Submitted:
14 February 2026
Posted:
17 February 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
2. Methods
2.1. Dataset
2.2. Signal Preprocessing
2.3. Feature Extraction
2.4. Classification Models
2.5. Peak Detection Algorithm Based on PTC
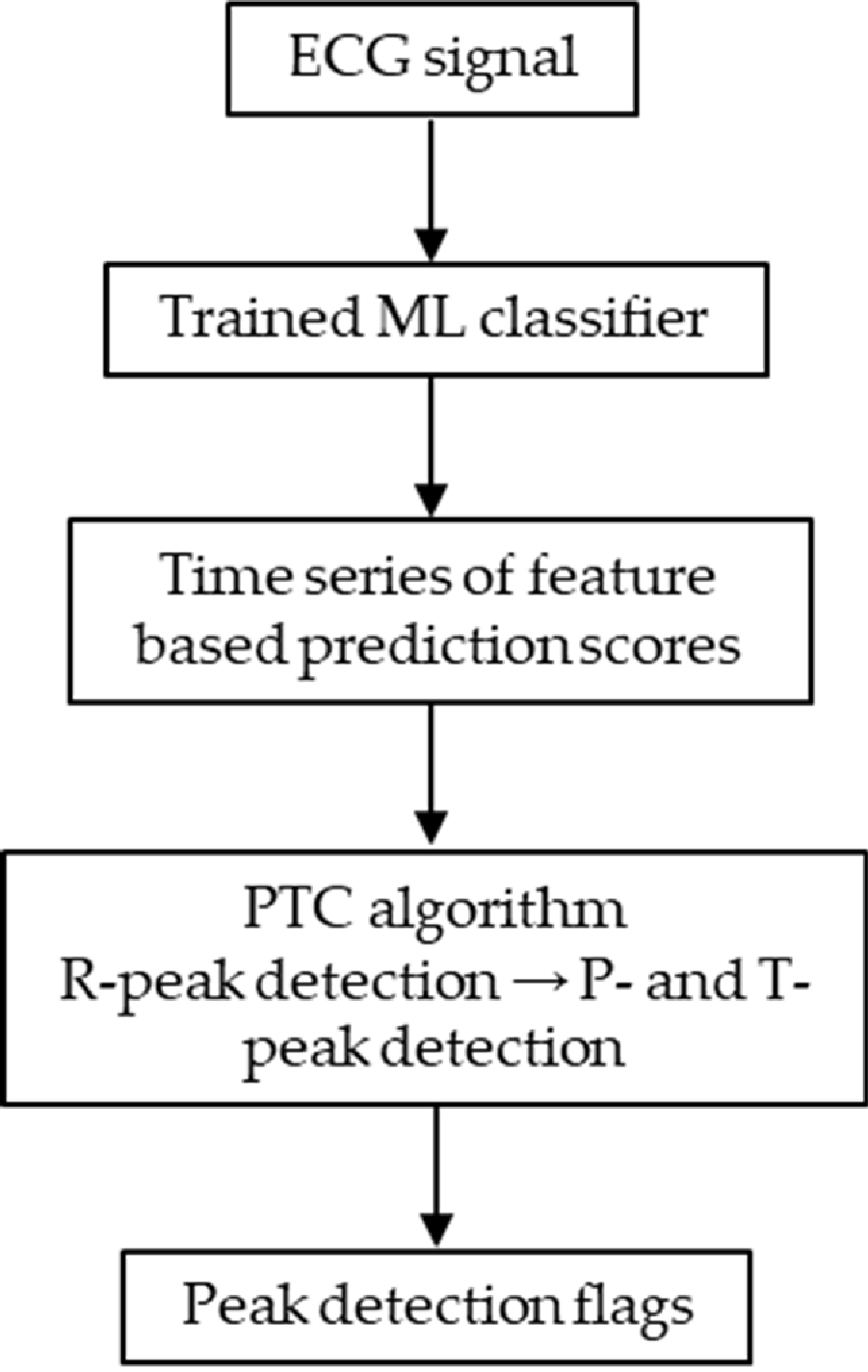
2.5.1. Algorithm Overview
Generation and Score-Based Ordering of R-Peak Candidates
R-peak Candidate Selection Using a Refractory Period
Visualization of the Sequential Selection Process Using a Table
R-Peak–Landmarked P- and T-Peak Detection
2.5.2. R-Peak Detection and Parameter Tuning
2.5.3. P-Peak Detection Within R-Centered Windows
2.5.4. T-Peak Detection
2.6. Evaluation Protocol
2.6.1. Baseline Classification Performance (Prior to Applying PTC)
2.6.2. Final Peak Detection Performance Evaluation (with PTC)
2.6.3. Handling of True Negatives (TNs) in Evaluation
2.7. Summary of Parameter Statistics
2.8. Application to Arrhythmic Data in the LUDB
2.9. Standardization for Implementation and Application to the PTB-XL ECG Dataset
2.9.1. Algorithm Design with Implementation in Mind
2.9.2. Standardization of Sampling Frequency and Preprocessing
2.9.3. Application to the PTB-XL ECG Dataset and Data Characteristics
2.10. Implementation Details
3. Results
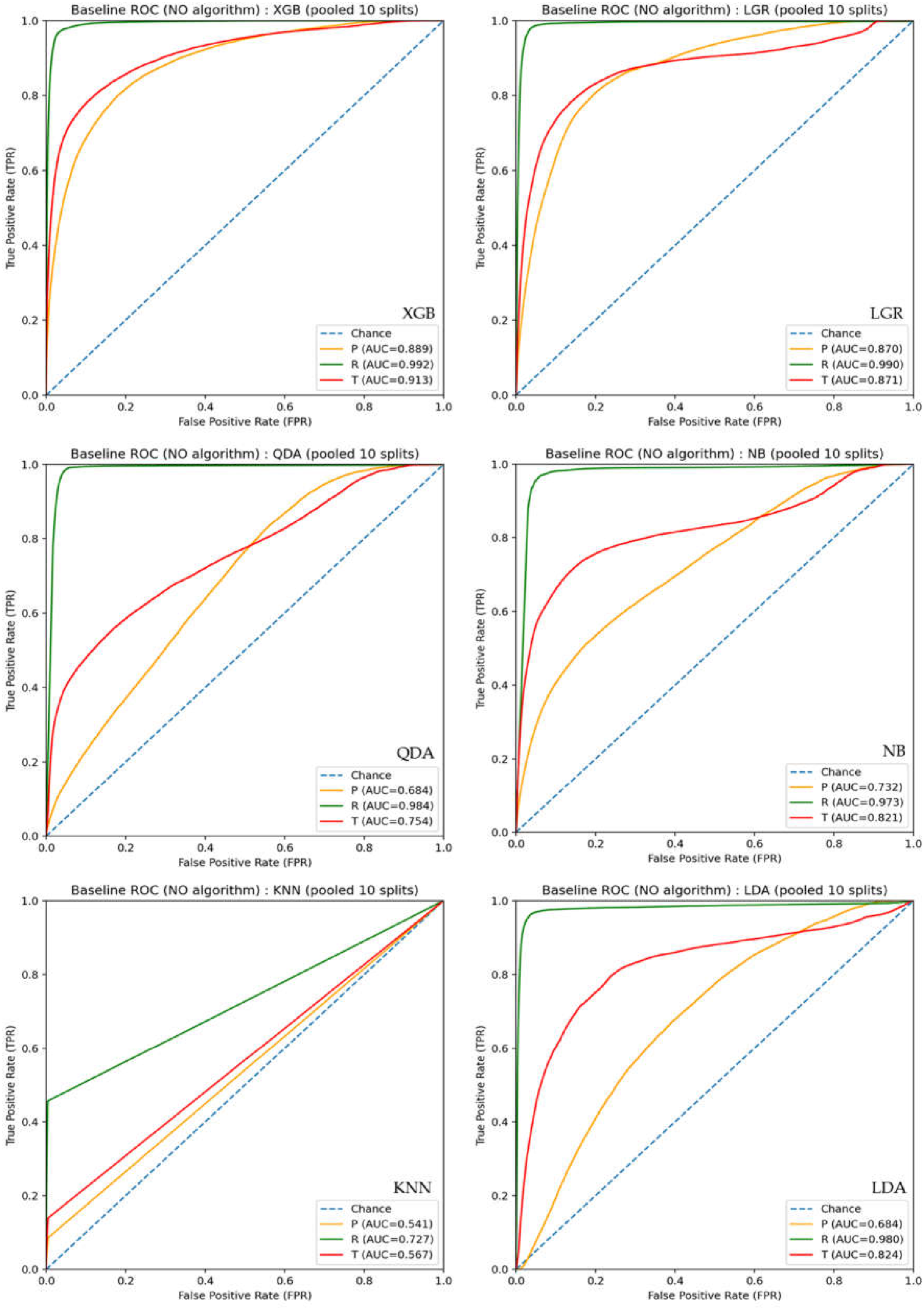
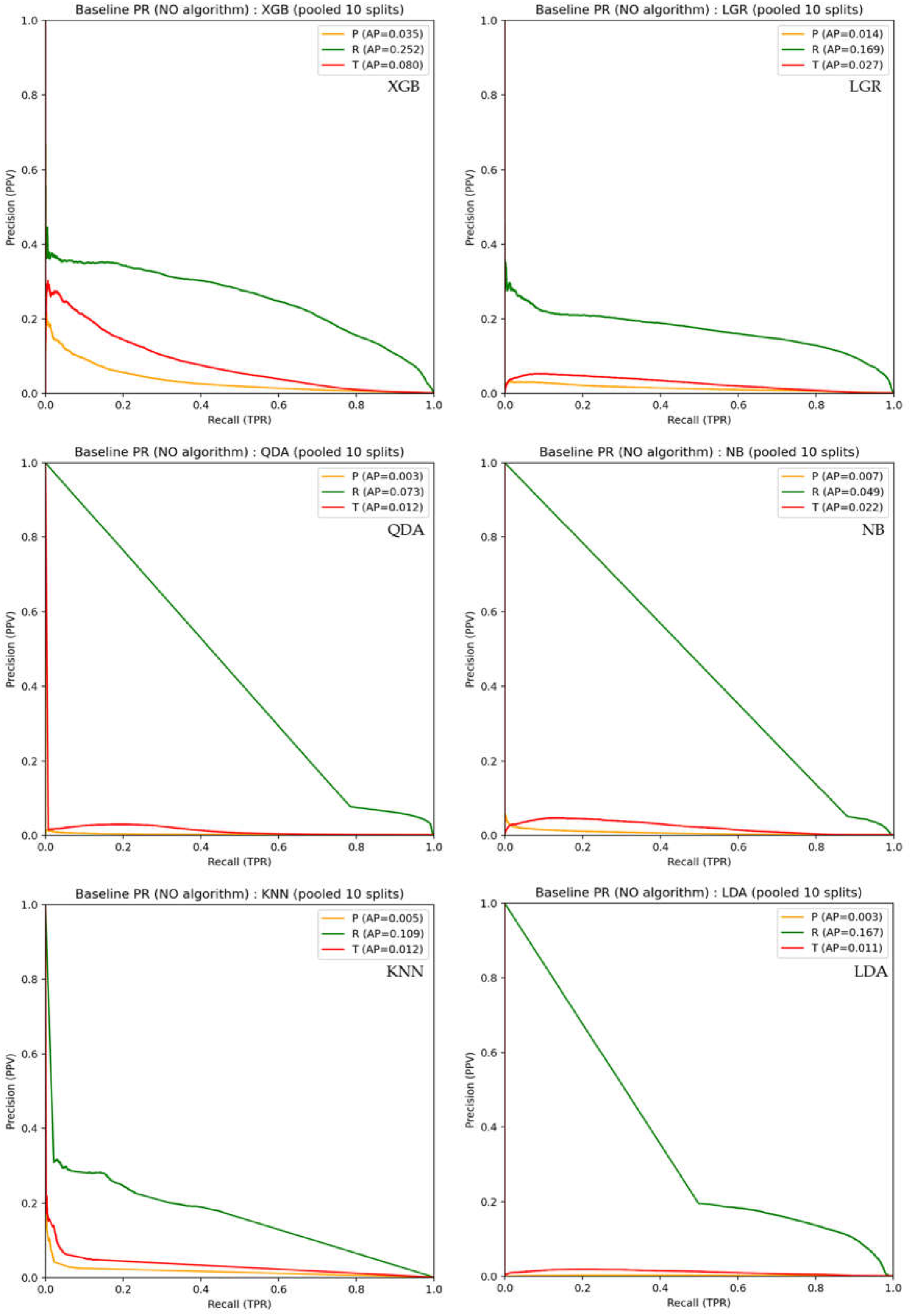
3.1. Baseline Classification Performance Prior to PTC
3.2. Peak Detection Performance with PTC
3.3. Effect of Temporal Tolerance on Peak Detection Accuracy
3.4. Stability of Optimized Algorithmic Parameters
3.5. Stability of Classifier Parameters
3.6. Performance on Arrhythmic Data in the LUDB
3.7. Robustness to Preprocessing Conditions and Practical Implementation Results
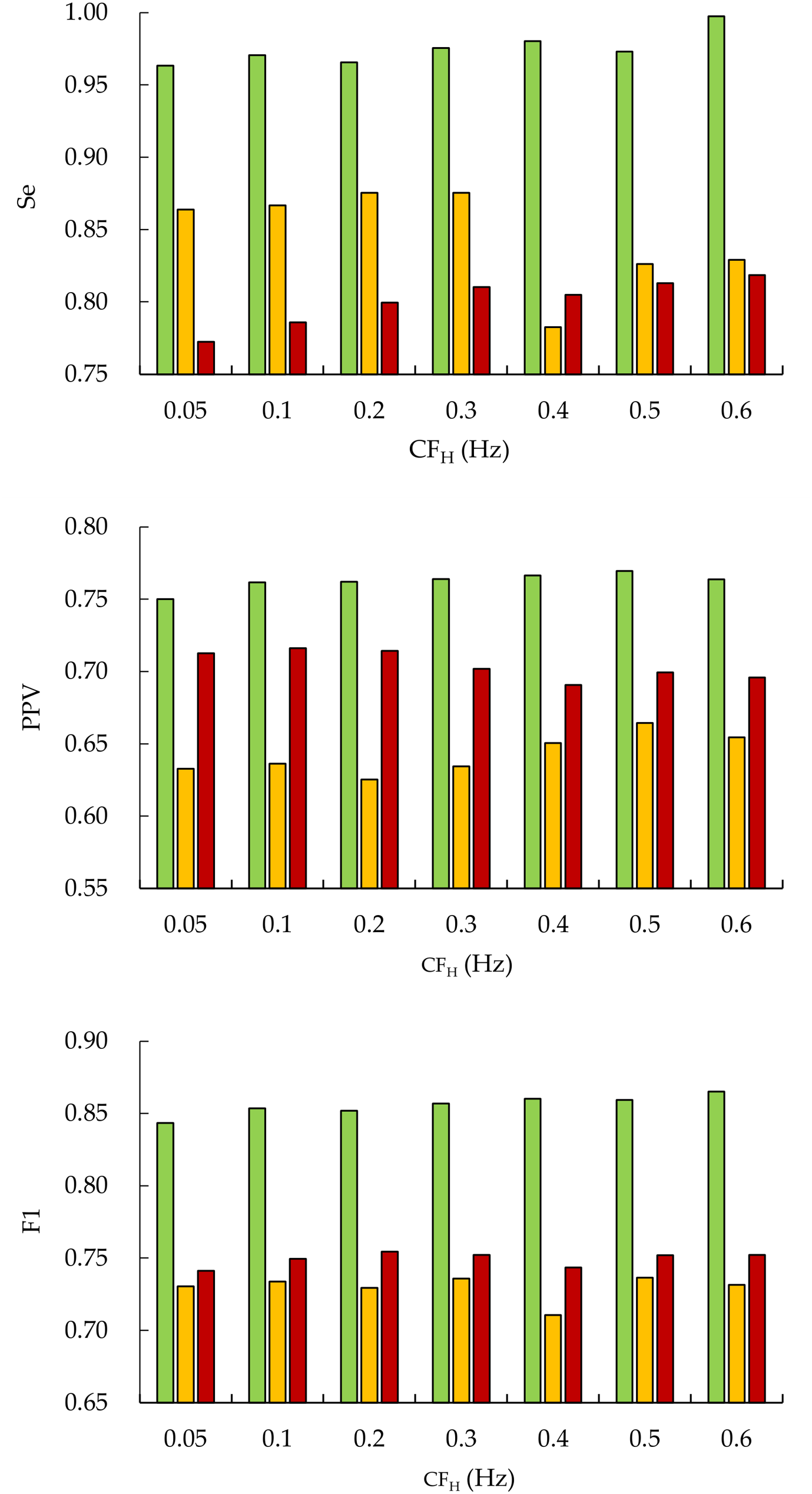
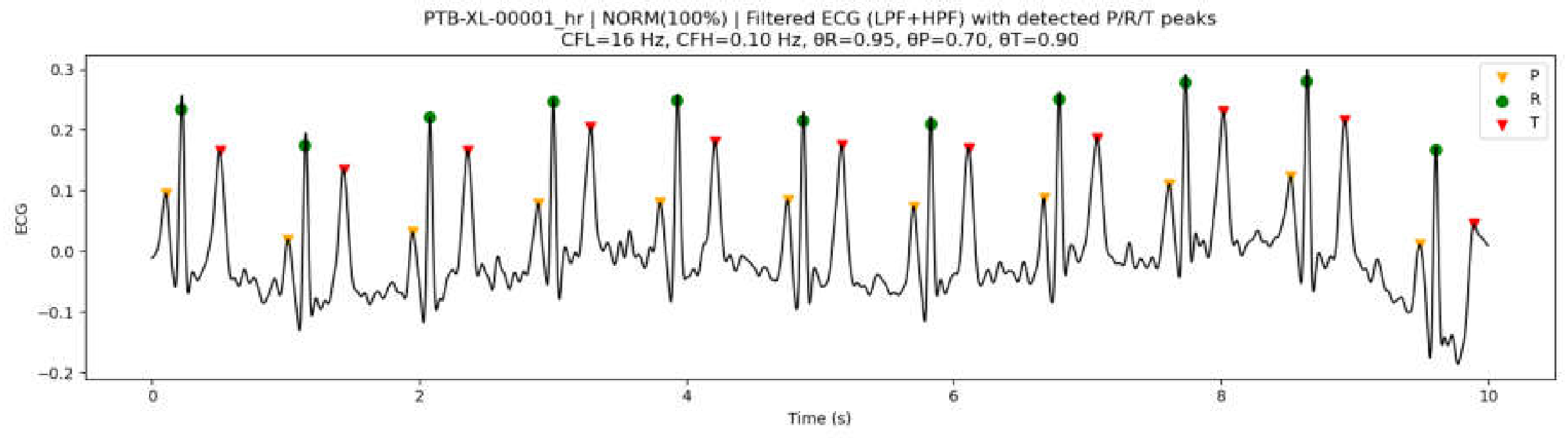
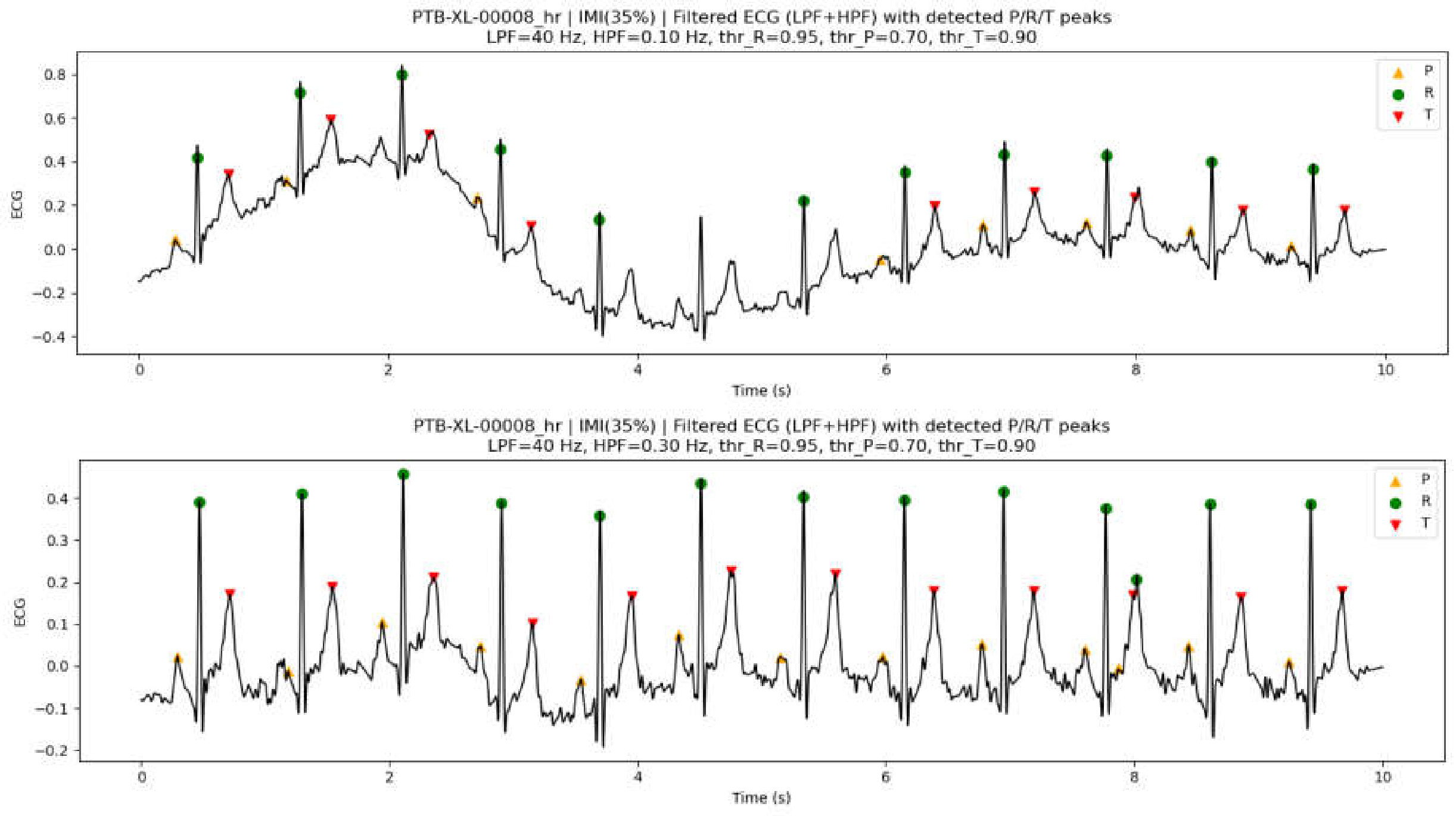
3.7.1. Effect of Preprocessing Parameters on Peak Detection Performance
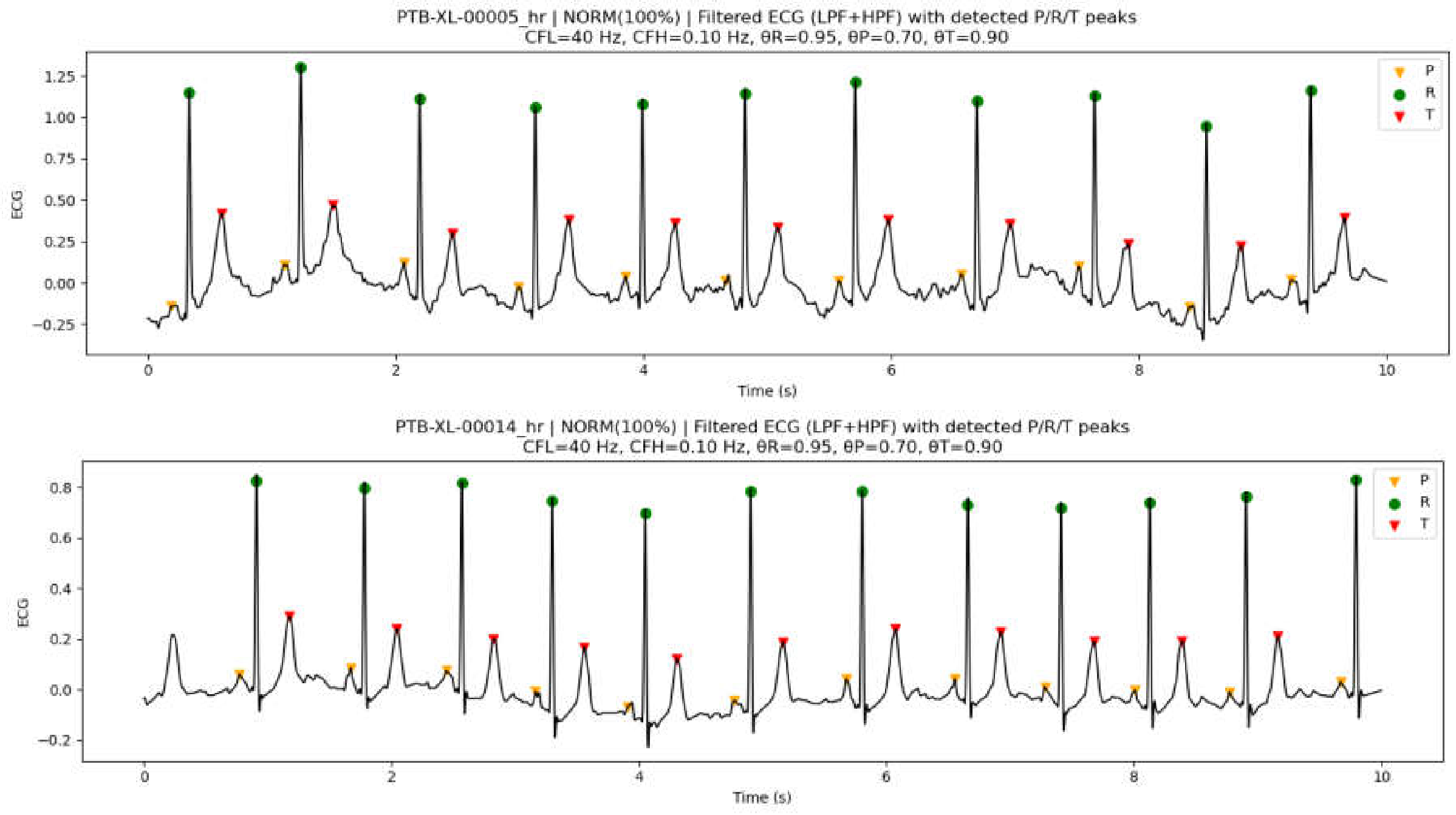
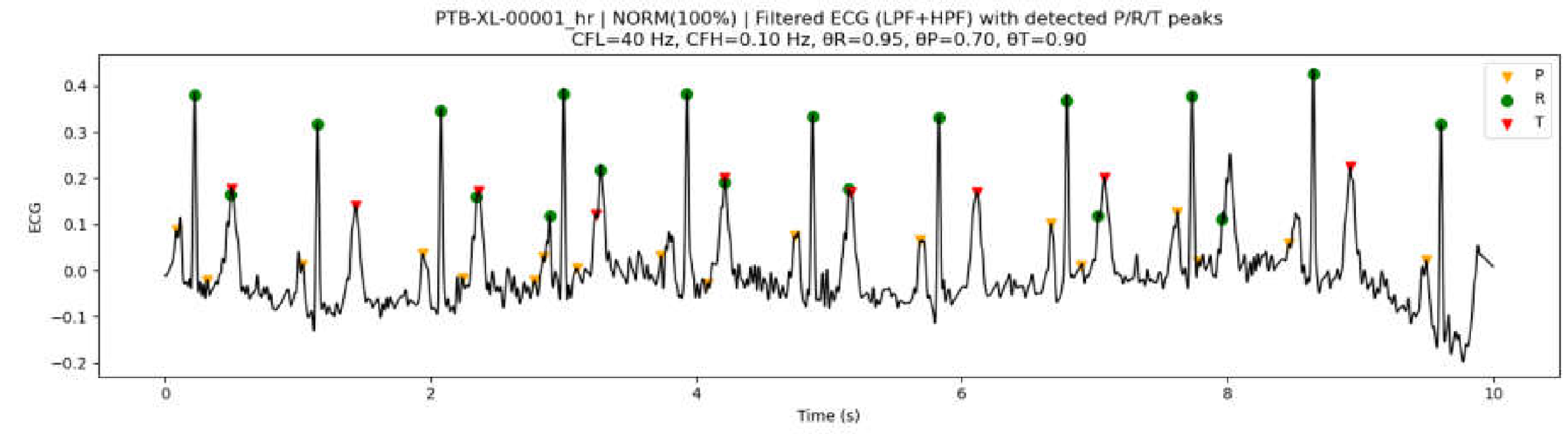
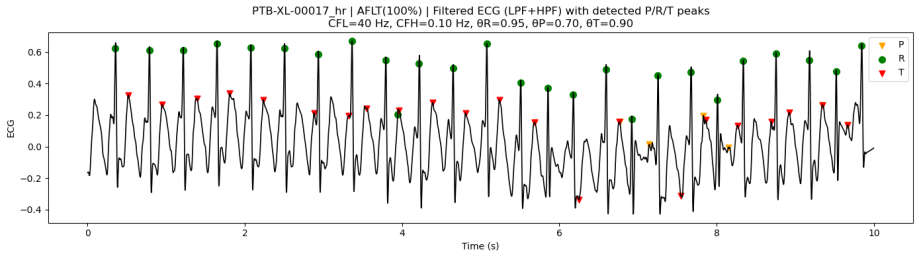
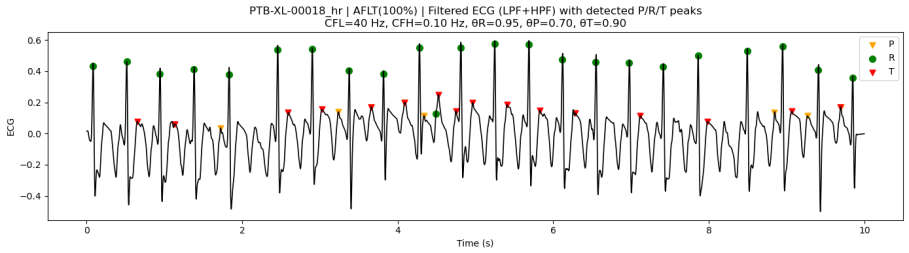
3.7.2. Peak Detection Examples on PTB-XL ECG Data
4. Discussion
4.1. Interpretation of Baseline Classification Performance and Limitations of AUC-Based Evaluation
4.2. Interpretation of PPV under Extreme Class Imbalance
4.3. PTC as a Human-Inspired Interpretation Model
4.4. Interpretation of Algorithm Behavior Under Arrhythmic Conditions
4.5. Lightweight Design and Practical Implications Compared with Deep Learning Approaches
4.6. Possible Applications to Other Biological Signals
4.7. Study Limitations
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
Abbreviations
| AF | atrial fibrillation |
| AFA | atrial fibrillation with aberrant conduction |
| AFLT | atrial flutter |
| AP | average precision |
| AUC | area under the ROC curve |
| CNN | convolutional neural network |
| ECG | electrocardiogram |
| FN | false-negative |
| FP | false-positive |
| GUI | graphical user interface |
| IMI | inferior myocardial infarction |
| ISN | irregular sinus rhythm |
| LUDB | Lobachevsky University Electrocardiography Database |
| PR | precision–recall |
| PPV | positive predictive value |
| PTC | physiological temporal constraint |
| ROC | receiver operating characteristic |
| RR interval | RRI |
| SAW | sinus arrhythmia with wandering atrial pacemaker |
| SBW | sinus bradycardia with wandering atrial pacemaker |
| Se | sensitivity |
| SNB | sinus bradycardia |
| SNA | sinus arrhythmia |
| SR | sinus rhythm |
| SNT | sinus tachycardia |
| SRW | sinus rhythm with wandering atrial pacemaker |
| TP | true-positive |
| TN | true-negative |
References
- Thakor, N.V.; Webster, J.G.; Tompkins, W.J. Optimal QRS detector. Med. Biol. Eng. Comput. 1983, 21, 343–350.
- Pan, J.; Tompkins, W.J. A Real-Time QRS Detection Algorithm. IEEE Trans. Biomed. Eng. 1985, 32, 230–236.
- Hamilton, P.S.; Tompkins, W.J. Quantitative Investigation of QRS Detection Rules Using the MIT/BIH Arrhythmia Database. IEEE Trans. Biomed. Eng. 1986, 33, 1157–1165.
- Yun, D.; Lee, H.-C.; Jung, C.-W.; Kwon, S.; Lee, S.-R.; Kim, K.; Kim, S.U.; Han, S.S. Robust R-Peak Detection in an Electrocardiogram with Stationary Wavelet Transformation and Separable Convolution. Sci. Rep. 2022, 12, 19638.
- Nurmaini, S.; Darmawahyuni, A.; Rachmatullah, M.N.; Firdaus, F.; Sapitri, A.I.; Tutuko, B.; Tondas, A.E.; Putra, M.H.P.; Islami, A. Robust Electrocardiogram Delineation Model for Automatic Morphological Abnormality Interpretation. Sci. Rep. 2023, 13, 13736.
- Su, X.; Wang, X.; Ge, H. Exercise ECG Classification Based on Novel R-Peak Detection Using BILSTM-CNN and Multi-Feature Fusion Method. Electronics 2025, 14, 281.
- Moskalenko, V.; Zolotykh, N.; Osipov, G.V. Deep learning for ECG segmentation. In Advances in Neural Computation, Machine Learning, and Cognitive Research III; Studies in Computational Intelligence; Springer: Cham, Switzerland, 2020, 246–254.
- Peimankar, A.; Puthusserypady, S. DENS-ECG: A deep learning approach for ECG signal delineation. Expert Syst. Appl. 2021, 165, 113911.
- Wu, W.; Huang, Y.; Wu, X. A New Deep Learning Method with Self-Supervised Learning for Delineation of the Electrocardiogram. Entropy 2022, 24, 1828.
- Krasteva, V.; Stoyanov, T.; Schmid, R.; Jekova, I. Delineation of 12-Lead ECG Representative Beats Using Convolutional Encoder–Decoders with Residual and Recurrent Connections. Sensors 2024, 24, 4645.
- Niu, Y.; Lin, N.; Tian, Y.; Tang, K.; Liu, B. ECG Waveform Segmentation via Dual-Stream Network with Selective Context Fusion. Electronics 2025, 14, 3925.
- Kalyakulina, A.; Yusipov, I.; Moskalenko, V.; Nikolskiy, A.; Kosonogov, K.; Zolotykh, N.; Ivanchenko, M. Lobachevsky University Electrocardiography Database (Version 1.0.1). PhysioNet 2021.
- Kalyakulina, A.I.; Yusipov, I.I.; Moskalenko, V.A.; Nikolskiy, A.V.; Kosonogov, K.A.; Osipov, G.V.; Zolotykh, N.Y.; Ivanchenko, M.V. LUDB: A New Open-Access Validation Tool for Electrocardiogram Delineation Algorithms. IEEE Access 2020, 8, 186181–186190.
- Goldberger, A.L.; Amaral, L.A.N.; Glass, L.; Hausdorff, J.M.; Ivanov, P.C.; Mark, R.G.; Stanley, H.E. PhysioBank, PhysioToolkit, and PhysioNet: Components of a New Research Resource for Complex Physiologic Signals. Circulation 2000, 101, e215–e220.
- Hnatkova, K.; Andršová, I.; Toman, O.; Smetana, P.; Huster, K.M.; Šišáková, M.; Barthel, P.; Novotný, T.; Schmidt, G.; Malik, M. Spatial distribution of physiologic 12-lead QRS complex. Sci. Rep. 2021, 11, 4289.
- Sattar, Y.; Chhabra, L. Electrocardiogram. In StatPearls [Internet]; StatPearls Publishing: Treasure Island, FL, USA, 2023. Available online: https://www.ncbi.nlm.nih.gov/books/NBK549803/(accessed on 1 January 2026).
- Surawicz, B.; Childers, R.; Deal, B.J.; Gettes, L.S. AHA/ACCF/HRS Recommendations for the Standardization and Interpretation of the Electrocardiogram: Part III: Intraventricular Conduction Disturbances. J. Am. Coll. Cardiol. 2009, 53, 976–981.
- Wagner, P.; Strodthoff, N.; Bousseljot, R.; Samek, W.; Schaeffter, T. PTB-XL, a Large Publicly Available Electrocardiography Dataset (Version 1.0.3). PhysioNet 2022.
- Wagner, P.; Strodthoff, N.; Bousseljot, R.-D.; Kreiseler, D.; Lunze, F.I.; Samek, W.; Schaeffter, T. PTB-XL: A Large Publicly Available ECG Dataset. Sci. Data 2020, 7, 154.
- Dotsinsky, I. ECG Baseline Wander Removal through the Use of the Least Mean Squares Adaptive Filtering Technique. BioMed. Eng. OnLine 2007, 6, 18.
- Lenis, G.; Pilia, N.; Loewe, A.; Schulze, W.H.; Dössel, O. Comparison of baseline wander removal techniques considering the preservation of ST changes in the ischemic ECG: A simulation study. Comput. Math. Methods Med. 2017, 2017, 9295029.
- Kher, R. Signal processing techniques for removing noise from ECG signals. J. Biomed. Eng. Res. 2019, 3, 101.
- Dobrev, D.; Neycheva, T.; Krasteva, V.; Jekova, I. Design of High-Pass and Low-Pass Active Inverse Filters to Compensate for Distortions in RC-Filtered Electrocardiograms. Technologies 2025, 13, 159.
- Davis, J.; Goadrich, M. The Relationship between Precision–Recall and ROC Curves. Proceedings of the 23rd International Conference on Machine Learning (ICML 2006), Pittsburgh, PA, USA, 25–29 June 2006; pp. 233–240.
- Saito, T.; Rehmsmeier, M. The Precision–Recall Plot Is More Informative than the ROC Plot When Evaluating Binary Classifiers on Imbalanced Datasets. PLoS ONE 2015, 10, e0118432.
- Ashley, E.A.; Niebauer, J. Cardiology Explained; Remedica: London, UK, 2004; Chapter 3: Conquering the ECG.
- Nesheiwat, Z.; Goyal, A.; Jagtap, M. Atrial Fibrillation. StatPearls [Internet]; StatPearls Publishing: Treasure Island, FL, USA, 2025. Available online: https://www.ncbi.nlm.nih.gov/books/NBK526072/(accessed on 1 January 2026).
- Kiranyaz, S.; Ince, T.; Gabbouj, M. Real-Time Patient-Specific ECG Classification by 1-D Convolutional Neural Networks. IEEE Trans. Biomed. Eng. 2016, 63, 664–675.
- Acharya, U.R.; Oh, S.L.; Hagiwara, Y.; Tan, J.H.; Adam, M.; Gertych, A.; Tan, R.S. A deep convolutional neural network model to classify heartbeats. Comput. Biol. Med. 2017, 89, 389–396.
- Hannun, A.Y.; Rajpurkar, P.; Haghpanahi, M.; Tison, G.H.; Bourn, C.; Turakhia, M.P.; Ng, A.Y. Cardiologist-Level Arrhythmia Detection and Classification in Ambulatory Electrocardiograms Using a Deep Neural Network. Nat. Med. 2019, 25, 65–69.
- Yao, Q.; Wang, R.; Fan, X.; Liu, J.; Li, Y. Multi-Class Arrhythmia Detection from 12-Lead Varied-Length ECG Using Attention-Based Time-Incremental Convolutional Neural Network. Inf. Fusion 2020, 53, 174–182.
- Xia, Y.; Wulan, N.; Wang, K.; Zhang, H. Detecting Atrial Fibrillation by Deep Convolutional Neural Networks. Comput. Biol. Med. 2018, 93, 84–92.
- Oh, S.L.; Ng, E.Y.K.; Tan, R.S.; Acharya, U.R. Automated diagnosis of arrhythmia using combination of CNN and LSTM techniques with variable length heart beats. Comput. Biol. Med. 2018, 102, 278–287.
- Pauker, S.G.; Kassirer, J.P. The threshold approach to clinical decision making. N. Engl. J. Med. 1980, 302, 1109–1117.
- Julien, C. The enigma of Mayer waves: Facts and models. Cardiovasc. Res. 2006, 70, 12–21.
- Stauss, H.M. Identification of blood pressure control mechanisms by power spectral analysis. Clin. Exp. Pharmacol. Physiol. 2007, 34, 362–368.
- Westerhof, N.; Lankhaar, J.W.; Westerhof, B.E. The arterial Windkessel. Med. Biol. Eng. Comput. 2009, 47, 131–141.
- Feldman, J.L.; Del Negro, C.A. Looking for inspiration: new perspectives on respiratory rhythm. Nat. Rev. Neurosci. 2006, 7, 232–242.
- Boukens, B.J.; Rivaud, M.R.; Rentschler, S.; Coronel, R. Misinterpretation of the mouse ECG: ‘mammalian’ ECG waves are species dependent. J. Physiol. 2014, 592, 4613–4626.
| Step | Candidate time (ms) |
Score | Previously selected R-peaks (ms) |
Distance to nearest selected peak (ms) |
Decision | Reason |
|---|---|---|---|---|---|---|
| 1 | 1600 | 0.99 | — | — | Adopt | First peak, highest score |
| 2 | 800 | 0.97 | [1600] | 800 (>50) | Adopt | Far from 1600 ms |
| 3 | 1635 | 0.94 | [800,1600] | 35 (≤50) | Reject | Within refractory of 1600 ms |
| 4 | 2400 | 0.92 | [800,1600] | 800 (>50) | Adopt | New beat |
| 5 | 770 | 0.88 | [800,1600,2400] | 30 (≤50) | Reject | Within refractory of 800 ms |
| 6 | 3220 | 0.85 | [800,1600,2400] | 820 (>50) | Adopt | New beat |
| 7 | 3270 | 0.8 | [800,1600,2400,3220] | 50 (≤50) | Reject | Within refractory of 3220 ms |
| 8 | 4020 | 0.76 | [800,1600,2400,3220] | 800 (>50) | Adopt | Distant new beat |
| 9 | 2370 | 0.71 | [800,1600,2400,3220,4020] | 30 (≤50) | Reject | Within refractory of 2400 ms |
| 10 | 4820 | 0.68 | [800,1600,2400,3220,4020] | 800 (>50) | Adopt | New beat |
| Classifier | P-peak | R-peak | T-peak | |||
|---|---|---|---|---|---|---|
| AUC | AP | AUC | AP | AUC | AP | |
| XGB | 0.889 | 0.035 | 0.992 | 0.252 | 0.913 | 0.080 |
| LGR | 0.870 | 0.014 | 0.990 | 0.169 | 0.871 | 0.027 |
| QDA | 0.684 | 0.003 | 0.984 | 0.073 | 0.754 | 0.012 |
| NB | 0.732 | 0.007 | 0.973 | 0.049 | 0.821 | 0.022 |
| KNN | 0.541 | 0.005 | 0.727 | 0.109 | 0.567 | 0.012 |
| LDA | 0.684 | 0.003 | 0.980 | 0.167 | 0.824 | 0.011 |
| Classifier | R-peak (tol=±20 ms) | ||||||
|---|---|---|---|---|---|---|---|
| N (Total ECG time points) |
Se | PPV | F1 | TP | FP | FN | |
| XGB | 225,000 | 0.963±0.038 | 0.787±0.025 | 0.866±0.026 | 397±24 | 108±14 | 15±16 |
| LGR | 225,000 | 0.955±0.046 | 0.785±0.029 | 0.861±0.035 | 394±25 | 111±19 | 18±17 |
| QDA | 225,000 | 0.932±0.064 | 0.755±0.033 | 0.834±0.041 | 384±32 | 124±18 | 28±26 |
| NB | 225,000 | 0.918±0.040 | 0.750±0.031 | 0.826±0.033 | 378±24 | 126±15 | 34±16 |
| KNN | 225,000 | 0.524±0.056 | 0.796±0.023 | 0.63±0.042 | 224±16 | 58±8 | 188±11 |
| LDA | 225,000 | 0.945±0.056 | 0.813±0.01 | 0.873±0.026 | 389±29 | 89±6 | 23±23 |
| Classifier | P-peak (tol=±20 ms) | ||||||
| N (Total ECG time points) |
Se | PPV | F1 | TP | FP | FN | |
| XGB | 225,000 | 0.794±0.055 | 0.626±0.025 | 0.699±0.032 | 275±22 | 177±24 | 81±24 |
| LGR | 225,000 | 0.804±0.048 | 0.588±0.041 | 0.679±0.042 | 283±20 | 198±23 | 73±13 |
| QDA | 225,000 | 0.669±0.062 | 0.489±0.048 | 0.565±0.052 | 245±26 | 244±20 | 110±20 |
| NB | 225,000 | 0.631±0.05 | 0.46±0.031 | 0.532±0.037 | 235±16 | 254±18 | 121±26 |
| KNN | 225,000 | 0.162±0.034 | 0.485±0.08 | 0.242±0.043 | 49±10 | 71±24 | 307±22 |
| LDA | 225,000 | 0.000 | – | 0.000 | 0 | 0 | 356±23 |
| Classifier | T-peak (tol=±20 ms) | ||||||
| N (Total ECG time points) |
Se | PPV | F1 | TP | FP | FN | |
| XGB | 225,000 | 0.766±0.064 | 0.629±0.057 | 0.69±0.055 | 279±23 | 177±38 | 90±23 |
| LGR | 225,000 | 0.769±0.071 | 0.607±0.059 | 0.678±0.062 | 279±32 | 170±27 | 91±28 |
| QDA | 225,000 | 0.670±0.074 | 0.503±0.047 | 0.575±0.057 | 258±26 | 233±28 | 111±27 |
| NB | 225,000 | 0.755±0.058 | 0.568±0.048 | 0.648±0.051 | 282±23 | 207±20 | 87±19 |
| KNN | 225,000 | 0.253±0.041 | 0.523±0.041 | 0.339±0.041 | 91±8 | 95±9 | 279±18 |
| LDA | 225,000 | 0.041±0.025 | 0.568±0.131 | 0.075±0.044 | 15±9 | 10±4 | 355±20 |
| Classifier | R-peak (tol_10ms) | R-peak (tol_30ms) | ||||
|---|---|---|---|---|---|---|
| Se | PPV | F1 | Se | PPV | F1 | |
| XGB | 0.946±0.050 | 0.773±0.033 | 0.85±0.037 | 0.964±0.037 | 0.787±0.026 | 0.867±0.026 |
| LGR | 0.937±0.059 | 0.769±0.037 | 0.845±0.045 | 0.966±0.033 | 0.794±0.026 | 0.872±0.026 |
| Classifier | P-peak (tol_10ms) | P-peak (tol_30ms) | ||||
| Se | PPV | F1 | Se | PPV | F1 | |
| XGB | 0.698±0.047 | 0.551±0.025 | 0.615±0.029 | 0.844±0.061 | 0.666±0.023 | 0.744±0.034 |
| LGR | 0.692±0.043 | 0.506±0.031 | 0.585±0.033 | 0.87±0.047 | 0.636±0.037 | 0.734±0.037 |
| Classifier | T-peak (tol_10ms) | T-peak (tol_30ms) | ||||
| Se | PPV | F1 | Se | PPV | F1 | |
| XGB | 0.681±0.057 | 0.559±0.054 | 0.613±0.051 | 0.799±0.061 | 0.656±0.054 | 0.719±0.051 |
| LGR | 0.675±0.065 | 0.533±0.051 | 0.595±0.055 | 0.809±0.069 | 0.639±0.059 | 0.714±0.061 |
| Classifier | win_len | refR (ms) | θR |
|---|---|---|---|
| XGB | 15 [13–15] (15) | 60 [40–80] (40) | 0.95 [0.60–0.95] (0.95) |
| LGR | 7 [7–15] (7) | 55 [30–80] (40) | 0.85 [0.65–0.90] (0.90) |
| QDA | 11 [9–15] (11) | 80 [40–80] (80) | 0.60 [0.45–1.00] (0.60) |
| NB | 7 [7–11] (7/9) | 75 [40–80] (80) | 0.55 [0.40–0.90] (0.55) |
| KNN | 9 [7–15] (9) | 30 [30–30] (30) | 0.40 [0.40–0.40] (0.40) |
| LDA | 7 [7–15] (7) | 40 [30–80] (40) | 0.40 [0.40–0.55] (0.40) |
| Classifier | θP | θT | |
| XGB | 0.70 [0.40–0.80] (0.70) | 0.70 [0.05–0.90] (0.80) | |
| LGR | 0.50 [0.50–0.60] (0.50) | 0.45 [0.05–0.50] (0.50) | |
| QDA | 0.05 [0.05–0.90] (0.05) | 0.05 [0.05–0.90] (0.05) | |
| NB | 0.80 [0.05–0.90] (0.90) | 0.05 [0.05–0.90] (0.05) | |
| KNN | 0.05 [0.05–0.05] (0.05) | 0.05 [0.05–0.05] (0.05) | |
| LDA | — | 0.05 [0.05–0.05] (0.05) |
| Classifier | Ppre (ms) | Ppost (ms) | Tpre (ms) | Tpost (ms) |
|---|---|---|---|---|
| XGB | 200 [180–240] (200) | 100 [80–100] (100) | 120 [100–120] (120) | 350 [300–450] (350) |
| LGR | 240 [200–240] (200) | 100 [100–100] (100) | 120 [60–120] (120) | 350 [300–450] (350) |
| QDA | 180 [160–200] (180) | 100 [100–100] (100) | 120 [80–120] (120) | 350 [300–450] (350) |
| NB | 220 [180–220] (220) | 100 [80–100] (100) | 120 [80–120] (120) | 350 [300–400] (350) |
| KNN | 220 [180–260] (200) | 40 [40–100] (40) | 40 [40–120] (40) | 350 [350–450] (350) |
| LDA | — | — | 40 [40–120] (40) | 300 [300–400] (300) |
| Classifier | Parameter | P-peak | R-peak | T-peak |
|---|---|---|---|---|
| XGB | n_estimators | 300 [300–300] (300) | 300 [300–300] (300) | 300 [300–300] (300) |
| max_depth | 4 [4–4] (4) | 4 [4–4] (4) | 4 [4–4] (4) | |
| 0.1 [0.1–0.1] (0.1) | 0.1 [0.1–0.1] (0.1) | 0.1 [0.1–0.1] (0.1) | ||
| LGR | C | 10 [1–10] (10) | 0.1 [0.1–1] (0.1/1) | 10 [0.1–10] (10) |
| KNN | n_neighbors | 3 [3–8] (3) | 3 [3–8] (3) | 3 [3–3] (3) |
| Peak | N (Total ECG time points) |
Se | PPV | F1 | TP | FP | FN |
|---|---|---|---|---|---|---|---|
| R | 867,564 | 0.931 | 0.764 | 0.839 | 1509 | 466 | 111 |
| P | 867,564 | 0.786 | 0.414 | 0.542 | 684 | 970 | 186 |
| T | 867,564 | 0.645 | 0.582 | 0.612 | 937 | 672 | 515 |
| Arrhythmia | R-peak | Arrhythmia | P-peak | Arrhythmia | T-peak |
|---|---|---|---|---|---|
| SNT | 0.935 | SNT | 0.867 | SRW | 0.882 |
| SRW | 0.917 | SNA | 0.786 | SAW | 0.842 |
| AFA | 0.905 | SAW | 0.750 | SNA | 0.807 |
| SNA | 0.904 | SR | 0.727 | SR | 0.769 |
| SR | 0.902 | SNB | 0.625 | SNT | 0.740 |
| SAW | 0.900 | ISN | 0.598 | ISN | 0.729 |
| ISN | 0.870 | SRW | 0.324 | TAF | 0.677 |
| AFLT | 0.857 | SBW | 0.320 | SNB | 0.655 |
| SNB | 0.844 | AF | — | AFLT | 0.352 |
| AF | 0.746 | AFA | — | AFA | 0.206 |
| SBW | 0.727 | AFLT | — | SBW | 0.000 |
| Arrhythmia | N (Total ECG time points) |
PPV | Se | F1 | TP | FP | FN | FP/N (%) |
|---|---|---|---|---|---|---|---|---|
| AF | 209,412 | 0.000 | — | — | 0 | 353 | 0 | 0.169 |
| AFA | 14,958 | 0.000 | — | — | 0 | 43 | 0 | 0.287 |
| AFLT | 44,874 | 0.000 | — | — | 0 | 136 | 0 | 0.303 |
| Parameters | CFH (Hz) | |||
|---|---|---|---|---|
| 0.05 | 0.1 | 0.2 | 0.3 | |
| win_len | 15 | 15 | 13 | 15 |
| refR (ms) | 50 | 50 | 40 | 30 |
| θR | 0.95 | 0.95 | 0.95 | 0.95 |
| θP | 0.7 | 0.7 | 0.6 | 0.6 |
| θT | 0.9 | 0.9 | 0.9 | 0.9 |
| Ppre (ms) | 200 | 200 | 200 | 200 |
| Ppost (ms) | 100 | 100 | 100 | 100 |
| Tpre (ms) | 120 | 120 | 120 | 60 |
| Tpost (ms) | 450 | 450 | 450 | 450 |
| n_estimators | 300 | 300 | 300 | 300 |
| max_depth | 4 | 4 | 4 | 4 |
| 0.1 | 0.1 | 0.1 | 0.1 | |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).




