Submitted:
06 February 2026
Posted:
09 February 2026
You are already at the latest version
Abstract
Keywords:
1. Introduction
- A strategy for correcting non-healthy misclassified pixels is introduced, showing its effectiveness for improving the tumor detection capability of any given segmentator in the HS domain.
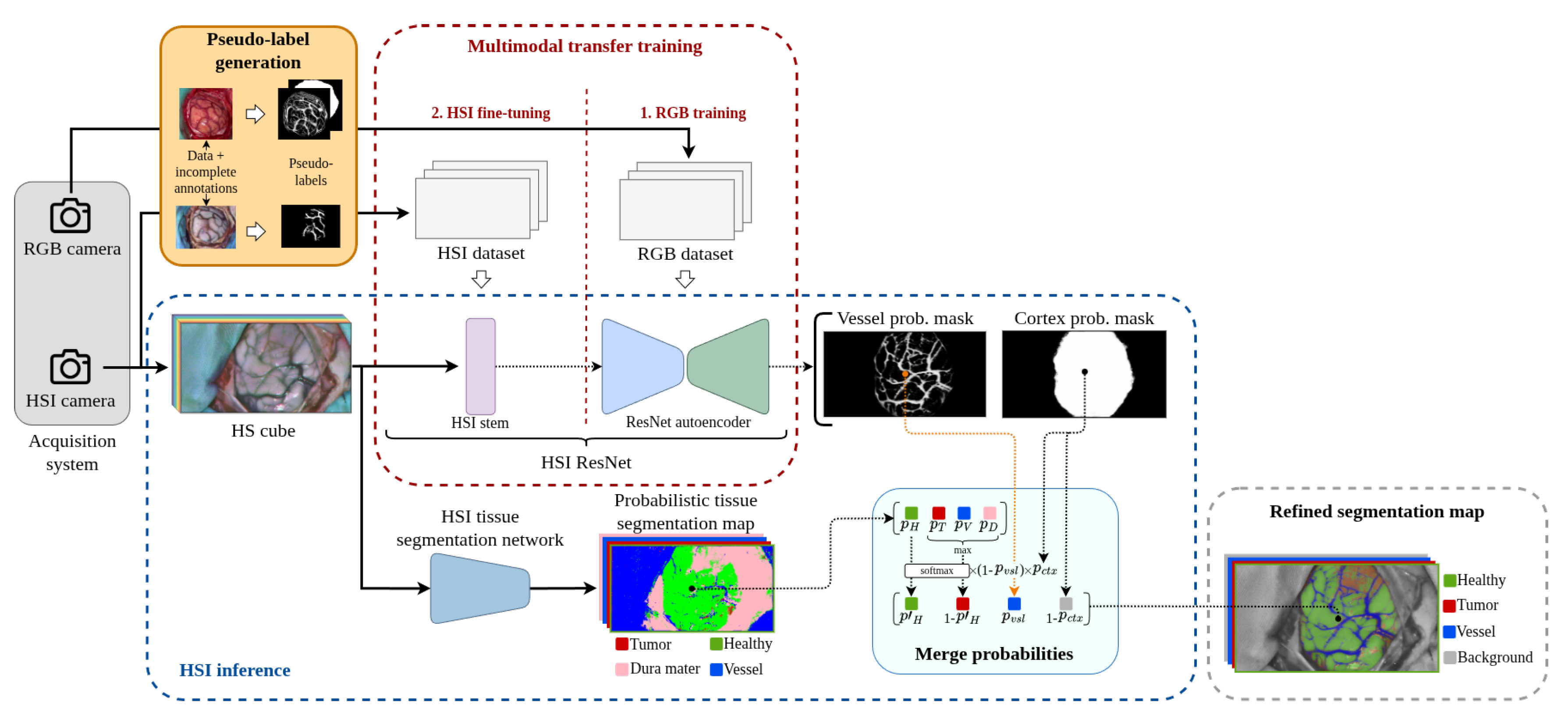
- To propose an RGB and HSI multimodal training methodology based on incomplete annotations capable of producing an accurate segmentation of the brain parenchyma and its blood vessels.
- To suggest the complementation of an HSI-NIR image source with an RGB image modality as a factor of improvement for brain tumor detection by enabling the proposed misclassification error correction strategy.
2. Related Work
2.1. Hyperspectral Imaging in Brain Tumor Detection
2.2. In-Vivo Brain Cortex Segmentation
2.3. Cortical Blood Vessel Segmentation
2.4. Limited Supervision in Medical Image Segmentation
2.5. Pseudo-Label Based Supervision
2.6. Multimodal Learning for Medical Image Segmentation
3. Materials and Methods
3.1. Data Acquisition
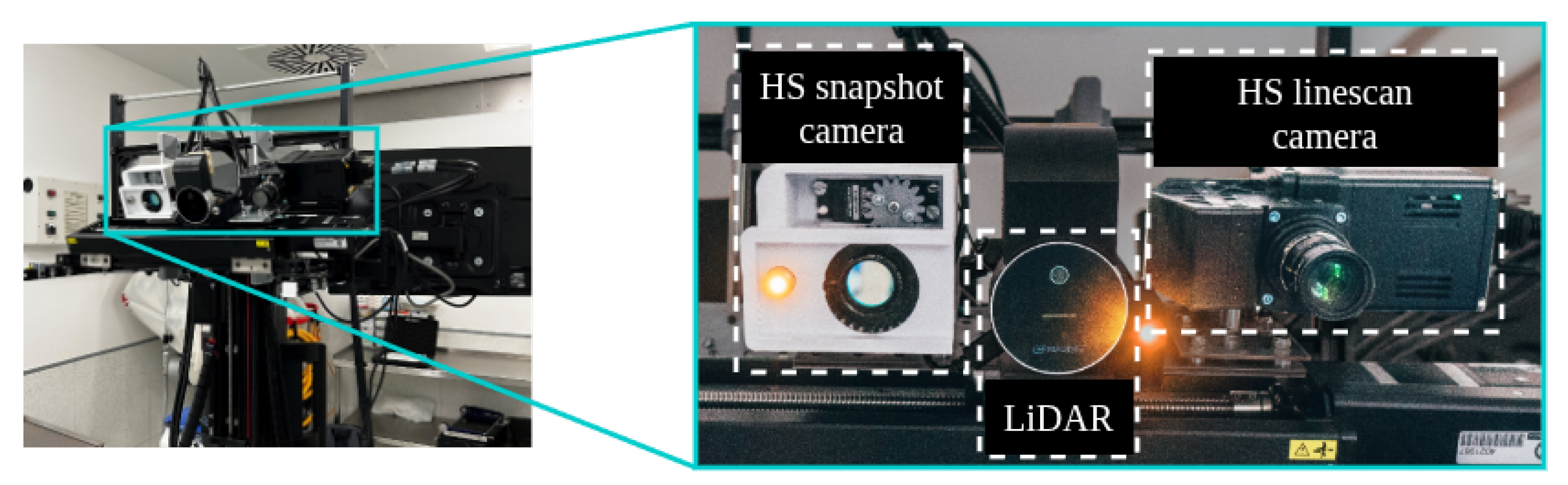
3.1.1. Acquisition Systems
3.1.2. Capturing Procedure
3.1.3. Data Preprocessing
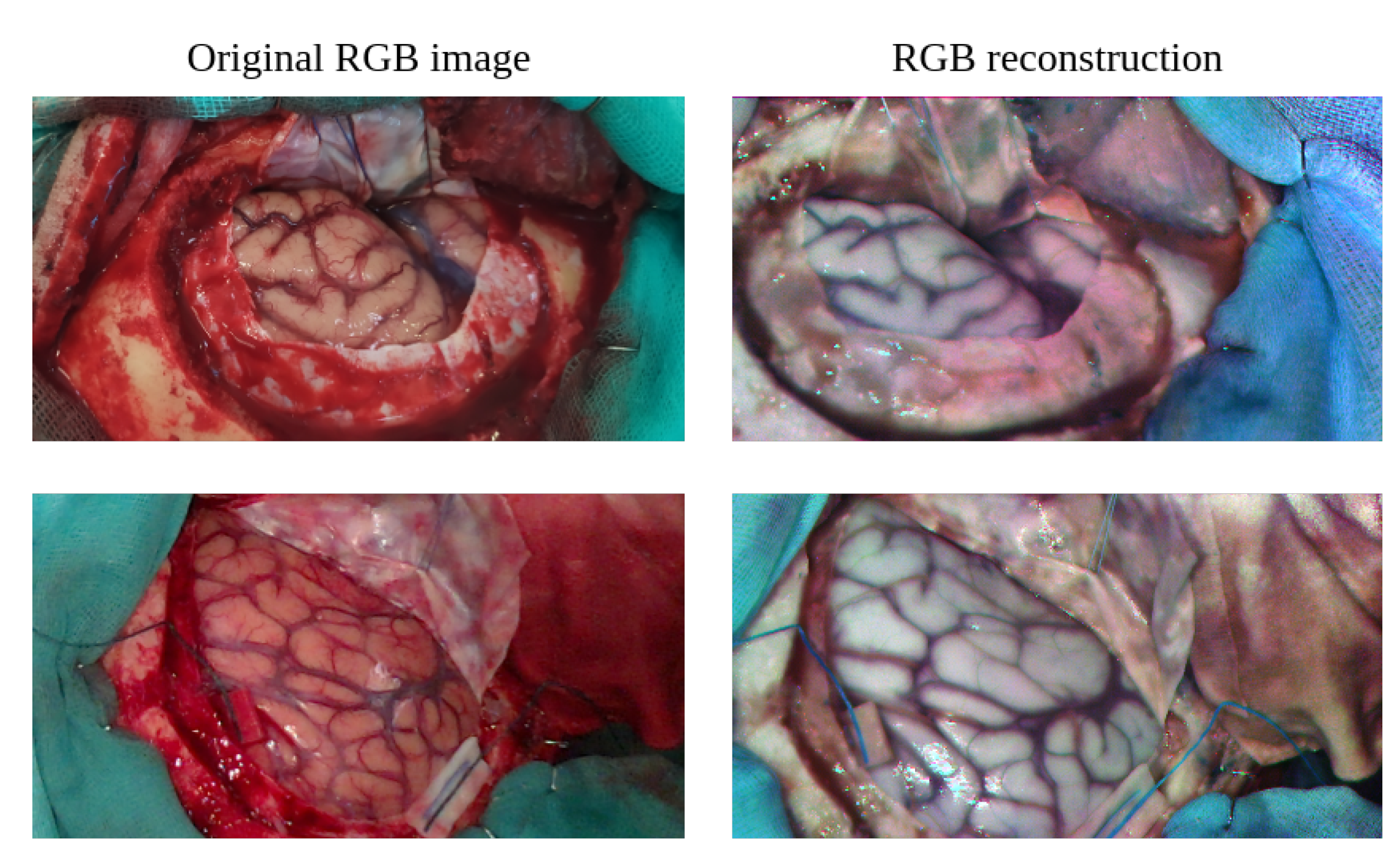
3.1.4. RGB Image Reconstruction from HSI
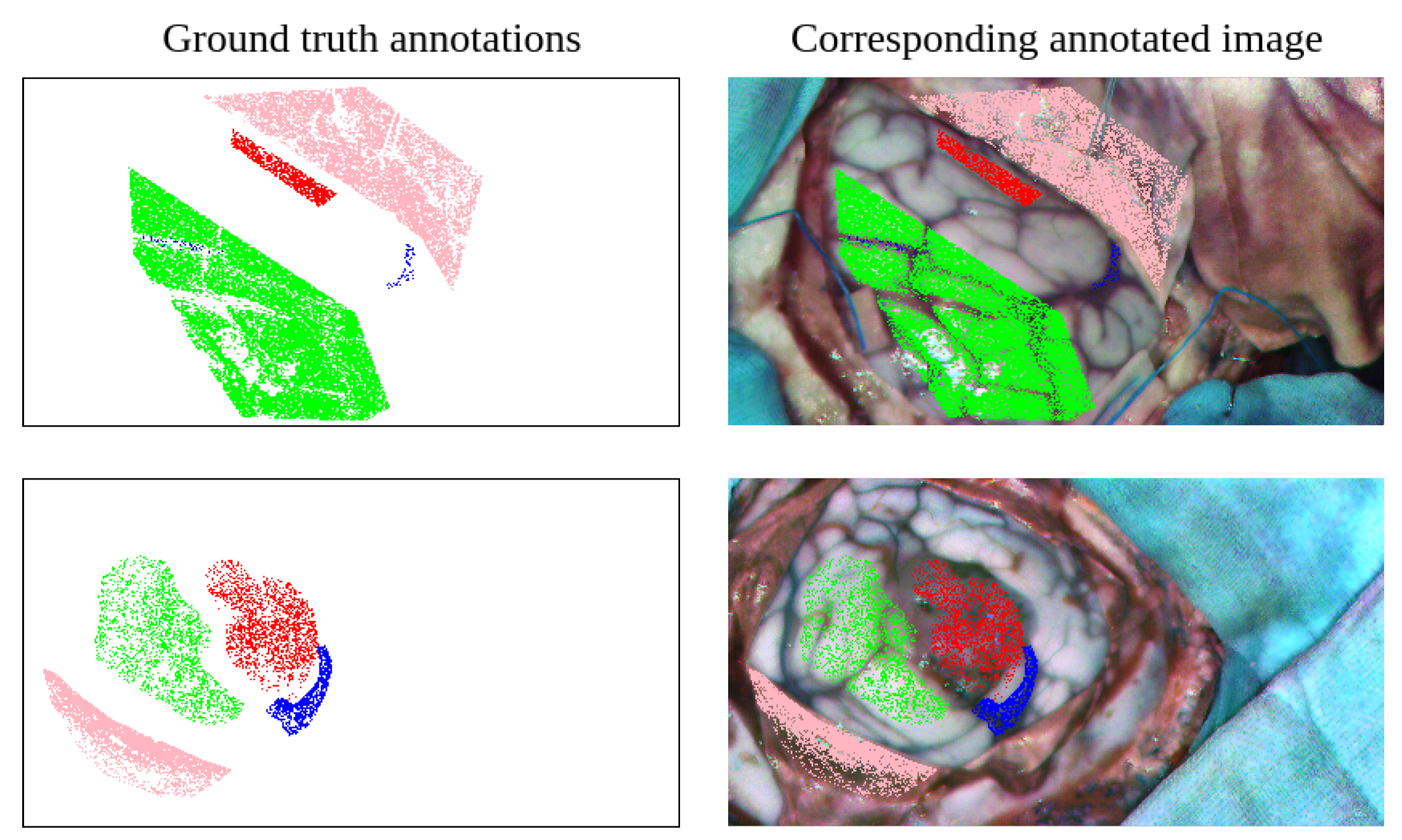
3.1.5. Hyperspectral Image Labeling Procedure
3.1.6. Dataset Composition
3.1.7. RGB Simplified Annotations
3.2. Pseudo-Label Generation
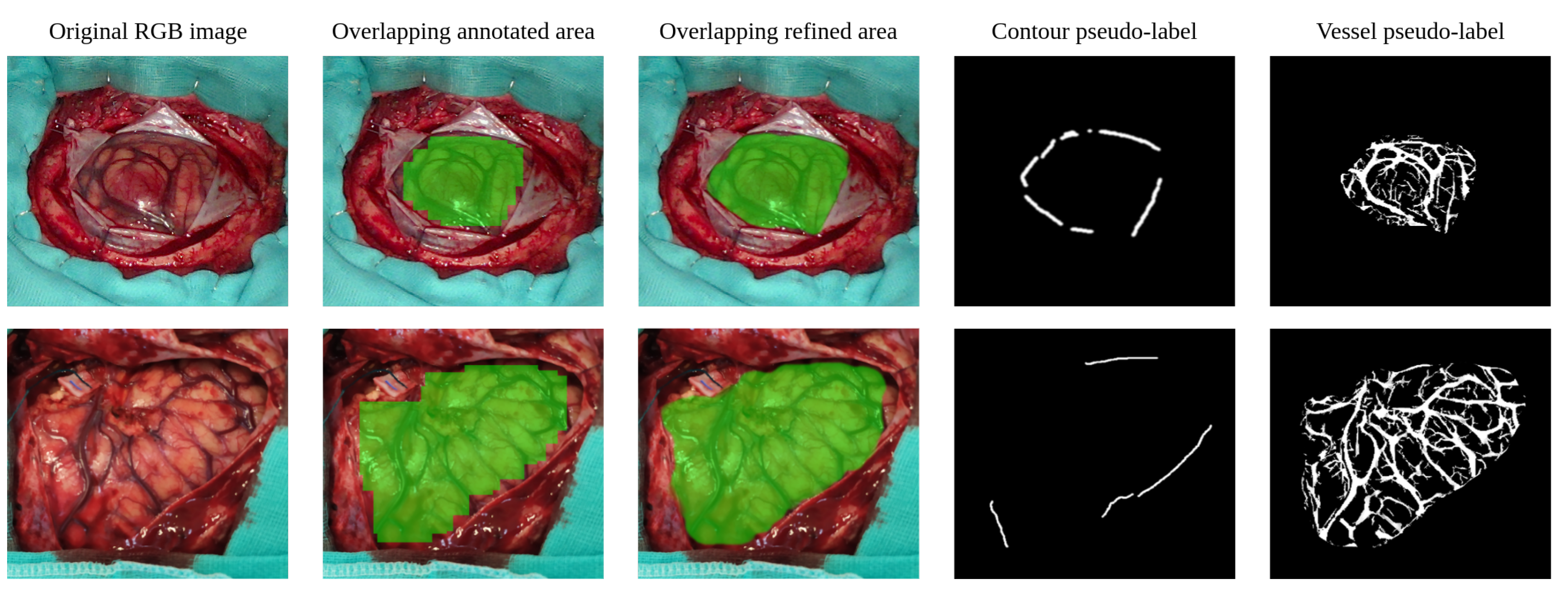
3.2.1. RGB Brain Cortex Annotation Refining
| Algorithm 1: Expansion of manual annotations |
 |
- Local clustering, in which the labeled region is divided into two clusters with the intention of separating the pixels of the cerebral cortex from the pixels of the blood vessels.
- Global clustering, by segmenting the entire image using a number of clusters N estimated using the Calinski-Harabasz (CH) score [42]. For each image, the interval between 5 and 20 clusters is evaluated, selecting the number of clusters that yields the highest CH score. In order to cautiously expand the manually annotated regions by these clusters, the number N suggested by the CH score is multiplied by a given factor f, thus performing an intentional over-segmentation. In this work, the factor f is empirically found to produce excessively atomised clusters above a value of 3, which makes mask expansion problematic. It is therefore set to 3.
3.2.2. RGB Brain Surface Perimeter Approximation
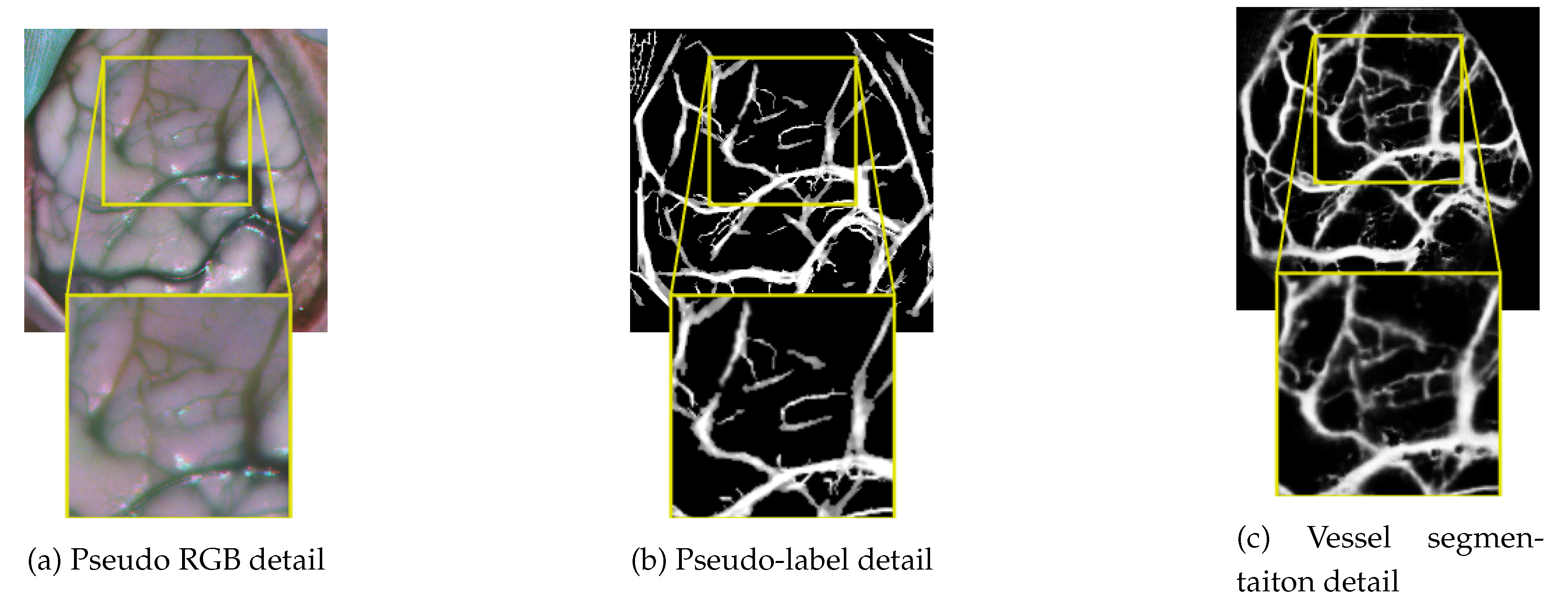
3.2.3. Cortical Vessel Pseudo-Label Generation
3.2.4. HSI Ground Truth Densification and Background Complementation
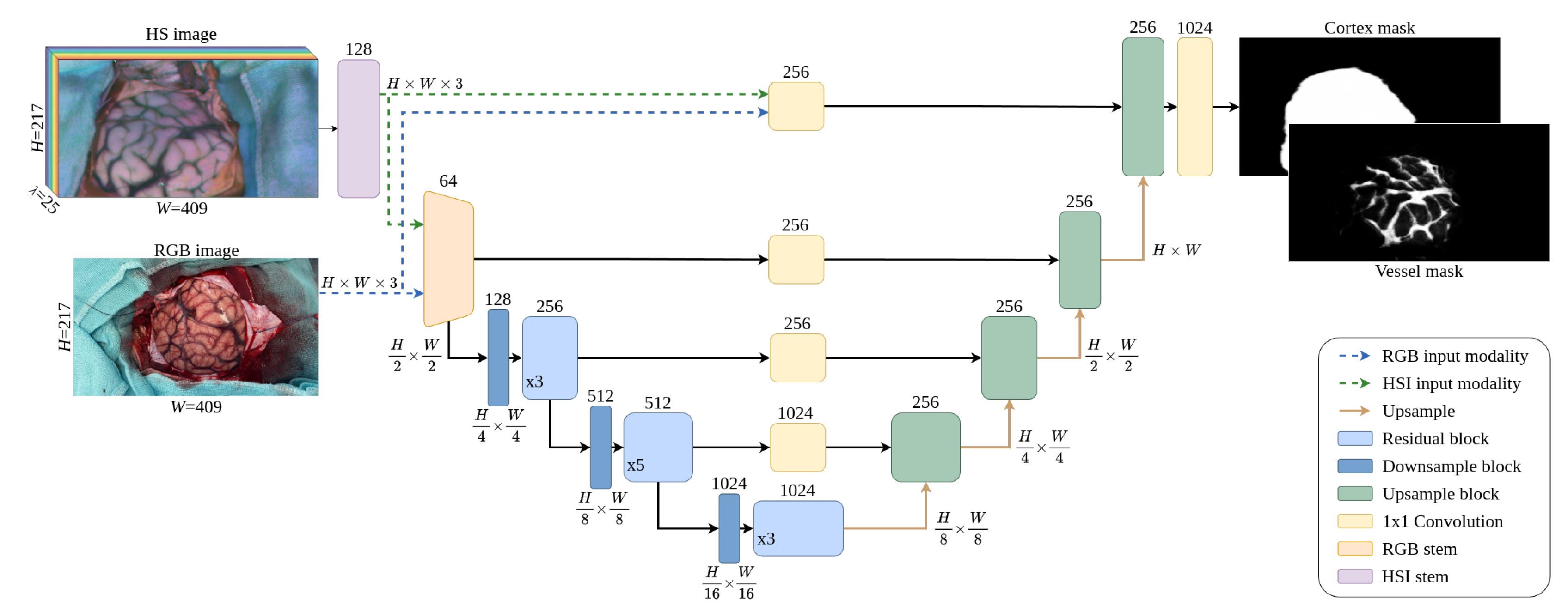
3.3. Neural Network Architecture
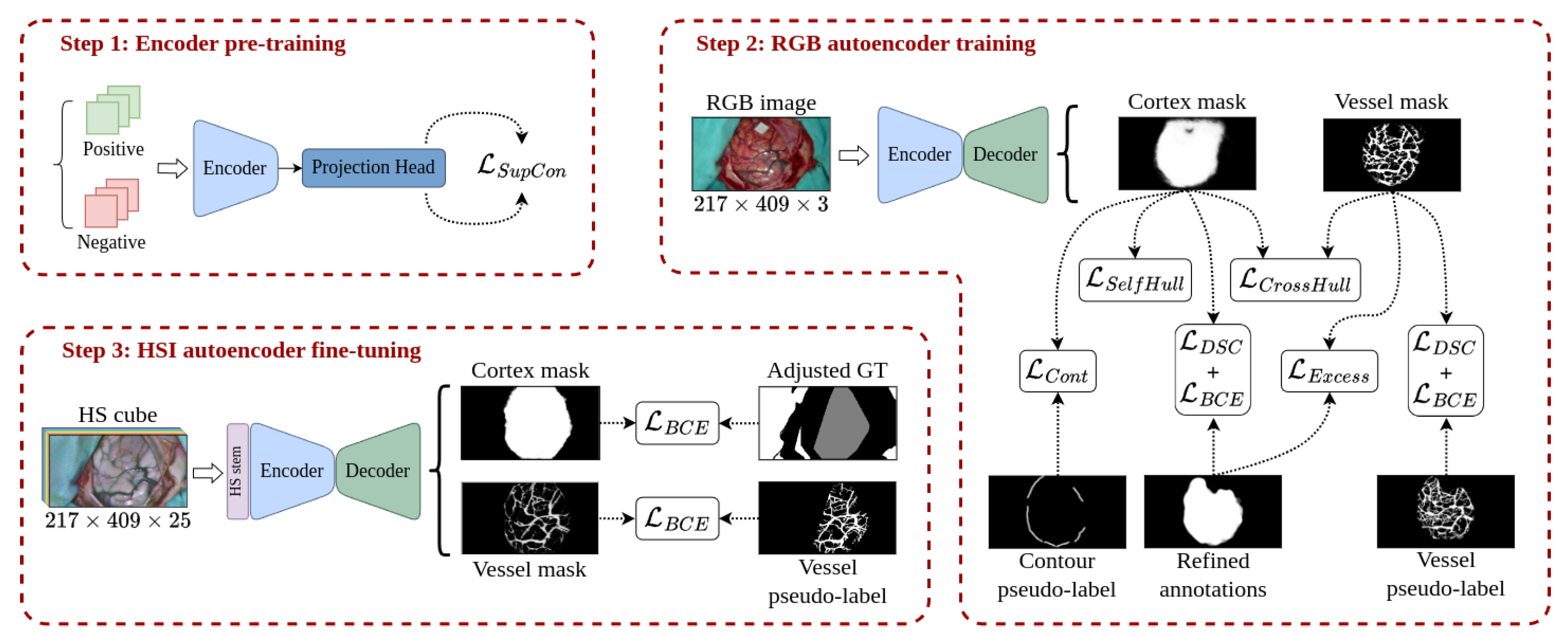
3.4. Multimodal Training Methodology
3.4.1. Encoder Pre-Training
3.4.2. RGB Domain Training
- : similar to the technique presented in [55], the contour loss aims to guide the limits of the cortex segmentation mask so that it matches the boundaries of the brain surface produced in Section 3.2.1. To do so, the limits of the cortex mask generated by the network are extracted using the Canny algorithm and dilated with an elliptical kernel. Then, the BCE is calculated between the extracted edges and the contour pseudo-labels only for the pixels where the contour pseudo-labels are greater than zero. Thus, the term penalizes the predicted cortical mask when it is not adjusting to the contour pseudo-labels.
- : since the exposed cerebral cortex is integrated by a single region, the segmentation mask of the cortex cannot be composed of multiple unconnected areas. If this occurs, it might be indicative that the model has partially detected the cortical area or has marked elements that do not belong to it. To force the generation of a single solid mask, the self-hull loss term computes the complement of the DSC between the predicted cortex mask and the area enclosed by its own concave hull [56]2, following a similar idea that the one suggested by Guo et al. in [57]. Hence, sparse segmentation of the cortical area or scattered activations in external zones produce empty areas within the concave hull of the predicted mask, leading to high loss values. It is important to note that this term must be used in conjunction with the constraint to avoid the cortex mask to adjust to a bad concave hull perimeter.
- : in a similar manner as in self-hull loss, the segmentation mask for blood vessels cannot be active outside the bounds of the predicted brain cortex mask and, on the other hand, the brain cortex mask should be confined within the limits of the detected vessels. Hence, the perimeter of both masks should be as close as possible. To constrain the consistency between both network outputs, the cross-hull loss calculates the complement of the DSC between the areas contained within the concave hulls of the predicted cortex and vessel masks.
- : complementary to the element applied to the blood vessel segmentation, excess loss term penalizes the activation of the predicted blood vessel mask outside the bounds of the refined annotations. This penalization is implemented as a minimization of the overlapping between the predicted vessel mask and the complement of the vessel refined annotations through the following equation:where DSC represents the Dice similarity coefficient between the complement of the refined annotation and the predicted vessel mask for the image inside a batch with size B. The factor is set to 10 for greater penalization when is close to one, whilst the logarithmic function smooths the slope of the loss function.
3.4.3. HSI Model Fine-Tuning
3.5. HS Image Combined Inference
4. Experiments and Results
4.1. Experimental Setting
4.2. Comparison with Other Methods
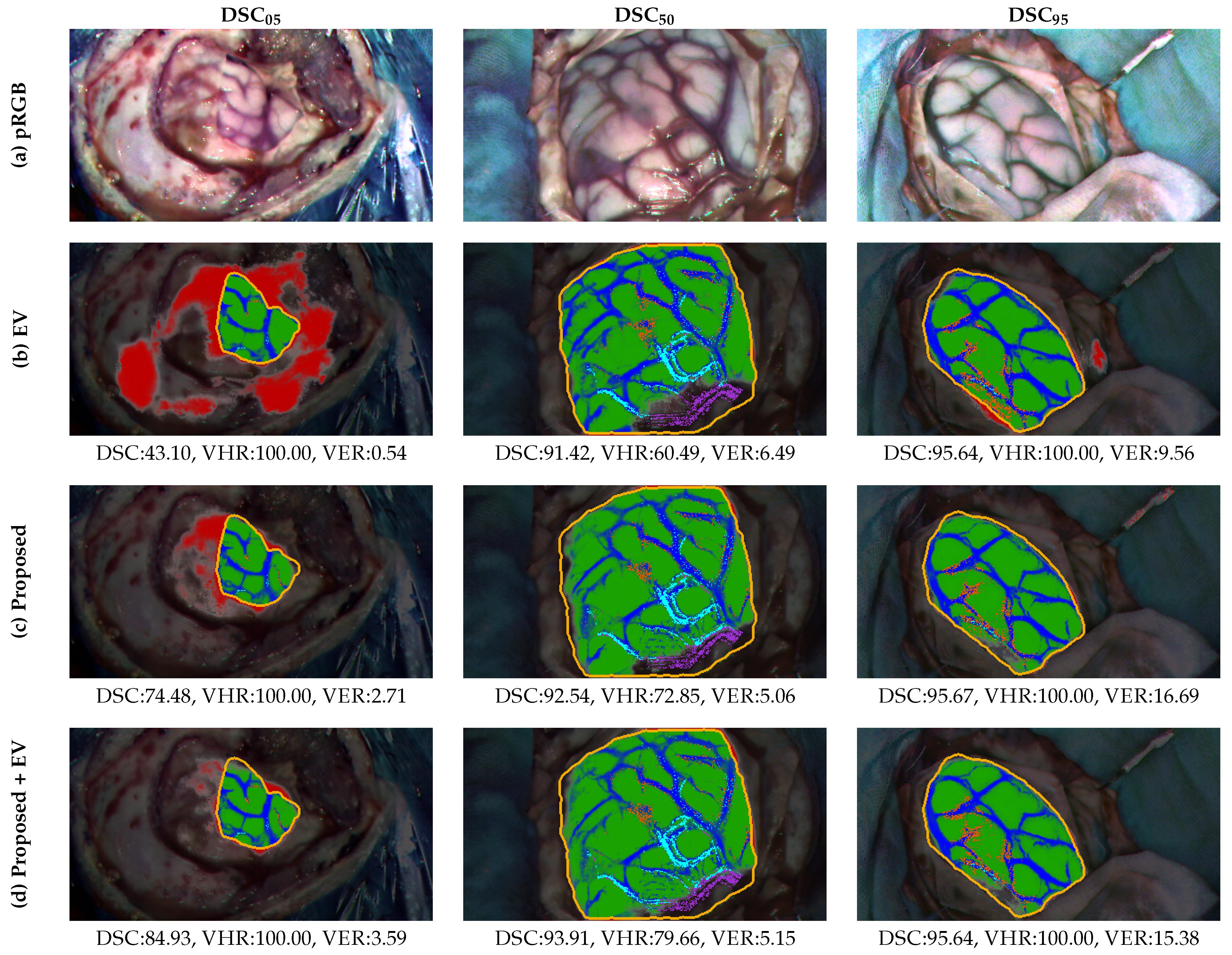
- Structural and shape regularization helps compensate for the lack of complete annotations achieving guiding the model towards a more coherent representations during training. In particular, the equivariance (EV) constraint approach [58] is commonly used to facilitate the model learning by ensuring the consistency between predictions of the transformed versions of the same image. In particular, the weakly supervised tumor segmentation methodology in PET/CT images proposed by G. Patel and J. Dolz [25] is adjusted to test the performance of the equivariance property as a regularization term in the training of the HSI ResNet. To do so, the BCE loss described in Section 3.4.3 is complemented with the mean squared error (MSE) loss between the prediction of the transformed images and the transformed prediction of the original set. The collection of applied transformations is made up of random flips and rotations.
- The second strategy aims to enforce coherence among embeddings corresponding to the same class prior to the generation of the segmentation mask. Therefore, the cosine similarity (CS)-based regularization approach developed by Huang et al. [24] for lymphoma segmentation in weak-annotated PET/CT images is also adopted as a complement to the base BCE loss explained in Section 3.4.3. It is of special interest for this work the self-supervised term of the loss function proposed in [24], which enforces the extracted features of the predicted tumor samples to be similar to each other but dissimilar to the non-tumor samples in terms of cosine similarity. This mechanism is adapted so that it can be applied to discriminate brain cortical pixels from the rest. The same idea is transferred to be used with the adjusted GT, so the base BCE loss function includes the self-supervised element just described and a weakly supervised regularization term.
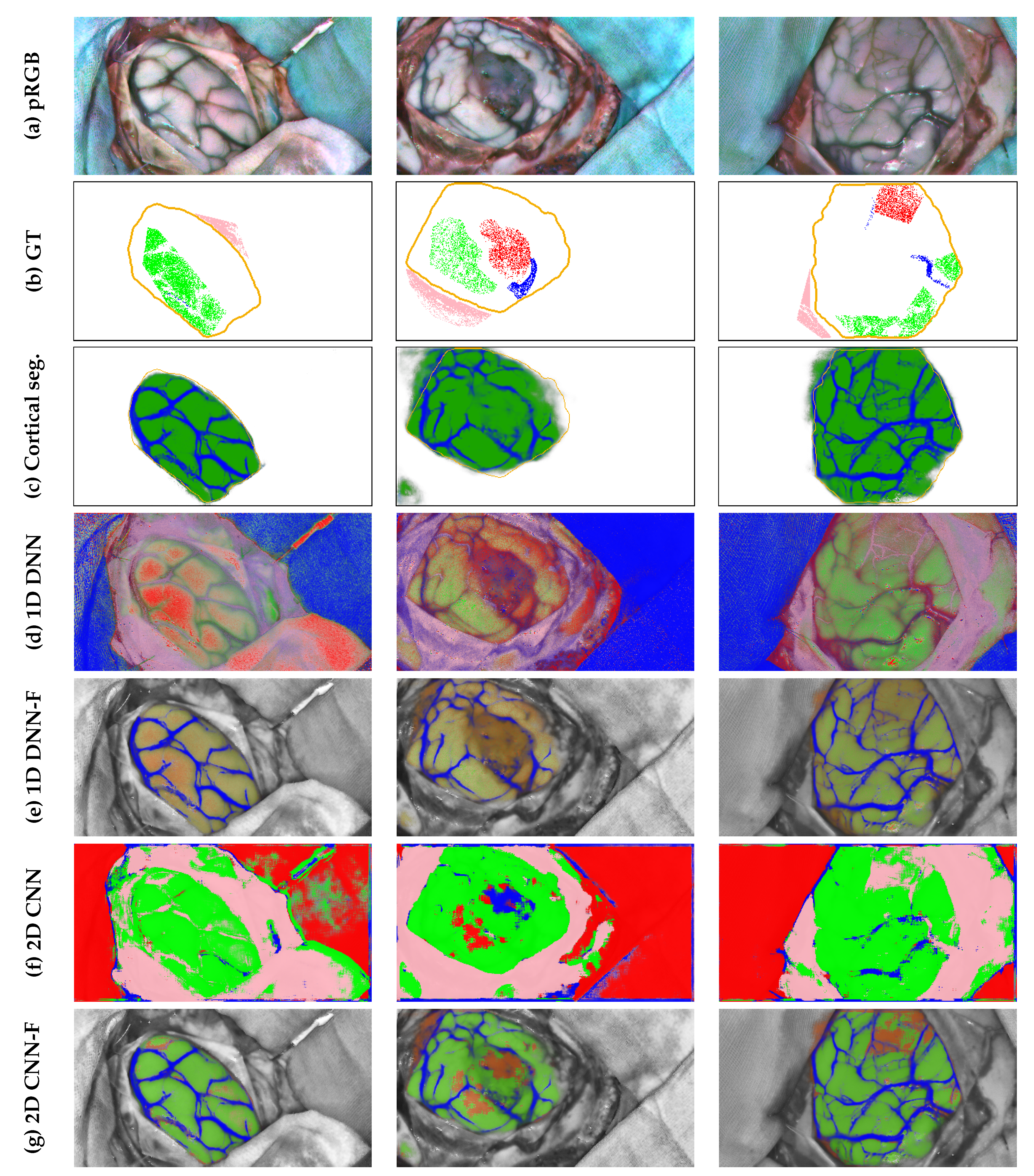
- One-dimensional deep neural network (1D_DNN), proposed by Fabelo et al. [59] and designed to work at HS single pixel level through a two hidden layers structure with 28 and 40 neurons respectively and a final output layer providing 4 different probabilities associated to each of the 4 tissues by softmax activation.
-
Two-dimensional convolutional neural network(2D_CNN), presented by Hao et al. [12] which implements a ResNet-18 architecture for processing 11 × 11 overlapping patches extracted from the HS cube to obtain the probabilities belonging to the 4 tissues to be segmented also using softmax activation.
4.3. Evaluation Metrics
4.4. Implementation Details
4.4.1. Brain Cortex and Vessels Segmentation Training Details
4.4.2. HS Tissue Segmentation Network Training Details
4.4.3. Software and Hardware Used
4.5. Quantitative Results
4.5.1. Comparative Analysis of Neural Network Architectures for Cortical Segmentation
4.5.2. Brain Surface and Cortical Vessels Segmentation
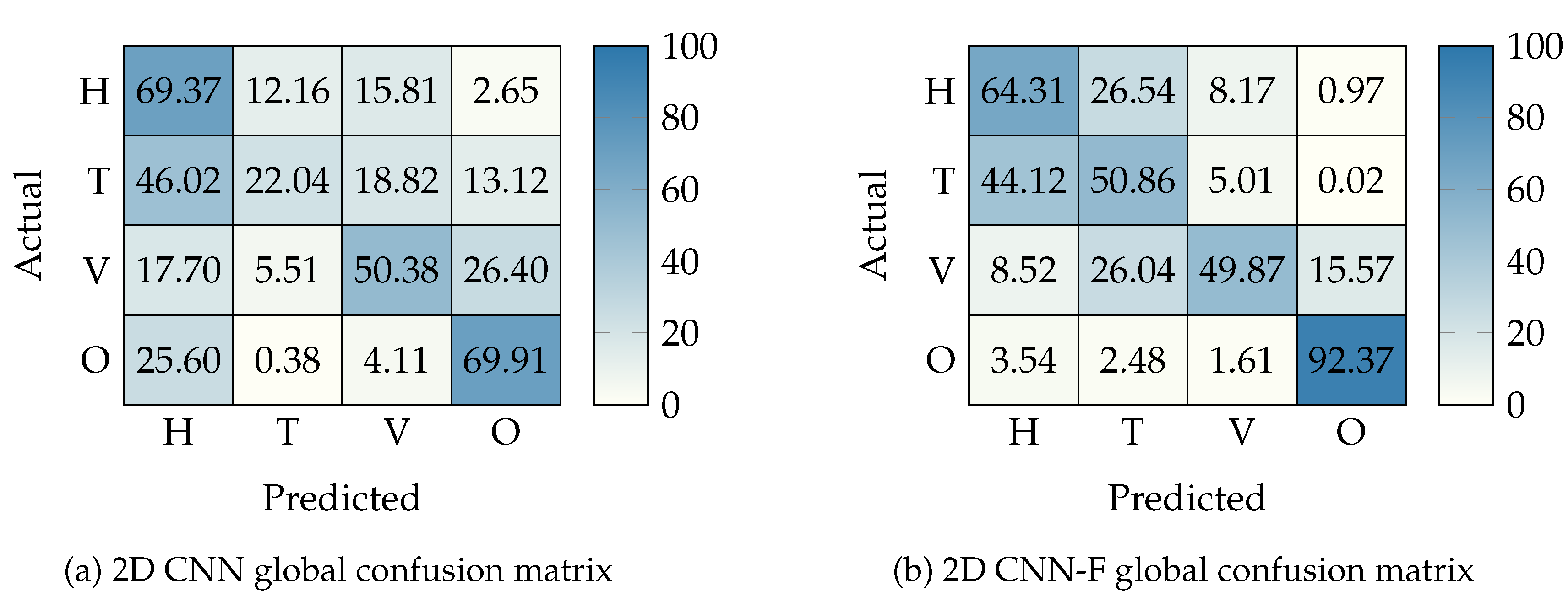
4.5.3. HS Tissue Segmentation
4.6. Qualitative Results
5. Discussion
5.1. Limitations
6. Conclusion
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Chan, H.P.; Hadjiiski, L.M.; Samala, R.K. Computer-aided diagnosis in the era of deep learning. Medical physics 2020, 47, e218–e227. [Google Scholar] [CrossRef]
- Lu, G.; Fei, B. Medical hyperspectral imaging: a review. Journal of Biomedical Optics 2014, 19, 010901. [Google Scholar] [CrossRef]
- Leon, R.; Gelado, S.H.; Fabelo, H.; Ortega, S.; Quintana, L.; Szolna, A.; Piñeiro, J.F.; Balea-Fernandez, F.; Morera, J.; Clavo, B.; et al. Hyperspectral imaging for in-vivo/ex-vivo tissue analysis of human brain cancer. In Proceedings of the Medical Imaging 2022: Image-Guided Procedures, Robotic Interventions, and Modeling; Linte, C.A., Siewerdsen, J.H., Eds.; International Society for Optics and Photonics, SPIE, 2022; Vol. 12034, p. 1203429. [Google Scholar] [CrossRef]
- Tajbakhsh, N.; Jeyaseelan, L.; Li, Q.; Chiang, J.N.; Wu, Z.; Ding, X. Embracing imperfect datasets: A review of deep learning solutions for medical image segmentation. Medical Image Analysis 2020, 63, 101693. [Google Scholar] [CrossRef]
- Seidlitz, S.; Sellner, J.; Odenthal, J.; Özdemir, B.; Studier-Fischer, A.; Knödler, S.; Ayala, L.; Adler, T.J.; Kenngott, H.G.; Tizabi, M.; et al. Robust deep learning-based semantic organ segmentation in hyperspectral images. Medical Image Analysis 2022, 80, 102488. [Google Scholar] [CrossRef]
- Leon, R.; Fabelo, H.; Ortega, S.; Cruz-Guerrero, I.A.; Campos-Delgado, D.U.; Szolna, A.; Piñeiro, J.F.; Espino, C.; O’Shanahan, A.J.; Hernandez, M.; et al. Hyperspectral imaging benchmark based on machine learning for intraoperative brain tumour detection. NPJ Precision Oncology 2023, 7, 119. [Google Scholar] [CrossRef] [PubMed]
- Urbanos, G.; Martín, A.; Vázquez, G.; Villanueva, M.; Villa, M.; Jimenez-Roldan, L.; Chavarrías, M.; Lagares, A.; Juárez, E.; Sanz, C. Supervised machine learning methods and hyperspectral imaging techniques jointly applied for brain cancer classification. Sensors 2021, 21, 3827. [Google Scholar] [CrossRef] [PubMed]
- Martín-Pérez, A.; Villa, M.; Rosa Olmeda, G.; Sancho, J.; Vazquez, G.; Urbanos, G.; Martinez de Ternero, A.; Chavarrías, M.; Jimenez-Roldan, L.; Perez-Nuñez, A.; et al. SLIMBRAIN database: A multimodal image database of in vivo human brains for tumour detection. Scientific Data 2025, 12, 836. [Google Scholar] [CrossRef] [PubMed]
- Fabelo, H.; Ortega, S.; Kabwama, S.; Callico, G.M.; Bulters, D.; Szolna, A.; Pineiro, J.F.; Sarmiento, R. HELICoiD project: A new use of hyperspectral imaging for brain cancer detection in real-time during neurosurgical operations. Proceedings of the Hyperspectral Imaging Sensors: Innovative Applications and Sensor Standards 2016. SPIE 2016, Vol. 9860, 986002. [Google Scholar]
- UPM; IMAS12. NEMESIS-3D-CM: clasificacióN intraopEratoria de tuMores cErebraleS mediante modelos InmerSivos 3D. Accessed. 2019.
- LeCun, Y.; Bottou, L.; Bengio, Y.; Haffner, P. Gradient-based learning applied to document recognition. Proceedings of the IEEE 1998, 86, 2278–2324. [Google Scholar] [CrossRef]
- Hao, Q.; Pei, Y.; Zhou, R.; Sun, B.; Sun, J.; Li, S.; Kang, X. Fusing multiple deep models for in vivo human brain hyperspectral image classification to identify glioblastoma tumor. IEEE Transactions on Instrumentation and Measurement 2021, 70, 1–14. [Google Scholar] [CrossRef]
- Luo, Y.W.; Chen, H.Y.; Li, Z.; Liu, W.P.; Wang, K.; Zhang, L.; Fu, P.; Yue, W.Q.; Bian, G.B. Fast instruments and tissues segmentation of micro-neurosurgical scene using high correlative non-local network. Computers in Biology and Medicine 2023, 153, 106531. [Google Scholar] [CrossRef]
- Fabelo, H.; Halicek, M.; Ortega, S.; Shahedi, M.; Szolna, A.; Piñeiro, J.F.; Sosa, C.; O’Shanahan, A.J.; Bisshopp, S.; Espino, C.; et al. Deep Learning-Based Framework for In Vivo Identification of Glioblastoma Tumor using Hyperspectral Images of Human Brain. Sensors 2019, 19. [Google Scholar] [CrossRef]
- Goni, M.R.; Ruhaiyem, N.I.R.; Mustapha, M.; Achuthan, A.; Che Mohd Nassir, C.M.N. Brain Vessel Segmentation Using Deep Learning—A Review. IEEE Access 2022, 10, 111322–111336. [Google Scholar] [CrossRef]
- Tetteh, G.; Efremov, V.; Forkert, N.D.; Schneider, M.; Kirschke, J.; Weber, B.; Zimmer, C.; Piraud, M.; Menze, B.H. Deepvesselnet: Vessel segmentation, centerline prediction, and bifurcation detection in 3-d angiographic volumes. Frontiers in Neuroscience 2020, 14, 592352. [Google Scholar] [CrossRef] [PubMed]
- Galdran, A.; Anjos, A.; Dolz, J.; Chakor, H.; Lombaert, H.; Ayed, I.B. State-of-the-art retinal vessel segmentation with minimalistic models. Scientific Reports 2022, 12, 6174. [Google Scholar] [CrossRef]
- Frangi, A.F.; Niessen, W.J.; Vincken, K.L.; Viergever, M.A. Multiscale vessel enhancement filtering. In Proceedings of the Medical Image Computing and Computer-Assisted Intervention—MICCAI’98: First International Conference, Cambridge, MA, USA, October 11–13, 1998 Proceedings; Springer, 1998; 1, pp. 130–137. [Google Scholar]
- Longo, A.; Morscher, S.; Najafababdi, J.M.; Jüstel, D.; Zakian, C.; Ntziachristos, V. Assessment of hessian-based Frangi vesselness filter in optoacoustic imaging. Photoacoustics 2020, 20, 100200. [Google Scholar] [CrossRef]
- Vazquez, G.; Villa, M.; Martín-Pérez, A.; Sancho, J.; Rosa, G.; Cebrián, P.L.; Sutradhar, P.; Ternero, A.M.d.; Chavarrías, M.; Lagares, A.; et al. Brain Blood Vessel Segmentation in Hyperspectral Images Through Linear Operators. In Proceedings of the International Workshop on Design and Architecture for Signal and Image Processing, 2023; Springer; pp. 28–39. [Google Scholar]
- Haouchine, N.; Nercessian, M.; Juvekar, P.; Golby, A.; Frisken, S. Cortical Vessel Segmentation for Neuronavigation Using Vesselness-Enforced Deep Neural Networks. IEEE Transactions on Medical Robotics and Bionics 2022, 4, 327–330. [Google Scholar] [CrossRef]
- Wu, Y.; Oda, M.; Hayashi, Y.; Takebe, T.; Nagata, S.; Wang, C.; Mori, K. Blood Vessel Segmentation From Low-Contrast and Wide-Field Optical Microscopic Images of Cranial Window by Attention-Gate-Based Network. In Proceedings of the Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR) Workshops, June 2022; pp. 1864–1873. [Google Scholar]
- Tajbakhsh, N.; Jeyaseelan, L.; Li, Q.; Chiang, J.N.; Wu, Z.; Ding, X. Embracing imperfect datasets: A review of deep learning solutions for medical image segmentation. Medical Image Analysis 2020, 63, 101693. [Google Scholar] [CrossRef]
- Huang, Z.; Guo, Y.; Zhang, N.; Huang, X.; Decazes, P.; Becker, S.; Ruan, S. Multi-scale feature similarity-based weakly supervised lymphoma segmentation in PET/CT images. Computers in Biology and Medicine 2022, 151, 106230. [Google Scholar] [CrossRef] [PubMed]
- Patel, G.; Dolz, J. Weakly supervised segmentation with cross-modality equivariant constraints. Medical image analysis 2022, 77, 102374. [Google Scholar] [CrossRef]
- Luo, X.; Hu, M.; Liao, W.; Zhai, S.; Song, T.; Wang, G.; Zhang, S. Scribble-supervised medical image segmentation via dual-branch network and dynamically mixed pseudo labels supervision. In Proceedings of the International Conference on Medical Image Computing and Computer-Assisted Intervention, 2022; Springer; pp. 528–538. [Google Scholar]
- Zhang, J.; Wang, G.; Xie, H.; Zhang, S.; Huang, N.; Zhang, S.; Gu, L. Weakly supervised vessel segmentation in X-ray angiograms by self-paced learning from noisy labels with suggestive annotation. Neurocomputing 2020, 417, 114–127. [Google Scholar] [CrossRef]
- Dang, V.N.; Galati, F.; Cortese, R.; Di Giacomo, G.; Marconetto, V.; Mathur, P.; Lekadir, K.; Lorenzi, M.; Prados, F.; Zuluaga, M.A. Vessel-CAPTCHA: An efficient learning framework for vessel annotation and segmentation. Medical Image Analysis 2022, 75, 102263. [Google Scholar] [CrossRef] [PubMed]
- Galati, F.; Falcetta, D.; Cortese, R.; Casolla, B.; Prados, F.; Burgos, N.; Zuluaga, M.A. A2V: A Semi-Supervised Domain Adaptation Framework for Brain Vessel Segmentation via Two-Phase Training Angiography-to-Venography Translation. In Proceedings of the 34th British Machine Vision Conference 2023, BMVC 2023, Aberdeen, UK, November 20-24, 2023; BMVA, 2023. [Google Scholar]
- Gu, R.; Zhang, J.; Wang, G.; Lei, W.; Song, T.; Zhang, X.; Li, K.; Zhang, S. Contrastive semi-supervised learning for domain adaptive segmentation across similar anatomical structures. IEEE Transactions on Medical Imaging 2022, 42, 245–256. [Google Scholar] [CrossRef] [PubMed]
- Berger, A.H.; Lux, L.; Shit, S.; Ezhov, I.; Kaissis, G.; Menten, M.J.; Rueckert, D.; Paetzold, J.C. Cross-Domain and Cross-Dimension Learning for Image-to-Graph Transformers. In Proceedings of the 2025 IEEE/CVF Winter Conference on Applications of Computer Vision (WACV), 2025; IEEE; pp. 64–74. [Google Scholar]
- Galati, F.; Cortese, R.; Prados, F.; Lorenzi, M.; Zuluaga, M.A. Federated multi-centric image segmentation with uneven label distribution. In Proceedings of the International Conference on Medical Image Computing and Computer-Assisted Intervention, 2024; Springer; pp. 350–360. [Google Scholar]
- Butoi, V.I.; Ortiz, J.J.G.; Ma, T.; Sabuncu, M.R.; Guttag, J.; Dalca, A.V. Universeg: Universal medical image segmentation. In Proceedings of the Proceedings of the IEEE/CVF International Conference on Computer Vision, 2023; pp. 21438–21451. [Google Scholar]
- Ma, J.; He, Y.; Li, F.; Han, L.; You, C.; Wang, B. Segment Anything in Medical Images. Nature Communications 2024, 15, 654. [Google Scholar] [CrossRef]
- Yuan, Y.; Zheng, X.; Lu, X. Hyperspectral Image Superresolution by Transfer Learning. IEEE Journal of Selected Topics in Applied Earth Observations and Remote Sensing 2017, 10, 1963–1974. [Google Scholar] [CrossRef]
- Ye, S.; Li, N.; Xue, J.; Long, Y.; Jia, S. HSI-DETR: A DETR-based Transfer Learning from RGB to Hyperspectral Images for Object Detection of Live and Dead Cells: To achieve better results, convert models with the fewest changes from RGB to HSI. In Proceedings of the Proceedings of the 2022 11th International Conference on Computing and Pattern Recognition, New York, NY, USA, 2023; ICCPR ’22, pp. 102–107. [Google Scholar] [CrossRef]
- GmbH, X. Manual for smallest Hyperspectral USB3 based camera family xiSpec; XIMEA GmbH.
- Fabelo, H.; Ortega, S.; Szolna, A.; Bulters, D.; Piñeiro, J.F.; Kabwama, S.; J-O’Shanahan, A.; Bulstrode, H.; Bisshopp, S.; Kiran, B.R.; et al. In-Vivo Hyperspectral Human Brain Image Database for Brain Cancer Detection. IEEE Access 2019, 7, 39098–39116. [Google Scholar] [CrossRef]
- Kruse, F.; Lefkoff, A.; Boardman, J.; Heidebrecht, K.; Shapiro, A.; Barloon, P.; Goetz, A. The spectral image processing system (SIPS)—interactive visualization and analysis of imaging spectrometer data. Remote Sensing of Environment;Airbone Imaging Spectrometry 1993, 44, 145–163, Airbone Imaging Spectrometry. [Google Scholar] [CrossRef]
- Lloyd, S. Least squares quantization in PCM. IEEE Transactions on Information Theory 1982, 28, 129–137. [Google Scholar] [CrossRef]
- Canny, J. A Computational Approach to Edge Detection. IEEE Transactions on Pattern Analysis and Machine Intelligence 1986, PAMI-8, 679–698. [Google Scholar] [CrossRef]
- Caliński, T.; Harabasz, J. A dendrite method for cluster analysis. Communications in Statistics 1974, 3, 1–27. Available online: https://www.tandfonline.com/doi/pdf/10.1080/03610927408827101. [CrossRef]
- Szegedy, C.; Vanhoucke, V.; Ioffe, S.; Shlens, J.; Wojna, Z. Rethinking the Inception Architecture for Computer Vision. CoRR 2015. abs/1512.00567, [1512.00567. [Google Scholar]
- Achanta, R.; Shaji, A.; Smith, K.; Lucchi, A.; Fua, P.; Süsstrunk, S. SLIC Superpixels Compared to State-of-the-Art Superpixel Methods. IEEE Transactions on Pattern Analysis and Machine Intelligence 2012, 34, 2274–2282. [Google Scholar] [CrossRef] [PubMed]
- Ricci, E.; Perfetti, R. Retinal Blood Vessel Segmentation Using Line Operators and Support Vector Classification. IEEE Transactions on Medical Imaging 2007, 26, 1357–1365. [Google Scholar] [CrossRef]
- Akiba, T.; Sano, S.; Yanase, T.; Ohta, T.; Koyama, M. Optuna: A Next-generation Hyperparameter Optimization Framework. 2019. [Google Scholar] [CrossRef]
- Scheeren, T.; Schober, P.; Schwarte, L. Monitoring tissue oxygenation by near infrared spectroscopy (NIRS): background and current applications. Journal of clinical monitoring and computing 2012, 26, 279–287. [Google Scholar] [CrossRef]
- He, T.; Zhang, Z.; Zhang, H.; Zhang, Z.; Xie, J.; Li, M. Bag of Tricks for Image Classification with Convolutional Neural Networks. In Proceedings of the Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR), June 2019. [Google Scholar]
- Ioffe, S.; Szegedy, C. Batch Normalization: Accelerating Deep Network Training by Reducing Internal Covariate Shift, 2015. arXiv. [arXiv:cs.LG/1502.03167].
- Maas, A.L.; Hannun, A.Y.; Ng, A.Y.; et al. Rectifier nonlinearities improve neural network acoustic models. In Proceedings of the Proc. icml. Atlanta, GA, 2013; Vol. 30, p. 3. [Google Scholar]
- He, K.; Zhang, X.; Ren, S.; Sun, J. Deep Residual Learning for Image Recognition. In Proceedings of the Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition (CVPR), June 2016. [Google Scholar]
- Khosla, P.; Teterwak, P.; Wang, C.; Sarna, A.; Tian, Y.; Isola, P.; Maschinot, A.; Liu, C.; Krishnan, D. Supervised contrastive learning. Advances in neural information processing systems 2020, 33, 18661–18673. [Google Scholar]
- Chen, T.; Kornblith, S.; Norouzi, M.; Hinton, G. A simple framework for contrastive learning of visual representations. In Proceedings of the International conference on machine learning. PMLR, 2020; pp. 1597–1607. [Google Scholar]
- Dice, L.R. Measures of the Amount of Ecologic Association Between Species. Ecology 1945, 26, 297–302. Available online: https://esajournals.onlinelibrary.wiley.com/doi/pdf/10.2307/1932409. [CrossRef]
- Marmanis, D.; Schindler, K.; Wegner, J.D.; Galliani, S.; Datcu, M.; Stilla, U. Classification with an edge: Improving semantic image segmentation with boundary detection. ISPRS Journal of Photogrammetry and Remote Sensing 2018, 135, 158–172. [Google Scholar] [CrossRef]
- Park, J.S.; Oh, S.J. A new concave hull algorithm and concaveness measure for n-dimensional datasets. Journal of Information science and engineering 2012, 28, 587–600. [Google Scholar]
- Guo, Z.; Liu, C.; Zhang, X.; Jiao, J.; Ji, X.; Ye, Q. Beyond Bounding-Box: Convex-Hull Feature Adaptation for Oriented and Densely Packed Object Detection. In Proceedings of the Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR), June 2021; pp. 8792–8801. [Google Scholar]
- Cohen, T.; Welling, M. Group equivariant convolutional networks. In Proceedings of the International conference on machine learning. PMLR, 2016; pp. 2990–2999. [Google Scholar]
- Fabelo, H.; Halicek, M.; Ortega, S.; Shahedi, M.; Szolna, A.; Piñeiro, J.F.; Sosa, C.; O’Shanahan, A.J.; Bisshopp, S.; Espino, C.; et al. Deep learning-based framework for in vivo identification of glioblastoma tumor using hyperspectral images of human brain. Sensors 2019, 19, 920. [Google Scholar] [CrossRef]
- Heimann, T.; van Ginneken, B.; Styner, M.A.; Arzhaeva, Y.; Aurich, V.; Bauer, C.; Beck, A.; Becker, C.; Beichel, R.; Bekes, G.; et al. Comparison and Evaluation of Methods for Liver Segmentation From CT Datasets. IEEE Transactions on Medical Imaging 2009, 28, 1251–1265. [Google Scholar] [CrossRef]
- Fawcett, T. An introduction to ROC analysis. Pattern Recognition Letters;ROC Analysis in Pattern Recognition 2006, 27, 861–874. [Google Scholar] [CrossRef]
- Loshchilov, I.; Hutter, F. Decoupled weight decay regularization. arXiv preprint arXiv:1711.05101. [CrossRef]
- Loshchilov, I.; Hutter, F. Sgdr: Stochastic gradient descent with warm restarts. arXiv preprint arXiv:1608.03983. [CrossRef]
- You, Y.; Gitman, I.; Ginsburg, B. Large batch training of convolutional networks. arXiv arXiv:1708.03888. [CrossRef]
- Paszke, A.; Gross, S.; Massa, F.; Lerer, A.; Bradbury, J.; Chanan, G.; Killeen, T.; Lin, Z.; Gimelshein, N.; Antiga, L.; et al. Pytorch: An imperative style, high-performance deep learning library. Advances in neural information processing systems 2019, 32. [Google Scholar]
- Jha, D.; Smedsrud, P.H.; Riegler, M.A.; Johansen, D.; Lange, T.D.; Halvorsen, P.; Johansen, D.H. ResUNet++: An Advanced Architecture for Medical Image Segmentation. In Proceedings of the Proceedings of the IEEE International Symposium on Multimedia (ISM), 2019; pp. 225–230. [Google Scholar]
- Roy, S.; Koehler, G.; Ulrich, C.; Baumgartner, M.; Petersen, J.; Isensee, F.; Jaeger, P.F.; Maier-Hein, K.H. Mednext: transformer-driven scaling of convnets for medical image segmentation. In Proceedings of the International Conference on Medical Image Computing and Computer-Assisted Intervention, 2023; Springer; pp. 405–415. [Google Scholar]
- Isensee, F.; Wald, T.; Ulrich, C.; Baumgartner, M.; Roy, S.; Maier-Hein, K.; Jaeger, P.F. nnu-net revisited: A call for rigorous validation in 3d medical image segmentation. In Proceedings of the International Conference on Medical Image Computing and Computer-Assisted Intervention, 2024; Springer; pp. 488–498. [Google Scholar]
- Frangi, A.F.; Niessen, W.J.; Vincken, K.L.; Viergever, M.A. Multiscale vessel enhancement filtering. In Proceedings of the International conference on medical image computing and computer-assisted intervention, 1998; Springer; pp. 130–137. [Google Scholar]
- Ibtehaz, N.; Rahman, M.S. MultiResUNet: Rethinking the U-Net architecture for multimodal biomedical image segmentation. Neural networks 2020, 121, 74–87. [Google Scholar] [CrossRef] [PubMed]
| 1 | |
| 2 |

| Healthy | Tumor | Blood vessels | Dura mater | Total | |
|---|---|---|---|---|---|
| Annotated pixels | 182 314 | 32 702 | 18 662 | 80 312 | 313 990 |
| Band | Kernel 1 | Kernel 2 | Thresh. 1 | Thresh. 2 |
|---|---|---|---|---|
| 7 | 7 | 25 | 40 | 900 |
| Class | Total pix. | Val-train pix. | Test pix. |
|---|---|---|---|
| Healthy | 182 314 | 146 986 | 35 328 |
| Tumor | 32 702 | 28 702 | 4 000 |
| Blood vessels | 18 662 | 13 587 | 5 075 |
| Dura mater | 80 312 | 65 944 | 14 368 |
| Cortex | Vessel | ||||
|---|---|---|---|---|---|
| NN model | DSC | ASSD | VHR | VER | |
| ResUNet [66] |
|
|
|
|
|
| MedNeXt [67] |
|
|
|
|
|
| HSI-ResNet |
|
|
|
|
|
| Cortex | Vessel | ||||
|---|---|---|---|---|---|
| Method | DSC | ASSD | VHR | VER | |
| Vessel pseudo-labels | - | - | |||
| HSI refined annotations | - | - | |||
| (a) | HSI solo training | ||||
| Encoder pre-training + HSI fine-tuning | |||||
| Encoder pre-training + RGB training | |||||
| RGB training + HSI fine-tuning | |||||
| (b) | HSI solo training with refined annotations | ||||
| Fully pre-trained with HSI refined annotations | |||||
| Fully pre-trained + pRGB fine-tuning | |||||
| Vessel-CAPTCHA [28] | |||||
| Frangi [69] | |||||
| (c) | UniverSeg [33] | ||||
| MultiResUNet [70] | |||||
| Cosine similarity [24] | |||||
| CS-CADA [30] | |||||
| Equivariance [25] | |||||
| MedSAM [34] | |||||
| Fully pre-trained segmentation (proposed) | |||||
| (d) | Proposed + CS | ||||
| Proposed + EV | |||||
| Method | F1 | AUC | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| H | T | V | O | mF1 | H | T | V | O | mAUC | ||
| 1D DNN | |||||||||||
| 1D DNN-F | |||||||||||
| 1D Difference | |||||||||||
| 2D CNN | |||||||||||
| 2D CNN-F | |||||||||||
| 2D Difference | |||||||||||
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.




