]
- Dimitrov, I. AllerTOP v.2--a server for in silico prediction of allergens . J Mol Model 2014, 20(6), p. 2278. [Google Scholar] [CrossRef]
- Yan, Y. The HDOCK server for integrated protein–protein docking . Nature Protocols 2020, 15(5), 1829–1852. [Google Scholar] [CrossRef] [PubMed]
- Gupta, S. In silico approach for predicting toxicity of peptides and proteins . PLoS One 2013, 8(9), e73957. [Google Scholar] [CrossRef] [PubMed]
- Krissinel, E.; Henrick, K. Inference of Macromolecular Assemblies from Crystalline State . Journal of Molecular Biology 2007, 372(3), 774–797. [Google Scholar] [CrossRef]
- Hallgren, J. DeepTMHMM predicts alpha and beta transmembrane proteins using deep neural networks. 2022.04.08.487609. bioRxiv 2022. [Google Scholar]
- Rapin, N. Computational Immunology Meets Bioinformatics: The Use of Prediction Tools for Molecular Binding in the Simulation of the Immune System . PLOS ONE 2010, 5(4), e9862. [Google Scholar] [CrossRef]
- Blom, N. Prediction of post-translational glycosylation and phosphorylation of proteins from the amino acid sequence . Proteomics 2004, 4(6), 1633–49. [Google Scholar] [CrossRef] [PubMed]
- Blom, N.; Gammeltoft, S.; Brunak, S. Sequence and structure-based prediction of eukaryotic protein phosphorylation sites . J Mol Biol 1999, 294(5), 1351–62. [Google Scholar] [CrossRef] [PubMed]
- Wilkins, M.R. Protein identification and analysis tools in the ExPASy server . Methods Mol Biol 1999, 112, 531–52. [Google Scholar]
- Gupta, R.; Brunak, S. Prediction of glycosylation across the human proteome and the correlation to protein function . In Pac Symp Biocomput; 2002; pp. 310–22. [Google Scholar]
- Dhanda, S.K. Prediction of IL4 inducing peptides . Clin Dev Immunol 2013, 263952. [Google Scholar] [CrossRef] [PubMed]
- Dhanda, S.K.; Vir, P.; Raghava, G.P. Designing of interferon-gamma inducing MHC class-II binders . Biol Direct 2013, 8, 30. [Google Scholar] [CrossRef]
- Kenyon, J.C.; Lever, A.M. The molecular biology of feline immunodeficiency virus (FIV) . Viruses 2011, 3(11), 2192–213. [Google Scholar] [CrossRef]
- González, S.A.; Affranchino, J.L. Properties and Functions of Feline Immunodeficiency Virus Gag Domains in Virion Assembly and Budding . Viruses 2018, 10. [Google Scholar] [CrossRef]
- Cervera, L. Production of HIV-1-based virus-like particles for vaccination: achievements and limits . Appl Microbiol Biotechnol 2019, 103(18), 7367–7384. [Google Scholar] [CrossRef]
- Chawla, M. Immunoinformatics-aided rational design of a multi-epitope vaccine targeting feline infectious peritonitis virus . Front Vet Sci 2023, 10, 1280273. [Google Scholar] [CrossRef]
- Romagnani, S. Th1/Th2 cells . Inflamm Bowel Dis 1999, 5(4), 285–94. [Google Scholar] [CrossRef] [PubMed]
- Alexander, J. Development of high potency universal DR-restricted helper epitopes by modification of high affinity DR-blocking peptides . Immunity 1994, 1(9), 751–61. [Google Scholar] [CrossRef] [PubMed]
- Krogh, A. Predicting transmembrane protein topology with a hidden Markov model: application to complete genomes . J Mol Biol 2001, 305(3), 567–80. [Google Scholar] [CrossRef]
- Zhao, T. Vaccine adjuvants: mechanisms and platforms . Signal Transduction and Targeted Therapy 2023, 8(1), 283. [Google Scholar] [CrossRef] [PubMed]
- He, M.; Zhou, X.; Wang, X. Glycosylation: mechanisms, biological functions and clinical implications . Signal Transduction and Targeted Therapy 2024, 9(1), p. 194. [Google Scholar] [CrossRef]
- Dunker, A.K. Intrinsically disordered protein . J Mol Graph Model 2001, 19(1), 26–59. [Google Scholar] [CrossRef]
- Barouch, D.H. Challenges in the development of an HIV-1 vaccine . Nature 2008, 455(7213), 613–9. [Google Scholar] [CrossRef]
- Aranyos, A.M. An initial examination of the potential role of T-cell immunity in protection against feline immunodeficiency virus (FIV) infection . Vaccine 2016, 34(12), 1480–8. [Google Scholar] [CrossRef]
- Collins, D.R.; Gaiha, G.D.; Walker, B.D. CD8(+) T cells in HIV control, cure and prevention . Nat Rev Immunol 2020, 20(8), 471–482. [Google Scholar] [CrossRef]
- Murphy, B.G. The Late Asymptomatic and Terminal Immunodeficiency Phases in Experimentally FIV-Infected Cats-A Long-Term Study . Viruses 2023, 15(8). [Google Scholar] [CrossRef]
- Plotkin, S.A. Correlates of protection induced by vaccination . Clin Vaccine Immunol 2010, 17(7), 1055–65. [Google Scholar] [CrossRef]
- del Guercio, M.F. Potent immunogenic short linear peptide constructs composed of B cell epitopes and Pan DR T helper epitopes (PADRE) for antibody responses in vivo . Vaccine 1997, 15(4), 441–8. [Google Scholar] [CrossRef]
- Alexander, J. The optimization of helper T lymphocyte (HTL) function in vaccine development . Immunol Res 1998, 18(2), 79–92. [Google Scholar] [CrossRef]
- Rani, N.A. Development of multi epitope subunit vaccines against emerging carp viruses Cyprinid herpesvirus 1 and 3 using immunoinformatics approach . Scientific Reports 2024, 14(1), p. 11783. [Google Scholar] [CrossRef]
- Rizarullah. Designing a Novel Multiepitope Vaccine from the Human Papilloma Virus E1 and E2 Proteins for Indonesia with Immunoinformatics and Molecular Dynamics Approaches . ACS Omega 2024, 9(14), 16547–16562. [Google Scholar] [CrossRef]
- Rani, N.A. Development of multi epitope subunit vaccines against emerging carp viruses Cyprinid herpesvirus 1 and 3 using immunoinformatics approach . Sci Rep 2024, 14(1), p. 11783. [Google Scholar] [CrossRef] [PubMed]
- Uhl, E.W. FIV vaccine development and its importance to veterinary and human medicine: a review FIV vaccine 2002 update and review . Vet Immunol Immunopathol 2002, 90(3-4), 113–32. [Google Scholar] [CrossRef] [PubMed]
- Robert-Tissot, C. Stimulation with a class A CpG oligonucleotide enhances resistance to infection with feline viruses from five different families . Vet Res 2012, 43(1), p. 60. [Google Scholar] [CrossRef] [PubMed]
- Wheat, W. Activation of upper respiratory tract mucosal innate immune responses in cats by liposomal toll-like receptor ligand complexes delivered topically . J Vet Intern Med 2019, 33(2), 838–845. [Google Scholar] [CrossRef] [PubMed]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




